Abstract
Suicides have increased to over 48,000 deaths yearly in the United States. Major depressive disorder (MDD) is the most common diagnosis among suicides, and identifying those at the highest risk for suicide is a pressing challenge. The objective of this study is to identify changes in gene expression associated with suicide in brain and blood for the development of biomarkers for suicide. Blood and brain were available for 45 subjects (53 blood samples and 69 dorsolateral prefrontal cortex (DLPFC) samples in total). Samples were collected from MDD patients who died by suicide (MDD-S), MDDs who died by other means (MDD-NS) and non-psychiatric controls. We analyzed gene expression using RNA and the NanoString platform. In blood, we identified 14 genes which significantly differentiated MDD-S versus MDD-NS. The top six genes differentially expressed in blood were: PER3, MTPAP, SLC25A26, CD19, SOX9, and GAR1. Additionally, four genes showed significant changes in brain and blood between MDD-S and MDD-NS; SOX9 was decreased and PER3 was increased in MDD-S in both tissues, while CD19 and TERF1 were increased in blood but decreased in DLPFC. To our knowledge, this is the first study to analyze matched blood and brain samples in a well-defined population of MDDs demonstrating significant differences in gene expression associated with completed suicide. Our results strongly suggest that blood gene expression is highly informative to understand molecular changes in suicide. Developing a suicide biomarker signature in blood could help health care professionals to identify subjects at high risk for suicide.
Similar content being viewed by others
Introduction
Suicide is a serious global public health problem that accounts for close to 800,000 deaths per year [1]. In the United States alone, suicide rates increased by more than 35% over the past 20 years, and in the past year, over 48,000 Americans died by suicide, making it the 10th leading cause of death [2]. Suicide prevention strategies and current medications, although helpful, have not stemmed the increase in self-inflicted deaths. Many individuals do not disclose suicidal intentions despite frequent contact with health care professionals. An estimated 30% of suicides visit a healthcare provider within a month of the suicide event [3, 4]. A dramatic rise in suicide also occurs in the days to weeks following discharge from psychiatric hospitals [5,6,7]. Thus, there is a critical opportunity for healthcare providers to evaluate at-risk individuals with a blood biomarker test to assess serious suicide intent.
Major depressive disorder (MDD) is the most common psychiatric diagnosis in suicide although the underlying biological determinants have yet to be identified. Converging evidence suggests that suicide is associated with a dysregulated stress response system and concurrently, an increase in inflammatory responses and their relevant downstream effects [8]. Additional clues to suicidality come from postmortem investigations in brain from depressed suicides showing disruptions in glucocorticoid and inflammatory responses [9,10,11,12,13], excitatory and inhibitory neurotransmitters (glutamate and GABA) [11, 14, 15], as well as dysregulation of polyamine stress response [16,17,18,19,20,21,22,23,24]. Expression of polyamine related genes in particular has consistently been shown to be altered in suicides compared to controls. The first study to specifically evaluate gene expression in DLPFC from depressed suicides (MDD-S, N = 21), depressed non-suicides (MDD-NS, N = 9) and controls (N = 29) using RNA-Seq found significant alterations of the spermidine/spermine N1-Acetyltransferase1 (SAT1) gene in both MDD groups [16]. A similar study reported dysregulation of SAT1 and significant alterations in glial cells in MDD-S [25]. In translational studies, SAT1 expression in blood was shown to be a potential peripheral suicide biomarker in males [26, 27], but not in females [28] suggesting a sex-specific effect that is not often taken into account when studying suicide.
Although many investigations focused on postmortem brain, to our knowledge there are no studies comparing gene expression in both postmortem blood and brain in samples taken from the same individuals. In this study we analyzed gene expression in MDD patients who died by suicide (MDD-S), in MDD patients who died of other causes (MDD-NS) and in non-psychiatric controls (C). The objective of our study is to provide preliminary information for the development of a blood biomarker signature that can be used to identify MDD patients at high risk for suicide.
Materials and methods
Subjects and samples
Brain and blood samples were obtained from the University of California, Irvine (UCI)-Pritzker Brain Bank per the Institutional Review Board (IRB). Consent was obtained from next-of-kin. Samples consisted of archival blood and dorsolateral prefrontal cortex (DLPFC) from MDD-suicides (N = 24 brain, 19 blood), MDD-non-suicides (N = 24 brain, 18 blood), and non-psychiatric controls (N = 21 brain, 16 blood). A psychological autopsy, through an extensive review of multiple sources of information, was performed by trained clinicians as described elsewhere [15, 29, 30]. Table 1 and Supplementary Table 1 list demographics of the subjects, including gender, age, diagnosis, race, blood/brain availability, cause of death, and clinical characteristics.
Postmortem blood samples were collected in EDTA tubes from the heart or femoral vein and stored in −20 °C freezers until RNA extraction. In some cases, only brain tissue was available (coagulation sometimes prevents blood draw) while only blood was available for some subjects due to consent preferences by the next-of-kin (Supplementary Table 1). The human brain dissection and freezing protocol is described in detail in Supplementary Information. Briefly, the whole brain is first inspected and photographed before the brainstem and cerebellum are separated from the cerebrum. The cerebrum is coronally sliced from anterior to posterior and slices are placed on steel plates, labeled, photographed, and flash-frozen using cooled 2-methyl butane (−40 °C). Once frozen the brain slices are bagged and vacuum-sealed, then stored in −80 °C freezers until time for further dissections or extractions.
RNA extractions
RNA extraction from blood was performed using the QIAamp RNA blood mini kit (Qiagen, Carlsbad, CA) and RNA extractions for brain samples were carried out using the Qiagen AllPrep DNA/RNA/Protein mini kit (Qiagen, Carlsbad, CA), both according to the manufacturer’s protocol. RNA concentrations were determined using Qubit fluorometric quantification (Thermo Fisher Scientific, Carlsbad, CA). Integrity of total RNA was evaluated using 18S and 28S ratios and the RNA integrity number (RIN) from the 2100 Agilent Bioanalyzer (Agilent technologies, Santa Clara, CA). RINs for both brain and blood are illustrated in Supplementary Fig. 1. RNA quality measures [i.e., pH, RIN, and postmortem interval (PMI)] for the brain samples are outlined in Supplementary Table 2 showing averages and standard deviations per group. Additionally, FOV (fields of view) were compared between brain and blood samples. The FOV is an imaging quality control measure from NanoString, which is indicative of the number of mRNA counts successfully imaged per lane in the array. Samples having low assay efficiency due to low quality of RNA will have a low number of counts and will result in a quality flag. The FOV threshold is set at 75% and any sample below this threshold is flagged and should not be used for downstream analysis.
NanoString assays for blood and brain
RNA (5 µl at a concentration of 20 ng/µl) was directly quantified using the NanoString nCounter analysis system (NanoString, Seattle, WA). The NanoString platform allows, without reverse transcription, to directly and digitally quantify the number of mRNA molecules present in a sample in a highly multiplexed manner within a single reaction using colored molecular barcodes. NanoString is less sensitive to RNA degradation and factors affecting RNA quality, such as sample degradation, fixing procedures, postmortem interval, and pH because RNA is directly assayed without reverse transcription or amplification [31,32,33]. This platform was therefore particularly useful to assay gene expression in postmortem archival blood samples. Samples were processed at the Genomics High-Throughput Facility of the University of California, Irvine. NanoString probes were custom designed to include house-keeping genes and 114 genes of interest for depression and suicide, including genes within the polyamine and metallothionein systems, stress response, glutamatergic and GABAergic neurotransmission, immune response, circadian rhythm, mitochondria function, and telomere length. Gene targets, accession number, probe position, and supporting references are outlined in Supplementary Table 3.
Statistical analysis
NanoString mRNA counts were processed using nSolver analysis software version 4.0 from NanoString for quality control (probe binding, counts, and image quality). The housekeeping genes included in our custom assay, detailed in Supplementary Table 3, included GAPDH, CYC1, and SDHA. Normalization procedures were done using the nCounter Analysis system global method, suggested by NanoString, that uses all probes or features to calculate a normalization factor instead of using a small number of housekeeping genes. Counts were logged (base 2) and imported into Partek Genomics Suite (Partek Inc, St-Louis, USA) for statistical analysis. ANOVA was used to investigate the effect of demographic variables, pH, and PMI on log2 gene expression levels. Gene expression and clinical phenotypic data were analyzed using multiple linear regression correcting for gender, postmortem factors, and age. The effects of suicide status, diagnosis and demographic variables (age, gender, and PMI) on gene expression were investigated using a 2-way ANCOVA model. For the DLPFC and blood, the influence of the time of death (TOD) on suicide and gene expression was assessed by adjusting the TOD to hours from sunrise as a continuous variable in the model to measure possible circadian effects. The adjusted TOD had a significant effect in the DLPFC, particularly for the expression of clock genes and was therefore included in the final analyses reported in the results section. Pair-wise post-hoc analyses were further performed to determine genes associated with suicide in MDD (MDD-S vs. MDD-NS). Other comparisons included the control group and both MDD groups (Control vs. MDD-NS and Control vs. MDD-S) as well as a comparison between the control and both MDD groups regardless of the suicide status Control vs. MDDs (non-suicides and suicides). To correct for multiple testing, a Benjamini–Hochberg false discovery rate (FDR) correction was used to generate adjusted q-values [34]. An estimate of the rate of the probability of a significant gene being a false discovery was calculated taking into account the number of comparisons (3 comparisons × 78 genes that were present both in blood and brain = 234 comparisons). A threshold of q < 0.1 was used for significance based on previous similar gene expression studies [16, 25] but focus on p-values was favored as this is an exploratory analysis of potential biomarkers to be confirmed in future studies.
We also sought to cross-validate our differentially expressed gene results with previous studies aiming to find suicide biomarkers in patients with various psychiatric disorders (MDDs, bipolar disorder and schizophrenia) using post-mortem and clinical blood [35]. We pooled our list of differently expressed genes between MDD-S and MDD-NS in blood/brain and compared our results to Niculescu and colleagues [35] Bonferroni-corrected lists of suicide biomarkers, which they validated in post-mortem samples. These analyses have a caveat in that our results are specific to MDD patients and we used a targeted multiplexed gene expression assay, while Niculescu et al. [35] used high throughput microarrays to study a number of psychiatric diagnoses (attention deficit disorder, anxiety disorder, bipolar disorder, MDD, mood disorder, not otherwise specified, psychosis not otherwise specified, post-traumatic stress disorder, schizoaffective disorder, and schizophrenia).
Results
No significant differences were observed for pH or RIN between MDD-S, MDD-NS, and Controls for blood and DLPFC. Gender and age varied between groups and were included as covariates. The DLPFC (but not blood) showed an effect of time of death (“hours since sunrise”) and was included as a covariate in the DLPFC analyses (Supplementary Table 4). The quality of the RNA from post-mortem blood samples was lower than the brain RNA quality as expected. RINs for the blood RNA were low for a majority of the subjects and high in brain for most of the subjects (Supplementary Fig. 1) unsurprisingly as we used RNA extracted from non-preserved blood. The NanoString platform and additional data quality measures were used to make sure blood RNA could be used to obtain reliable gene expression data. Specifically, the fields of view (FOV), an imaging quality control measure indicative of the number of mRNA counts imaged per lane of the array were compared between brain and blood samples (Supplemental Table 5). The percent FOV was comparable between brain and blood and exceeded the 75% threshold set by NanoString, thus demonstrating the lack of effect of low RIN in the blood on mRNA measures.
Suicide blood biomarkers
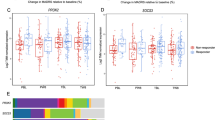
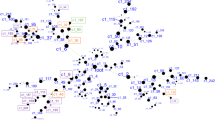
Our main objective was to identify possible blood biomarkers for suicide comparing MDD-S and MDD-NS. As shown in Table 2, 14 genes significantly differentiated MDD-S from MDD-NS, at p ≤ 0.05, and eleven of these genes survived FDR correction at q ≤ 0.1, which is depicted in Fig. 1. Of these genes, two circadian genes, period circadian regulator 3 (PER3) and PER2 were significantly upregulated in MDD-S vs. MDD-NS (PER3: p = 0.001, Q = 0.1; FC = 2.650; PER2: (p = 0.041, q = 0.207; FC = 1.871). Additional differentially expressed genes (DEGs) included MTPAP (Mitochondrial Poly(A) Polymerase), SLC25A26 (Solute Carrier Family 25 Member 26), CD19 (B-Lymphocyte Antigen CD19), and GAR1 (GAR1 Ribonucleoprotein) while SOX9 (SRY-Box Transcription Factor 9) was decreased in suicides vs non-suicide MDDs.
Graphical representation of eleven genes significantly differentially expressed between MDD-S and MDD-NS in blood, at p-value ≤ 0.05, and also passing FDR correction for multiple testing at q ≤ 0.1. Four of the 11 genes; MTPAP (Mitochondrial Poly(A) Polymerase), SLC25A26 (Solute Carrier Family 25 Member 26), NR3C1 (Nuclear Receptor Subfamily 3 Group C Member 1), and POT1 (Protection of Telomeres 1) were also significantly differentially expressed between MDD-S and control groups.
Two lymphocyte markers, CD19 and CD6, were upregulated MDD-S vs. MDD-NS (Fig. 1 and Table 2). CD19 is a surface membrane antigen of B-Lymphocytes used as a biomarker for lymphocyte development [36, 37]. Increased CD19+ B cells have been previously reported to be associated with depression [38]. CD6 is a surface membrane antigen of T-lymphocytes involved in the activation of T cells [39], and considered a promising target for autoimmunity and neuroinflammation [40]. CD6 is implicated in T cell infiltration into the central nervous system due its role in T cell trafficking across endothelial cell barriers [41]. Genetic variants of CD6 are associated with clinical response to TNF-alpha (Tumor Necrosis Factor-alpha) [42].
Twenty genes in blood (p-value ≤ 0.05; Q-value ≤ 0.1) were differentially expressed between MDD-S and controls (Supplementary Table 6). Interestingly, many of these genes overlapped with those identified between MDD-S and MDD-NS, including SLC25A26, GAR1, MTPAP, and NR3C1 (glucocorticoid receptor; Nuclear Receptor Subfamily 3 Group C Member 1). Also, of note is the significant difference in SAT1 gene expression in suicides. This gene has been associated with suicide and depression in postmortem brains and clinical populations with suicidal ideation [16, 24, 26, 43]. Comparisons were also carried out between MDD-NS and Controls (Supplementary Table 7), and an overall comparison between MDDs (MDD-S and MDD-NS) and Controls (Supplementary Table 8).
Postmortem DLPFC gene expression
A secondary objective of the current study was to investigate gene expression differences relevant to suicide in postmortem brain. Of the genes analyzed in the DLPFC, seventeen genes were downregulated, while only one gene (AGMAT) was upregulated in MDD-S compared to MDD-NS (at p-value ≤ 0.01, Q-value ≤ 0.1 (for 16 of the genes)) (Fig. 2 and Table 3). The top gene differentially expressed between MDD-S and MDD-NS was SAT1. In addition to the main SAT1 probe (SAT1_all), measuring all splice variants of the SAT1 gene, two additional splice variant specific probes (SAT1-002 and SAT1-001) were also significantly reduced in MDD-S (Fig. 2 and Table 3). The second most significant gene between MDD-S and MDD-NS was the brain-specific, MT3 (Metallothionein 3) gene, which was decreased in suicides. A total of four metallothioneins (MT3, MT2A, MT1A, and MT1X) were significantly reduced in MDD-S, although these same genes did not reach significance in blood. Metallothioneins are known to be involved in neuroprotection against, among other things, the deleterious effects of long-term increases in cortisol levels [44]. The decreased expression of SAT1 and MT3, as well as other members of the polyamine and metallothionein families, were previously reported to be differentially expressed in MDD-S by our group [15, 24].
Comparisons were also carried out between suicide (MDD-S) and controls (Supplementary Table 9). Five genes were significantly differentially expressed in MDD-S (p ≤ 0.05), with the most significant, and only gene passing FDR, being SAT1. This finding provides additional evidence for the involvement of the polyamine stress response in suicide. Gene expression differences were also observed between MDD-NS and controls (Supplementary Table 10), and both MDD groups (MDD-S + MDD-NS) versus controls (Supplementary Table 11). Top genes in both these comparisons included the upregulation of stress-related genes (FKBP5, ODC1, and a splice variant of SAT1).
Blood and brain concordant changes
Most expression changes associated with suicide were tissue specific, meaning they were observed either in blood or brain only. However, four genes showed significant changes in both brain and blood (at p-value ≤ 0.05, Supplementary Fig. 1). Two of these genes were concordant with regards to directionality (PER3 and SOX9). The circadian rhythm gene PER3 was significantly increased in MDD-S compared to MDD-NS (Fig. 1 and Table 3). PER3 is an important timekeeping gene involved in the regulation of circadian rhythms and is associated with delayed sleep phase disturbances in humans [45]. The other gene concordant between blood and brain was SOX9. This gene was significantly reduced in both blood and brain of MDD-S compared to MDD-NS (Figs. 1 and 2). SOX9, a marker of B cells in blood and astrocytes in brain, has previously been shown to be decreased in the frontal cortex in MDD, by our group [46], and in depressed suicides [47]. The two remaining genes, CD19 and TERF1 (Telomere Regulation factor 1), were increased in suicides in blood, while being decreased in the brain. CD19 was differentially increased in MDD-S suggesting B cell mediated inflammation in suicides. TERF1 codes for a protein that binds telomere ends and is part of the shelterin nucleoprotein complex. We [48] and others [49] have previously reported aberrant telomere length in brain and blood associated with MDD and suicide.
Cross-validation of our results with previous studies aiming to identify blood biomarkers to predict suicide risk revealed some overlapping genes. We found increased expression of SAT1 in the blood of MDD-S versus MDD-NS, although this association did not reach significance. However, when comparing MDD-S to controls we observed a significant increase of SAT1 expression associated with suicide (Supplementary Table 6). Through liquid biopsies and post-mortem validation, Niculescu et al. [35], also showed increased SAT1 levels to be associated with suicide in blood.
In our current study, and in a previous one using a cohort of mood disorder patients (MDDs and bipolar disorder) [30], we observed significantly decreased levels of the metallothionein family of genes to be associated with suicide. Similarly, Niculescu and colleagues found Metallothionein 1E (MT1E) among their universal Bonferroni validated biomarkers for suicide [35]. Lastly, the SKA2 (Spindle and Kinetochore Associated Complex Subunit 2) gene was found to be down-regulated in suicide, in both our blood and brain samples, as well as in Niculescu’s blood samples [35].
Discussion
A targeted gene expression approach was developed to assess relevant suicide specific biomarkers in postmortem blood and brain using a platform designed to overcome any bias introduced due to sample degradation (NanoString). We used frozen blood, not preserved in specialized RNA stabilizing buffers, because of its availability and common use by Coroners and Medical Examiners for toxicology. Our results suggest non-preserved blood can be used to discover suicide specific biomarkers that can also be assessed in brain tissue from the same subjects when available. Little is known about how gene expression levels in brain directly correlate with expression changes in blood, because brain and blood samples are not usually obtained from the same subject. To our knowledge, this is the first study to determine suicide specific gene expression changes both in blood and brain tissue from the same MDD subjects to develop a molecular signature in blood to predict individuals at highest risk for suicide. We clearly show that among MDD patients, suicide victims have a gene expression signature distinct from non-suicides that includes key genes involved in stress response and polyamine metabolism, immune response, and telomere maintenance. Genes identified as dysregulated in suicide could be potential targets for future pharmacological interventions to prevent suicide in MDD patients but could also be used to develop a molecular test to identify patients at high risk for suicide.
Previous studies have predominantly focused on subjects with suicidal behaviors, but not on suicide victims, to identify state (epigenetic, expression) or trait markers for suicide by performing genome wide association studies (GWAS) in samples of patients with suicidal ideation or suicide attempts [50,51,52,53,54,55,56,57,58,59]. GWAS studies in particular have identified single nucleotide polymorphisms (SNPs) associated with suicide attempts and suicidal behaviors [50,51,52,53, 56, 58, 60]. These SNP associations while informative for suicidal behaviors are not necessarily helpful for the identification of subjects about to commit suicide. Subjects with suicidal behaviors, including attempters might have a different clinical presentation and represent a distinct phenotype that does not always predict future suicide. This makes the study of actual suicide patients particularly important for the identification, understanding and prediction of at-risk individuals.
Recent studies have shown suicide specific molecular signatures can be found in blood and brain tissue. Four recent genome wide gene association studies (GWAS) of suicide were conducted using blood [58, 61,62,63]. These studies, while showing a high SNP-based heritability, in large admixed populations across psychiatric diagnoses, revealed no overlap in the identification of genes associated with completed suicide. However, the association of the HTR2A (5-Hydroxytryptamine Receptor 2A) gene with death by suicide identified by Coon and colleagues [63] is corroborated by our observation of decreased HTR2A expression in MDD-S blood when compared to controls (Supplementary Table 6). Several recent investigations of gene expression were also conducted in brain tissue from suicide victims. Nagy et al. [64] performed single nuclei gene expression analysis in DLPFC from MDD suicide completers compared to controls, and found 96 differentially expressed genes and major cellular dysregulation in deep layer excitatory neurons and immature oligodendrocyte precursor cells (OPCs). Mahajan et al. [65] investigated expression changes in the hippocampus of MDDs and found significant dysregulation of immune-related genes (inflammation, cytokines) in MDDs although they did not directly compare MDD-S to MDD-NS. Finally, Jabbi et al. [66] investigated gene expression changes in suicide patients with mixed diagnoses (MDD, bipolar disorder [BD]) in the insula. Their results also showed changes in suicides vs. controls mainly related to immune function and cytokines. Overall, these recent GWAS and gene expression studies did not directly compare patients, who died by suicide to patients who died of other causes.
Comparisons have previously been made between gene expression in the brain and peripheral blood, with both tissues sharing significant transcriptomic expression [67, 68]. However, the correlation is approximately 0.5, indicating that not all genes are expressed in the same direction between both tissues. For example, SAT1, which has been consistently reported to be significantly down-regulated in the suicide brain [16, 20, 24], was found to be increased in living bipolar subjects with suicidal ideation [27], and in two groups of post-mortem blood from all-male suicide completers with mixed diagnoses (schizophrenia, bipolar, and MDD) [26, 35]. Additional evidence also shows isoform-specific differential expression of SAT1 and risk allele association with suicide [16, 20, 24]. In our study we minimized diagnostic variability by using only MDD subjects and observed a significant decrease in SAT1 expression in the brain, although SAT1 was increased in MDD-S (vs. MDD-NS), in blood, it did not reach significance. Altogether, this evidence suggests that SAT1 has the potential to be both an indicator of suicidal ideation (SI) across certain psychiatric mood disorders, and be a predictor of suicide completion that is amenable to pharmacological intervention.
Our previous work shows significantly decreased MT1E expression in both the Nucleus Accumbens and anterior cingulate cortex of mood disorder patients who died by suicide compared to non-suicides [30], while Niculescu and colleagues have observed increased MT1E expression to be among their validated universal biomarkers associated with suicidality in blood [35]. This suggests metallothionein genes could be involved in suicide biology; however, further investigation is needed to determine if select MT genes can identify patients at risk for suicide and/or predict suicide completion. Lastly, another gene characterized by Niculescu and colleagues [35] as a universal biomarker for suicide is SKA2, which was downregulated. In our cohort, this gene had decreased expression in MDD-S compared to MDD-NS but, did not reach statistical significance. Overall, there is some validation of the blood-based findings between our study and that by Niculescu and colleagues [35] (SAT1, SKA2, and metallothionein family genes) even with the methodological differences between the studies.
As described in the “Results” section, four genes showed highly significant differential expression in blood and brain. Two of these genes had similar directionality (PER3 and SOX9) and two had opposing directions (CD19 and TERF1) between blood and brain (Supplementary Fig. 1). The variability in direction between the two tissues is understandable because blood and brain may not be exposed to the same metabolic environment or stress related factors, which could influence the expression levels for some genes. SOX9 is an astrocytic marker in the brain and B cell marker in blood that has previously been shown to be decreased in MDD-S compared to controls in the prefrontal cortex [47]. Our results show that SOX9 expression is significantly reduced both in blood and brain in MDD-S compared to MDD-NS suggesting similar immune/astrocytic dysregulations in suicide that could be further investigated.
We observed a significant effect of time of death in the DLPFC. Therefore, we corrected for time of death based on hours since sunrise (adjusted for daylight savings time). The adjustment of the time of death to hours from sunrise was done by counting the number of hours between sunrise and time of death taking into account daylight saving time. Despite the correction for multiple testing, the increase in the circadian gene, PER3, still remained. Dysregulated circadian rhythms in MDD involve significant disruptions in 24 h rhythms of mood, temperature, hormone secretion and sleep [45]. The circadian system is regulated by a set of core clock genes. The period genes are central to circadian timing and provide negative feedback to shut off their own transcription through a set of complex feedback loops. Mutations in the period gene, PER3, are implicated in sleep disorders associated with shifts in circadian rhythms and are thought to increase susceptibility to MDD [69]. In postmortem brain, we studied ~12,000 genes and observed a dramatic disruption in sinusoidal rhythms of core clock genes across six brain regions of MDDs compared to non-psychiatric controls [70], with PER3 as one of the top dysregulated genes. PER3 contains a glucocorticoid recognition element (GRE) on its promoter [71] and relevant to MDD, SNPs in the PER3 gene can alter multiple systems including response to antidepressants [72]. Additional evidence for the involvement of period genes with suicide has been found by Levy et al. [28], where they identified increased blood expression of PER1 to be related to suicidality in females.
Inflammation is a frequent feature of the pathophysiology of depression with markers of inflammation observed in blood [73,74,75,76] and brain [77]. However, it is not clear if inflammation is a mediatory mechanism of depression or suicide because other factors such as obesity and smoking also play a role [73]. Our results show that peripheral postmortem blood of MDD-S has significantly higher levels of two inflammatory markers compared to MDD-NS. The expression of CD19 and CD6 genes, markers for B and T cells respectively, are upregulated (Fig. 1 and Table 2). Similarly, expression of the glucocorticoid receptor is also increased in MDD-S compared to MDD-NS (Table 2), underlining stress/inflammation upregulation in suicides. Our results are indicative of a significant relationship between suicide and an overactive immune system in MDD patients leading to the hypothesis that inflammation is playing a role in the suicidal tendencies of depressed individuals. The DLPFC also showed a similar pattern of immune dysregulation and stress response alterations in suicides. Several polyamine (PAOX, ODC1, SAT1, and AGMAT) and metallothionein (MT2A and MT1A) genes were altered in MDD-S (Table 3) confirming previous reports [16, 17, 24,25,26,27, 78, 79]. Furthermore, key inflammation and stress response genes were similarly differentially expressed in the DLPFC of suicides including cortisol related genes (CRH, HSD11B2, and FKBP5) as well as CD19 and the astrocytic marker, SOX9 [80].
Another significant finding is the involvement of several mitochondrial genes in suicide. MAOA, a gene that encodes mitochondrial enzymes, was significantly lower in the DLPFC of MDD-S compared to MDD-NS. In blood, three genes were differentially expressed between MDD-S and MDD-NS. Two nuclear genes coding for mitochondria located proteins MTPAP (a mitochondrial Poly(A) polymerase) and the mitochondrial polyamine transporter SLC25A26 were increased in blood in suicides, while expression of ND6 (a mitochondria encoded subunit of the respiratory chain Complex I) was decreased in suicides. MTPAP and SLC25A26 were also significantly increased in MDD-S compared to controls suggesting that mitochondrial alterations could be used as potential signatures to differentiate MDD-S from MDD-NS patients and also from controls.
Our study results have some limitations. First, our sample has a higher proportion of males than females. To minimize the effects of gender bias, we balanced the genders between diagnostic groups. However, it is known that a greater number of women suffer from depression and future studies could benefit from increasing the number of females. Additionally, while this a small study, it clearly shows that this approach can be useful in identifying suicide (state) biomarkers that can be used in a clinical setting to identify MDD patients in a highly suicidal state for additional interventions. In the future, larger studies of SNP (trait) markers could be combined with investigations of gene expression suicide phenotype signatures (state) to provide a more comprehensive overview of the contribution of these factors to an increased risk for suicide.
We have performed a targeted gene expression investigation of 117 genes. Although there was substantial a priori reasons for gene selection, it is quite possible that additional suicide biomarkers may have been missed because a higher throughput technology, such as RNA-Seq, was not employed. This would be a next step in future studies to confirm our current findings and to identify additional suicide-related gene expression differences. Lastly, we have only analyzed one brain region. Although the DLPFC is implicated in MDD and suicide there may be other regions, which could provide additional gene expression information (e.g., the anterior cingulate or hippocampus), to help refine our blood suicide signature. In addition, validation of the identified blood-based suicide marker could benefit from testing in a clinical sample of MDDs with severe suicidal behaviors and ideation. Multiple testing correction using methods such as FDR is useful to determine the most likely results to be differentially expressed given the number of comparisons. Because this is an exploratory study, we present and discuss both the top significant genes and the adjusted q-values as potential biomarkers to be tested in future clinical studies. Validation would confirm the utility of the biomarker to identify at-risk individuals who would benefit from immediate interventions to prevent suicide.
Additional studies to compare the immune cell phenotype and function in the blood of depressed suicidal and depressed non-suicidal individuals are warranted. Activation of antigen presenting cells macrophages and dendritic cells (DCs), responsible for activation of T and B lymphocytes and the phenotype and expression of activation markers particularly CD6 and B lymphocytes could be analyzed in highly suicidal MDD patients to further investigate the role of immune dysregulation. The T activation cells can also be studied and secretion of cytokines determined by multiplexing. There has also been a link proposed between increased inflammation in MDD and decreased telomere length in depression, although this has not yet been extended to encompass suicide. The relationship between inflammation and telomere length in our current sample needs to be further explored and compared between blood and brain, particularly in view of the increased levels of TERF1 in blood from patients who died by suicide (Table 2). TERF1 is of particular interest because it plays a direct role in telomere length as part of a protein complex (shelterin) responsible for protecting telomeres [81].
Overall, brain and blood differentially expressed genes were observed between MDD-S and MDD-NS. We included comparisons, in brain and blood, with the Control group and both MDD groups (Control vs. MDD-NS and Control vs. MDD-S) as well as a comparison between Controls and both MDD groups regardless of the suicide status (Control vs. MDD (S + NS) to assess gene expression changes associated with depression (Supplementary Tables 6–11).
In conclusion, our results suggest that blood expression profiles could help to identify individuals at the highest risk of death by suicide and be highly informative to understand molecular changes in suicide. Our study design, which includes age and gender matched comparison groups, allowed for the identification of suicide specific expression changes either unique to or in common between DLPFC and blood. Specifically, stress response changes, including polyamine metabolism, circadian rhythm, immune dysregulation, and telomere maintenance are shared among suicides in blood suggesting future studies in large clinical populations investigating these systems might help in the identification of acutely suicidal MDD patients.
References
Suicide [Internet]. (2019). https://www.who.int/health-topics/suicide#tab=tab_1.
Hedegaard H, Curtin SC, Warner M. Increase in suicide mortality in the United States, 1999–2018. NCHS Data Brief. 2020;362:1–8.
Ahmedani BK, Simon GE, Stewart C, Beck A, Waitzfelder BE, Rossom R, et al. Health care contacts in the year before suicide death. J Gen Intern Med. 2014;29:870–7.
Luoma JB, Martin CE, Pearson JL. Contact with mental health and primary care providers before suicide: a review of the evidence. Am J Psychiatry 2002;159:909–16.
Forte A, Buscajoni A, Fiorillo A, Pompili M, Baldessarini RJ. Suicidal risk following hospital discharge: a review. Harv Rev Psychiatry. 2019;27:209–16.
Chung D, Hadzi-Pavlovic D, Wang M, Swaraj S, Olfson M, Large M. Meta-analysis of suicide rates in the first week and the first month after psychiatric hospitalisation. BMJ Open. 2019;9:e023883.
Large M, Sharma S, Cannon E, Ryan C, Nielssen O. Risk factors for suicide within a year of discharge from psychiatric hospital: a systematic meta-analysis. Aust N Z J Psychiatry 2011;45:619–28.
Oquendo MA, Sullivan GM, Sudol K, Baca-Garcia E, Stanley BH, Sublette ME, et al. Toward a biosignature for suicide. Am J Psychiatry. 2014;171:1259–77.
Yoshino Y, Dwivedi Y. Elevated expression of unfolded protein response genes in the prefrontal cortex of depressed subjects: effect of suicide. J Affect Disord. 2020;262:229–36.
Shinko Y, Otsuka I, Okazaki S, Horai T, Boku S, Takahashi M, et al. Chemokine alterations in the postmortem brains of suicide completers. J Psychiatr Res. 2020;120:29–33.
Zhao J, Verwer RWH, Gao SF, Qi XR, Lucassen PJ, Kessels HW, et al. Prefrontal alterations in GABAergic and glutamatergic gene expression in relation to depression and suicide. J Psychiatr Res. 2018;102:261–74. https://doi.org/10.1016/j.jpsychires.2018.04.020.
Chen GG, Almeida D, Fiori L, Turecki G. Evidence of reduced agmatine concentrations in the cerebral cortex of suicides. Int J Neuropsychopharmacol. 2018;21:895–900.
Yin H, Pantazatos SP, Galfalvy H, Huang YY, Rosoklija GB, Dwork AJ, et al. A pilot integrative genomics study of GABA and glutamate neurotransmitter systems in suicide, suicidal behavior, and major depressive disorder. Am J Med Genet B. 2016;171B:414–26.
Cabello-Arreola A, Ho AM, Ozerdem A, Cuellar-Barboza AB, Kucuker MU, Heppelmann CJ, et al. Differential dorsolateral prefrontal cortex proteomic profiles of suicide victims with mood disorders. Genes. 2020;11:256.
Sequeira A, Mamdani F, Ernst C, Vawter MP, Bunney WE, Lebel V, et al. Global brain gene expression analysis links glutamatergic and GABAergic alterations to suicide and major depression. PLoS ONE. 2009;4:e6585.
Pantazatos SP, Andrews SJ, Dunning-Broadbent J, Pang J, Huang YY, Arango V, et al. Isoform-level brain expression profiling of the spermidine/spermine N1-Acetyltransferase1 (SAT1) gene in major depression and suicide. Neurobiol Dis. 2015;79:123–34.
Sokolowski M, Ben-Efraim YJ, Wasserman J, Wasserman D. Glutamatergic GRIN2B and polyaminergic ODC1 genes in suicide attempts: associations and gene-environment interactions with childhood/adolescent physical assault. Mol Psychiatry. 2012;18:985–92.
Guipponi M, Deutsch S, Kohler K, Perroud N, Le GF, Vessaz M, et al. Genetic and epigenetic analysis of SSAT gene dysregulation in suicidal behavior. Am J Med Genet B. 2009;150B:799–807.
Fiori LM, Bureau A, Labbe A, Croteau J, Noel S, Merette C, et al. Global gene expression profiling of the polyamine system in suicide completers. Int J Neuropsychopharmacol. 2011;14:595–605.
Fiori LM, Mechawar N, Turecki G. Identification and characterization of spermidine/spermine N1-acetyltransferase promoter variants in suicide completers. Biol Psychiatry. 2009;66:460–7.
Fiori LM, Turecki G. Association of the SAT1 in/del polymorphism with suicide completion. Am J Med Genet B. 2010;153B:825–9.
Fiori LM, Wanner B, Jomphe V, Croteau J, Vitaro F, Tremblay RE, et al. Association of polyaminergic loci with anxiety, mood disorders, and attempted suicide. PLoS ONE 2010;5:e15146.
Klempan TA, Rujescu D, Merette C, Himmelman C, Sequeira A, Canetti L, et al. Profiling brain expression of the spermidine/spermine N1-acetyltransferase 1 (SAT1) gene in suicide. Am J Med Genet B. 2009;150B:934–43.
Sequeira A, Gwadry FG, Ffrench-Mullen JM, Canetti L, Gingras Y, Casero RA Jr., et al. Implication of SSAT by gene expression and genetic variation in suicide and major depression. Arch Gen Psychiatry. 2006;63:35–48.
Pantazatos SP, Huang YY, Rosoklija GB, Dwork AJ, Arango V, Mann JJ. Whole-transcriptome brain expression and exon-usage profiling in major depression and suicide: evidence for altered glial, endothelial and ATPase activity. Mol Psychiatry. 2017;22:760–73.
Niculescu AB, Levey DF, Phalen PL, Le-Niculescu H, Dainton HD, Jain N, et al. Understanding and predicting suicidality using a combined genomic and clinical risk assessment approach. Mol Psychiatry. 2015;20:1266–85.
Le-Niculescu H, Levey DF, Ayalew M, Palmer L, Gavrin LM, Jain N, et al. Discovery and validation of blood biomarkers for suicidality. Mol Psychiatry. 2013;18:1249–64.
Levey DF, Niculescu EM, Le-Niculescu H, Dainton HL, Phalen PL, Ladd TB, et al. Towards understanding and predicting suicidality in women: biomarkers and clinical risk assessment. Mol Psychiatry. 2016;21:768–85.
Kelly TM, Mann JJ. Validity of DSM-III-R diagnosis by psychological autopsy: a comparison with clinician ante-mortem diagnosis. Acta Psychiatr Scand. 1996;94:337–43.
Sequeira A, Morgan L, Walsh DM, Cartagena PM, Choudary P, Li J, et al. Gene expression changes in the prefrontal cortex, anterior cingulate cortex and nucleus accumbens of mood disorders subjects that committed suicide. PLoS ONE. 2012;7:e35367.
Geiss GK, Bumgarner RE, Birditt B, Dahl T, Dowidar N, Dunaway DL, et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol. 2008;26:317–25.
Govindpani K, Turner C, Waldvogel HJ, Faull RLM, Kwakowsky A. Impaired expression of GABA signaling components in the Alzheimer’s Disease middle temporal gyrus. Int J Mol Sci. 2020;21:8704.
Reis PP, Waldron L, Goswami RS, Xu W, Xuan Y, Perez-Ordonez B, et al. mRNA transcript quantification in archival samples using multiplexed, color-coded probes. BMC Biotechnol. 2011;11:46.
Benjamini Y, Drai D, Elmer G, Kafkafi N, Golani I. Controlling the false discovery rate in behavior genetics research. Behav Brain Res. 2001;125:279–84.
Niculescu AB, Le-Niculescu H, Levey DF, Phalen PL, Dainton HL, Roseberry K, et al. Precision medicine for suicidality: from universality to subtypes and personalization. Mol Psychiatry. 2017;22:1250–73.
Scheuermann RH, Racila E. CD19 antigen in leukemia and lymphoma diagnosis and immunotherapy. Leuk Lymphoma. 1995;18:385–97.
Wang K, Wei G, Liu D. CD19: a biomarker for B cell development, lymphoma diagnosis and therapy. Exp Hematol Oncol. 2012;1:36.
Maes M, Stevens WJ, DeClerck LS, Bridts CH, Peeters D, Schotte C, et al. A significantly increased number and percentage of B cells in depressed subjects: results of flow cytometric measurements. J Affect Disord. 1992;24:127–34.
Brown MH. CD6 as a cell surface receptor and as a target for regulating immune responses. Curr Drug Targets. 2016;17:619–29.
Santos RF, Oliveira L, Carmo AM. Tuning T cell activation: the function of CD6 at the immunological synapse and in T cell responses. Curr Drug Targets. 2016;17:630–9.
Kofler DM, Farkas A, von Bergwelt-Baildon M, Hafler DA. The link between CD6 and autoimmunity: genetic and cellular associations. Curr Drug Targets. 2016;17:651–65.
Krintel SB, Essioux L, Wool A, Johansen JS, Schreiber E, Zekharya T, et al. CD6 and syntaxin binding protein 6 variants and response to tumor necrosis factor alpha inhibitors in Danish patients with rheumatoid arthritis. PLoS ONE. 2012;7:e38539.
Fiori LM, Turecki G. Gene expression profiling of suicide completers. Eur Psychiatry. 2010;25:287–90.
Martinho A, Gonçalves I, Santos CR. Glucocorticoids regulate metallothionein-1/2 expression in rat choroid plexus: effects on apoptosis. Mol Cell Biochem. 2013;376:41–51.
Bunney BG, Li JZ, Walsh DM, Stein R, Vawter MP, Cartagena P, et al. Circadian dysregulation of clock genes: clues to rapid treatments in major depressive disorder. Mol Psychiatry. 2015;20:48–55.
Hagenauer MH, Schulmann A, Li JZ, Vawter MP, Walsh DM, Thompson RC, et al. Inference of cell type content from human brain transcriptomic datasets illuminates the effects of age, manner of death, dissection, and psychiatric diagnosis. PLoS ONE. 2018;13:e0200003.
Nagy C, Suderman M, Yang J, Szyf M, Mechawar N, Ernst C, et al. Astrocytic abnormalities and global DNA methylation patterns in depression and suicide. Mol Psychiatry. 2015;20:320–8.
Mamdani F, Rollins B, Morgan L, Myers RM, Barchas JD, Schatzberg AF, et al. Variable telomere length across post-mortem human brain regions and specific reduction in the hippocampus of major depressive disorder. Transl Psychiatry. 2015;5:e636.
Otsuka I, Izumi T, Boku S, Kimura A, Zhang Y, Mouri K, et al. Aberrant telomere length and mitochondrial DNA copy number in suicide completers. Sci Rep. 2017;7:3176.
Mullins N, Bigdeli TB, Børglum AD, Coleman JRI, Demontis D, Mehta D, et al. GWAS of suicide attempt in psychiatric disorders and association with major depression polygenic risk scores. Am J Psychiatry. 2019;176:651–60.
Levey DF, Polimanti R, Cheng Z, Zhou H, Nuñez YZ, Jain S, et al. Genetic associations with suicide attempt severity and genetic overlap with major depression. Transl Psychiatry. 2019;9:22.
González-Castro TB, Martínez-Magaña JJ, Tovilla-Zárate CA, Juárez-Rojop IE, Sarmiento E, Genis-Mendoza AD, et al. Gene-level genome-wide association analysis of suicide attempt, a preliminary study in a psychiatric Mexican population. Mol Genet Genom Med. 2019;7:e983.
Erlangsen A, Appadurai V, Wang Y, Turecki G, Mors O, Werge T, et al. Genetics of suicide attempts in individuals with and without mental disorders: a population-based genome-wide association study. Mol Psychiatry. 2018;25:2410–21.
Stein MB, Ware EB, Mitchell C, Chen CY, Borja S, Cai T, et al. Genomewide association studies of suicide attempts in US soldiers. Am J Med Genet Part B. 2017;174:786–97.
Willour VL, Seifuddin F, Mahon PB, Jancic D, Pirooznia M, Steele J, et al. A genome-wide association study of attempted suicide. Mol Psychiatry. 2012;17:433–44.
Kimbrel NA, Garrett ME, Dennis MF, Hauser MA, Ashley-Koch AE, Beckham JC. A genome-wide association study of suicide attempts and suicidal ideation in U.S. military veterans. Psychiatry Res. 2018;269:64–9.
Perlis RH, Huang J, Purcell S, Fava M, Rush AJ, Sullivan PF, et al. Genome-wide association study of suicide attempts in mood disorder patients. Am J Psychiatry. 2010;167:1499–507.
Strawbridge RJ, Ward J, Ferguson A, Graham N, Shaw RJ, Cullen B, et al. Identification of novel genome-wide associations for suicidality in UK Biobank, genetic correlation with psychiatric disorders and polygenic association with completed suicide. EBioMedicine. 2019;41:517–25.
Sokolowski M, Wasserman J, Wasserman D. Polygenic associations of neurodevelopmental genes in suicide attempt. Mol Psychiatry. 2016;21:1381–90.
Sokolowski M, Wasserman J, Wasserman D. Gene-level associations in suicide attempter families show overrepresentation of synaptic genes and genes differentially expressed in brain development. Am J Med Genet Part B. 2018;177:774–84.
Otsuka I, Akiyama M, Shirakawa O, Okazaki S, Momozawa Y, Kamatani Y, et al. Genome-wide association studies identify polygenic effects for completed suicide in the Japanese population. Neuropsychopharmacology. 2019;44:2119–24.
Docherty AR, Shabalin AA, DiBlasi E, Monson E, Mullins N, Adkins DE, et al. Genome-wide association study of suicide death and polygenic prediction of clinical antecedents. Am J Psychiatry. 2020;177:917–27.
Coon H, Darlington TM, DiBlasi E, Callor WB, Ferris E, Fraser A, et al. Genome-wide significant regions in 43 Utah high-risk families implicate multiple genes involved in risk for completed suicide. Mol Psychiatry. 2020;25:3077–90.
Nagy C, Maitra M, Tanti A, Suderman M, Théroux JF, Davoli MA, et al. Single-nucleus transcriptomics of the prefrontal cortex in major depressive disorder implicates oligodendrocyte precursor cells and excitatory neurons. Nat Neurosci. 2020;23:771–81.
Mahajan GJ, Vallender EJ, Garrett MR, Challagundla L, Overholser JC, Jurjus G, et al. Altered neuro-inflammatory gene expression in hippocampus in major depressive disorder. Prog Neuro-Psychopharmacol Biol Psychiatry. 2018;82:177–86.
Jabbi M, Arasappan D, Eickhoff SB, Strakowski SM, Nemeroff CB, Hofmann HA. Neuro-transcriptomic signatures for mood disorder morbidity and suicide mortality. J Psychiatr Res. 2020;127:62–74.
Sullivan PF, Fan C, Perou CM. Evaluating the comparability of gene expression in blood and brain. Am J Med Genet B. 2006;141B:261–8.
Rollins B, Martin MV, Morgan L, Vawter MP. Analysis of whole genome biomarker expression in blood and brain. Am J Med Genet B. 2010;153B:919–36.
Artioli P, Lorenzi C, Pirovano A, Serretti A, Benedetti F, Catalano M, et al. How do genes exert their role? Period 3 gene variants and possible influences on mood disorder phenotypes. Eur Neuropsychopharmacol. 2007;17:587–94.
Li JZ, Bunney BG, Meng F, Hagenauer MH, Walsh DM, Vawter MP, et al. Circadian patterns of gene expression in the human brain and disruption in major depressive disorder. Proc Natl Acad Sci USA. 2013;110:9950–5.
Archer SN, Carpen JD, Gibson M, Lim GH, Johnston JD, Skene DJ, et al. Polymorphism in the PER3 promoter associates with diurnal preference and delayed sleep phase disorder. Sleep. 2010;33:695–701.
Xu Y, Ma H, Zhao T, Wen D, Wen Y, Qiao D, et al. Association between period 3 gene polymorphisms and adverse effects of antidepressants for major depressive disorder. Genet Test Mol Biomark. 2019;23:843–9.
Fried EI, von Stockert S, Haslbeck JMB, Lamers F, Schoevers RA, Penninx B. Using network analysis to examine links between individual depressive symptoms, inflammatory markers, and covariates. Psychol Med. 2019;50:1–9.
Haapakoski R, Ebmeier KP, Alenius H, Kivimäki M. Innate and adaptive immunity in the development of depression: an update on current knowledge and technological advances. Prog Neuro-Psychopharmacol Biol Psychiatry. 2016;66:63–72.
Haapakoski R, Mathieu J, Ebmeier KP, Alenius H, Kivimäki M. Cumulative meta-analysis of interleukins 6 and 1β, tumour necrosis factor α and C-reactive protein in patients with major depressive disorder. Brain Behav Immun. 2015;49:206–15.
Lamers F, Milaneschi Y, de Jonge P, Giltay EJ, Penninx B. Metabolic and inflammatory markers: associations with individual depressive symptoms. Psychol Med. 2018;48:1102–10.
Holmes SE, Hinz R, Conen S, Gregory CJ, Matthews JC, Anton-Rodriguez JM, et al. Elevated translocator protein in anterior cingulate in major depression and a role for inflammation in suicidal thinking: a positron emission tomography study. Biol Psychiatry. 2018;83:61–9.
Desautels A, Turecki G, Montplaisir J, Brisebois K, Sequeira A, Adam B, et al. Evidence for a genetic association between monoamine oxidase A and restless legs syndrome. Neurology. 2002;59:215–9.
Limon A, Mamdani F, Hjelm BE, Vawter MP, Sequeira A. Targets of polyamine dysregulation in major depression and suicide: activity-dependent feedback, excitability, and neurotransmission. Neurosci Biobehav Rev. 2016;66:80–91.
Sun W, Cornwell A, Li J, Peng S, Osorio MJ, Aalling N, et al. SOX9 is an astrocyte-specific nuclear marker in the adult brain outside the neurogenic regions. J Neurosci. 2017;37:4493–507.
Schmutz I, de Lange T. Shelterin. Curr Biol. 2016;26:R397–9.
Acknowledgements
This work was supported by NIMH research grant R01MH097082 (to AS), American Foundation for Suicide Prevention Standard Research Grant SRG-0-121-18 (to AS), and Hope for Depression Research Foundation (HDRF) (to HA). The collection of brain tissue was supported by funding from the Pritzker Family Philanthropic Fund. The authors are members of the Pritzker Neuropsychiatric Disorders Research Consortium, which is supported by the Pritzker Neuropsychiatric Disorders Research Fund L.L.C. A shared intellectual property agreement exists between this philanthropic fund and the University of Michigan, Stanford University, the Weill Medical College of Cornell University, Hudson Alpha Institute of Biotechnology, and the University of California at Irvine, to encourage the development of appropriate findings for research and clinical applications. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. We would also like to thank the Orange County Coroner Division personnel and the families without whom this research project would not be possible.
Author information
Authors and Affiliations
Contributions
AS and FM conceived and designed the experiments. BB, KB, PC, DW, and WEB were involved in collection of post-mortem brain and blood, and performance of psychological autopsies. AS, FM, and MDW carried out brain dissections and experiments. AS and FM analyzed the data and wrote the manuscript. MDW, BB, MPV, WEB, FSL, JB, AFS, RMM, SJW, and HA contributed to manuscript preparation, reviewing, and editing. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mamdani, F., Weber, M.D., Bunney, B. et al. Identification of potential blood biomarkers associated with suicide in major depressive disorder. Transl Psychiatry 12, 159 (2022). https://doi.org/10.1038/s41398-022-01918-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-022-01918-w
This article is cited by
-
Next-generation precision medicine for suicidality prevention
Translational Psychiatry (2024)
-
Elevated SCN11A concentrations associated with lower serum lipid levels in patients with major depressive disorder
Translational Psychiatry (2024)
-
Identification of Potential Biomarkers for Major Depressive Disorder: Based on Integrated Bioinformatics and Clinical Validation
Molecular Neurobiology (2024)
-
Blood-based biomarkers for suicide in depression: moving a step closer to prediction
Neuropsychopharmacology (2023)
-
MANF/EWSR1/ANXA6 pathway might as the bridge between hypolipidemia and major depressive disorder
Translational Psychiatry (2022)