Abstract
Liver, as an immune and detoxification organ, represents an important line of defense against bacteria and infection and a vulnerable organ that is easily injured during sepsis. Artesunate (ART) is an anti-malaria agent, that also exhibits broad pharmacological activities including anti-inflammatory, immune-regulation and liver protection. In this study, we investigated the cellular responses in liver to sepsis infection and ART hepatic-protective mechanisms against sepsis. Cecal ligation and puncture (CLP)-induced sepsis model was established in mice. The mice were administered ART (10 mg/kg, i.p.) at 4 h, and sacrificed at 12 h after the surgery. Liver samples were collected for preparing single-cell RNA transcriptome sequencing (scRNA-seq). The scRNA-seq analysis revealed that sepsis-induced a dramatic reduction of hepatic endothelial cells, especially the subtypes characterized with proliferation and differentiation. Macrophages were recruited during sepsis and released inflammatory cytokines (Tnf, Il1b, Il6), chemokines (Ccl6, Cd14), and transcription factor (Nfkb1), resulting in liver inflammatory responses. Massive apoptosis of lymphocytes and abnormal recruitment of neutrophils caused immune dysfunction. ART treatment significantly improved the survival of CLP mice within 96 h, and partially relieved or reversed the above-mentioned pathological features, mitigating the impact of sepsis on liver injury, inflammation, and dysfunction. This study provides comprehensive fundamental proof for the liver protective efficacy of ART against sepsis infection, which would potentially contribute to its clinical translation for sepsis therapy.

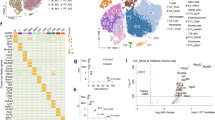
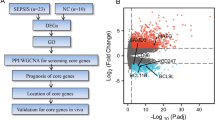
Single cell transcriptome reveals the changes of various hepatocyte subtypes of CLP-induced liver injury and the potential pharmacological effects of artesunate on sepsis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, et al. The third international consensus definitions for sepsis and septic shock (sepsis-3). JAMA. 2016;315:801–10.
Evans L, Rhodes A, Alhazzani W, Antonelli M, Coopersmith CM, French C, et al. Executive summary: surviving sepsis campaign: international guidelines for the management of sepsis and septic shock 2021. Crit Care Med. 2021;49:1974–82.
Van der Poll T, van de Veerdonk FL, Scicluna BP, Netea MG. The immunopathology of sepsis and potential therapeutic targets. Nat Rev Immunol. 2017;17:407–20.
Pierrakos C, Velissaris D, Bisdorff M, Marshall JC, Vincent JL. Biomarkers of sepsis: time for a reappraisal. Crit Care. 2020;24:287.
Schnabl B, Brenner DA. Interactions between the intestinal microbiome and liver diseases. Gastroenterology. 2014;146:1513–24.
Jenne CN, Kubes P. Immune surveillance by the liver. Nat Immunol. 2013;14:996–1006.
Protzer U, Maini MK, Knolle PA. Living in the liver: hepatic infections. Nat Rev Immunol. 2012;12:201–13.
Strnad P, Tacke F, Koch A, Trautwein C. Liver-guardian, modifier and target of sepsis. Nat Rev Gastroenterol Hepatol. 2017;14:55–66.
Heymann F, Tacke F. Immunology in the liver-from homeostasis to disease. Nat Rev Gastroenterol Hepatol. 2016;13:88–110.
Chambers DC, Carew AM, Lukowski SW, Powell JE. Transcriptomics and single-cell RNA-sequencing. Respirology. 2019;24:29–36.
Zhu Y, Ouyang Z, Du H, Wang M, Wang J, Sun H, et al. New opportunities and challenges of natural products research: When target identification meets single-cell multiomics. Acta Pharm Sin B. 2022;12:4011–39.
Papalexi E, Satija R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat Rev Immunol. 2018;18:35–45.
Xiong X, Kuang H, Liu T, Lin JD. A single-cell perspective of the mammalian liver in health and disease. Hepatology. 2020;71:1467–73.
Sun Y, Wu L, Zhong Y, Zhou K, Hou Y, Wang Z, et al. Single-cell landscape of the ecosystem in early-relapse hepatocellular carcinoma. Cell. 2021;184:404–21.e16.
Reyes M, Filbin MR, Bhattacharyya RP, Billman K, Eisenhaure T, Hung DT, et al. An immune-cell signature of bacterial sepsis. Nat Med. 2020;26:333–40.
Wen M, Cai G, Ye J, Liu X, Ding H, Zeng H. Single-cell transcriptomics reveals the alteration of peripheral blood mononuclear cells driven by sepsis. Ann Transl Med. 2020;8:125.
Darden DB, Dong X, Brusko MA, Kelly L, Fenner B, Rincon JC, et al. A novel single-cell RNA-seq analysis of non-myeloid circulating cells in late sepsis. Front Immunol. 2021;12:696536.
Wei C, Zhao CX, Liu S, Zhao JH, Ye Z, Wang H, et al. Activity-based protein profiling reveals that secondary-carbon-centered radicals of synthetic 1,2,4-trioxolanes are predominately responsible for modification of protein targets in malaria parasites. Chem Commun. 2019;55:9535–8.
Zyad A, Tilaoui M, Jaafari A, Oukerrou MA, Mouse HA. More insights into the pharmacological effects of artemisinin. Phytother Res. 2018;32:216–29.
Efferth T, Oesch F. The immunosuppressive activity of artemisinin-type drugs towards inflammatory and autoimmune diseases. Med Res Rev. 2021;41:3023–61.
Wong YK, Xu C, Kalesh KA, He Y, Lin Q, Wong WSF, et al. Artemisinin as an anticancer drug: Recent advances in target profiling and mechanisms of action. Med Res Rev. 2017;37:1492–517.
Yang Y, Liu Q, Shi X, Zheng Q, Chen L, Sun Y. Advances in plant-derived natural products for antitumor immunotherapy. Arch Pharm Res. 2021;44:987–1011.
Cao TH, Jin SG, Fei DS, Kang K, Jiang L, Lian ZY, et al. Artesunate protects against sepsis-induced lung injury via Heme Oxygenase-1 modulation. Inflammation. 2016;39:651–62.
Zhang E, Wang J, Chen Q, Wang Z, Li D, Jiang N, et al. Artesunate ameliorates sepsis-induced acute lung injury by activating the mTOR/AKT/PI3K axis. Gene. 2020;759:144969.
Lin SP, Wei JX, Hu JS, Bu JY, Zhu LD, Li Q, et al. Artemisinin improves neurocognitive deficits associated with sepsis by activating the AMPK axis in microglia. Acta Pharmacol Sin. 2021;42:1069–79.
Yan Z, Luo H, Xie B, Tian T, Li S, Chen Z, et al. Targeting adaptor protein SLP76 of RAGE as a therapeutic approach for lethal sepsis. Nat Commun. 2021;12:308.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i90.
Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM 3rd, et al. Comprehensive integration of single-cell data. Cell. 2019;177:1888–902.e21.
Ramachandran P, Matchett KP, Dobie R, Wilson-Kanamori JR, Henderson NC. Single-cell technologies in hepatology: new insights into liver biology and disease pathogenesis. Nat Rev Gastroenterol Hepatol. 2020;17:457–72.
Trapnell C, Cacchiarelli D, Grimsby J, Pokharel P, Li S, Morse M, et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol. 2014;32:381–6.
Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan CH, et al. Inference and analysis of cell-cell communication using CellChat. Nat Commun. 2021;12:1088.
Wang X, Yang L, Wang YC, Xu ZR, Feng Y, Zhang J, et al. Comparative analysis of cell lineage differentiation during hepatogenesis in humans and mice at the single-cell transcriptome level. Cell Res. 2020;30:1109–26.
Xiong X, Kuang H, Ansari S, Liu T, Gong J, Wang S, et al. Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol Cell. 2019;75:644–60 e5.
Zhu D, Rostami MR, Zuo WL, Leopold PL, Crystal RG. Single-cell transcriptome analysis of mouse liver cell-specific tropism and transcriptional dysregulation following intravenous administration of AAVrh.10 Vectors. Hum Gene Ther. 2020;31:590–604.
Liu N, Jiang C, Cai P, Shen Z, Sun W, Xu H, et al. Single-cell analysis of COVID-19, sepsis, and HIV infection reveals hyperinflammatory and immunosuppressive signatures in monocytes. Cell Rep. 2021;37:109793.
Qiu X, Li J, Bonenfant J, Jaroszewski L, Mittal A, Klein W, et al. Dynamic changes in human single-cell transcriptional signatures during fatal sepsis. J Leukoc Biol. 2021;110:1253–68.
Chousterman BG, Swirski FK, Weber GF. Cytokine storm and sepsis disease pathogenesis. Semin Immunopathol. 2017;39:517–28.
Poisson J, Lemoinne S, Boulanger C, Durand F, Moreau R, Valla D, et al. Liver sinusoidal endothelial cells: Physiology and role in liver diseases. J Hepatol. 2017;66:212–27.
DeLeve LD, Maretti-Mira AC. Liver sinusoidal endothelial cell: an update. Semin Liver Dis. 2017;37:377–87.
Hutchins NA, Chung CS, Borgerding JN, Ayala CA, Ayala A. Kupffer cells protect liver sinusoidal endothelial cells from Fas-dependent apoptosis in sepsis by down-regulating gp130. Am J Pathol. 2013;182:742–54.
Thomas MR, Storey RF. The role of platelets in inflammation. Thromb Haemost. 2015;114:449–58.
Xu H, Ye X, Steinberg H, Liu SF. Selective blockade of endothelial NF-kappaB pathway differentially affects systemic inflammation and multiple organ dysfunction and injury in septic mice. J Pathol. 2010;220:490–8.
Matejovic M, Krouzecky A, Martinkova V, Rokyta R Jr., Kralova H, Treska V, et al. Selective inducible nitric oxide synthase inhibition during long-term hyperdynamic porcine bacteremia. Shock. 2004;21:458–65.
Escobar DA, Botero-Quintero AM, Kautza BC, Luciano J, Loughran P, Darwiche S, et al. Adenosine monophosphate-activated protein kinase activation protects against sepsis-induced organ injury and inflammation. J Surg Res. 2015;194:262–72.
Krenkel O, Tacke F. Liver macrophages in tissue homeostasis and disease. Nat Rev Immunol. 2017;17:306–21.
Chen X, Liu Y, Gao Y, Shou S, Chai Y. The roles of macrophage polarization in the host immune response to sepsis. Int Immunopharmacol. 2021;96:107791.
Wang N, Liang H, Zen K. Molecular mechanisms that influence the macrophage M1-M2 polarization balance. Front Immunol. 2014;5:614.
Gaddam RR, Fraser R, Badiei A, Chambers S, Cogger VC, Le Couteur DG, et al. Differential effects of Kupffer cell inactivation on inflammation and the liver sieve following caecal-ligation and luncture-induced sepsis in mice. Shock. 2017;47:480–90.
Cao C, Yu M, Chai Y. Pathological alteration and therapeutic implications of sepsis-induced immune cell apoptosis. Cell Death Dis. 2019;10:782.
Venet F, Monneret G. Advances in the understanding and treatment of sepsis-induced immunosuppression. Nat Rev Nephrol. 2018;14:121–37.
Crinier A, Narni-Mancinelli E, Ugolini S, Vivier E. SnapShot: natural killer cells. Cell. 2020;180:1280– e1.
Wilson JK, Zhao Y, Singer M, Spencer J, Shankar-Hari M. Lymphocyte subset expression and serum concentrations of PD-1/PD-L1 in sepsis—pilot study. Crit Care. 2018;22:95.
Zhang Y, Zhou Y, Lou J, Li J, Bo L, Zhu K, et al. PD-L1 blockade improves survival in experimental sepsis by inhibiting lymphocyte apoptosis and reversing monocyte dysfunction. Crit Care. 2010;14:R220.
Van der Poll T, Shankar-Hari M, Wiersinga WJ. The immunology of sepsis. Immunity. 2021;54:2450–64.
Rungelrath V, Kobayashi SD, DeLeo FR. Neutrophils in innate immunity and systems biology-level approaches. Wiley Interdiscip Rev Syst Biol Med. 2020;12:e1458.
Sabroe I, Dower SK, Whyte MK. The role of Toll-like receptors in the regulation of neutrophil migration, activation, and apoptosis. Clin Infect Dis. 2005;41:S421–6.
Mao JY, Zhang JH, Cheng W, Chen JW, Cui N. Effects of neutrophil extracellular traps in patients with septic coagulopathy and their interaction with autophagy. Front Immunol. 2021;12:757041.
Armingol E, Officer A, Harismendy O, Lewis NE. Deciphering cell-cell interactions and communication from gene expression. Nat Rev Genet. 2021;22:71–88.
Sun J, Zhang J, Wang X, Ji F, Ronco C, Tian J, et al. Gut-liver crosstalk in sepsis-induced liver injury. Crit Care. 2020;24:614.
Yan J, Li S, Li S. The role of the liver in sepsis. Int Rev Immunol. 2014;33:498–510.
Ince C, Mayeux PR, Nguyen T, Gomez H, Kellum JA, Ospina-Tascon GA, et al. The endothelium in sepsis. Shock. 2016;45:259–70.
Roszer T. Understanding the mysterious M2 macrophage through activation markers and effector mechanisms. Mediat Inflamm. 2015;2015:816460.
Liu YC, Zou XB, Chai YF, Yao YM. Macrophage polarization in inflammatory diseases. Int J Biol Sci. 2014;10:520–9.
Wasyluk W, Zwolak A. Metabolic alterations in Sepsis. J Clin Med. 2021;10:2412.
de Oliveira S, Rosowski EE, Huttenlocher A. Neutrophil migration in infection and wound repair: going forward in reverse. Nat Rev Immunol. 2016;16:378–91.
Wang JF, Li JB, Zhao YJ, Yi WJ, Bian JJ, Wan XJ, et al. Up-regulation of programmed cell death 1 ligand 1 on neutrophils may be involved in sepsis-induced immunosuppression: an animal study and a prospective case-control study. Anesthesiology. 2015;122:852–63.
Zhang Y, Li J, Lou J, Zhou Y, Bo L, Zhu J, et al. Upregulation of programmed death-1 on T cells and programmed death ligand-1 on monocytes in septic shock patients. Crit Care. 2011;15:R70.
Chen R, Zhou L. PD-1 signaling pathway in sepsis: does it have a future? Clin Immunol. 2021;229:108742.
Jones DM, Read KA, Oestreich KJ. Dynamic roles for IL-2-STAT5 signaling in effector and regulatory CD4+ T cell populations. J Immunol. 2020;205:1721–30.
Iba T, Levy JH. Inflammation and thrombosis: roles of neutrophils, platelets, and endothelial cells and their interactions in thrombus formation during sepsis. J Thromb Haemost. 2018;16:231–41.
Duerschmied D, Suidan GL, Demers M, Herr N, Carbo C, Brill A, et al. Platelet serotonin promotes the recruitment of neutrophils to sites of acute inflammation in mice. Blood. 2013;121:1008–15.
Wiersinga WJ, Leopold SJ, Cranendonk DR, van der Poll T. Host innate immune responses to sepsis. Virulence. 2014;5:36–44.
Zhang Q, Cao X. Epigenetic remodeling in innate immunity and inflammation. Annu Rev Immunol. 2021;39:279–311.
Acknowledgements
We gratefully acknowledge financial support from the Establishment of Sino-Austria “Belt and Road” Joint Laboratory on Traditional Chinese Medicine for Severe Infectious Diseases and Joint Research (2020YFE0205100); the National Key Research and Development Program of China (2020YFA0908000 and 2022YFC2303600); the Innovation Team and Talents Cultivation Program of National Administration of Traditional Chinese Medicine (ZYYCXTD-C-202002); the National Natural Science Foundation of China (82141001, 82274182, 82074098 and 82173914); the CACMS Innovation Fund (CI2021A05101 and CI2021A05104); the Scientific and Technological Innovation Project of China Academy of Chinese Medical Sciences (CI2021B014); the Science and Technology Foundation of Shenzhen (JCYJ20210324115800001); the Fundamental Research Funds for the Central Public Welfare Research Institutes (ZZ14-YQ-050, ZZ14-YQ-051, ZZ14-YQ-052, ZZ14-FL-002, ZZ14-ND-010 and ZZ15-ND-10); the Shenzhen Governmental Sustainable Development Fund (KCXFZ20201221173612034); the Shenzhen Key Laboratory of Kidney Diseases (ZDSYS201504301616234); the Shenzhen Fund for Guangdong Provincial High-level Clinical Key Specialties (SZGSP001).
Author information
Authors and Affiliations
Contributions
JGW, YJ, and ZJL conceived and designed the experiments. XLH, JYC, YLF, and PS performed the major experiment and main data analysis. LLX, QZ, PG, and PL assisted with animal experiments and tissue dissociation. YKW, CW, YMB, and QL executed other experiments and bioinformatic analyses. XLH, JYC, and YKW wrote the manuscript. FLL and ZJL revised the manuscript. All authors have reviewed and edited the manuscript and agreed with the content of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, Xl., Chen, Jy., Feng, Yl. et al. Single-cell RNA sequencing deciphers the mechanism of sepsis-induced liver injury and the therapeutic effects of artesunate. Acta Pharmacol Sin 44, 1801–1814 (2023). https://doi.org/10.1038/s41401-023-01065-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-023-01065-y