Abstract
Previous genome-wide association studies (GWAS) have identified potential genetic variants associated with the risk of major depressive disorder (MDD), but the underlying biological interpretation remains largely unknown. We aimed to prioritize genes that were pleiotropically or potentially causally associated with MDD. We applied the summary data-based Mendelian randomization (SMR) method integrating GWAS and gene expression quantitative trait loci (eQTL) data in 13 brain regions to identify genes that were pleiotropically associated with MDD. In addition, we repeated the analysis by using the meta-analyzed version of the eQTL summary data in the brain (brain-eMeta). We identified multiple significant genes across different brain regions that may be involved in the pathogenesis of MDD. The prime-specific gene BTN3A2 (corresponding probe: ENSG00000186470.9) was the top hit showing pleiotropic association with MDD in 9 of the 13 brain regions and in brain-eMeta, after correction for multiple testing. Many of the identified genes are located in the human major histocompatibility complex (MHC) region on chromosome 6 and are mainly involved in the immune response. Our SMR analysis indicated that multiple genes showed pleiotropic association with MDD across the brain regions. These findings provided important leads to a better understanding of the mechanism of MDD and revealed potential therapeutic targets for the prevention and effective treatment of MDD.
Similar content being viewed by others
Introduction
Major depressive disorder (MDD) is a significant medical condition impacting an individual’s mood, behavior, appetite, and sleep as well as thoughts of suicide1. MDD is a leading cause of disability and morbidity worldwide2, with an estimated lifetime prevalence of around 15%3. MDD is a complex multifactorial disorder, with contributions from both genetic and environmental factors4. However, the exact etiology of MDD remains to be unclear, and there is pressing urgency to further explore the pathological mechanisms underlying MDD to facilitate the design and implementation of efficient prevention strategies or novel treatments.
Previous twin studies found the heritability of MDD to be ~30–40%5,6. Genome-wide association studies (GWASs) have been successful in identifying genetic variants associated with MDD7,8,9,10,11, such as single-nucleotide polymorphisms (SNPs) in/near sortilin-related VPS10-domain-containing receptor 3 (SORCS3), transcription factor 4 (TCF4), and neuronal growth regulator 1 (NEGR1). However, the biological interpretation of the identified genetic variants remains largely unclear. Because many of the genetic variants identified in GWASs are located in non-coding regions, it is likely that these genetic variants exert their effects on diseases/disorders via gene expression12. Therefore, it is important to explore the relationship between genetic variation and gene expression to better understand the regulatory pathways underlying the pathogenesis of MDD.
Mendelian randomization (MR) is a method for exploring the potential causal association between an exposure and an outcome by using genetic variants as the instrumental variables (IVs) for exposure13. Compared with traditional statistical methods used in the association studies, MR reduces confounding and reverse causation and is becoming increasingly popular in the exploration of etiological mechanisms14,15. A novel analytical framework through a summary data-based MR (SMR) approach integrating cis-expression quantitative trait loci (cis-eQTL) or cis-DNA methylation QTL (cis-mQTL) and GWAS data have been successful in identifying gene expressions or DNA methylation loci that are pleiotropically or potentially causally associated with various phenotypes, such as cardiovascular diseases, systemic lupus erythematosus, inflammatory bowel disease, and educational attainment16,17,18,19,20, indicating that it is a promising tool to explore genes pleiotropically associated with complex traits.
Previous research that adopted the SMR approach to examine possible causal genes for MDD found three candidate genes (NEGR1, BAG6, and HLG-B). The GWAS data used in the study were based on 42,455 subjects (16,823 MDD cases and 25,632 controls), and the cis-eQTL data were based on a meta-analyzed version of the eQTL summary data instead of specific brain regions21. In this study, we adopted the SMR approach by leveraging GWAS summarized results for MDD which were based on a much larger sample size (807,553 subjects), and cis-eQTL data in 13 different brain regions to prioritize genes that are pleiotropically or potentially causally associated with MDD across different brain regions.
Materials and methods
In the SMR analysis, cis-eQTL genetic variants were used as the IVs for gene expression. We performed SMR analysis for different regions in the brain. We used the Version 7 release of the eQTL summarized data from the Genotype Tissue Expression (GTEx)22 project, which included 13 different regions: amygdala, anterior cingulate cortex, caudate nucleus, cerebellar hemisphere, cerebellum, cortex, frontal cortex, hippocampus, hypothalamus, nucleus accumbens, putamen, spinal cord, and substantia nigra22. In addition, we repeated the analysis by using the meta-analyzed version of the eQTL summary data (named brain-eMeta hereafter), which included results from the GTEx data of brain tissues22, the Common Mind Consortium23, and the Religious Orders Study and the Rush Memory and Aging Project24. Results from these three studies were meta-analyzed using the MeCS method (meta-analysis of cis-eQTL in correlated samples) to increase the power of detecting brain eQTLs25. Only SNPs within 1 Mb distance from each individual probe are available. The eQTL data can be downloaded at https://cnsgenomics.com/data/SMR/#eQTLsummarydata.
The GWAS summarized data for MDD were provided by the Psychiatric Genomics Consortium10. The results were based on three large genome-wide association studies8,9,11, including a total of 807,553 individuals (246,363 cases and 561,190 controls, after excluding overlapping samples) and 8,098,588 genetic variants. The GWAS summarized data can be downloaded at https://www.med.unc.edu/pgc/download-results/mdd/.
MR was carried out considering cis-eQTL genetic variants as the IVs, gene expression as the exposure, and MDD as the outcome. MR analysis was performed using the method as implemented in the software SMR. Detailed information regarding the SMR method has been described previously16. Briefly, SMR uses the principles of MR integrating GWAS and eQTL summary statistics to test for pleiotropic association between gene expression and MDD due to a shared and potentially causal variant at a locus. The heterogeneity in dependent instruments (HEIDI) test was done to explore the existence of linkage in the observed association. Rejection of the null hypothesis (i.e., PHEIDI < 0.05) indicates that the observed association might be due to two distinct genetic variants in high linkage disequilibrium with each other. We adopted the default settings in SMR (e.g., PeQTL < 5 × 10−8, minor allele frequency [MAF] > 0.01, excluding SNPs in very strong linkage disequilibrium [LD, r2 > 0.9] with the top associated eQTL, and removing SNPs in low LD or not in LD [r2 < 0.05] with the top associated eQTL), and used false discovery rate (FDR) to adjust for multiple testing.
Annotations of the transcripts were based on the Affymetrix exon array S1.0 platforms. To functionally annotate putative transcripts, we conducted functional enrichment analysis using the functional annotation tool “Metascape” for the significant genes in different brain regions and in brain-eMeta. Gene symbols corresponding to putative genes (FDR P < 0.05) were used as the input of the gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis.
Data cleaning and statistical/bioinformatical analysis were performed using R version 4.0.2 (https://www.r-project.org/), PLINK 1.9 (https://www.cog-genomics.org/plink/1.9/) and SMR (https://cnsgenomics.com/software/smr/).
Results
The number of participants used for generating the eQTL data varied across the brain regions, ranging from 114 to 209, so did the number of eligible probes involved in the final SMR analysis, ranging from 661 to 3765. The brain-eMeta analysis involved more subjects (n = 1194) and more probes (n = 7421). The GWAS meta-analysis data involved roughly 800,000 subjects. The detailed information was shown in Table 1.
We identified multiple genes showing pleiotropic association with MDD across the different brain regions (Table 2 and Supplementary Fig. S1). Out of the 13 brain regions, the human major histocompatibility complex (MHC) gene BTN3A2 (ENSG00000186470.9) was the top hit showing pleiotropic association with MDD in 9 regions, after correction for multiple testing. Each of the other two genes, RPL31P12 (ENSG00000227207.2) and RP1-265C24.5 (ENSG00000219392.1) was the top gene pleiotropically associated with MDD in two brain regions (Table 2).
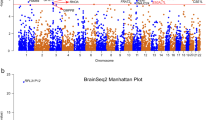
Specifically, for BTN3A2, the most significantly pleiotropic associations with MDD were detected in two brain regions: caudate nucleus and spinal cord (β [SE] = 0.043 [0.008], P = 7.76 × 10−8; β [SE] = 0.042 [0.008], P = 1.72 × 10−7, respectively; Fig. 1). It also showed a significantly pleiotropic association with MDD in the four brain regions where it was not the top gene (Supplementary Table S1). RPL31P12 showed the most significantly pleiotropic association with MDD in cerebellar hemisphere and cerebellum (β [SE] = −0.037 [0.006], P = 7.53 × 10−11; β [SE] = −0.033 [0.005], P = 1.34 × 10−12, respectively; Fig. 2). RP1-265C24.5 showed significantly pleiotropic association in cortex and nucleus accumbens (β [SE] = 0.036 [0.007], P = 6.13 × 10−8; β [SE] = 0.036 [0.006], P = 1.63 × 10−8, respectively; Fig. 3).
A Caudate nucleus. B Spinal cord. Top plot, gray dots represent the −log10(P values) for SNPs from the GWAS of MDD, and rhombuses represent the −log10(P values) for probes from the SMR test with solid rhombuses indicating that the probes pass HEIDI test and hollow rhombuses indicating that the probes do not pass the HEIDI test. Middle plot, eQTL results for the probe ENSG000001864770.9 tagging BTN3A2. Bottom plot, location of genes tagged by the probe. Highlighted in maroon indicates probes that pass the SMR threshold. GWAS genome-wide association study, MDD major depressive disorder, SMR summary data-based Mendelian randomization, HEIDI heterogeneity in dependent instruments, eQTL expression quantitative trait loci.
A Cerebellar hemisphere. B Cerebellum. Top plot, gray dots represent the −log10(P values) for SNPs from the GWAS of MDD, and rhombuses represent the −log10(P values) for probes from the SMR test with solid rhombuses indicating that the probes pass HEIDI test and hollow rhombuses indicating that the probes do not pass the HEIDI test. Middle plot, eQTL results for the probe ENSG00000227207.2 tagging RPL31P12. Bottom plot, location of genes tagged by the probe. Highlighted in maroon indicates probes that pass the SMR threshold. GWAS genome-wide association studies, MDD major depressive disorder, SMR summary data-based Mendelian randomization, HEIDI heterogeneity in dependent instruments, eQTL expression quantitative trait loci.
A Cortex. B Nucleus accumbens. Top plot, gray dots represent the −log10(P values) for SNPs from the GWAS of MDD, and rhombuses represent the −log10(P values) for probes from the SMR test with solid rhombuses indicating that the probes pass HEIDI test and hollow rhombuses indicating that the probes do not pass the HEIDI test. Middle plot, eQTL results for the probe ENSG00000219392.1 tagging RP1-265C24.5. Bottom plot, location of genes tagged by the probe. Highlighted in maroon indicates probes that pass the SMR threshold. GWAS genome-wide association studies, MDD major depressive disorder, SMR summary data-based Mendelian randomization, HEIDI heterogeneity in dependent instruments, eQTL expression quantitative trait loci.
The complement gene C4A (ENSG00000244731.3) was significantly associated with MDD in 7 different brain regions, after correction for multiple testing (Supplementary Table S1). Of note, both BTN3A2, C4A and RP1-265C24.5 are on chromosome 6 while RPL31P12 is on chromosome 1. Two brain regions, the cerebellar hemisphere and cerebellum, have a relatively large number of significant genes (21 genes and 30 genes, respectively; Table 2).
GO enrichment analysis of biological process and molecular function showed that the significant genes across the different brain regions were involved in four GO terms, including negative regulation of endopeptidase activity (GO:0010951), adaptive immune response (GO:0002250), platelet degranulation (GO:0002576), and negative regulation of defense response (GO:0031348; Supplementary Fig. S2A). Concept network analysis of the identified genes revealed multiple domains related to immune response (Supplementary Fig. S2B). More information could be found in Supplementary Table S2.
Using brain-eMeta eQTL data, we found 75 genes that showed pleiotropic association with MDD, after correction for multiple testing. Specifically, we identified BTN3A2 (ENSG00000186470) that showed the most significantly pleiotropic association with MDD (β [SE] = 0.027 [0.004], P = 3.44 × 10−12; Supplementary Table S3), followed by RPL31P12 (ENSG00000227207, β [SE] = −0.039 [0.006], P = 3.43 × 10−11). We found that C4A and RP1-265C24.5 also showed significant pleiotropic association with MDD (β [SE] = 0.031 [0.005], P = 1.58 × 10−8 and β [SE] = 0.047 [0.008], P = 2.11 × 10−9, respectively).
GO enrichment analysis of biological process and molecular function showed that the significant genes in brain-eMeta were involved in eight GO terms, including allograft rejection (ko05330), butyrophilin (BTN) family interactions (R-HSA-8851680), platelet degranulation (GO:0002576), immunoregulatory interactions between a lymphoid and a non-lymphoid cell (R-HSA-198933), nuclear chromosome segregation (GO:0098813), telomere maintenance (GO:0000723), organelle localization by membrane tethering (GO:0140056) and lipid transport (GO:0006869; Supplementary Fig. S3A). Similar to the findings for the different brain regions, concept network analysis in brain-eMeta also revealed multiple domains related to immune response (Supplementary Fig. S3B). More information could be found in Supplementary Table S4.
Discussion
In this study, we integrated GWAS and eQTL data in the MR analysis to explore putative genes that showed pleiotropic/potentially causal association with MDD. Across the different brain regions, we identified multiple significant genes that may be involved in the pathogenesis of MDD. The identified genes were mainly involved in the immune response. Our findings provided important leads to a better understanding of the mechanisms underlying MDD and revealed potential therapeutic targets for the effective treatment of MDD.
Compared to a previous study that adopted a similar SMR approach by integrating GWAS results for MDD and a meta-analyzed version of the eQTL summary data (brain eMeta) to explore causal genes for MDD, our study used the GWAS summary data which were based on a much larger sample size (807,553 vs. 42,455), and we explored the potential pleiotropic association across 13 brain regions21. Using brain eMeta, we not only confirmed the significantly pleiotropic association of the three candidate genes (NEGR1, BAG6, and HLG-B) as reported by the study but also identified many other genes (Supplementary Table S3). Moreover, we found many genes showing pleiotropic association with MDD across different brain regions (Table 2 and Supplementary Table S1), representing putative novels genes underlying the pathogenesis of MDD. Our findings suggested that the etiology of MDD involved different genes across different brain regions.
Several of the identified genes in our study, such as BTN3A2, BTN3A3, PRSS16, HLA-C, C4A, and HLA-DMA, are located in or around the human major histocompatibility complex (MHC) region on chromosome 6. MHC represents the most complex genomic region due to its unintelligible linkage disequilibrium26. Many genes in or around MHC play an important role in immune response and immune regulation and are involved in a variety of inflammatory and autoimmune diseases27,28,29,30,31. The MHC regions can be roughly divided into three classes that are functionally distinct, with class I and II regions containing highly polymorphic human leukocyte antigen (HLA) genes associated with autoimmune disease risk32,33 and class III region containing complement component 4 regions associated with schizophrenia risk34. Recent GWASs identified a number of genetic variants in the MHC region associated with depression risk, with the strongest association observed in or near the class I region9,10,11.
We found that BTN3A2 was significantly associated with MDD across many brain regions. BTN3A2, which encodes a member of the immunoglobulin superfamily, resides in the juxta-telomeric region (class I) of MHC35. The BTN3A2 protein may be involved in adaptive immune response36. Previous studies showed that BTN3A2 was a potential risk gene for Alzheimer’s disease, schizophrenia, and intellectual disability37,38,39. A meta-analysis of GWAS found that BTN3A2 was associated with neuroticism40, an important risk factor for MDD41. Overexpression of BTN3A2 suppressed the excitatory synaptic activity onto CA1 pyramidal neurons, most likely through the interaction with the presynaptic adhesion molecule neurexins37,42. Previous research showed that BTN3A2 was expressed in multiple cell types in the brain, including astrocyte, neuron, oligodendrocyte, and microglia43. These findings, together with ours, demonstrated the important role of BTN3A2 in the nervous system and highlighted the potential of this gene as a promising target for the prevention and treatment of MDD.
A previous GWAS of MDD highlighted the importance of the prefrontal brain regions10. In the prefrontal cortex, we found four significant genes, including BTN3A2, C4A, RP1-265C24.5, and CYP21A1P, that were associated with MDD after correction for multiple testing. The gene C4A was significant in seven brain regions and in the analysis using brain-eMeta. C4A localizes to the MHC class III region and encodes the acidic form of complement factor 4. In the mouse brain, C4A gene is mainly expressed in astrocytes and neurons44. C4A is involved in the classical complement activation pathway45 and was reported to be associated with schizophrenia, aging, and Alzheimer’s disease34,46,47. Moreover, genetic variants in BTN3A2 and C4A were in different LD blocks, suggesting that both genes might be independent risk factors for mental disorders such as schizophrenia and MDD37.
Both MDD and schizophrenia are mental illnesses contributing substantially to the global disease burden. It was reported that depressed patients had a higher risk of developing psychosis. Moreover, even prior to the emergence of psychotic symptoms, patients with a high risk of schizophrenia had a higher risk for developing depressive symptoms48. In consistent with previous findings49, some of the identified genes showing pleiotropic association with MDD were also associated with schizophrenia, such as BTN3A2, BTN3A3, PRSS16, HLA-C, C4A, and HLA-DMA, indicating a potential overlapped mechanism between schizophrenia and MDD.
Our study has some limitations. The number of probes used in our SMR analysis was limited for some brain regions (Table 1), and we may have missed some important genes. The HEIDI test was significant for some of the identified genes, indicating the possibility of horizontal pleiotropy (Supplementary Tables S1 and S3), i.e., the identified association might be due to two distinct genetic variants in high linkage disequilibrium with each other. In addition, we only included study participants of European ethnicity, and our findings might not be generalized to other ethnicities. More studies are needed to validate our findings in independent populations. We adopted correction for multiple testing to reduce the false-positive rate; however, we may have missed important genes. Due to a lack of individual eQTL data, we could not quantify the changes in gene expression in subjects with MDD in comparison with the control.
In conclusion, our SMR analysis revealed that multiple genes showed pleiotropic association with MDD across the brain regions. More studies are needed to explore the underlying physiological mechanisms in the etiology of MDD.
Code availability
The R and shell scripts used for the analyses are available from the corresponding authors on reasonable request.
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary Information files.
References
Smith, K. Mental health: a world of depression. Nature 515, 181 (2014).
GBD 2017 Disease and Injury Incidence and Prevalence Collaborators. Global, regional, and national incidence, prevalence, and years lived with disability for 354 diseases and injuries for 195 countries and territories, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 392, 1789–1858 (2018).
Kessler, R. C. et al. Lifetime prevalence and age-of-onset distributions of DSM-IV disorders in the National Comorbidity Survey Replication. Arch. Gen. Psychiatry 62, 593–602 (2005).
Otte, C. et al. Major depressive disorder. Nat. Rev. Dis. Prim. 2, 16065 (2016).
Flint, J. & Kendler, K. S. The genetics of major depression. Neuron 81, 484–503 (2014).
Sullivan, P. F., Neale, M. C. & Kendler, K. S. Genetic epidemiology of major depression: review and meta-analysis. Am. J. Psychiatry 157, 1552–1562 (2000).
Lewis, C. M. et al. Genome-wide association study of major recurrent depression in the U.K. population. Am. J. Psychiatry 167, 949–957 (2010).
Hyde, C. L. et al. Identification of 15 genetic loci associated with risk of major depression in individuals of European descent. Nat. Genet. 48, 1031–1036 (2016).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Howard, D. M. et al. Genome-wide association study of depression phenotypes in UK Biobank identifies variants in excitatory synaptic pathways. Nat. Commun. 9, 1470 (2018).
Visscher, P., Brown, M., McCarthy, M. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Davey Smith, G. & Hemani, G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet. 23, R89–R98 (2014).
Thanassoulis, G. & O’Donnell, C. J. Mendelian randomization: nature’s randomized trial in the post-genome era. JAMA 301, 2386–2388 (2009).
Burgess, S., Timpson, N. J., Ebrahim, S. & Davey Smith, G. Mendelian randomization: where are we now and where are we going? Int J. Epidemiol. 44, 379–388 (2015).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Pavlides, J. M. et al. Predicting gene targets from integrative analyses of summary data from GWAS and eQTL studies for 28 human complex traits. Genome Med. 8, 84 (2016).
Porcu, E. et al. Mendelian randomization integrating GWAS and eQTL data reveals genetic determinants of complex and clinical traits. Nat. Commun. 10, 3300 (2019).
Mo, X. et al. Mendelian randomization analysis revealed potential causal factors for systemic lupus erythematosus. Immunology 159, 279–288 (2020).
Wu, Y. et al. Integrative analysis of omics summary data reveals putative mechanisms underlying complex traits. Nat. Commun. 9, 918 (2018).
Wang, X. et al. Integrating genome-wide association study and expression quantitative trait loci data identifies NEGR1 as a causal risk gene of major depression disorder. J. Affect. Disord. 265, 679–686 (2020).
Aguet, F. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Ng, B. et al. An xQTL map integrates the genetic architecture of the human brain’s transcriptome and epigenome. Nat. Neurosci. 20, 1418–1426 (2017).
Qi, T. et al. Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood. Nat. Commun. 9, 2282 (2018).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Lang, M. L. How do natural killer T cells help B cells? Expert Rev. Vaccines 8, 1109–1121 (2009).
Neely, J. et al. Gene expression meta-analysis reveals concordance in gene activation, pathway, and cell-type enrichment in dermatomyositis target tissues. ACR Open Rheumatol. 1, 657–666 (2019).
Whitehead, A. S., Colten, H. R., Chang, C. C. & Demars, R. Localization of the human MHC-linked complement genes between HLA-B and HLA-DR by using HLA mutant cell lines. J. Immunol. 134, 641–643 (1985).
Zakharova, M. Y., Belyanina, T. A., Sokolov, A. V., Kiselev, I. S. & Mamedov, A. E. The contribution of major histocompatibility complex class II genes to an association with autoimmune diseases. Acta Nat. 11, 4–12 (2019).
Horton, R. et al. Gene map of the extended human MHC. Nat. Rev. Genet. 5, 889–899 (2004).
Dendrou, C. A., Petersen, J., Rossjohn, J. & Fugger, L. HLA variation and disease. Nat. Rev. Immunol. 18, 325–339 (2018).
Gough, S. C. & Simmonds, M. J. The HLA region and autoimmune disease: associations and mechanisms of action. Curr. genomics 8, 453–465 (2007).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Afrache, H., Pontarotti, P., Abi-Rached, L. & Olive, D. Evolutionary and polymorphism analyses reveal the central role of BTN3A2 in the concerted evolution of the BTN3 gene family. Immunogenetics 69, 379–390 (2017).
Rhodes, D. A., Stammers, M., Malcherek, G., Beck, S. & Trowsdale, J. The cluster of BTN genes in the extended major histocompatibility complex. Genomics 71, 351–362 (2001).
Wu, Y. et al. Identification of the primate-specific gene BTN3A2 as an additional schizophrenia risk gene in the MHC loci. EBioMedicine 44, 530–541 (2019).
Anazi, S. et al. Clinical genomics expands the morbid genome of intellectual disability and offers a high diagnostic yield. Mol. Psychiatry 22, 615–624 (2017).
Shi, J. et al. Common variants on chromosome 6p22.1 are associated with schizophrenia. Nature 460, 753–757 (2009).
Nagel, M. et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat. Genet. 50, 920–927 (2018).
Speed, D., Hemani, G., Speed, M. S., Børglum, A. D. & Østergaard, S. D. Investigating the causal relationship between neuroticism and depression via Mendelian randomization. Acta Psychiatr. Scand. 139, 395–397 (2019).
Südhof, T. C. Synaptic neurexin complexes: a molecular code for the logic of neural circuits. Cell 171, 745–769 (2017).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Chung, E. K. et al. Determining the one, two, three, or four long and short loci of human complement C4 in a major histocompatibility complex haplotype encoding C4A or C4B proteins. Am. J. Hum. Genet. 71, 810–822 (2002).
Bennett, S. et al. Plasma levels of complement 4a protein are increased in Alzheimer’s disease. Alzheimer Dis. Assoc. Disord. 26, 329–334 (2012).
Cao, W. et al. Modelling biological age based on plasma peptides in Han Chinese adults. Aging 12, 10676–10686 (2020).
Samsom, J. N. & Wong, A. H. Schizophrenia and depression co-morbidity: what we have learned from animal models. Front. psychiatry 6, 13 (2015).
O’Brien, H. E. et al. Expression quantitative trait loci in the developing human brain and their enrichment in neuropsychiatric disorders. Genome Biol. 19, 194 (2018).
Acknowledgements
The study was supported by NIH/NIA grants P30AG10161, R01AG15819, R01AG17917, R01AG36042, U01AG61356, and 1RF1AG064312-01. Huiquan Jing’s research was supported by the National Key Research and Development Program of China (2018YFC2000400). D.L. was supported by China Scholarship Council (CSC 201908110339). We confirmed that all authors have reviewed the contents of the article being submitted, approved its contents, and validated the accuracy of the data.
Author information
Authors and Affiliations
Contributions
H.J. and J.Y. designed the study. H.Y., B.F., and W.L. analyzed data and performed data interpretation. D.L., C.Z., and J.Y. wrote the initial draft, and B.F., W.L., X.Y., M.X., W.Z., and H.J. contributed to writing the subsequent versions of the manuscript. All authors reviewed the study findings and read and approved the final version before submission.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, H., Liu, D., Zhao, C. et al. Mendelian randomization integrating GWAS and eQTL data revealed genes pleiotropically associated with major depressive disorder. Transl Psychiatry 11, 225 (2021). https://doi.org/10.1038/s41398-021-01348-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-021-01348-0
This article is cited by
-
New Insights on the Therapeutic Potential of Runt-Related Transcription Factor 2 for Osteoarthritis: Evidence from Mendelian Randomization
Rheumatology and Therapy (2024)
-
Dihydroergotamine and Bromocriptine: Potential Drugs for the Treatment of Major Depressive Disorder and Alzheimer’s Disease Comorbidity
Molecular Neurobiology (2024)
-
Integrating genetics and transcriptomics to study major depressive disorder: a conceptual framework, bioinformatic approaches, and recent findings
Translational Psychiatry (2023)
-
CADM2 is implicated in impulsive personality and numerous other traits by genome- and phenome-wide association studies in humans and mice
Translational Psychiatry (2023)
-
Mendelian Randomization Analysis Identified Potential Genes Pleiotropically Associated with Polycystic Ovary Syndrome
Reproductive Sciences (2022)