Abstract
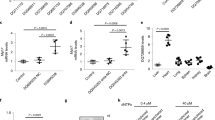
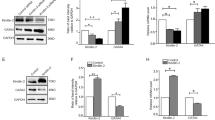
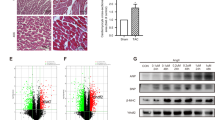
Histone modification plays an important role in pathological cardiac hypertrophy and heart failure. In this study we investigated the role of a histone arginine demethylase, Jumonji C domain-containing protein 6 (JMJD6) in pathological cardiac hypertrophy. Cardiac hypertrophy was induced in rats by subcutaneous injection of isoproterenol (ISO, 1.2 mg·kg−1·d−1) for a week. At the end of the experiment, the rats underwent echocardiography, followed by euthanasia and heart collection. We found that JMJD6 levels were compensatorily increased in ISO-induced hypertrophic cardiac tissues, but reduced in patients with heart failure with reduced ejection fraction (HFrEF). Furthermore, we demonstrated that JMJD6 overexpression significantly attenuated ISO-induced hypertrophy in neonatal rat cardiomyocytes (NRCMs) evidenced by the decreased cardiomyocyte surface area and hypertrophic genes expression. Cardiac-specific JMJD6 overexpression in rats protected the hearts against ISO-induced cardiac hypertrophy and fibrosis, and rescued cardiac function. Conversely, depletion of JMJD6 by single-guide RNA (sgRNA) exacerbated ISO-induced hypertrophic responses in NRCMs. We revealed that JMJD6 interacted with NF-κB p65 in cytoplasm and reduced nuclear levels of p65 under hypertrophic stimulation in vivo and in vitro. Mechanistically, JMJD6 bound to p65 and demethylated p65 at the R149 residue to inhibit the nuclear translocation of p65, thus inactivating NF-κB signaling and protecting against pathological cardiac hypertrophy. In addition, we found that JMJD6 demethylated histone H3R8, which might be a new histone substrate of JMJD6. These results suggest that JMJD6 may be a potential target for therapeutic interventions in cardiac hypertrophy and heart failure.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Virani SS, Alonso A, Benjamin EJ, Bittencourt MS, Callaway CW, Carson AP, et al. Heart disease and stroke statistics-2020 update: a report from the American Heart Association. Circulation. 2020;141:e139–e596.
Frey N, Katus HA, Olson EN, Hill JA. Hypertrophy of the heart: a new therapeutic target? Circulation. 2004;109:1580–9.
Maillet M, van Berlo JH, Molkentin JD. Molecular basis of physiological heart growth: fundamental concepts and new players. Nat Rev Mol Cell Biol. 2013;14:38–48.
MacDonald MR, Petrie MC, Hawkins NM, Petrie JR, Fisher M, McKelvie R, et al. Diabetes, left ventricular systolic dysfunction, and chronic heart failure. Eur Heart J. 2008;29:1224–40.
Mano H. Epigenetic abnormalities in cardiac hypertrophy and heart failure. Environ Health Prev Med. 2008;13:25–29.
Lin Z, Li Z, Guo Z, Cao Y, Li J, Liu P, et al. Epigenetic reader bromodomain containing protein 2 facilitates pathological cardiac hypertrophy via regulating the expression of citrate cycle genes. Front Pharmacol. 2022;13:887991.
Liu CF, Tang WHW. Epigenetics in cardiac hypertrophy and heart failure. JACC Basic Transl Sci. 2019;4:976–93.
Li Q, Li ZM, Sun SY, Wang LP, Wang PX, Guo Z, et al. PARP1 interacts with HMGB1 and promotes its nuclear export in pathological myocardial hypertrophy. Acta Pharmacol Sin. 2019;40:589–98.
Li Z, Guo Z, Lan R, Cai S, Lin Z, Li J, et al. The poly(ADP-ribosyl)ation of BRD4 mediated by PARP1 promoted pathological cardiac hypertrophy. Acta Pharm Sin B. 2021;11:1286–99.
Greer EL, Shi Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nat Rev Genet. 2012;13:343–57.
Bannister AJ, Kouzarides T. Reversing histone methylation. Nature. 2005;436:1103–6.
Bannister AJ, Kouzarides T. Regulation of chromatin by histone modifications. Cell Res. 2011;21:381–95.
Zhao Z, Shilatifard A. Epigenetic modifications of histones in cancer. Genome Biol. 2019;20:245.
Peter CJ, Akbarian S. Balancing histone methylation activities in psychiatric disorders. Trends Mol Med. 2011;17:372–9.
Cai S, Wang P, Xie T, Li Z, Li J, Lan R, et al. Histone H4R3 symmetric di-methylation by prmt5 protects against cardiac hypertrophy via regulation of Filip1L/beta-catenin. Pharmacol Res. 2020;161:105104.
Pyun JH, Kim HJ, Jeong MH, Ahn BY, Vuong TA, Lee DI, et al. Cardiac specific PRMT1 ablation causes heart failure through CaMKII dysregulation. Nat Commun. 2018;9:5107.
Zhang QJ, Liu ZP. Histone methylations in heart development, congenital and adult heart diseases. Epigenomics. 2015;7:321–30.
Zhang QJ, Chen HZ, Wang L, Liu DP, Hill JA, Liu ZP. The histone trimethyllysine demethylase JMJD2A promotes cardiac hypertrophy in response to hypertrophic stimuli in mice. J Clin Invest. 2011;121:2447–56.
Guo Z, Lu J, Li J, Wang P, Li Z, Zhong Y, et al. JMJD3 inhibition protects against isoproterenol-induced cardiac hypertrophy by suppressing beta-MHC expression. Mol Cell Endocrinol. 2018;477:1–14.
Chang B, Chen Y, Zhao Y, Bruick RK. JMJD6 is a histone arginine demethylase. Science. 2007;318:444–7.
Liu W, Ma Q, Wong K, Li W, Ohgi K, Zhang J, et al. Brd4 and JMJD6-associated anti-pause enhancers in regulation of transcriptional pause release. Cell. 2013;155:1581–95.
Unoki M, Masuda A, Dohmae N, Arita K, Yoshimatsu M, Iwai Y, et al. Lysyl 5-hydroxylation, a novel histone modification, by Jumonji domain containing 6 (JMJD6). J Biol Chem. 2013;288:6053–62.
Webby CJ, Wolf A, Gromak N, Dreger M, Kramer H, Kessler B, et al. Jmjd6 catalyses lysyl-hydroxylation of U2AF65, a protein associated with RNA splicing. Science. 2009;325:90–93.
Poulard C, Rambaud J, Hussein N, Corbo L, Le Romancer M. JMJD6 regulates ERalpha methylation on arginine. PLoS One. 2014;9:e87982.
Tikhanovich I, Kuravi S, Artigues A, Villar MT, Dorko K, Nawabi A, et al. Dynamic arginine methylation of tumor necrosis factor (TNF) receptor-associated factor 6 regulates toll-like receptor signaling. J Biol Chem. 2015;290:22236–49.
Tsai WC, Reineke LC, Jain A, Jung SY, Lloyd RE. Histone arginine demethylase JMJD6 is linked to stress granule assembly through demethylation of the stress granule-nucleating protein G3BP1. J Biol Chem. 2017;292:18886–96.
Vrankova S, Barta A, Klimentova J, Dovinova I, Liskova S, Dobesova Z, et al. The regulatory role of nuclear factor kappa B in the heart of hereditary hypertriglyceridemic rat. Oxid Med Cell Longev. 2016;2016:9814038.
Huxford T, Huang DB, Malek S, Ghosh G. The crystal structure of the IkappaBalpha/NF-kappaB complex reveals mechanisms of NF-kappaB inactivation. Cell. 1998;95:759–70.
Jacobs MD, Harrison SC. Structure of an IkappaBalpha/NF-kappaB complex. Cell. 1998;95:749–58.
Huang B, Yang XD, Lamb A, Chen LF. Posttranslational modifications of NF-kappaB: another layer of regulation for NF-kappaB signaling pathway. Cell Signal. 2010;22:1282–90.
Lu T, Stark GR. NF-kappaB: regulation by methylation. Cancer Res. 2015;75:3692–5.
Ea CK, Baltimore D. Regulation of NF-kappaB activity through lysine monomethylation of p65. Proc Natl Acad Sci USA. 2009;106:18972–7.
Levy D, Kuo AJ, Chang Y, Schaefer U, Kitson C, Cheung P, et al. Lysine methylation of the NF-kappaB subunit RelA by SETD6 couples activity of the histone methyltransferase GLP at chromatin to tonic repression of NF-kappaB signaling. Nat Immunol. 2011;12:29–36.
Lu T, Jackson MW, Wang B, Yang M, Chance MR, Miyagi M, et al. Regulation of NF-kappaB by NSD1/FBXL11-dependent reversible lysine methylation of p65. Proc Natl Acad Sci USA. 2010;107:46–51.
Yang XD, Huang B, Li M, Lamb A, Kelleher NL, Chen LF. Negative regulation of NF-kappaB action by Set9-mediated lysine methylation of the RelA subunit. EMBO J. 2009;28:1055–66.
Wei H, Wang B, Miyagi M, She Y, Gopalan B, Huang DB, et al. PRMT5 dimethylates R30 of the p65 subunit to activate NF-kappaB. Proc Natl Acad Sci USA. 2013;110:13516–21.
Hu Y, Guo Z, Lu J, Wang P, Sun S, Zhang Y, et al. sFRP1 has a biphasic effect on doxorubicin-induced cardiotoxicity in a cellular location-dependent manner in NRCMs and Rats. Arch Toxicol. 2019;93:533–46.
Liang L, Tu Y, Lu J, Wang P, Guo Z, Wang Q, et al. Dkk1 exacerbates doxorubicin-induced cardiotoxicity by inhibiting the Wnt/beta-catenin signaling pathway. J Cell Sci. 2019;132:jcs228478.
Sun S, Hu Y, Zheng Q, Guo Z, Sun D, Chen S, et al. Poly(ADP-ribose) polymerase 1 induces cardiac fibrosis by mediating mammalian target of rapamycin activity. J Cell Biochem. 2019;120:4813–26.
Wang P, Lan R, Guo Z, Cai S, Wang J, Wang Q, et al. Histone demethylase JMJD3 mediated doxorubicin-induced cardiomyopathy by suppressing SESN2 expression. Front Cell Dev Biol. 2020;8:548605.
Li J, Huang J, Lu J, Guo Z, Li Z, Gao H, et al. Sirtuin 1 represses PKC-zeta activity through regulating interplay of acetylation and phosphorylation in cardiac hypertrophy. Br J Pharmacol. 2019;176:416–35.
Lu J, Li J, Hu Y, Guo Z, Sun D, Wang P, et al. Chrysophanol protects against doxorubicin-induced cardiotoxicity by suppressing cellular PARylation. Acta Pharm Sin B 2019;9:782–93.
Guo Z, Liao Z, Huang L, Liu D, Yin D, He M. Kaempferol protects cardiomyocytes against anoxia/reoxygenation injury via mitochondrial pathway mediated by SIRT1. Eur J Pharmacol. 2015;761:245–53.
Li Z, Zhang X, Guo Z, Zhong Y, Wang P, Li J, et al. SIRT6 suppresses NFATc4 expression and activation in cardiomyocyte hypertrophy. Front Pharmacol. 2018;9:1519.
Guo Z, Valenzuela Ripoll C, Picataggi A, Rawnsley D, Ozcan M, Chirinos J, et al. Apolipoprotein M attenuates anthracycline cardiotoxicity and lysosomal injury. JACC Basic Transl Sci. 2023;8:340–55.
Ozcan M, Guo Z, Valenzuela Ripoll C, Diab A, Picataggi A, Rawnsley D, et al. Sustained alternate day fasting potentiates doxorubicin cardiotoxicity. Cell Metab. 2023. Online ahead of print. https://doi.org/10.1016/j.cmet.2023.02.006.
Guo Z, Zhang Y, Liu C, Youn JY, Cai H. Toll-like receptor 2 (TLR2) knockout abrogates diabetic and obese phenotypes while restoring endothelial function via inhibition of NOX1. Diabetes. 2021;70:2107–19.
Luo W, Wang Y, Yang H, Dai C, Hong H, Li J, et al. Heme oxygenase-1 ameliorates oxidative stress-induced endothelial senescence via regulating endothelial nitric oxide synthase activation and coupling. Aging (Albany NY). 2018;10:1722–44.
Chen H, Xue Y, Huang N, Yao X, Sun Z. MeMo: a web tool for prediction of protein methylation modifications. Nucleic Acids Res. 2006;34:W249–W253.
Javan H, Szucsik AM, Li L, Schaaf CL, Salama ME, Selzman CH. Cardiomyocyte p65 nuclear factor-kappaB is necessary for compensatory adaptation to pressure overload. Circ Heart Fail. 2015;8:109–18.
Gupta S, Young D, Maitra RK, Gupta A, Popovic ZB, Yong SL, et al. Prevention of cardiac hypertrophy and heart failure by silencing of NF-kappaB. J Mol Biol. 2008;375:637–49.
Hong HQ, Lu J, Fang XL, Zhang YH, Cai Y, Yuan J, et al. G3BP2 is involved in isoproterenol-induced cardiac hypertrophy through activating the NF-kappaB signaling pathway. Acta Pharmacol Sin. 2018;39:184–94.
Yu SS, Cai Y, Ye JT, Pi RB, Chen SR, Liu PQ, et al. Sirtuin 6 protects cardiomyocytes from hypertrophy in vitro via inhibition of NF-kappaB-dependent transcriptional activity. Br J Pharmacol. 2013;168:117–28.
Sakurai H, Chiba H, Miyoshi H, Sugita T, Toriumi W. IkappaB kinases phosphorylate NF-kappaB p65 subunit on serine 536 in the transactivation domain. J Biol Chem. 1999;274:30353–6.
Sasaki CY, Barberi TJ, Ghosh P, Longo DL. Phosphorylation of RelA/p65 on serine 536 defines an I{kappa}B{alpha}-independent NF-{kappa}B pathway. J Biol Chem. 2005;280:34538–47.
Backs J, Olson EN. Control of cardiac growth by histone acetylation/deacetylation. Circ Res. 2006;98:15–24.
Ooi JY, Tuano NK, Rafehi H, Gao XM, Ziemann M, Du XJ, et al. HDAC inhibition attenuates cardiac hypertrophy by acetylation and deacetylation of target genes. Epigenetics. 2015;10:418–30.
Papait R, Cattaneo P, Kunderfranco P, Greco C, Carullo P, Guffanti A, et al. Genome-wide analysis of histone marks identifying an epigenetic signature of promoters and enhancers underlying cardiac hypertrophy. Proc Natl Acad Sci USA. 2013;110:20164–9.
Zang R, Tan Q, Zeng F, Wang D, Yu S, Wang Q. JMJD1A represses the development of cardiomyocyte hypertrophy by regulating the expression of catalase. Biomed Res Int. 2020;2020:5081323.
Yu S, Li Y, Zhao H, Wang Q, Chen P. The Histone demethylase JMJD1C regulates CAMKK2-AMPK signaling to participate in cardiac hypertrophy. Front Physiol. 2020;11:539.
Rosales W, Lizcano F. The histone demethylase JMJD2A modulates the induction of hypertrophy markers in iPSC-derived cardiomyocytes. Front Genet. 2018;9:14.
Bristow MR, Kao DP, Breathett KK, Altman NL, Gorcsan J 3rd, Gill EA, et al. Structural and functional phenotyping of the failing heart: Is the left ventricular ejection fraction obsolete? JACC Heart Fail. 2017;5:772–81.
Yancy CW, Jessup M, Bozkurt B, Butler J, Casey DE Jr, Drazner MH, et al. 2013 ACCF/AHA guideline for the management of heart failure: executive summary: a report of the American College of Cardiology Foundation/American Heart Association Task Force on practice guidelines. Circulation. 2013;128:1810–52.
Purcell NH, Tang G, Yu C, Mercurio F, DiDonato JA, Lin A. Activation of NF-kappa B is required for hypertrophic growth of primary rat neonatal ventricular cardiomyocytes. Proc Natl Acad Sci USA. 2001;98:6668–73.
Zelarayan L, Renger A, Noack C, Zafiriou MP, Gehrke C, van der Nagel R, et al. NF-kappaB activation is required for adaptive cardiac hypertrophy. Cardiovasc Res. 2009;84:416–24.
Zou J, Li H, Chen X, Zeng S, Ye J, Zhou C, et al. C/EBPbeta knockdown protects cardiomyocytes from hypertrophy via inhibition of p65-NFkappaB. Mol Cell Endocrinol. 2014;390:18–25.
Sizemore N, Lerner N, Dombrowski N, Sakurai H, Stark GR. Distinct roles of the Ikappa B kinase alpha and beta subunits in liberating nuclear factor kappa B (NF-kappa B) from Ikappa B and in phosphorylating the p65 subunit of NF-kappa B. J Biol Chem. 2002;277:3863–9.
Acknowledgements
ZG was supported by the American Heart Association Postdoctoral Fellowship (898679). JL was supported by National Natural Science Foundation of China (82173808), Natural Science Foundation of Guangdong Province (2021B1515020100) and Guangzhou Basic and Applied Basic Research Project (202102020173). WWL was supported by National Natural Science Foundation of China (82003746). PQL was supported by National Natural Science Foundation of China (U21A20419, 81872860), Local Innovative and Research Teams Project of Guangdong Pearl River Talents Program (2017BT01Y093), and National Engineering and Technology Research Center for New drug Druggability Evaluation (Seed Program of Guangdong Province, 2017B090903004). JGZ was supported by Academic Promotion Program of Shandong First Medical University (2019LJ003). LW was supported by the Project of Shandong Medical and Health Science and Technology (202002040923).
Author information
Authors and Affiliations
Contributions
ZG and YHH performed the study, analyzed the data, wrote and revised the manuscript. GSF, ZZL, SDC, QQW and WWL contributed to the animal experiments. GSF, QL, LYL, LW and JGZ contributed to the acquisition of data and data interpretation. CVR contributed to the language polish. AJ provided the TAC mice cardiac tissues. ZKW provided heart tissues from humans with heart failure. ZG, JL and PQL made the hypothesis and participated in the experimental design, and manuscript preparation. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, Z., Hu, Yh., Feng, Gs. et al. JMJD6 protects against isoproterenol-induced cardiac hypertrophy via inhibition of NF-κB activation by demethylating R149 of the p65 subunit. Acta Pharmacol Sin 44, 1777–1789 (2023). https://doi.org/10.1038/s41401-023-01086-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-023-01086-7