Abstract
The microbiome-gut-brain axis plays a role in anxiety, the stress response and social development, and is of growing interest in neuropsychiatric conditions. The gut microbiota shows compositional alterations in a variety of psychiatric disorders including depression, generalised anxiety disorder (GAD), autism spectrum disorder (ASD) and schizophrenia but studies investigating the gut microbiome in social anxiety disorder (SAD) are very limited. Using whole-genome shotgun analysis of 49 faecal samples (31 cases and 18 sex- and age-matched controls), we analysed compositional and functional differences in the gut microbiome of patients with SAD in comparison to healthy controls. Overall microbiota composition, as measured by beta-diversity, was found to be different between the SAD and control groups and several taxonomic differences were seen at a genus- and species-level. The relative abundance of the genera Anaeromassillibacillus and Gordonibacter were elevated in SAD, while Parasuterella was enriched in healthy controls. At a species-level, Anaeromassilibacillus sp An250 was found to be more abundant in SAD patients while Parasutterella excrementihominis was higher in controls. No differences were seen in alpha diversity. In relation to functional differences, the gut metabolic module ‘aspartate degradation I’ was elevated in SAD patients. In conclusion, the gut microbiome of patients with SAD differs in composition and function to that of healthy controls. Larger, longitudinal studies are warranted to validate these preliminary results and explore the clinical implications of these microbiome changes.
Similar content being viewed by others
Introduction
Social anxiety disorder (SAD) is one of the most common psychiatric conditions with estimated lifetime prevalence rates as high as 13% reported in the United States [1] and similarly high prevalence rates across Europe [2]. The current Diagnostic and Statistical Manual of Mental Disorders – Fifth Edition (DSM-V) describes SAD as a condition characterised by ‘marked fear or anxiety about one or more social situations in which the individual is exposed to possible scrutiny by others.’ These fears generally extend across a variety of situations, including social interactions (e.g., having a conversation, meeting unfamiliar people), being observed (e.g., eating or drinking), and performing in front of others (e.g., giving a speech) [3]. SAD typically begins early in life and tends to run a chronic, often lifelong, course [4]. It is associated with serious functional disability and markedly reduced quality of life [5] with up to 69% of sufferers experiencing another lifetime major comorbid disorder [6]. In particular, SAD markedly increases the risk of subsequent depression [7] which is associated with a poorer prognosis and greater risk of suicide attempts [8]. Thus, timely intervention in this early-onset disorder has the potential to not only reduce substantial disability, but to markedly reduce the psychiatric disease burden later in life. Current first-line treatments include selective serotonin reuptake inhibitors (SSRIs), serotonin and norepinephrine reuptake inhibitors (SNRIs) and cognitive behavioural therapy [9]. Unfortunately, a significant proportion of patients fail to adequately respond to first-line pharmacotherapy [10] and even fewer patients will respond to subsequent second-line treatments. In a large randomised controlled trial (RCT) investigating augmentation and switch strategies for refractory SAD (defined as more than two unsuccessful adequate pharmacological treatment trials), only 46% of patients demonstrated a response to treatment, while only 21% of all patients achieved remission at the 12-week endpoint [11]. Thus, there is a great necessity for an improved understanding of the neurobiological basis for this condition and the development of alternative therapeutic strategies. The microbiome-gut-brain axis may represent one such potential avenue for investigation.
The human gastrointestinal tract (GIT) harbours a vast assembly of microorganisms, predominantly bacteria but also fungi, viruses, protozoa and archaea. While the term gut microbiota refers to the assemblage of living organisms present within the gut, the term microbiome encompasses the micro-organisms and their ‘theatre of activity’ i.e., their structural elements, genomes and metabolites, as well as the surrounding environmental conditions [12, 13]. It is estimated that the number of bacteria in the human gut is slightly in excess of the total number of human cells, at approximately 3.8 ×10 13 [14] and that the collective genome of these bacterial cells vastly exceeds the amount of human DNA present in the body [15]. Given this enormous, modifiable reservoir of genetic potential, it is unsurprising that there is keen interest in the potential role of the gut microbiome in the aetiology and treatment of many disease processes. The microbiome is recognised as a key player in bidirectional signalling between the gut and brain, with the term ‘microbiome-gut-brain’ (MGB) axis describing this communication network. Many physiological systems relevant to psychiatric disorders come under the influence of the gut microbiome, including the immune system, vagal neurotransmission, tryptophan metabolism, endocrine function and the stress response system, making the MGB axis an attractive new therapeutic target in psychiatry [16].
Despite a growing interest in the role of the gut microbiome in the neurobiology of the stress response [17] and social behaviour [18, 19], there has been very limited investigation of the gut microbiome in relation to SAD. Indeed, apart from a few small studies in generalised anxiety disorder (GAD) [20,21,22], and in co-morbid irritable bowel syndrome (IBS) [23], gut microbiome composition or function has remained largely unexplored in patients with clinical anxiety disorders, including SAD, panic disorder and agoraphobia. This is despite an abundance of preclinical studies demonstrating that anxiety-like behaviours are altered in animal models following a variety of microbiota manipulations [24] and clear evidence that certain probiotics can reduce self-reported anxiety levels in healthy human volunteers [25] The past decade has also seen a spate of preclinical studies revealing a role for the gut microbiome in social development and social behaviour in animals [26, 27] and this has extended to human studies in recent years. Differences in gut microbiota composition and diversity have recently been linked to personality traits such as sociability in the general population [28]. Additionally, altered gut microbiota composition has been demonstrated in adults experiencing social exclusion [29], a psychological phenomenon to which those with SAD are particularly sensitive [30]. Interestingly, a potential therapeutic role for the microbiota in SAD is supported by a cross-sectional study which reported that higher intake of fermented, probiotic-containing foods by healthy students appeared to be protective against developing SAD in those who were at higher genetic risk, as measured by trait neuroticism [31].
Here we report on compositional and functional differences in the gut microbiome of patients with SAD using whole-genome shotgun analysis of 49 faecal samples (31 cases and 18 controls). The functional differences, based on Kyoto Encyclopedia of Genes and Genomes (KEGG) orthologues, are explored using the recently described gut-brain module (GBM) [32] and gut metabolic module (GMM) [33] analysis. We hypothesise that the gut microbiome is compositionally and functionally altered in those with SAD.
Methods
Participants
Patients with a clinical diagnosis of SAD were recruited through local general practitioners, psychologists and outpatient psychiatric clinics. The study was also advertised through local and national SAD support groups and by an online website (www.sadgut.ie). Eligibility was limited to men and women aged between 18–65 years with a clinical diagnosis of SAD. Exclusion criteria included any significant acute or chronic medical illness (including functional gastrointestinal disorders such as irritable bowel syndrome); the presence of any condition or medication which the investigator believed would interfere with the objectives of the study or confound the interpretation of the study, including anticonvulsants, centrally acting corticosteroids, opioid pain relievers, laxatives, enemas, anti-coagulants, over-the counter non-steroidal anti-inflammatories (NSAIDS); the use of probiotics, prebiotics or antibiotics in the previous 4 weeks; females who were pregnant or breastfeeding; subjects who were vegetarian or adhering to a strict specific diet. Specific psychiatric exclusion criteria included a lifetime diagnosis of psychotic disorder, intellectual disability, bipolar disorder, dementia or ASD and a current diagnosis of major depressive disorder (MDD), eating disorder, alcohol or substance abuse or dependence. A past history of depression was permitted, as was the presence of a comorbid anxiety disorder, provided the clinician was satisfied that SAD was the primary diagnosis. SAD participants were permitted to continue taking their regular psychotropic medication. Healthy controls were recruited through email and print advertising in University College Cork. Controls were required to have no past or current psychiatric diagnosis along with the other general exclusion criteria outlined for the SAD participants.
Procedures
All study procedures were approved by the Clinical Research Ethics Committee of the Cork Teaching Hospitals (Study number APC085) and the study was conducted in accordance with the ICH Guidelines on Good Clinical Practice, and the Declaration of Helsinki. All participants provided written informed consent.
All participants were interviewed by an experienced psychiatrist using the MINI International Neuropsychiatric Interview (Version 7.0) [34] to confirm the diagnosis of SAD based on DSM-V criteria and assess for any relevant comorbidities. No patient met the criteria for the ‘performance only’ specifier and all experienced social anxiety symptoms across a range of situations.
Social anxiety symptoms were assessed using the Liebowitz Social Anxiety Scale-Self Report (LSAS), a 24-item scale which was initially developed as a clinician-rated instrument [35, 36] but was later shown to have excellent psychometric properties as a self-report scale [37,38,39]
To quantify nutrient intake, participants completed the self-administered 152-item SLAN-06 (Survey of Lifestyle, Attitudes and Nutrition in Ireland) food frequency questionnaire (FFQ), which is adapted from the EPIC Norfolk questionnaire [40] and validated to be used in an Irish population [41]. Participants were asked to estimate the frequency with which they consumed a specified portion size of each of the foods listed over the preceding year. The FFQs were analysed for nutrient intake using the FETA software [42]. Stool consistency was assessed using the Bristol Stool Chart (BSC) [43]. Exercise levels were measured using the International Physical Activity Questionnaire (self-administered short form) [44] and sleep using the Pittsburgh Sleep Quality Index [45].
Biological/faecal samples
Freshly voided faecal samples were collected from study participants into plastic containers containing an AnaeroGen sachet (Oxoid AGS AnaeroGen Compact, Fischer Scientific, Dublin) to generate anaerobic conditions within the container. Participants were instructed to collect the faecal sample as close to the study visit as possible and to keep the sample containers in a refrigerator at 4 °C until delivery to the study site. A cool pack was used to transport the sample to the study site, where it was immediately stored at − 80 °C for later analysis.
Microbiome sample preparation and whole genome shotgun sequencing
Total bacterial metagenomics DNA was extracted using the QIAmp Fast DNA Stool Mini kit (Qiagen, UK) with a modified protocol combined with repeated bead beating method (Zhongtang Yu & Mark Morrison 2018). Briefly, 1 ml of lysis buffer (500 mM NaCl, 50 mM Tris-HCl pH8.0, 50 mM EDTA and 4% sodium dodecyl sulphate) was added to the stool sample in the bead-beating tube. The samples were homogenized using a mini beadbeater (BioSpec) and incubated at 70 °C for 15 minutes (for cell lysis) followed by centrifugation at 4 °C. The supernatant was removed and the bead-beating step was repeated. Ammonium acetate (Sigma Aldrich, Ireland) was added to the pooled supernatant and incubated on ice. Following a centrifugation step the supernatant was transferred to Eppendorf tubes containing iso-propanol. The following day, DNA was pelleted and washed with 70% ethanol and dissolved in Tris-EDTA. The DNA was then RNAse and proteinase-K treated and purified according to the manufacturer’s instructions (QIAmp Fast DNA Stool Mini kit; Qiagen, UK). The DNA was quantified using Qubit and stored at −30 °C.
Whole genome shotgun sequencing was performed using Nextera XT kit. Library prep was done following the Nextera XT DNA Library Preparation Guide from Illumina. Quality of the library was evaluated using the Agilent High Sensitivity DNA chip and running it on the Bioanalyzer and the DNA was quantified using Qubit DNA High sensitivity kit read on a qubit fluorometer 3.0. The samples were pooled and sequencing was carried out on the NextSeq500 using a 300 cycle High Output v2 kit.
Taxanomic and functional analysis
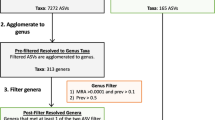
We performed quality checks on raw sequences from all faecal samples using FastQC [46]. Shotgun metagenomic sequencing data were then processed through analysis workflow that utilizes Huttenhower Biobakery pipeline [47], including Kneaddata [48], MetaPhlAn3 [49] and HUMAnN3 [50] to obtain species, genes and pathways abundance matrix. Briefly, quality filtering and host genome decontamination (human) was performed using Trimmomatic [51] and Bowtie2 [52] via Kneaddata wrapper program with following parameters: ILLUMINACLIP:/NexteraPE-PE.fa:2:30:10, SLIDINGWINDOW:5:25, MINLEN:60, LEADING:3, TRAILING:3. Taxonomic and functional profiling of the microbial community was performed using MetaPhlan3 and HUMAnN3 using default parameter. Next, gene abundance matrix was further collapsed by KEGG Orthology (KO) term and Gene Ontology (GO) term mapping via “humann_regroup_table” function provided within HUMAnN3.
Further data-handling was undertaken in R (version 4.03) using the Rstudio GUI (version 1.4.1103). In all microbiome analysis with the exception of alpha diversity, taxa with a prevalence of <5% of samples at the genus level were excluded from analysis as ratios are invariant to sub-setting and this study employs compositional data analysis techniques [53, 54]. Principal component analysis was performed on centred log-ratio transformed (clr) values using the ALDEx2 library [55]. The number of permutations was always set to 1000. Beta diversity was computed in terms of Aitchison distance, or Euclidean distance between clr-transformed data. Alpha diversity was computed using the iNEXT library [56]. KEGG orthologues were used as features to compute functional alpha diversity. Gut-Brain Modules (GBMs) and Gut-Metabolic Modules (GMMs) were calculated from HUMAnN3 output using the R version of the Gomixer tool [32]. Stacked barplots were generated by normalising counts to 1, generating proportions. Genera that were never detected at a 10% relative abundance or higher were aggregated and defined as rare taxa for the purposes of the stacked barplots. Differential abundance of both microbes and functional modules were calculated using implementations of the ALDEx2 library. Effect sizes were calculated using Cohen’s D statistic. A p value of <0.05 was deemed significant in all cases. To correct for multiple testing in tests involving microbiome features, the Benjamini-Hochberg (BH) post-hoc was performed with a q-value of 0.1 used as a cut-off for species and 0.2 for functional modules. Plotting was done using the ggplot2 [57] and patchwork [58] libraries in R. Custom R scripts and functions are available online at https://github.com/thomazbastiaanssen/Tjazi [59]. A linear modelling approach was used to test for a group effect on taxonomic and functional differences, whilst adjusting for covariates including age, sex, BMI, exercise and dietary differences.
Statistical analysis of metadata
All metadata were analysed using SPSS 25 (IBM, Armonk, NY, USA). Visual inspection of box plots was used to identify outliers and consideration given to removal of those lying more than three times the interquartile range (IQR) below the first quartile or above the third quartile. Missing values were excluded from analysis. Normality of data was assessed by visual inspection of histograms along with examination of skewness and the Shapiro-Wilk statistic. Differences in demographic data, FFQ and LSAS scale scores between the SAD and control group were assessed using Chi-squared or Fisher’s Exact test for categorical variables, and independent t-tests or non-parametric Mann-Whitney U tests for continuous variables. Data are presented as mean ± SD unless stated otherwise.
Results
Demographics
Based on previous microbiota studies from our laboratory [60], a sample size of 30 participants was estimated to achieve significant changes in microbiota composition. Thirty-one patients with social anxiety disorder (SAD) and eighteen healthy controls participated in the study. There were no significant differences between patients and controls in relation to age, sex, race, years of education, birth delivery mode, alcohol consumption, smoking status, or stool consistency (Table 1). Individuals in the SAD group had higher BMI scores compared to controls (t(46)=2.65, p = 0.01). SAD patients had significantly lower exercise levels than controls, based on mean IPAQ scores (t(45) = −2.125, p = 0.04), the difference when looking at exercise categories being at the level of a trend (X2(2) = 5.822, p = 0.054). Almost three-quarters (74.2%) of SAD patients had a past history of MDD and 35.5% had a comorbid secondary anxiety disorder. Just over two-thirds (67.7%) of patients were taking psychotropic medication. The majority of these patients were prescribed an SSRI (48.4%) (8 taking Escitalopram, 2 taking Vortioxetine, 2 taking Sertraline, 1 taking Citalopram, 1 taking Paroxetine, 1 taking Fluoxetine) or SNRI (9.7%) (3 taking Venlafaxine) with 22.6% (n = 7) taking an alternative regular psychotropic medication (1 patient taking Pregabalin, 1 taking Agomelatine, 1 taking Bupropion, 2 taking Trazadone and 2 taking low-dose (50 mg) Quetiapine).
Dietary intake
Based on FFQ analysis, the only significant difference in nutrient intake seen between the SAD patients and controls was in relation to carbohydrates (Table 2). SAD patients had greater intake of total carbohydrates (U = 154, z = −1.983, p = 0.047), which appeared to be driven by higher total sugar intake (U = 140, z = −2.31, p = 0.021) as other carbohydrate groups, starch and fibre, were equivalent in patients and controls. No other differences in nutrient groups, vitamins or minerals were seen between patients and controls.
Compositional differences in the gut microbiota of SAD patients
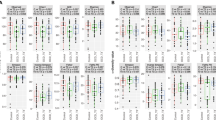
The gut microbiota of patients with SAD differed from those of healthy controls in terms of overall composition as well as in relation to specific genus- and species-level differentially abundant features. Beta diversity was found to be different between the two groups as measured by PERMANOVA (p = 0.038, R2 = 0.028) using the compositionally appropriate Aitchison distance metric (Fig. 1A). No differences were found between groups in alpha diversity, based on the Chao1, Shannon or Simpson indices (Fig. 1B).
A Beta diversity between SAD and healthy control groups, as measured by Aitchison Distance. p-value based on PERMANOVA test. B Alpha-diversity between SAD and healthy controls, as measured by Chao1, Simpson and Shannon indices. p-values based on Student’s t-tests. C Relative abundance of species-level taxa for each participant. Each column represents one participant. Genera that were never detected at a 10% relative abundance or higher are aggregated and defined as rare taxa for the purposes of the stacked barplots. (* p = <0.05) (HC: Healthy Control, SAD: Social Anxiety Disorder).
A total of 73 genera and 159 species were identified (Fig. 1C). Of these, three genera and two species were found to show significant differences in relative abundance after false discovery rate (FDR) correction using the Benjamini-Hochberg procedure. At the genus level, Anaeromassilibacillus and Gordonibacter were enriched in SAD while Parasutterella was more abundant in the control samples (Fig. 2A). At the species level, Anaeromassilibacillus sp An250 was found to be more abundant in SAD patients (padj = 0.024, effect size = −1.036). Specifically, this species was present in 48.4% (15/31) of the SAD samples but only found in 5.6% (1/18) of the control samples. Conversely, the bacterial species, Parasutterella excrementihominis was found to be enriched in healthy controls (padj = 0.042, effect size = 1.120) (Fig. 2B). After adjusting for age, sex, BMI, exercise and dietary differences (total carbohydrates), these genus- and species-level relative abundance differences remained significant. We found no statistically significant differences in the relative abundance of any microbial taxa between unmedicated SAD patients and those taking psychotropic medication or between those SAD participants with or without a history of MDD.
A Genus-level differences in relative abundance between SAD and controls seen in three genera; Anaeromassillibacillus and Gordonibacter are enriched in SAD while Parasutterella is enriched in healthy controls. B Species-level differences in relative abundance between SAD and controls; Anaeromassilibacillus sp An250 is increased in SAD while Parasuterella excrementihominis is enriched in healthy controls. (*p = <0.05) (Clr centred log-ratio transformed, HC Healthy Control, SAD Social Anxiety Disorder).
Functional differences in the gut microbiome of SAD patients
No differences were found in functional diversity between the two groups (Fig. 3A). We identified 69 of the 103 Gut-Metabolic Modules (GMMs) curated in the current database [33] and 26 of the 56 Gut-Brain Modules (GBMs) characterised by Valles-Colomer and colleagues [32]. No GBMs reached our threshold for significance after FDR correction. However, one GMM, Aspartate Degradation I, describing the capacity of the microbiome to degrade aspartate by the enzyme aspartate aminotransferase (AspAT), was found to be significantly more abundant in patients with SAD (padj = 0.150, effect size = −1.032) (Fig. 3B). This functional difference remained between the two groups after adjusting for age, sex, BMI, exercise and dietary differences (total carbohydrates).
A One gut metabolic module, Aspartate Degradation I, was found to be increased in SAD patients. B Functional diversity, between SAD and healthy controls, as measured by Chao1, Simpson and Shannon indices. p values based on Student’s t-test. No differences seen between the groups. (*p = <0.05) (Clr centred log-ratio transformed, HC Healthy Control, SAD Social Anxiety Disorder).
We then set out to investigate whether any microbial taxa or functional modules were associated with LSAS scores. After controlling the FDR, we did not detect any such associations (data not shown).
Discussion
This study demonstrates, for the first time, that the gut microbiome is compositionally and functionally altered in people with social anxiety disorder (SAD) compared with healthy controls. Moreover, it increases the growing evidence linking social brain function and the microbiome [27]. Firstly, we show that beta diversity, an indicator of overall microbiota composition, was significantly different between the two groups. The relative abundance of three genera, Anaeromassilibacillus, Gordonibacter and Parasutterella, and two corresponding species, Anaeromassilibacillus sp An250 and Parasutterella excrementihominis differed significantly between SAD patients and controls. Additionally, functional differences were evident with the microbiome of SAD patients enriched for the gut metabolic pathway, aspartate degradation I.
Strikingly, we found Anaeromassilibacillus sp An250 to be present in almost half of our SAD group but in only one healthy control. Anaeromassilibacillus is a newly-discovered genus, which was first isolated in 2017 from the faecal sample of a 1-yo Senegalese patient with kwashiorkor [61]. Several strains of Anaeromassilibacillus, including sp An250 have since been identified from the caecal microbiome of chickens [62, 63]. Anaeromassilibacillus is a member of the Clostridiales order and Clostridiaceae family of bacteria [61], taxonomic groups which are increased in the gut microbiota of patients with autism spectrum disorder (ASD) [64], depression [65] and schizophrenia [66]. Conversely, various genera from the Clostridiales order were found to be reduced in the faecal microbiota of people with GAD [21] although Clostridiales was positively correlated with anxiety scores in a study analysing serum microbial DNA composition in patients with mood disorders [67]. Despite such inconsistencies, significant shifts in the abundance of Clostridiales taxa appears to be common to many psychiatric disorders and may represent disease-shared microbial responses [68]. Furthermore, preclinical studies suggest a link between Clostridiales and social behaviour. In a recent study, mice subjected to early life social isolation stress showed a significantly increased abundance of Clostridiales. These mice subsequently demonstrated reductions in sociability and social novelty behaviours, which negatively correlated with Clostridiales abundance [69]. In another study, mice exposed to a social stressor had increased relative abundance of the genus Clostridium [70]. However, it is difficult to extrapolate findings from animal studies to humans [71] especially with regards to a process as complex as social behaviour.
Given the very recent addition of Anaeromassilibacillus to human microbiome databases, there is little in the existing literature about its role in human health and disease. It was one of several genera found to be enriched in untreated patients with MDD compared to those receiving antidepressant treatment, suggesting that it could be altered by psychotropic medication or be an indicator of treatment response [72]. Additionally, the relative abundance of Anaeromassilibacillus reduced significantly in the faeces of children with ASD after guar gum prebiotic supplementation, which was associated with reduced irritability and improved constipation [73], again suggesting that reduction of this genus may be associated with improved psychopathology. Thus, although the literature is sparse, Anaeromassilibacillus appears be of relevance in ASD and depression, psychiatric conditions which are highly comorbid with SAD [74, 75] and which may involve shared pathophysiological processes. Of note, we did not see a difference in the relative abundance of Anaeromassilibacillus in medicated, compared to, unmedicated SAD patients; although given the small sample size, this should be interpreted with caution.
Gordonibacter is another genus about which relatively little is known. It is a member of the Eggerthellaceae family and Coriobacteriia class [76] and is notable in its ability to produce urolithins from the metabolism of polyphenols [77], which may have an impact on mental health [78]. Parasutterella has been more extensively studied. It is a member of the Sutterellaceae family and in humans, is largely represented by a single species, Parasutterella excrementihominis [79]. Similar to our finding of lower Parasutterella levels in SAD, members of this genus have also been found to be reduced in ASD [80]. Weight and dietary factors appear to be important influences. Parasutterella is negatively associated with BMI and waist circumference [81] and conversely, can be induced by high sugar [82] and high-fat diets [83]. Our SAD group had elevated sugar intake and did not differ in terms of fat intake but, although the group difference for Parasutterella excrementihominis remained after adjusting for BMI, increased BMI in the SAD group could contribute to its reduced abundance in SAD patients.
It is difficult to interpret the importance and relevance of specific bacterial taxa differences in a patient group. The gut microbiome is a highly complex and dynamic ecosystem where microbes continuously interact with, and impact, one another and the host [84]. Attempts are underway to characterise microbial community structures and gain insights into the many complex microbe-microbe and host-microbe networks and interactions [85, 86]. Some human gut microbial groups appear to be highly influential and exert a metabolic impact on a substantial number of other microbial entities, so-called ‘network influencers’ [85]. None of our differentially expressed genera or species have been reported as being such core taxa or ‘influencers’ and it is unclear what these over- and under-represented taxa mean in the overall context of the gut microbial environment of SAD patients. With this in mind, exploring microbial function may offer deeper insights than relying on composition alone in an ever-changing ecosystem.
Using GBMs and GMMs, which are microbiome-related functional pathways that have been manually curated from existing literature [32, 33], we identified one functional pathway that was enriched in the SAD group – aspartate degradation I. According to MetCyc, a comprehensive reference database of metabolic pathways and enzymes [87], this pathway involves the conversion of the amino acid, L-aspartate to the corresponding keto acid, oxaloacetate, by the enzyme, aspartate aminotransferase (AspAT). Several bacteria and archaea have demonstrated this enzymatic ability including Haloferax mediterranei [88], Pseudoalteromonas translucida TAC125 [89], Saccharolobus solfataricus [90] and Escherichia coli [91, 92]. Interestingly, bacterial AspAT enzyme activity may represent a link between gut microbiome function and the tryptophan-kynurenine pathway, a key physiological system in psychiatric disorders. There are significant interactions between tryptophan metabolism and the MGB axis [93,94,95,96] and the gut microbiome may influence host diet selection behaviour by mediating the availability of essential amino acids such as tryptophan [97]. Tryptophan metabolism involves the downstream conversion of kynurenine to kynurenic acid (KYNA) by the enzyme kynurenine aminotransferase (KAT). KYNA is an important neuroactive substance, which is elevated by chronic stress in animal models [98, 99] as well as in psychiatric conditions such as schizophrenia [100, 101] and SAD [102]. Notably, KAT activity has been detected in E. coli cells in vitro, and authors suggested that the source of KYNA detected in the rat small intestine could be the gut bacteria [103]. This bacterial KAT enzyme protein has been identified as being identical to the bacterial AspAT enzyme [104] and thus, an elevation in the ‘aspartate degradation I’ functional pathway may represent increased KAT, as well as AspAT potential, by the microbiome. While currently a hypothetical supposition, it is possible that the elevated peripheral KYNA which we previously reported in SAD patients [102] may be linked with the key functional difference seen in the microbiome of this group. In support of this hypothesis is the fact that D-cycloserine, an orally-administered, broad-spectrum antibiotic, has been found to enhance the treatment response to exposure therapy for SAD [105, 106], an effect which could plausibly be related to its ability to inhibit KAT activity and lower KYNA [107].
This is, to our knowledge, the first study to investigate the composition and function of the gut microbiome in patients with SAD and has several notable strengths. Firstly, our sample consisted of carefully selected patients with a pre-existing clinical diagnosis of SAD who had sought treatment from a mental health professional. Secondly, we used a whole genome shotgun sequencing method, providing information on the functional capacity of the microbiome, as well as offering greater resolution of bacterial species identification, than the more commonly used 16 S rRNA gene sequencing [108, 109]. Thirdly, we took into account many of the important host variables known to confound gut microbiota studies in human disease [110]. Stool quality is a particularly strong source of gut microbiota variance [110, 111] which has often been neglected in psychiatric microbiota studies. Stool consistency, as measured by the BSC, was matched between our groups, as were other important variables, including age, sex, birth delivery mode, smoking status and alcohol. Our groups were not matched in terms of BMI and exercise levels, variables which may be of relevance to the gut microbiome [112, 113]. Although adjusting for these variables did not affect group differences, it would, of course, be preferable to have samples with equivalent BMI and exercise scores. Additionally, we collected detailed dietary information, which has often been lacking in studies of the microbiota in psychiatric conditions. Our groups were well-matched in terms of overall dietary intake. The only difference seen was in relation to carbohydrate consumption, driven by higher sugar intake in the SAD group, and this was adjusted for in our statistical analyses.
Study limitations include the small sample size and the single-time point nature of the study, which prevents the establishment of any causal relationships. Additionally, two thirds of our SAD patients were taking psychotropic medication, which may have had an impact on microbiota composition [114, 115]. The majority of our medicated patients were taking an SSRI antidepressant, escitalopram being the most commonly prescribed. Escitalopram has antibacterial activity against some gut commensal strains in vitro [116, 117] although this effect did not translate to an in vivo animal model [117]. Other prescribed SSRIs in our patient group included fluoxetine, citalopram, sertraline, paroxetine and vortioxetine, all of which have shown varying levels of antibacterial activity in vitro [117,118,119,120], with in vivo evidence available for fluoxetine [121,122,123]. The SNRI venlafaxine, conversely, does not appear to impact common gut commensals in vitro [116, 117], although an influence on the microbial richness and on the abundance of certain genera were seen in a mouse model [122]. Thus, the translatability of studies using isolated in-vitro strains to animal models is unclear, with even more uncertainty in relation to their applicability to the human gut microbiome. Limited human data in relation to the effect of antidepressants on the microbiome is available. A small study of 17 depressed patients commenced on escitalopram, found no significant differences in beta-diversity or in any taxa levels between pre-treatment and 6-week post-treatment time-points, although increased alpha diversity was evident [124]. Furthermore, a longitudinal study of 40 patients with depression and/or anxiety revealed no difference in beta diversity between those taking, and not taking, antidepressant medications and no change in alpha diversity in antidepressant-treated patients between baseline and endpoint timepoints. Antipsychotic medications, conversely, did appear to exert an effect on the gut microbiome [125], consistent with previous findings [126]. Two of our patients were prescribed low-dose Quetiapine, a second-generation antipsychotic which thus may have had an impact.
Aside from impacting microbiota composition, it is also of course possible that psychotropic medications could have influenced functional pathways. A recent study demonstrated that oral intake of fluoxetine or amitriptyline by rats exposed to chronic unpredictable mild stress resulted in alterations in KEGG metabolic pathways, particularly those pathways concerning carbohydrate metabolism, membrane transport, and signal transduction [127]. However, no such alterations in KEGG pathways were seen in a longitudinal follow-up of psychiatric patients taking antipsychotics, antidepressants and/or anxiolytic medications [125]. In an approach similar to ours, these authors also chose to analyse GBMs in the psychiatric group, although they specifically investigated only 6 of the 56 available GBMs, namely those involving GABA and tryptophan synthesis or degradation. They found alterations in certain GBMs in patients taking antipsychotics and antidepressants but not in those taking anxiolytic medications. All in all, although there is clear evidence that many antidepressants have antibacterial effects, this evidence is based primarily on in-vitro and animal studies, and the impact of these medications on the human gut microbiome structure and function remain largely unknown. Although we did not find any differences in the relative abundance of any taxa between medicated and unmedicated patients, we cannot rule out a potential influence.
Finally, some of our SAD patient group had a past history of depression and/or a comorbid anxiety disorder. However, patients with a current depressive episode were excluded and in all, the primary diagnosis was SAD with any comorbid anxiety disorder representing a secondary diagnosis. Although we did not find a difference in the relative abundance of any taxa in those SAD patients with or without a past history of MDD, this must be interpreted with caution given the small numbers of such sub-groups. While it is not possible to disentangle the currently reported observations from the past psychiatric history of study participants, this was a clinically representative sample and we believe that including such patients makes our findings more generalizable considering the significant overlap between depression and anxiety disorders. Given the paucity of studies exploring the gut microbiome in any clinical anxiety disorders, our findings, despite the limitations, are important in generating a foundational base for larger, prospective and interventional microbiome studies in these highly prevalent and disabling psychiatric conditions. Additionally, future preclinical studies and secondary validation experiments would offer a complementary approach to confirm the presence and role of these differentially expressed bacterial taxa and functional pathways. Earlier compositional microbiota studies in depression [60, 128] and ASD [129, 130] have paved the way for a variety of successful interventional studies utilising probiotics [25], dietary change [131] and faecal microbiota transplantation (FMT) [132] in these conditions. It is hopeful that microbiota-based therapeutic interventions may also be realised for patients with clinical anxiety disorders. Indeed, a previous cross-sectional study in university students has suggested that consumption of fermented foods may be protective against the development of social anxiety [31].
In conclusion, the gut microbiome of patients with SAD differs in composition and function to that of healthy controls, raising the possibility that the MGB axis may represent a biomarker reservoir and potential therapeutic target for this early-onset, chronic disorder. Further preclinical studies focussing on mechanistic pathways and larger, longitudinal studies in SAD patients are needed to validate our preliminary results, understand the clinical implications (if any) and investigate the impact of psychotropic medication and treatment on the gut microbiome in SAD.
Data availability
Whole genome sequences are available at: European Nucleotide Archive, Accession ID PRJEB58864.
References
Kessler RC, Petukhova M, Sampson NA, Zaslavsky AM, Wittchen HU. Twelve-month and lifetime prevalence and lifetime morbid risk of anxiety and mood disorders in the United States. Int J Methods Psychiatr Res. 2012;21:169–84.
Fehm L, Pelissolo A, Furmark T, Wittchen H-U. Size and burden of social phobia in Europe. Eur Neuropsychopharmacol. 2005;15:453–62.
APA, APA. Diagnostic and statistical manual for mental disorders, 5th edition(DSM-V). Arlington, VA: American Psychiatric Publishing, 2013.
Keller MB. The lifelong course of social anxiety disorder: a clinical perspective. Acta Psychiatr Scand Suppl. 2003;2003:85–94.
Stein MB, Kean YM. Disability and quality of life in social phobia: epidemiologic findings. Am J Psychiatry. 2000;157:1606–13.
Schneier FR, Johnson J, Hornig CD, Liebowitz MR, Weissman MM. Social phobia. Comorbidity and morbidity in an epidemiologic sample. Arch Gen Psychiatry. 1992;49:282–8.
Beesdo K, Bittner A, Pine DS, Stein MB, Höfler M, Lieb R, et al. Incidence of Social Anxiety Disorder and the Consistent Risk for Secondary Depression in the First Three Decades of Life. Arch Gen Psychiatry. 2007;64:903–12.
Stein MB, Fuetsch M, Müller N, Höfler M, Lieb R, Wittchen HU. Social anxiety disorder and the risk of depression: a prospective community study of adolescents and young adults. Arch Gen Psychiatry. 2001;58:251–6.
Baldwin DS, Anderson IM, Nutt DJ, Allgulander C, Bandelow B, den Boer, et al. Evidence-based pharmacological treatment of anxiety disorders, post-traumatic stress disorder and obsessive-compulsive disorder: a revision of the 2005 guidelines from the British Association for Psychopharmacology. J Psychopharmacol. 2014;28:403–39.
Stein MB, Stein DJ. Social anxiety disorder. Lancet. 2008;371:1115–25.
Pollack MH, Van Ameringen M, Simon NM, Worthington JW, Hoge EA, Keshaviah A, et al. A double-blind randomized controlled trial of augmentation and switch strategies for refractory social anxiety disorder. Am J Psychiatry. 2014;171:44–53.
Berg G, Rybakova D, Fischer D, Cernava T, Vergès M-CC, Charles T, et al. Microbiome definition re-visited: old concepts and new challenges. Microbiome. 2020;8:103.
Marchesi JR, Ravel J. The vocabulary of microbiome research: a proposal. Microbiome. 2015;3:31.
Sender R, Fuchs S, Milo R. Revised Estimates for the Number of Human and Bacteria Cells in the Body. PLoS Biol. 2016;14:e1002533–e1002533.
Backhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI. Host-bacterial mutualism in the human intestine. Science. 2005;307:1915–20.
Butler MI, Cryan JF, Dinan TG. Man and the Microbiome: A New Theory of Everything? Annu Rev Clin Psychol. 2019;15:371–98.
Dinan TG, Cryan JF. Regulation of the stress response by the gut microbiota: implications for psychoneuroendocrinology. Psychoneuroendocrinology. 2012;37:1369–78.
Wu W-L, Adame MD, Liou C-W, Barlow JT, Lai T-T, Sharon G, et al. Microbiota regulate social behaviour via stress response neurons in the brain. Nature. 2021;595:409–14.
Cryan JF, Dinan TG. Decoding the role of the microbiome on amygdala function and social behaviour. Neuropsychopharmacol: Off Publ Am Coll Neuropsychopharmacol. 2019;44:233–4.
Chen YH, Bai J, Wu D, Yu SF, Qiang XL, Bai H, et al. Association between fecal microbiota and generalized anxiety disorder: Severity and early treatment response. J Affect Disord. 2019;259:56–66.
Jiang HY, Zhang X, Yu ZH, Zhang Z, Deng M, Zhao JH, et al. Altered gut microbiota profile in patients with generalized anxiety disorder. J Psychiatr Res. 2018;104:130–6.
Mason BL, Li Q, Minhajuddin A, Czysz AH, Coughlin LA, Hussain SK, et al. Reduced anti-inflammatory gut microbiota are associated with depression and anhedonia. J Affect Disord. 2020;266:394–401.
De Palma G, Lynch MD, Lu J, Dang VT, Deng Y, Jury J, et al. Transplantation of fecal microbiota from patients with irritable bowel syndrome alters gut function and behavior in recipient mice. Sci Transl Med. 2017;9:eaaf6397.
Cryan JF, O’Riordan KJ, Cowan CSM, Sandhu KV, Bastiaanssen TFS, Boehme M, et al. The Microbiota-Gut-Brain Axis. Physiol Rev. 2019;99:1877–2013.
Liu RT, Walsh RFL, Sheehan AE. Prebiotics and probiotics for depression and anxiety: A systematic review and meta-analysis of controlled clinical trials. Neurosci Biobehav Rev. 2019;102:13–23.
Sarkar A, Harty S, Johnson KVA, Moeller AH, Carmody RN, Lehto SM, et al. The role of the microbiome in the neurobiology of social behaviour. Biol Rev. 2020;95:1131–66.
Sherwin E, Bordenstein SR, Quinn JL, Dinan TG, Cryan JF. Microbiota and the social brain. Science. 2019;366:eaar2016.
Johnson KVA. Gut microbiome composition and diversity are related to human personality traits. Hum Microbiome J. 2020;15:100069.
Kim C-S, Shin G-E, Cheong Y, Shin JH, Shin D-M, Chun WY. Experiencing social exclusion changes gut microbiota composition. Transl Psychiatry. 2022;12:254.
Heeren A, Dricot L, Billieux J, Philippot P, Grynberg D, de Timary P, et al. Correlates of Social Exclusion in Social Anxiety Disorder: An fMRI study. Sci Rep. 2017;7:260.
Hilimire MR, DeVylder JE, Forestell CA. Fermented foods, neuroticism, and social anxiety: An interaction model. Psychiatry Res. 2015;228:203–8.
Valles-Colomer M, Falony G, Darzi Y, Tigchelaar EF, Wang J, Tito RY, et al. The neuroactive potential of the human gut microbiota in quality of life and depression. Nat Microbiol. 2019;4:623–32.
Vieira-Silva S, Falony G, Darzi Y, Lima-Mendez G, Garcia Yunta R, Okuda S, et al. Species–function relationships shape ecological properties of the human gut microbiome. Nat Microbiol. 2016;1:16088.
Sheehan DV, Lecrubier Y, Sheehan KH, Amorim P, Janavs J, Weiller E, et al. The Mini-International Neuropsychiatric Interview (M.I.N.I.): the development and validation of a structured diagnostic psychiatric interview for DSM-IV and ICD-10. J Clin Psychiatry. 1998;59:22–33. quiz 34-57
Heimberg RG, Horner KJ, Juster HR, Safren SA, Brown EJ, Schneier FR, et al. Psychometric properties of the Liebowitz Social Anxiety Scale. Psychol Med. 1999;29:199–212.
Liebowitz MR. Social phobia. Mod Probl Pharmacopsychiatry. 1987;22:141–73.
Fresco DM, Coles ME, Heimberg RG, Liebowitz MR, Hami S, Stein MB, et al. The Liebowitz Social Anxiety Scale: a comparison of the psychometric properties of self-report and clinician-administered formats. Psychol Med. 2001;31:1025–35.
Baker SL, Heinrichs N, Kim HJ, Hofmann SG. The liebowitz social anxiety scale as a self-report instrument: a preliminary psychometric analysis. Behav Res Ther. 2002;40:701–15.
Oakman J, Van Ameringen M, Mancini C, Farvolden P. A confirmatory factor analysis of a self-report version of the Liebowitz Social Anxiety Scale. J Clin Psychol. 2003;59:149–61.
Riboli E, Kaaks R. The EPIC Project: rationale and study design. European Prospective Investigation into Cancer and Nutrition. Int J Epidemiol. 1997;26:S6–14.
Harrington J, PI, Lutomski J, Morgan K, McGee H, Shelley E, et al. SLÁN 2007: Survey of Lifestyle, Attitudes and Nutrition in Ireland. Dietary Habits of the Irish Population. Dublin: Department of Health and Children, 2008.
Mulligan AA, Luben RN, Bhaniani A, Parry-Smith DJ, O’Connor L, Khawaja AP, et al. A new tool for converting food frequency questionnaire data into nutrient and food group values: FETA research methods and availability. BMJ Open. 2014;4:e004503.
Lewis SJ, Heaton KW. Stool form scale as a useful guide to intestinal transit time. Scand J Gastroenterol. 1997;32:920–4.
Craig CL, Marshall AL, Sjöström M, Bauman AE, Booth ML, Ainsworth BE, et al. International physical activity questionnaire: 12-country reliability and validity. Med Sci Sports Exerc. 2003;35:1381–95.
Buysse DJ, Reynolds CF 3rd, Monk TH, Berman SR, Kupfer DJ. The Pittsburgh Sleep Quality Index: a new instrument for psychiatric practice and research. Psychiatry Res. 1989;28:193–213.
Andrews, S (2010) FastQC: a quality control tool for high throughput sequence data.
McIver LJ, Abu-Ali G, Franzosa EA, Schwager R, Morgan XC, Waldron L, et al. bioBakery: a meta’omic analysis environment. Bioinformatics. 2018;34:1235–7.
Huttenhower Lab, T. ([cited 2017 Dec 19]) “KneadData”.
Truong DT, Franzosa EA, Tickle TL, Scholz M, Weingart G, Pasolli E, et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat Methods. 2015;10:902–3. United States
Franzosa EA, McIver LJ, Rahnavard G, Thompson LR, Schirmer M, Weingart G, et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat Methods. 2018;15:962–8.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Aitchison J. The statistical analysis of compositional data. J Royal Stat Soc:Series B (Methodological). 1982;44:139–77.
Gloor GB, Macklaim JM, Pawlowsky-Glahn V, Egozcue JJ. Microbiome Datasets Are Compositional: And This Is Not Optional. Front Microbiol. 2017;8:2224.
Fernandes AD, Reid JN, Macklaim JM, McMurrough TA, Edgell DR, Gloor GB. Unifying the analysis of high-throughput sequencing datasets: characterizing RNA-seq, 16S rRNA gene sequencing and selective growth experiments by compositional data analysis. Microbiome. 2014;2:15.
Hsieh TC, Ma KH, Chao A. iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol Evol. 2016;7:1451–6.
Wickham H. ggplot2. WIREs Comp Stat. 2011;3:180–5.
Pedersen, TL 2019. “Patchwork: The composer of plots.” R package version 1.0.
Bastiaanssen, TF, Quinn, TP, Loughman, A. Treating Bugs as Features: A compositional guide to the statistical analysis of the microbiome-gut-brain axis. arXiv preprint arXiv:2207.12475, 2022.
Kelly JR, Borre Y, C OB, Patterson E, El Aidy S, Deane J, et al. Transferring the blues: Depression-associated gut microbiota induces neurobehavioural changes in the rat. J Psychiatr Res. 2016;82:109–18.
Guilhot E, Alou MT, Lagier JC, Labas N, Couderc C, Delerce J, et al. Genome sequence and description of Anaeromassilibacillus senegalensis gen. nov., sp. nov., isolated from the gut of patient with kwashiorkor. N. Microbes N. Infect. 2017;17:54–64.
Glendinning L, Stewart RD, Pallen MJ, Watson KA, Watson M. Assembly of hundreds of novel bacterial genomes from the chicken caecum. Genome Biol. 2020;21:34.
Medvecky M, Cejkova D, Polansky O, Karasova D, Kubasova T, Cizek A, et al. Whole genome sequencing and function prediction of 133 gut anaerobes isolated from chicken caecum in pure cultures. BMC Genom. 2018;19:561.
Ho LKH, Tong VJW, Syn N, Nagarajan N, Tham EH, Tay SK, et al. Gut microbiota changes in children with autism spectrum disorder: a systematic review. Gut Pathog. 2020;12:6.
Barandouzi ZA, Starkweather AR, Henderson WA, Gyamfi A, Cong XS. Altered Composition of Gut Microbiota in Depression: A Systematic Review. Front Psychiatry. 2020;11:541–541.
Zhu F, Ju Y, Wang W, Wang Q, Guo R, Ma Q, et al. Metagenome-wide association of gut microbiome features for schizophrenia. Nat Commun. 2020;11:1612.
Rhee SJ, Kim H, Lee Y, Lee HJ, Park CHK, Yang J, et al. The association between serum microbial DNA composition and symptoms of depression and anxiety in mood disorders. Sci Rep. 2021;11:13987.
Li J, Ma Y, Bao Z, Gui X, Li AN, Yang Z, et al. Clostridiales are predominant microbes that mediate psychiatric disorders. J Psychiatr Res. 2020;130:48–56.
Usui N, Matsuzaki H, Shimada S. Characterization of Early Life Stress-Affected Gut Microbiota. Brain Sci. 2021;11:913.
Bailey MT, Dowd SE, Galley JD, Hufnagle AR, Allen RG, Lyte M. Exposure to a social stressor alters the structure of the intestinal microbiota: implications for stressor-induced immunomodulation. Brain Behav Immun. 2011;25:397–407.
Hugenholtz F, de Vos WM. Mouse models for human intestinal microbiota research: a critical evaluation. Cell Mol Life Sci. 2018;75:149–60.
Fontana A, Manchia M, Panebianco C, Paribello P, Arzedi C, Cossu E, et al. Exploring the Role of Gut Microbiota in Major Depressive Disorder and in Treatment Resistance to Antidepressants. Biomedicines. 2020;8:311.
Inoue R, Sakaue Y, Kawada Y, Tamaki R, Yasukawa Z, Ozeki M, et al. Dietary supplementation with partially hydrolyzed guar gum helps improve constipation and gut dysbiosis symptoms and behavioral irritability in children with autism spectrum disorder. J Clin Biochem Nutr. 2019;64:217–23.
Chartier MJ, Walker JR, Stein MB. Considering comorbidity in socialphobia. Soc Psychiatry Psychiatr Epidemiol. 2003;38:728–34.
Maddox BB, White SW. Comorbid Social Anxiety Disorder in Adults with Autism Spectrum Disorder. J Autism Dev Disord. 2015;45:3949–60.
Würdemann D, Tindall BJ, Pukall R, Lünsdorf H, Strömpl C, Namuth T, et al. Gordonibacter pamelaeae gen. nov., sp. nov., a new member of the Coriobacteriaceae isolated from a patient with Crohn’s disease, and reclassification of Eggerthella hongkongensis Lau et al. 2006 as Paraeggerthella hongkongensis gen. nov., comb. nov. Int J Syst Evol Microbiol. 2009;59:1405–15.
Selma MV, Beltrán D, García-Villalba R, Espín JC, Tomás-Barberán FA. Description of urolithin production capacity from ellagic acid of two human intestinal Gordonibacter species. Food Funct. 2014;5:1779–84.
Westfall S, Pasinetti GM. The Gut Microbiota Links Dietary Polyphenols With Management of Psychiatric Mood Disorders. Front Neurosci. 2019;13:1196–1196.
Morotomi, M (2014) The family Sutterellaceae., in Rosenberg E, DE, Lory S, Stackebrandt E, Thompson F (ed.) The Prokaryotes. Berlin, Heidelberg: Springer, 1005–12.
Ding X, Xu Y, Zhang X, Zhang L, Duan G, Song C, et al. Gut microbiota changes in patients with autism spectrum disorders. J Psychiatr Res. 2020;129:149–59.
Zeng Q, Li D, He Y, Li Y, Yang Z, Zhao X, et al. Discrepant gut microbiota markers for the classification of obesity-related metabolic abnormalities. Sci Rep. 2019;9:13424.
Noble EE, Hsu TM, Jones RB, Fodor AA, Goran MI, Kanoski SE. Early-Life Sugar Consumption Affects the Rat Microbiome Independently of Obesity. J Nutr. 2017;147:20–28.
Kreutzer C, Peters S, Schulte DM, Fangmann D, Türk K, Wolff S, et al. Hypothalamic Inflammation in Human Obesity Is Mediated by Environmental and Genetic Factors. Diabetes. 2017;66:2407–15.
Coyte KZ, Schluter J, Foster KR. The ecology of the microbiome: Networks, competition, and stability. Science. 2015;350:663–6.
Sung J, Kim S, Cabatbat JJT, Jang S, Jin YS, Jung GY, et al. Global metabolic interaction network of the human gut microbiota for context-specific community-scale analysis. Nat Commun. 2017;8:15393.
Lim R, Cabatbat JJT, Martin TLP, Kim H, Kim S, Sung J, et al. Large-scale metabolic interaction network of the mouse and human gut microbiota. Sci Data. 2020;7:204.
Caspi R, Billington R, Keseler IM, Kothari A, Krummenacker M, Midford PE, et al. The MetaCyc database of metabolic pathways and enzymes - a 2019 update. Nucleic Acids Res. 2020;48:D445–d453.
Muriana FJ, Alvarez-Ossorio MC, Relimpio AM. Purification and characterization of aspartate aminotransferase from the halophile archaebacterium Haloferax mediterranei. Biochem J. 1991;278:149–54.
Birolo L, Tutino ML, Fontanella B, Gerday C, Mainolfi K, Pascarella S, et al. Aspartate aminotransferase from the Antarctic bacterium Pseudoalteromonas haloplanktis TAC 125. Cloning, expression, properties, and molecular modelling. Eur J Biochem. 2000;267:2790–802.
Marino G, Nitti G, Arnone MI, Sannia G, Gambacorta A, De Rosa M. Purification and characterization of aspartate aminotransferase from the thermoacidophilic archaebacterium Sulfolobus solfataricus. J Biol Chem. 1988;263:12305–9.
Powell JT, Morrison JF. The purification and properties of the aspartate aminotransferase and aromatic-amino-acid aminotransferase from Escherichia coli. Eur J Biochem. 1978;87:391–400.
Fotheringham IG, Dacey SA, Taylor PP, Smith TJ, Hunter MG, Finlay ME, et al. The cloning and sequence analysis of the aspC and tyrB genes from Escherichia coli K12. Comparison of the primary structures of the aspartate aminotransferase and aromatic aminotransferase of E. coli with those of the pig aspartate aminotransferase isoenzymes. Biochem J. 1986;234:593–604.
Purton T, Staskova L, Lane MM, Dawson SL, West M, Firth J, et al. Prebiotic and probiotic supplementation and the tryptophan-kynurenine pathway: A systematic review and meta analysis. Neurosci Biobehav Rev. 2021;123:1–13.
Spichak S, Bastiaanssen TFS, Berding K, Vlckova K, Clarke G, Dinan TG, et al. Mining microbes for mental health: Determining the role of microbial metabolic pathways in human brain health and disease. Neurosci Biobehav Rev. 2021;125:698–761.
Gheorghe CE, Martin JA, Manriquez FV, Dinan TG, Cryan JF, Clarke G. Focus on the essentials: tryptophan metabolism and the microbiome-gut-brain axis. Curr Opin Pharm. 2019;48:137–45.
Kennedy PJ, Cryan JF, Dinan TG, Clarke G. Kynurenine pathway metabolism and the microbiota-gut-brain axis. Neuropharmacology. 2017;112:399–412.
Trevelline BK, Kohl KD. The gut microbiome influences host diet selection behavior. Proc Natl Acad Sci. 2022;119:e2117537119.
Fuertig R, Azzinnari D, Bergamini G, Cathomas F, Sigrist H, Seifritz E, et al. Mouse chronic social stress increases blood and brain kynurenine pathway activity and fear behaviour: Both effects are reversed by inhibition of indoleamine 2,3-dioxygenase. Brain, Behav, Immun. 2016;54:59–72.
Kiank C, Zeden JP, Drude S, Domanska G, Fusch G, Otten W, et al. Psychological stress-induced, IDO1-dependent tryptophan catabolism: implications on immunosuppression in mice and humans. PLoS One. 2010;5:e11825.
Chiappelli J, Pocivavsek A, Nugent KL, Notarangelo FM, Kochunov P, Rowland LM, et al. Stress-induced increase in kynurenic acid as a potential biomarker for patients with schizophrenia and distress intolerance. JAMA Psychiatry. 2014;71:761–8.
Plitman E, Iwata Y, Caravaggio F, Nakajima S, Chung JK, Gerretsen P, et al. Kynurenic Acid in Schizophrenia: A Systematic Review and Meta-analysis. Schizophr Bull. 2017;43:764–77.
Butler MI, Long-Smith C, Moloney GM, Morkl S, O’Mahony SM, Cryan JF, et al. The immune-kynurenine pathway in social anxiety disorder. Brain Behav Immun. 2022;99:317–26.
Kuc D, Zgrajka W, Parada-turska J, Urbanik-sypniewska T, Turski WA. Micromolar concentration of kynurenic acid in rat small intestine. Amino Acids. 2008;35:503–5.
Han Q, Fang J, Li J. Kynurenine aminotransferase and glutamine transaminase K of Escherichia coli: identity with aspartate aminotransferase. Biochemical J. 2001;360:617–23.
Guastella AJ, Richardson R, Lovibond PF, Rapee RM, Gaston JE, Mitchell P, et al. A randomized controlled trial of D-cycloserine enhancement of exposure therapy for social anxiety disorder. Biol Psychiatry. 2008;63:544–9.
Hofmann SG, Meuret AE, Smits JA, Simon NM, Pollack MH, Eisenmenger K, et al. Augmentation of exposure therapy with D-cycloserine for social anxiety disorder. Arch Gen Psychiatry. 2006;63:298–304.
Baran H, Kepplinger B. D-Cycloserine lowers kynurenic acid formation-new mechanism of action. Eur Neuropsychopharmacol. 2014;24:639–44.
Sunagawa S, Mende DR, Zeller G, Izquierdo-carrasco F, Berger SA, Kultima JR, et al. Metagenomic species profiling using universal phylogenetic marker genes. Nat Methods. 2013;10:1196–9.
Matias Rodrigues JF, Schmidt TSB, Tackmann J, von Mering C. MAPseq: highly efficient k-mer search with confidence estimates, for rRNA sequence analysis. Bioinformatics. 2017;33:3808–10.
Vujkovic-Cvijin I, Sklar J, Jiang L, Natarajan L, Knight R, Belkaid Y. Host variables confound gut microbiota studies of human disease. Nature. 2020;587:448–454.
Falony G, Joossens M, Vieira-Silva S, Wang J, Darzi Y, Faust K, et al. Population-level analysis of gut microbiome variation. Science. 2016;352:560–4.
John GK, Mullin GE. The Gut Microbiome and Obesity. Curr Oncol Rep. 2016;18:45.
Mailing LJ, Allen JM, Buford TW, Fields CJ, Woods JA. Exercise and the Gut Microbiome: A Review of the Evidence, Potential Mechanisms, and Implications for Human Health. Exerc Sport Sci Rev. 2019;47:75–85.
Cussotto S, Clarke G, Dinan TG, Cryan JF. Psychotropics and the Microbiome: a Chamber of Secrets…. Psychopharmacology. 2019;236:1411–32.
Maier L, Pruteanu M, Kuhn M, Zeller G, Telzerow A, Anderson EE, et al. Extensive impact of non-antibiotic drugs on human gut bacteria. Nature. 2018;555:623–8.
Ait Chait Y, Mottawea W, Tompkins TA, Hammami R. Unravelling the antimicrobial action of antidepressants on gut commensal microbes. Sci Rep. 2020;10:17878–17878.
Cussotto S, Strain CR, Fouhy F, Strain RG, Peterson VL, Clarke G, Stanton C, Dinan TG, Cryan JF. Differential effects of psychotropic drugs on microbiome composition and gastrointestinal function. Psychopharmacology (Berl). 2019;236:1671–85.
Younis W, AbdelKhalek A, Mayhoub AS, Seleem MN. In Vitro Screening of an FDA-Approved Library Against ESKAPE Pathogens. Curr Pharm Des. 2017;23:2147–57.
Bohnert JA, Szymaniak-Vits M, Schuster S, Kern WV. Efflux inhibition by selective serotonin reuptake inhibitors in Escherichia coli. J Antimicrob Chemother. 2011;66:2057–60.
Ayaz M, Subhan F, Ahmed J, Khan AU, Ullah F, Ullah I, et al. Sertraline enhances the activity of antimicrobial agents against pathogens of clinical relevance. J Biol Res (Thessalon). 2015;22:4.
Ramsteijn AS, Jašarević E, Houwing DJ, Bale TL, Olivier JD. Antidepressant treatment with fluoxetine during pregnancy and lactation modulates the gut microbiome and metabolome in a rat model relevant to depression. Gut Microbes. 2020;11:735–53.
Lukić I, Getselter D, Ziv O, Oron O, Reuveni E, Koren O, et al. Antidepressants affect gut microbiota and Ruminococcus flavefaciens is able to abolish their effects on depressive-like behavior. Transl Psychiatry. 2019;9:133–133.
Lyte M, Daniels KM, Schmitz-Esser S. Fluoxetine-induced alteration of murine gut microbial community structure: evidence for a microbial endocrinology-based mechanism of action responsible for fluoxetine-induced side effects. PeerJ. 2019;7:e6199.
Liśkiewicz P, Pełka-Wysiecka J, Kaczmarczyk M, Łoniewski I, Wroński M, Bąba-Kubiś A, et al. Fecal Microbiota Analysis in Patients Going through a Depressive Episode during Treatment in a Psychiatric Hospital Setting. J Clin Med. 2019;8:164.
Tomizawa Y, Kurokawa S, Ishii D, Miyaho K, Ishii C, Sanada K, et al. Effects of Psychotropics on the Microbiome in Patients With Depression and Anxiety: Considerations in a Naturalistic Clinical Setting. Int J Neuropsychopharmacol. 2021;24:97–107.
Flowers SA, Evans SJ, Ward KM, McInnis MG, Ellingrod VL. Interaction Between Atypical Antipsychotics and the Gut Microbiome in a Bipolar Disease Cohort. Pharmacotherapy. 2017;37:261–7.
Zhang W, Qu W, Wang H, Yan H. Antidepressants fluoxetine and amitriptyline induce alterations in intestinal microbiota and gut microbiome function in rats exposed to chronic unpredictable mild stress. Transl Psychiatry. 2021;11:131.
Naseribafrouei A, Hestad K, Avershina E, Sekelja M, Linlokken A, Wilson R, et al. Correlation between the human fecal microbiota and depression. Neurogastroenterol Motil. 2014;26:1155–62.
Song Y, Liu C, Finegold SM. Real-time PCR quantitation of clostridia in feces of autistic children. Appl Environ Microbiol. 2004;70:6459–65.
Parracho HM, Bingham MO, Gibson GR, McCartney AL. Differences between the gut microflora of children with autistic spectrum disorders and that of healthy children. J Med Microbiol. 2005;54:987–91.
Jacka FN, O’Neil A, Opie R, Itsiopoulos C, Cotton S, Mohebbi M, et al. A randomised controlled trial of dietary improvement for adults with major depression (the ‘SMILES’ trial). BMC Med. 2017;15:23.
Kang D-W, Adams JB, Gregory AC, Borody T, Chittick L, Fasano A, et al. Microbiota Transfer Therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: an open-label study. Microbiome. 2017;5:10.
Acknowledgements
The authors would like to sincerely thank the participants who volunteered for the study and without whose time and effort this research would not have been possible. APC Microbiome Ireland is a research centre funded by Science Foundation Ireland (Grant number SFI/12/RC/2273_P2).
Author information
Authors and Affiliations
Contributions
MIB conceptualized and designed the study, performed participant assessments, performed data analysis, interpreted the data and drafted the manuscript. TFSB performed the bioinformatics of the microbiota data. CLS contributed to study design and data interpretation. SM performed participant assessments and contributed to data interpretation. KB contributed to analysis and interpretation of dietary data. CS, DP, SP and CS contributed to DNA extraction, library preparation and microbiota sequencing. SMOM and NLR contributed to data interpretation. JFC, GC and TGD contributed to the design of the study, interpreted the data, and contributed to writing of the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
GC has received honoraria from Janssen, Probi, and Apsen as an invited speaker; is in receipt of research funding from Pharmavite, Tate and Lyle, Nestle and Fonterra; and is a paid consultant for Yakult, Zentiva, and Heel pharmaceuticals. JFC has been an invited speaker at conferences organized by Mead Johnson, Alkermes, Janssen, Ordesa, and Yakult. TGD and JC have received research funding from GlaxoSmithKline, Mead Johnson, Cremo Nutricia, Pharmavite, Dupont, and 4D Pharma. This support neither influenced nor constrained the contents of this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Butler, M.I., Bastiaanssen, T.F.S., Long-Smith, C. et al. The gut microbiome in social anxiety disorder: evidence of altered composition and function. Transl Psychiatry 13, 95 (2023). https://doi.org/10.1038/s41398-023-02325-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-023-02325-5
This article is cited by
-
Association between gut microbiota and anxiety disorders: a bidirectional two-sample mendelian randomization study
BMC Psychiatry (2024)
-
Stress-resilience impacts psychological wellbeing as evidenced by brain–gut microbiome interactions
Nature Mental Health (2024)
-
Bifidobacterium longum 1714 improves sleep quality and aspects of well-being in healthy adults: a randomized, double-blind, placebo-controlled clinical trial
Scientific Reports (2024)
-
Gut microbiome and psychiatric disorders
BMC Psychiatry (2023)