Abstract
Psychiatric and neurological disorders (PNDs) affect millions worldwide and only a few drugs achieve complete therapeutic success in the treatment of these disorders. Due to the high cost of developing novel drugs, drug repositioning represents a promising alternative method of treatment. In this manuscript, we used a network medicine approach to investigate the molecular characteristics of PNDs and identify novel drug candidates for repositioning. Using IBM Watson for Drug Discovery, a powerful machine learning text-mining application, we built knowledge networks containing connections between PNDs and genes or drugs mentioned in the scientific literature published in the past 50 years. This approach revealed several drugs that target key PND-related genes, which have never been used to treat these disorders to date. We validate our framework by detecting drugs that have been undergoing clinical trial for treating some of the PNDs, but have no published results in their support. Our data provides comprehensive insights into the molecular pathology of PNDs and offers promising drug repositioning candidates for follow-up trials.
Similar content being viewed by others
Introduction
Psychiatric and neurological disorders (PNDs) represent a burden for public health. The World Health Organization estimates that at least 450 million people suffer from PNDs (ref. 1). Depression (322 million affected)2, bipolar disorder (60 million)1, schizophrenia (23 million)3, dementia and Alzheimer’s disease (50 million)1, and anxiety (260 million)2 are the most prevalent PNDs in the world. Autism spectrum disorders (1 in 59 children)4 and PNDs, such as Huntington’s disease (5–7 in 100,000 affected)1, and Parkinson’s disease (1–4% of all elderly people)1 are also of great concern.
Five major classes of drugs are used to treat PNDs: antidepressants, antipsychotics, anxiolytics, mood stabilizers, and stimulants. However, disease remission is not always achieved5,6. This stems from an incomplete knowledge of the molecular mechanisms of both PNDs (ref. 7) and the psychiatric drugs8. In addition, PNDs share several clinical and genetic components9, which makes the precise treatment and a subsequent targeted drug development more challenging10. Specifically, drug repositioning, which relies on testing drugs already in use for a disease to treat another illness based on the shared molecular pathology of both11, may be applied to treat PNDs (ref. 12).
Network medicine13 is an emerging field that combines systems biology and network science to understand how genes interact in disease and health. For PNDs, co-expression networks14,15,16 and genome-wide association studies9,17,18 have unraveled molecular mechanisms and genomic variations related to these disorders. Many more small-scale studies have investigated the roles of specific genes in PNDs. The daunting task, now, is to make sense of all the published data, stored in millions of research papers, that describe the interplay among genes, drugs, and other variables in the development and outcomes of PNDs.
Here, we used a network medicine approach to dissect the molecular mechanisms of PNDs and identify novel drug candidates for repositioning. Using IBM Watson for Drug Discovery (WDD), a machine learning text-mining application, we built knowledge networks containing connections between PNDs and genes or drugs mentioned in the published scientific literature in the past 50 years. We found classic and potentially unexplored pathways associated to PNDs. We also identified several drugs that target key PND-related genes that have never been used to treat these disorders previously. Validating our approach, some of these drugs are currently being tested to treat PNDs in clinical trials, with no previously published results. Our data provides comprehensive insight into the molecular pathology of PNDs and offers promising drug repositioning candidates for follow-up trials.
Materials and methods
Construction of the knowledge networks
We used the IBM WDD, an online tool, to perform queries for major PNDs: Alzheimer’s disease, dementia, anxiety, depression, Huntington’s disease, Parkinson’s disease, schizophrenia, bipolar disorder, and autism. We performed two independent searches using WDD: one for genes associated with PNDs (gene–PND) and another for drugs associated with PNDs (drug–PND). WDD detects associations in original papers and reviews from PMC Open Access (full text), Pubmed (abstracts), and patents. Relationships are detected by a natural language-processing algorithm when two entities of interest (gene, disease, or drug) are present in the same sentence of a document and are connected through a verb or preposition. WDD gives relationships a confidence score (0–100%) based on the number of documents supporting the connection and on link meaningfulness. Only relations with at least two documents of evidence and a confidence score >50% were maintained in our study. Searches were performed from July to August of the year 2018.
Network analysis
The Louvain19 method was used to detect modules of highly connected genes or drugs. For each pair of PNDs in the gene and drug networks, we performed Fisher’s exact test to calculate the significance of the overlap of genes or drugs in PNDs. Fisher’s exact test p-values (<0.01) were considered significant. Results were presented as −log10p-value.
Functional gene enrichment
Genes from modules in the gene–PND network and those exclusively associated with each PND were submitted for functional gene enrichment using enrichR (ref. 20). Enrichment was performed against the Gene Ontology Biological Process and the KEGG databases. The enriched terms with an enrichR combined score of at least 20 and p < 0.01 were retained, and the most enriched terms were used to describe the results.
Drug repositioning
Gene co-expression modules of PNDs were obtained from the Supplementary material of Gandal15. Co-expression modules were detected and the first principal component of the module’s expression (eigengene) was used to determine module–disease association. Module membership (kME) was calculated for each gene. The kME is equal to the Pearson’s correlation R between the expression of the gene and the eigengene of the module. Module hubs are those that have kME > 0.5 in their module.
The repositioning with Open Targets21 database was performed using a custom R code. This script, which is available on GitHub (https://github.com/csbl-usp/OpenTargets_drug_repositioning), accesses the platform API client of Open Targets database and performs the same steps done for the drug repositioning with WDD. Only relationships with an overall association score >0.5 were utilized.
Results
Molecular characterization of PNDs using the scientific knowledge
IBM WDD is a cognitive computing, artificial intelligence platform that was used to extract existing connections between genes, diseases, and drugs from millions of published documents related to the medical sciences22. WDD uses a dictionary created by artificial intelligence to group terms that are used conversely in the literature (e.g., gene IDs from distinct databases). It identifies relations between the searched term and other terms of interest in the literature. A natural language-processing algorithm detects these relations. We used WDD to investigate the genes and drugs that were shared among different PNDs. A total of 1588 genes and 722 drugs was identified as associated with PNDs. The network constructed with the genes associated with PNDs separated the diseases into two groups: neurodegenerative disorders (Alzheimer’s, Parkinson’s and Huntington’s diseases, and dementia) and psychiatric disorders (depression, anxiety, bipolar disorder, schizophrenia, and autism; Fig. 1a). The gene network identified five clusters of disorders with similar clinical characteristics: Alzheimer’s disease and dementia, cognition hindering neurodegenerative illnesses (Fig. 1a—purple color); Huntington’s disease and Parkinson’s disease, disorders that affect movement due to basal ganglia degeneration23 (Fig. 1a—light blue color); depression and anxiety, fear/threat-related disorders24 (Fig. 1a—red color); and schizophrenia and bipolar disorder, which share a spectrum of psychotic symptoms25 (Fig. 1a—brown-yellow color). Autism (Fig. 1a—green color), the only developmental PND analyzed, did not cluster with any other disease. Fisher’s exact test confirmed the division between neurodegenerative and psychiatric disorders and the associations between clinically similar disorders (Fig. 1b). The highest similarity was observed between disorders of the same group (neurodegenerative or psychiatric) and between PNDs clustered within the same modules (Fig. 1b). However, the separation between neurodegenerative and psychiatric disorders was not as evident in the PND-drug network (Supplementary Fig. 1).
a A knowledge network for genes colored according to Louvain-defined modules: Alzheimer’s disease (AD) and dementia (DM) (purple), Huntington’s disease (HD) and Parkinson’s disease (PKD; light blue), depression (MDD) and anxiety (AX; red), schizophrenia (SCZ) and bipolar disorder (BD; green-yellow), and autism (ASD; green). The dashed line in a separates neurodegenerative disorders from psychiatric disorders in the network. b The significance of the gene overlap between PNDs. Larger nodes represent PNDs with more genes and thicker edges represent a more significant overlap between PNDs (proportional to −log10p-value of Fisher’s exact test).
Functional gene analysis: insights into PND molecular pathology
We performed functional enrichment analyses with the genes in each module and with those unique to each PND (Fig. 2a). For all the modules, we found well-known, hallmark molecular characteristics of PNDs (Fig. 2, Supplementary results). We also had potentially novel insights into the PNDs. The genes in the Alzheimer’s disease/dementia module (Fig. 2b) were strongly enriched for neutrophil degranulation (33 genes) and microRNAs in cancer (21 genes; Fig. 2b). Neutrophil phenotype alterations in Alzheimer’s disease correlate with disease progression26, and neutrophil depletion improves memory and slows disease progression in mice27. One miRNA involved in cancer and Alzheimer’s disease, miR-146a, regulates innate immune response through inflammation in both diseases28. Conversely, many miRNAs that stimulate proliferation in cancer seem to favor apoptosis in Alzheimer’s disease28. A better comprehension of how miRNAs regulate the cell cycle and the immune system can open new therapeutic opportunities for treating both cancer and Alzheimer’s disease.
a A summary of genes connected to PNDs enriched for relevant biological processes (GO) and pathways (KEGG). b–f The most significant gene enrichment categories for genes in the modules: Alzheimer’s and dementia (b), Huntington’s disease and Parkinson’s disease (c), depression and anxiety (d), schizophrenia and bipolar disorder (e), and autism (f).
Huntington’s disease and Parkinson’s disease present severe neuronal loss in the basal ganglia23. In this light, genes involved in insulin resistance were enriched in this module (Fig. 2c). A recent study showed that over 60% of nondiabetic Parkinson’s disease patients had insulin resistance29. Genes unique to Huntington’s disease were enriched for the regulation of microtubule-based movement (Fig. 2c). The huntingtin protein interacts with several cell motility proteins, including HAP1 (ref. 30) and HDAC6 (ref. 31). These interactions mediate organelle trafficking32, and energy production and consumption via the axonal bounding of GAPDH to synaptic vesicles33.
The inflammatory response was enriched in the depression/anxiety module (Fig. 2d): c-reactive protein levels have been known to predict the outcome of antidepressant treatment34,35 and disease severity36. In addition, immunomodulatory proteins were found to be altered in the postmortem brain37, blood, and plasma38 of depression patients. Genes uniquely connected to depression were enriched for striated and cardiac muscle hypertrophy (HTR2B, TCAP, and TTN; Fig. 2d). Anxiety shares most of its genes with depression and has only eight unique genes (Fig. 2d). Two of these, HCN4 and ADCY8, have also been associated with obsessive-compulsive disorder39. Activated cyclic nucleotide-gated channels (HCNs) have been investigated as targets for novel antidepressants40, although HCN4 was not one of these cases.
Genes in the bipolar disorder and schizophrenia module (Fig. 2e) were enriched for long-term potentiation (LTP) and circadian entrainment (Fig. 2e). Those unique to bipolar disorder were also enriched for circadian rhythm (PER3 and RORB; Fig. 2e). Insomnia and sleep disorders affect the majority of schizophrenia patients41, and are also present in both depressive and manic phases of bipolar disorder42. Bipolar disorder patients even show symptoms that are synchronized with the circadian rhythm43. Furthermore, genes uniquely connected to schizophrenia were enriched for glutamate receptor transmission and ErbB signaling pathways (Fig. 2e). Neural dysconnectivity, a hallmark of schizophrenia, likely stems from aberrant synaptic plasticity and the incorrect developmental wiring of neurons due to oligodendrocyte malfunction44. These processes likely depend most prominently on glutamatergic transmission, neuregulin1 (NRG1)-ErbB signaling, and LTP (refs. 45,46). Three schizophrenia-unique genes related to glial cell differentiation were identified (Fig. 2e): ERBB3, PTPRZ1, and SOX10. Indeed, schizophrenia patients present an altered co-expression of genes associated with NFκB signaling along with genes co-expressed in oligodendrocytes, astrocytes, and microglia16,47.
The genes in the autism module (Fig. 2f) were enriched for the KEGG term “Inflammatory Bowel Disease (IBD)”. Autism patients have high comorbidity with IBD along with Crohn’s disease48. A fecal transplant from healthy subjects, a prospective therapy for IBD (ref. 49), has been suggested to alleviate digestive symptoms, and aggressive or repetitive behaviors in some autism patients50. The mechanisms behind these effects remain unclear, but the genes related to autism that were found to overlap with those of IBD are associated with inflammatory response, allergy, and the response to helminth parasites (interleukin (IL)6, IL23A, IL13, IL5, HLA-DRB1, HLA-DRB4, IFNG, and TGFB1). Here, IL13 was found to be targeted by drugs that have never been tested for autism. The circadian cycle-related gene RORA, also shared by IBD and autism, perhaps affects immune function due to the disruption of daily rhythms51.
Network medicine framework
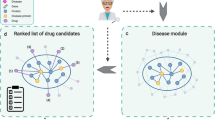
From the gene–PND network (Fig. 3a), we selected genes that were exclusively associated to each PND and that were coexpressed in brain tissue of PND patients according to Gandal et al.15 (Fig. 3b). Co-expression hubs are potential drug targets since they may influence the expression of several other genes52. To select the most relevant drugs, we kept only those targeting genes that are co-expressed in brain tissues of PND patients15. These co-expression networks included patients and healthy subjects from 700 microarray gene expression studies. The selected genes (Supplementary Table 1) were then submitted to a new round of WDD searches to find drugs associated with them (Fig. 3c). This resulted in 1305 drugs. Since our goal was drug repositioning, we removed from the network 782 drugs known to be associated with the PNDs or that were associated to more than one gene (Fig. 3d, Supplementary Table 2). This allowed a subsequent manual curation of 30% of the results (Fig. 3e). We read each document provided by WDD (Supplementary Table 2) that supports the drug–gene and the gene–PND relationships, and manually removed errors and any relationship that was not in fact described in the documents (examples in Supplementary Table 3). It is important to note that reviewing these 30% of interactions did not constitute a WDD performance evaluation, which was not the goal of our work. With the remaining drugs, we searched the Drug Bank (https://www.drugbank.ca/)53 for any ongoing or finished clinical trials, involving these drugs and the PNDs (Fig. 3e). Finally, we performed an open literature review in Pubmed for selected drug–gene–PND relationships (Fig. 3e). We aimed at explaining how each drug could potentially affect their target gene and how this effect could impact the disease. Drugs that presented promising evidence of a viable mechanism that could potentially promote disease altering effects were selected for discussion. The scripts used to perform these steps are available on GitHub (https://github.com/csbl-usp/WDD_drug_repositioning).
a WDD searches for gene–PND and drug–PND associations. All WDD results were filtered to keep only relationships supported by two documents or more and at least 50% confidence. b Selection of exclusive genes of each PND that are coexpressed in brain tissue according to Gandal et al.15. c WDD searches for drug–gene associations using exclusive coexpressed genes of PNDs. d Removal of drugs obtained in a and of drugs targeting more than one gene. e Drug prioritization through literature searches and selection of relevant cases.
Drug repositioning for PNDs
Our network medicine framework with WDD was applied to find suitable candidates for drug repositioning for PNDs (Fig. 4, Supplementary Fig. 2). We manually curated the resulting list and selected those drugs with potential for follow-up testing. We prioritized 63 drugs targeting 31 genes and eight PNDs that showed potential for follow-up testing (Supplementary Table 4). Literature search revealed that 18 of those drugs had already been associated with a PND, suggesting that our criteria for initial screening was stringent.
a–c The most promising repositioning drug candidates (blue) that target the genes (red) unique to Alzheimer’s disease (a), schizophrenia (b), and autism and bipolar disorder (c). The drugs discussed in the main text are highlighted here: monoclonal antibodies (green), inhibitors (magenta), and anticonvulsants (cyan).
We also used Open Targets21 to demonstrate that the drug repositioning framework proposed here can be successfully applied to any drug–gene–disease interaction database (see “Methods”). Applying the same framework described in Fig. 3a–d, we obtained 4670 drugs (Supplementary Table 2). Due to drug name synonyms (which were consolidated in WDD), Open Targets associations include many redundant drug–gene interactions. Nevertheless, we were able to detect 91 drugs in common between WDD and Open Targets (Supplementary Table 5). Promising cases of drugs that target genes connected to Alzheimer’s disease, schizophrenia, bipolar disorder, and autism are depicted in Fig. 4 and described below.
Two anti-CD33 monoclonal antibodies, Gemtuzumab Ozogamicin and Lintuzumab, were identified as promising repositioning candidates concerning Alzheimer’s disease (Fig. 4a). Lintuzumab (also found in Open Targets—Supplementary Table 5) has been reported to reduce the microglial cell surface CD33 by 80% (refs. 54,55). CD33 is involved in the inflammatory response related to the amyloid cascade in Alzheimer’s disease54. The CD33 gene harbors a single-nucleotide polymorphism (SNP) that protects against Alzheimer’s disease (rs12459419T). This SNP leads to the production of a nonfunctional splicing isoform of CD33, lacking exon 2 (refs. 54,55). Treatment with anti-CD33 antibodies might replicate this effect by reducing the CD33 protein levels and reduce microglia-associated neuroinflammation. Drugs targeting the colony stimulation factor 1 receptor (CSF1R) present similar potential (Fig. 4a). This particular gene is essential for microglia survival in the brain56. Also, the treatment with CSF1R inhibitors in mice leads to a reduced microglia-mediated accumulation of amyloid plaques56 and prevents the development of Alzheimer’s disease-like symptoms due to the anti-inflammatory effect of halting microglia proliferation57. We found two drugs, JNJ-40346527 (CSF1R inhibitor) and Emactuzumab (anti-CSF1R monoclonal antibody; Fig. 4a), which have been extensively used in oncology58 but never to treat Alzheimer’s disease. Ronacaleret, a calcium-sensing receptor (CASR) inhibitor, could also mitigate the deleterious effects of the amyloid cascade. CASR is expressed by astrocytes and is responsible for detecting synaptic cleft Ca2+ concentrations. Extracellular amyloid-β oligomers activate CASR, which induces the accumulation and secretion of more oligomers, nitric oxide release, and VEGF-A expression59. This leads to neuronal death, sparing the astrocytes, which continue the amyloid cascade60. Halting this pathway early could reduce neuronal death and slow down the progression of Alzheimer’s.
Some schizophrenia-related genes were also promising drug targets (Fig. 4b) found both in WDD and Open Targets (Supplementary Table 5). ERBB2, an essential gene in the NRG1/ErbB signaling pathway, is targeted by six drugs—three inhibitors (AC-480, Mubritinib, and CP 724714) and three monoclonal antibodies (Trastuzumab, Ertumaxomab, and MM-302; Fig. 4b). This pathway is involved in schizophrenia due to its relevance to synaptic transmission and plasticity45. Moreover, it is known that NRG1/ErbB signaling triggers myelination in oligodendrocytes61. ErbB1 inhibition has been proposed as a potential antipsychotic approach62, and the anti-ErB2 monoclonal antibody Trastuzumab has been suggested as a possible treatment tool for schizophrenia63. We also found SLC6A9 (glycine transporter 1—GlyT1) inhibitors (Fig. 4b). Bitopertin (NCT01116830) and PF-03463275 (NCT01911676) are actually going through clinical trials for schizophrenia64,65 (Fig. 4b). Two anticonvulsant drugs (Levetiracetam and Brivaracetam) targeting the synaptic vesicle glycoprotein 2A are also promising in this respect (Fig. 4b). Levetiracetam (Fig. 4b) was found to improve cognition in a rat model for schizophrenia66. Brivaracetam (Fig. 4b) has never been used to treat schizophrenia, which makes it an attractive drug repositioning candidate.
Among the drugs connected to the remaining PND-related genes (Fig. 4c, Supplementary Fig. 2), we highlight those that act upon IL13 for autism and CEACAM5 for bipolar disorder (Fig. 4c). Drugs that affect genes connected to depression, dementia, and Parkinson’s disease are depicted in Supplementary Fig. 2, and discussed in the Supplementary results section. IL13 has increased expression in the T lymphocytes of autism patients67 and exerts its inflammatory effects through the acidic mammalian chitinase (AMCase)68. In this research, we found two anti-IL13 monoclonal antibodies (Lebrikizumab and Tralokinumab) and an AMCase inhibitor (allosamidin; Fig. 4c). Neither drugs have been tested for autism. CEACAM5 (carcinoembryonic antigen-related cell adhesion molecule 5, OMIM 114890) has been reported to be a potential biomarker for bipolar disorder69. CEACAM5 levels are higher in the manic phase69, and lithium reduces its levels70. Two CEACAM5-specific antibodies and one bi-specific CEA/CD33 antibody with no prior connections to bipolar disorder were also identified using WDD and Open Targets (Fig. 4c, Supplementary Table 5). Although it is unclear whether alterations in CEA serum levels cause or arise due to bipolar disorder, reducing the CEACAM5 serum levels might be a promising approach.
Discussion
Our results provide robust evidence in favor of Barabási’s shared components hypothesis13, which states that “[…]diseases that share disease-associated cellular components (genes, proteins, metabolites, and miRNAs) show phenotypic similarity and comorbidity”. Since we used data obtained entirely from previously published works, none of the individual relationships between genes and drugs, and genes and PNDs, by definition, are novel. Nonetheless, the network medicine framework presented here was able to integrate this accumulated knowledge from the scientific literature to obtain several previously unknown associations between drugs and PNDs. We also showed that this framework can be applied with success to different drug–gene–diseases interaction databases.
PNDs are dimensional conditions with multiple overlapping layers of complexity71. We saw that PNDs that share more symptoms, also share more genes. These findings support the idea that PND-related genes are associated with brain networks involved in shared behavioral manifestations, such as cognition and fear-threatening reactions72. Our results also confirm the genetic separation between neurological and psychiatric disorders, seen recently using GWAS results9.
We were able to break down the inherent characteristics of PNDs to find particularities. Cornerstone biological pathways associated with neuropsychiatry were readily detectable in our data: amyloid beta plaque formation in Alzheimer’s disease and dementia, apoptosis for Parkinsons’s diseases and Hutington disease, synaptic transmission for depression, anxiety, bipolar disorder and schizophrenia, and synaptic organization and cell–cell adhesion for autism (Supplementary results). We were also able to detect consistent characteristics that are just being described in the literature and have not been fully explored yet, such as the involvement of cancer-related miRNAs in Alzheimer’s disease28, the regulation of dopamine transmission by the circadian cycle73, and the role of subcellular molecular trafficking in Huntington’s disease32,33. Our results also supported a genetic relation between depression and heart disease. Hypertrophic cardiomyopathy (HCM) and depression are also common comorbidities74, and patients with HCM are correlated with a higher prevalence of depression75. Finally, by looking at the complete scope of the literature published in the past 50 years, we were able to identify a consistent neuroimmune/inflammatory genetic signature in all PNDs.
Previous computer-based or experimental drug repositioning frameworks have relied on gene expression, drug–trarget binding or phenotypical screenings to find candidates76,77. WDD does not distinguish associations between genes and disorders that occur due to SNPs, epigenetic modifications, or differences in expression. If two PNDs share a gene, each relation could involve a different mechanism. No information on drug effect direction is provided either. Thus, we had to compromise in selecting potential candidates for drug repositioning; we decided to only keep drugs that were found to affect one PND gene. This reduced the amount of collected data, which allowed us to validate several drug–gene–PND connection individually and prioritize candidates for discussion. There was a caveat of increasing specificity and decreasing the potential reach of the drugs. Since PNDs are complex, omnigenic disorders71, solutions focused on individual genes may not be ideal. We selected drugs affecting genes that are coexpressed in brain tissue from PND patients15. Co-expressed genes usually play more critical roles in diseases, are more often targeted by drugs, and have influence over the expression of other genes52. We predict that the reach of the selected drugs will be broad due to the coexpressed nature of their targets. Using this approach, we found monoclonal antibodies with repositioning potential. Monoclonal antibodies are highly specific concerning their targets, but they are also large molecules with low permeability through the blood–brain barrier. This can be a challenge in the follow-up validation of the drugs found in this research. However, some of these antibodies are being discussed as viable alternatives for treatment of schizophrenia and Alzheimer’s disease54,63, which indicates that our approach is efficient in finding potential candidates for drug repositioning. We also found drugs that could be used in combination to treat PNDs. Six drugs that could treat schizophrenia were found to target ERBB2. Recently, the NRG1-ERBB4 signaling antagonist Spironolactone was identified in a cell-based drug repositioning screening as a candidate for clinical trials for schizophrenia77. These drugs have the potential to reduce dysconnectivity and hallucination by regulating the NRG1-ErbB signaling. Four drugs could be used synergistically to reduce microglia-mediated inflammation in Alzheimer’s disease, through the inhibition of CSF1R and CD33. Three other drugs were also found to target IL13, which could be used to reduce the inflammatory response in autism.
Our network medicine approach was able to successfully integrate the data obtained from millions of scientific papers using complex networks to generate new insights about PNDs. The network medicine framework proposed here can be applied to drug–gene–disease interaction databases, such as WDD and Open Targets. The drugs we selected here are highly promising candidates for repositioning that could be taken into consideration for follow-up in vitro and in vivo screenings.
References
World Health, O. The World Health Report 2001 (2001).
Who. Depression and Other Common Mental Disorders: Global Health Estimates, 24 (World Health Organization, 2017).
Kahn, R. S. et al. Schizophrenia. Nat. Rev. Dis. Prim. 1, 15067–15067 (2015).
Baio, J. et al. Prevalence of autism spectrum disorders in a total population sample-autism and developmental disabilities monitoring network, 11 Sites, United States, 2014. MMWR Surveill. Summ. 67, 1–25 (2018).
Cipriani, A. et al. Comparative efficacy and acceptability of 21 antidepressant drugs for the acute treatment of adults with major depressive disorder: a systematic review and network meta-analysis. Lancet 391, 1357–1366 (2018).
Gomez, A. F., Barthel, A. L. & Hofmann, S. G. Comparing the efficacy of benzodiazepines and serotonergic anti-depressants for adults with generalized anxiety disorder: a meta-analytic review. Expert Opin. Pharmacother. 19, 883–894 (2018).
Jauhar, S. & Young, A. H. Controversies in bipolar disorder; role of second-generation antipsychotic for maintenance therapy. Int. J. Bipolar Disord. 7, 10 (2019).
Gitlin, M. J. Antidepressants in bipolar depression: an enduring controversy. Int. J. Bipolar Disord. 6, 25 (2018).
Anttila, V. et al. Analysis of shared heritability in common disorders of the brain. Science 360, eaap8757 (2018).
Simon, N. G. & Brownstein, M. J. Challenges in developing drugs for neurological and psychiatric disorders. Prog. Neurobiol. 152, 1–2 (2017).
So, H.-c., Chau, C. K.-L., Lau, A., Wong, S.-Y. & Zhao, K. Translating GWAS findings into therapies for depression and anxiety disorders: gene-set analyses reveal enrichment of psychiatric drug classes and implications for drug repositioning. Psychol. Med. 49, 2692–2708 (2018).
Tian, Z., Teng, Z., Cheng, S. & Guo, M. Computational drug repositioning using meta-path-based semantic network analysis. BMC Syst. Biol. 12, 134 (2018).
Barabási, A.-L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12, 56–68 (2011).
Boloc, D. et al. Improving pharmacogenetic prediction of extrapyramidal symptoms induced by antipsychotics. Transl. Psychiatry 8, 276 (2018).
Gandal, M. J. et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science 359, 693–697 (2018).
Gandal, M. J. et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science 362, eaat8127 (2018).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Mistry, S., Escott-Price, V., Florio, A. D., Smith, D. J. & Zammit, S. Genetic risk for bipolar disorder and psychopathology from childhood to early adulthood. J. Affect. Disord. 246, 633–639 (2019).
Blondel, V. D., Guillaume, J. L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, 1–12 (2008).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Carvalho-Silva, D. et al. Open Targets Platform: new developments and updates two years on. Nucleic Acids Res. 47, D1056–D1065 (2019).
Chen, Y., Elenee Argentinis, J. D. & Weber, G. IBM Watson: how cognitive computing can be applied to big data challenges in life sciences research. Clin. Ther. 38, 688–701 (2016).
Schroll, H. & Hamker, F. H. Basal Ganglia dysfunctions in movement disorders: What can be learned from computational simulations. Mov. Disord. 31, 1591–1601 (2016).
Howe, T. R. et al. American Psychiatric Association (eds Sam, Goldstein & Jack A., Naglieri) Diagnostic and Statistical Manual of Mental Disorders 4 Text Revision (DSM-IV-TR). 84–85 (Springer: US, 2000).
Jablensky, A. The diagnostic concept of schizophrenia: its history, evolution, and future prospects. Dialogues Clin. Neurosci. 12, 271–287 (2010).
Dong, Y. et al. Neutrophil hyperactivation correlates with Alzheimer’s disease progression. Ann. Neurol. 83, 387–405 (2018).
Zenaro, E. et al. Neutrophils promote Alzheimer’s disease–like pathology and cognitive decline via LFA-1 integrin. Nat. Med. 21, 880–886 (2015).
Holohan, K. N., Lahiri, D. K., Schneider, B. P., Foroud, T. & Saykin, A. J. Functional microRNAs in Alzheimer’s disease and cancer: differential regulation of common mechanisms and pathway. Front. Genet. 3, 1–16 (2013).
Hogg, E. et al. High prevalence of undiagnosed insulin resistance in non-diabetic subjects with parkinson’s disease. J. Parkinson’s Dis. 8, 259–265 (2018).
Kalchman, M. A. et al. HIP1 interacts with membrane-associated huntingtin in the brain. Nat. Genet. 16, 44–53 (1997).
Iwata, A., Riley, B. E., Johnston, J. A. & Kopito, R. R. HDAC6 and microtubules are required for autophagic degradation of aggregated Huntingtin. J. Biol. Chem. 280, 40282–40292 (2005).
Caviston, J. P., Ross, J. L., Antony, S. M., Tokito, M. & Holzbaur, E. L. F. Huntingtin facilitates dynein/dynactin-mediated vesicle transport. Proc. Natl Acad. Sci. USA 104, 10045–10050 (2007).
Zala, D. et al. Vesicular glycolysis provides on-board energy for fast axonal transport. Cell 152, 479–491 (2013).
Martins-de-Souza, D., Guest, P. C. & Steiner, J. A proteomic signature associated to atypical antipsychotic response in schizophrenia patients: a pilot study. Eur. Arch. Psychiatry Clin. Neurosci. 270, 127–134 (2019).
Uher, R. et al. An inflammatory biomarker as a differential predictor of outcome of depression treatment with escitalopram and nortriptyline. Am. J. Psychiatry 171, 1278–1286 (2014).
Köhler-Forsberg, O. et al. Association between C-reactive protein (CRP) with depression symptom severity and specific depressive symptoms in major depression. Brain, Behav. Immun. 62, 344–350 (2017).
Martins-De-Souza, D. et al. Identification of proteomic signatures associated with depression and psychotic depression in post-mortem brains from major depression patients. Transl. Psychiatry 2, e87–13 (2012).
Martins-De-Souza, D. et al. Plasma fibrinogen: now also an antidepressant response marker? Transl. Psychiatry 4, e352–e354 (2014).
Kelmendi, B. et al. Association of polymorphisms in HCN4 with mood disorders and obsessive compulsive disorder. Neurosci. Lett. 496, 195–199 (2011).
Ku, S. M. & Han, M. H. HCN channel targets for novel antidepressant treatment. Neurotherapeutics 14, 698–715 (2017).
Palmese, L. B. et al. Insomnia is frequent in schizophrenia and associated with night eating and obesity. Schizophr. Res. 133, 238–243 (2011).
Bradley, A. J. et al. Sleep and circadian rhythm disturbance in bipolar disorder. Psychol. Med. 47, 1678–1689 (2017).
Wehr, T. A. Bipolar mood cycles associated with lunar entrainment of a circadian rhythm. Transl. Psychiatry 8, 151–151 (2018).
Stephan, K. E., Friston, K. J. & Frith, C. D. Dysconnection in schizophrenia: from abnormal synaptic plasticity to failures of self-monitoring. Schizophr. Bull. 35, 509–527 (2009).
Buonanno, A. The neuregulin signaling pathway and schizophrenia: from genes to synapses and neural circuits. Brain Res. Bull. 83, 122–131 (2010).
Ledonne, A. et al. Neuregulin 1/ErbB signalling modulates hippocampal mGluRI-dependent LTD and object recognition memory. Pharmacol. Res. 130, 12–24 (2018).
de Baumont, A. et al. Innate immune response is differentially dysregulated between bipolar disease and schizophrenia. Schizophr. Res. 161, 215–221 (2015).
Doshi-Velez, F. et al. Prevalence of inflammatory bowel disease among patients with autism spectrum disorders. Inflamm. Bowel Dis. 21, 1–1 (2015).
Weingarden, A. R. & Vaughn, B. P. Intestinal microbiota, fecal microbiota transplantation, and inflammatory bowel disease. Gut Microbes 8, 238–252 (2017).
Kang, D. W. et al. Long-term benefit of microbiota transfer therapy on autism symptoms and gut microbiota. Sci. Rep. 9, 1–9 (2019).
Palmieri, O. et al. Systematic analysis of circadian genes using genome-wide cDNA microarrays in the inflammatory bowel disease transcriptome. Chronobiol. Int. 32, 903–916 (2015).
van Dam, S., Võsa, U., van der Graaf, A., Franke, L. & de Magalhães, J. P. Gene co-expression analysis for functional classification and gene–disease predictions. Brief. Bioinform. 19, bbw139–bbw139 (2017).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Malik, M. et al. Genetics of CD33 in Alzheimer’s disease and acute myeloid leukemia. Hum. Mol. Genet. 24, 3557–3570 (2015).
Malik, M. et al. CD33 Alzheimer’s risk-altering polymorphism, CD33 expression, and exon 2 splicing. J. Neurosci. 33, 13320–13325 (2013).
Sosna, J. et al. Early long-term administration of the CSF1R inhibitor PLX3397 ablates microglia and reduces accumulation of intraneuronal amyloid, neuritic plaque deposition and pre-fibrillar oligomers in 5XFAD mouse model of Alzheimer’s disease. Mol. Neurodegener. 13, 1–11 (2018).
Olmos-Alonso, A. et al. Pharmacological targeting of CSF1R inhibits microglial proliferation and prevents the progression of Alzheimer’s-like pathology. Brain 139, 891–907 (2016).
Ries, C. H. et al. Targeting tumor-associated macrophages with anti-CSF-1R antibody reveals a strategy for cancer therapy. Cancer Cell 25, 846–859 (2014).
Dal Prà, I. et al. The Aβ peptides-activated calcium-sensing receptor stimulates the production and secretion of vascular endothelial growth factor-A by normoxic adult human cortical astrocytes. NeuroMolecular Med. 16, 645–657 (2014).
Chiarini, A. et al. Preventing the spread of Alzheimer’s disease neuropathology: a role for calcilytics? Curr. Pharm. Biotechnol. 16, 696–706 (2015).
Cassoli, J. S. et al. Disturbed macro-connectivity in schizophrenia linked to oligodendrocyte dysfunction: from structural findings to molecules. NPJ Schizophr. 1, 15034–15034 (2015).
Mizuno, M. et al. Antipsychotic potential of quinazoline ErbB1 inhibitors in a schizophrenia model established with neonatal hippocampal lesioning. J. Pharmacol. Sci. 114, 320–331 (2010).
Sastry, P. S. R. K. & Sita Ratna, W. Intrathecal therapy with trastuzumab may be beneficial in cases of refractory schizophrenia. Med. Hypotheses 62, 542–545 (2004).
Bugarski-Kirola, D. et al. Efficacy and safety of adjunctive bitopertin versus placebo in patients with suboptimally controlled symptoms of schizophrenia treated with antipsychotics: results from three phase 3, randomised, double-blind, parallel-group, placebo-controlled, multicent. Lancet Psychiatry 3, 1115–1128 (2016).
D’Souza, D. C. et al. Dose-related target occupancy and effects on circuitry, behavior, and neuroplasticity of the glycine transporter-1 inhibitor PF-03463275 in healthy and schizophrenia subjects. Biol. Psychiatry 84, 413–421 (2018).
Koh, M. T., Shao, Y., Rosenzweig-Lipson, S. & Gallagher, M. Treatment with levetiracetam improves cognition in a ketamine rat model of schizophrenia. Schizophr. Res. 193, 119–125 (2018).
Ashwood, P. et al. Altered T cell responses in children with autism. Brain, Behav. Immun. 25, 840–849 (2011).
Zhu, Z. et al. Acidic mammalian chitinase in asthmatic Th2 inflammation and IL-13 pathway activation. Science 304, 1678–1682 (2004).
Bulut, M. et al. Evaluation of serum inflammatory markers in treatment-resistant manic patients and adequate responder manic patients. Psychiatry Res. 272, 73–79 (2018).
Asad-Ur-Rahman, F. N. U. & Saif, M. W. Elevated level of serum carcinoembryonic antigen (CEA) and search for a malignancy: a case report. Cureus 8, 8–11 (2016).
Boyle, E. A., Li, Y. I. & Pritchard, J. K. An expanded view of complex traits: from polygenic to omnigenic. Cell 169, 1177–1186 (2017).
Insel, T. et al. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am. J. Psychiatry 167, 748–751 (2010).
Kim, J. et al. Implications of circadian rhythm in dopamine and mood regulation. Mol. Cells 40, 450–456 (2017).
Zahid, I. et al. Frequency and predictors of depression in congestive heart failure. Indian Heart J. 70, S199–S203 (2018).
Igoumenou, A. et al. Depression in patients with hypertrophic cardiomyopathy: is there any relation with the risk factors for sudden death? Heart Asia 4, 44–48 (2012).
Pushpakom, S. et al. Drug repurposing: progress, challenges and recommendations. Nat. Rev. Drug Discov. 18, 41–58 (2019).
Wehr, M. C. et al. Spironolactone is an antagonist of NRG 1‐ ERBB 4 signaling and schizophrenia‐relevant endophenotypes in mice. EMBO Mol. Med. 9, 1448–1462 (2017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lüscher Dias, T., Schuch, V., Beltrão-Braga, P.C.B. et al. Drug repositioning for psychiatric and neurological disorders through a network medicine approach. Transl Psychiatry 10, 141 (2020). https://doi.org/10.1038/s41398-020-0827-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-0827-5
This article is cited by
-
Drug targets for COVID-19 therapeutics: Ongoing global efforts
Journal of Biosciences (2020)