Abstract
Antidepressants exhibit similar efficacy, but varying tolerability, in randomized controlled trials. Predicting tolerability in real-world clinical populations may facilitate personalization of treatment and maximize adherence. This retrospective longitudinal cohort study aimed to determine the extent to which incorporating patient history from electronic health records improved prediction of unplanned treatment discontinuation at index antidepressant prescription. Clinical data were analyzed from individuals from health networks affiliated with two large academic medical centers between March 1, 2008 and December 31, 2014. In total, the study cohorts included 51,683 patients with at least one International Classification of Diseases diagnostic code for major depressive disorder or depressive disorder not otherwise specified who initiated antidepressant treatment. Among 70,121 total medication changes, 16,665 (23.77%) of them were followed by failure to return; maximum risk was observed with paroxetine (27.71% discontinuation), and minimum with venlafaxine (20.78% discontinuation); Mantel–Haenzel χ2 (8 df) = 126.44, p = 1.54e–23 <1e–6. Models incorporating diagnostic and procedure codes and medication prescriptions improved per-medication Areas Under the Curve (AUCs) to a mean of 0.69 [0.64–0.73] (ranging from 0.62 for paroxetine to 0.80 for escitalopram), with similar performance in the second, replication health system. Machine learning applied to coded electronic health records facilitates identification of individuals at high-risk for treatment dropout following change in antidepressant medication. Such methods may assist primary care physicians and psychiatrists in the clinic to personalize antidepressant treatment on the basis not solely of efficacy, but of tolerability.
Similar content being viewed by others
Introduction
While efforts at personalization of antidepressant (AD) treatment have focused on therapeutic efficacy, on average, modern ADs show greater differences in tolerability. For example, a recent large meta-analysis found significant differences for rates of acute discontinuation, but not efficacy, among 21 treatments investigated in randomized, controlled trials1. These authors highlighted the need to develop new strategies to distinguish individual-level differences in medication response, even where group-level differences are modest.
From a clinical perspective, personalization within medication class is challenging2,3. Treatment discontinuation may reflect a range of features, from depression-associated amotivation and hopelessness to failure to perceive a benefit to concerns about cost. However heterogeneous, the consequences of treatment discontinuation are substantial, contributing to poor treatment outcomes and depression chronicity4. Consistent with the importance of discontinuation as a clinical indicator, time to discontinuation has been used as an endpoint in large clinical trials5.
Previous work has demonstrated that electronic health records (EHRs) can be leveraged to generate sufficient sample sizes to facilitate machine learning studies of AD6. Here, we apply these methods to develop predictions of treatment discontinuation using large-scale EHR data from one health system and further characterize the performance of these models in a second academic medical center network. Specifically, we develop predictors of failure to return for a follow-up psychiatric visit of any kind after a change in antidepressant prescription as a face-valid, if nonspecific, adverse outcome readily detectable using coded EHR data.
Methods
Study overview and cohort description
The study cohort derivation began with 252,351 patients drawn from two academic medical center treatment networks in the Northeast United States (subsequently referred to as Site A and Site B) who received at least one antidepressant prescription between 2008 and 2014. The cohort included all individuals with at least one International Classification of Diseases, Ninth Revision (ICD9) diagnostic code for major depressive disorder (ICD9s 296.2x, 296.3x) or depressive disorder not otherwise specified (311). For these individuals, a datamart was generated using i2b2 server software (i2b2, Boston, MA, USA)7. Available patient data included sociodemographic information, diagnostic and procedure codes, as well as inpatient medication administrations and outpatient medication prescriptions extracted by ingredient and duration. The Partners HealthCare institutional review board approved the study protocol, waiving the requirement for informed consent as only de-identified data was utilized and no human subjects contact was required.
Inclusion criteria
For model development, we restricted the cohorts to patients of age 18–80 years who were affiliated with one primary site (either Site A or B) and had received at least one of the nine most commonly-prescribed antidepressants (see Supplementary Table 1 for the list of antidepressants and Supplementary Fig. 1 for an illustration of this process). Individuals were excluded from primary analysis if they had no encounter of any type in the EHR system after 90 days from the last prescription registered (i.e., were potentially lost to follow-up, as could be the case if they transferred their care to another health system).
Outcome definition
The primary outcome was treatment discontinuation following index prescription, defined as <90 days of prescription availability and no evidence of non-pharmacologic psychiatric treatment. This latter feature is defined as an absence of a psychiatric Current Procedural Terminology (CPT) code in 13 months following the last antidepressant prescription (see Supplementary Table 2 for a list of psychiatric codes), as a means of excluding individuals who, while not able to continue pharmacologic treatment, remain in ongoing psychiatric treatment. As noted above, we also excluded individuals who had no interaction with the health system in the 90 days following the last antidepressant treatment, as a means of decreasing the possibility that individuals were lost to follow-up because of (for example) change in treatment network or other transfer of care. Supplementary Fig. 2 illustrates the codes observed for an example individual discontinuing treatment after 225 days; of note, while there are no further antidepressant prescriptions or psychiatric visits, the continuation of non-psychiatric follow-up indicates that this individual has not been lost to follow-up.
For subsequent analysis, 27,366 eligible patients in the Site A cohort were randomly assigned to a training (80%), validation (10%), and testing (10%) data set. All 16,630 eligible patients from Site B were held out for testing.
Modeling approach
Prediction task
We sought to predict treatment discontinuation for a given antidepressant medication on the basis of sociodemographic features, diagnostic codes, procedures, and medication data available at the time of index prescription of any given antidepressant, censoring all subsequent data. We therefore built models and evaluated the prediction task for each of the nine most-prescribed medications (see Supplementary Table 1) individually. Given a specific patient encounter and a specific medication, each prediction method yields a score or probability that the specific medication in question would lead to treatment discontinuation. We evaluated our prediction models at every change in treatment prescription under the condition that there was sufficient prior history—i.e., at least one prior ‘fact’ of any kind, comprised of diagnostic code, procedure, or prescription.
Feature derivation
From an initial set of 23,949 possible ICD or CPT codes and medications, we applied frequency thresholding to select 3852 codes occurring in at least 100 patients in Site A. For each patient and each encounter date, we built a compact representation of past event history via a count vector indicating how often each of these codewords appeared in the patient’s past history. Sociodemographic variables, including self-reported gender, race/ethnicity, and age at treatment event (in fractional years, e.g., 36.7 years old) are recorded for all patients at time of prescription and represented via one-hot encoded vectors (i.e., using binary features). We also include the calendar date of the treatment date (in fractional years) to capture secular trends.
Classification methods and metrics
We applied two standard classifiers, logistic regression (LR) and random forests (RF). Logistic regression used the implementation provided in the open-source Scikit Learn (version 0.18.1) toolkit for Python8. We tuned two hyperparameters on the Site A validation set: the type of regularization (L1 or L2 norm penalty) and the regularization strength. Random forests used the “ExtraTreeClassifier” implementation in Scikit Learn and tuned three hyperparameters on validation data: the number of trees, the fraction of features used in each tree, and the minimum number of samples at leaf nodes. Using each method, a separate classifier for each of the nine target medications was trained on the Site A training set. Hyperparameters were tuned using grid search to find the parameter combination that performs best on the Area Under the Curve (AUC) metric.
In primary analysis, model performance was compared using AUC for each of the nine medication prediction tasks in the held-out testing set from Site A and then in the independent Site B, to understand the extent to which code-based prediction of treatment discontinuation generalize to different scenarios. Recognizing that there are numerous means of model comparison, we elected to compare AUC’s using Student’s t-tests for familiarity and simplicity9. In light of the large cohort as well as the need to test multiple hypotheses simultaneously, we conservatively chose to apply Bonferroni correction by considering a significance level alpha = 0.05/100 = 0.0005 for these comparisons. Statistical validation for AUC values is reported in Supplementary Table 3 using 100 bootstraps. In secondary analysis, we characterized the performance of the two classifiers in sub-cohorts. First, while primary analysis sought to model all providers jointly, we subsequently examined models restricted to psychiatric versus non-psychiatric prescribers in order to address the possibility that each setting might have more prescriber-specific predictors. Second, we examined performance of models stratified by number of prior failed treatments—i.e., to what extent does ability to predict dropout change with additional treatment data.
Results
Rates of dropout across provider types and demographic subgroups
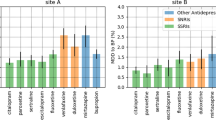
The two cohorts, spanning Site A and Site B, included 43,996 patients of which 14,975 (34.04%) were male, 34,544 (78.52%) were white, and 27,366 (62.20%) were cared for at Site A; mean age was 47.7 (SD 15.4) years. Demographics for the cohort by site are shown in Table 1. In total, 9502 of 27,366 (34.7%) patients from Site A discontinued treatment after a new antidepressant prescription, and 6274 of 16,630 (37.7%) at Site B. Figure 1 shows the proportion of dropouts among each antidepressant, organized by pharmacologic class. Among 70,121 total medication changes, 16,665 (23.77%) of them were followed by failure to return; maximum risk was observed with paroxetine (27.71% discontinuation), and minimum with venlafaxine (20.78% discontinuation). In general, greatest rates of discontinuation were observed with selective serotonin reuptake inhibitors across all types of providers (panel a), and discontinuation rates among non-psychiatrists (panel b) were substantially greater than psychiatrists (panel c); median Mantel–Haenzel χ2 (1 df) = 150.58, p = 1.29e–34 < 5e–4 (see Supplementary Table 4).
Model discrimination
Figure 2 shows the AUC for the LR and RF classifiers across individual medications and provider types. Both classifiers exhibit similar performances across medications; mean AUC was 0.67 [0.62–0.71] for LR and 0.69 [0.64–0.73] for RF, with best discrimination for escitalopram (AUC of 0.80), and poorest for paroxetine (AUC of 0.62). Compared with the baseline models (training with sociodemographic information alone, or sociodemographics plus the actual year of the prescription), incorporating EHR data increases discrimination substantially and significantly (median Welch’s t-test = −29.31; p = 5.34e–72 < 5e–4 for LR, and median Welch’s t-test = −26.08; p = 1.15e–65 < 5e–4 for RF) (Supplementary Table 3). AUC values are also greater in general for individuals treated by psychiatrists compared with non-psychiatrists: 0.66 vs 0.64 (median Welch’s t-test = −2.41; p = 3.79e–4 < 5e–4 for LR, and median Welch’s t-test = −6.38; p = 3.68e–15 < 5e–4 for RF). Detailed statistical validation can be found in Supplementary Table 3a, b. For categories of features most strongly associated with discontinuation in each model, see Supplementary Fig. 3.
Prediction accuracy with respect to number of previous prescriptions
We next examined the extent to which model performance depends on the number of prior treatment trials observed. Figure 3 presents this stratified analysis of AUC values according to prior treatments. In general, for models incorporating coded clinical data, AUC values are stable or improve with the number of previous documented treatments (see Supplementary Table 3c for statistical validation).
Stratification according to how many distinct treatments have already been tried in the past. AUC improves with the number of tried treatments in the past. Four different types of input data are considered: sociodemographic features (dem), date of prescription (date), and diagnostic/procedure codes (codes). Confidence intervals computed using 500 bootstraps.
Model replication at a second site
While Fig. 2 represents internal validation within Site A, Fig. 4 shows AUC values in the second health system, Site B, whose data were not used for training. AUCs using the best classifiers (RF) remain in the 0.67–0.70 range for all medications, with only modest change from Site A, providing support for the portability of these models. At this site, optimal model performance was achieved for venlafaxine with AUC of 0.70 [0.68–0.72], and poorest for paroxetine with AUC of 0.67 [0.65–0.70].
Model calibration and lift
Supplementary Figure 4 shows a lift histogram of discontinuation rate for different deciles of prescriptions sorted according to the LR or RF classifier, indicating goodness of fit in both the original health system and the replication site. The top risk decile identified by both classifiers was associated with treatment discontinuation for nearly half of the observed encounters (e.g., 43.97% and 49.41% for LR and RF, respectively), compared with an average discontinuation rate of 24.92%—i.e., lift was >2. Similarly, Supplementary Fig. 5 shows calibration curves for the LR and RF classifiers, indicating promising calibration in both the original network (Site A) and the replication network (Site B).
Discussion
In this investigation of 51,683 individuals with major depression across two academic medical center-based health systems, we identified predictive models that improve significantly on chance and on simple sociodemographic models in discriminating dropout risk by incorporating coded clinical features in LR and RF classifiers. Performance was only slightly diminished in a second health system, further supporting a lack of overfitting and the potential for portability. Models were also well-calibrated across both sites, with good correlation between predicted and observed discontinuation rates. We also observed that discrimination in these models was comparable among individuals with no prior treatment, one prior treatment, or two or more prior treatments; if anything, additional history tends to improve performance. Finally, we observed that discontinuation rates were greater among primary compared with specialty care settings, which likely reflects unmeasured differences in depression severity or other illness features, as multiple studies suggest that primary compared with specialty care for other diseases can yield similar outcomes (see, for example, Chai-Coetzer et al.10).
Our results are difficult to compare to prior work, including our own initial publication examining treatment-resistant depression in a large effectiveness study11. In particular, few studies have examined longitudinal health records and few have attempted to model dropout directly, despite the recognition that off-target effects of medications are useful in predicting clinical outcomes12,13. Other strategies aimed at personalized antidepressant treatment, including use of pharmacogenomic testing or other biomarkers, have yielded mixed results, among them a recent large negative study14. An important step in dissemination of any prediction model will be demonstration of improvement in outcomes using randomized, double-blind trials.
A key question is how models of this kind and accuracy might be applied to improve the care of patients with major depressive disorder; the models described here could have at least two possible applications. In the first case, a clinician is ready to prescribe a given medication based upon standard guidelines or algorithms. In this case, the risk for discontinuation predicted by the machine-learned model might help in prioritizing interventions aimed at retention in treatment and adherence, including making follow-up phone calls, deploying mobile applications to improve adherence, or simply scheduling an earlier return visit. That is, a clinician or case manager might see the predicted risk of failure to return and select from a range of strategies to increase follow-up. Alternatively, these models might be applied in settings where there are multiple reasonable next-step treatment options, in the form of a decision support tool that ranks such options in terms of predicted follow-up probability. Here, all other things being equal, the clinician might prefer the medication with the lowest risk of treatment discontinuation for that patient. Our models require no additional data collection, so they are straightforward to apply in real time at the point of care.
Beyond targeting treatments, these models may be useful in the design of a new generation of EHR-linked clinical trials15. For example, trials may be stratified by discontinuation risk, or interventions aimed at high-risk individuals could be studied in that subgroup16. The interpretability of machine-learned models would be particularly important in this regard, as it facilitates identification of modifiable risk factors specific to a given intervention.
Several important limitations merit consideration. The first reflects reliance on coded clinical data: undoubtedly, additional patient-level variables not captured in these data would further improve prediction. Most notably, the role of concomitant psychotherapy cannot be characterized; although such psychotherapy is captured in coded data and can be incorporated in prediction, the type of therapy being delivered cannot be determined. Models using coded data represent a starting point, a baseline to be improved upon. In addition, while the present study includes an independent replication site, both hospitals are located in the same region with overlapping catchment areas. Thus, it will be important to pursue further studies in other regions, including non-US health systems. An advantage of the code-only model is that it should be readily translatable to most US and European health systems.
We also note that while predictions are made for individual medications, we cannot exclude the possibility that some of the prediction is not treatment-specific per se. That is, some of the features may simply reflect overall dropout risk rather than risk with a given medication. More generally, these analyses do not seek to capture reasons for discontinuation, which are likely to be multi-factorial. They do not examine which adverse effects might contribute to dropout, or other relevant factors including cost, availability of transportation, and symptomatic worsening. We simply consider the face-valid question: which patients are most likely to receive a single antidepressant prescription, and then not return for follow-up? Undoubtedly, consideration of additional patient-level features could also improve prediction. As such, this work should be considered to complement, rather than replace, patient self-report and passive measures as outcome prediction tools.
With these limitations in mind, these results nonetheless provide a first step toward personalization of antidepressant treatment on the basis not of efficacy, but of tolerability. A key strength of the present study, beyond demonstrating portability, is a common model applicable to both primary and specialty care settings. Along with efforts to integrate other predictors, this work should encourage others to pursue modeling of this simple but important outcome. True personalization may be less likely to arrive with discovery of a single critical biomarker, but rather with incremental efforts to improve upon chance in the ability to stratify outcomes.
References
Cipriani, A. et al. Comparative efficacy and acceptability of 21 antidepressant drugs for the acute treatment of adults with major depressive disorder: a systematic review and network meta-analysis. Lancet 391, 1357–1366 (2018).
Warden, D., Rush, A. J., Trivedi, M. H., Fava, M. & Wisniewski, S. R. The STAR*D Project results: a comprehensive review of findings. Curr. Psychiatry Rep. 9, 449–459 (2007).
Jung, S.-H. et al. Factors affecting treatment discontinuation and treatment outcome in patients with schizophrenia in Korea: 10-year follow-up study. Psychiatry Investig. 8, 22–29 (2011).
Masand, P. S. Tolerability and adherence issues in antidepressant therapy. Clin. Ther. 25, 2289–2304 (2003).
Lieberman, J. A. et al. Effectiveness of antipsychotic drugs in patients with chronic schizophrenia. N. Engl. J. Med. 353, 1209–1223 (2005).
Perlis, R. H. et al. Using electronic medical records to enable large-scale studies in psychiatry: treatment resistant depression as a model. Psychol. Med. 42, 41–50 (2012).
Murphy, S. N. et al. Architecture of the open-source clinical research chart from Informatics for Integrating Biology and the Bedside. AMIA Annu. Symp. Proc. 548–552 (2007).
Pedregosa, F. et al. Scikit-learn: machine learning in python. J. Mach. Learn Res. 12, 2825–2830 (2011).
Steyerberg, E. W. et al. Assessing the performance of prediction models: a framework for some traditional and novel measures. Epidemiol. Camb. Mass 21, 128–138 (2010).
Chai-Coetzer, C. L. et al. Primary care vs specialist sleep center management of obstructive sleep apnea and daytime sleepiness and quality of life: a randomized trial. JAMA 309, 997–1004 (2013).
Perlis, R. H. A clinical risk stratification tool for predicting treatment resistance in major depressive disorder. Biol. Psychiatry 74, 7–14 (2013).
McCoy, Jr. T. H., Castro, V. M., Cagan, A., Roberson, A. M. & Perlis, R. H. Validation of a risk stratification tool for fall-related injury in a state-wide cohort. BMJ Open 7, e012189 (2017).
Castro, V. M. et al. Stratification of risk for hospital admissions for injury related to fall: cohort study. BMJ 349, g5863 (2014).
Greden, J. F. et al. Impact of pharmacogenomics on clinical outcomes in major depressive disorder in the GUIDED trial: a large, patient- and rater-blinded, randomized, controlled study. J. Psychiatr. Res. 111, 59–67 (2019).
Perlis, R. H., Fava, M. & McCoy, Jr. T. H. Can electronic health records revive central nervous system clinical trials? Mol. Psychiatry. https://doi.org/10.1038/s41380-018-0278-z (2018).
Perlis, R. H. Abandoning personalization to get to precision in the pharmacotherapy of depression. World Psychiatry 15, 228–235 (2016).
Acknowledgements
This study was funded by Oracle Labs, Harvard Center of Research on Computation and Society, the Harvard Data Science Initiative, and the National Institute of Mental Health (1R01MH106577-01). The funders had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the paper; or decision to submit the paper for publication.
Author information
Authors and Affiliations
Contributions
All authors have contributed meaningfully to this work and gave final approval to submit for publication. M.F.P. developed statistical models, conducted analysis and drafted the paper. T.H.M. contributed to study concept, design and critical revision of the paper. M.H. contributed to analysis and critical revision of the paper. R.H.P. conceived the study concept and design, aided in interpretation of results, and drafted the paper. F.D.V. conceived study design and contributed to development of statistical models, interpretation of results, and critical revision of the paper.
Corresponding authors
Ethics declarations
Conflict of interest
R.H.P. holds equity in Psy Therapeutics and Outermost Therapeutics; serves on the scientific advisory boards of Genomind and Takeda; and consults to RID Ventures. R.H.P. receives research funding from NIMH, NHLBI, NHGRI, and Telefonica Alfa. R.H.P. is an associate editor for JAMA Network Open. T.H.M. receives research funding from The Stanley Center at the Broad Institute, the Brain and Behavior Research Foundation, National Institute of Aging, and Telefonica Alfa. M.H. reports grants from Oracle Labs during the conduct of this study. The other authors have no disclosures to report. The authors thank Victor Castro for preparing the datamart and Partners Research Computing for computational resources.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pradier, M.F., McCoy Jr, T.H., Hughes, M. et al. Predicting treatment dropout after antidepressant initiation. Transl Psychiatry 10, 60 (2020). https://doi.org/10.1038/s41398-020-0716-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-0716-y
This article is cited by
-
A randomized sham-controlled trial of high-dosage accelerated intermittent theta burst rTMS in major depression: study protocol
BMC Psychiatry (2024)
-
Discontinuation of antidepressant treatment: a retrospective cohort study on more than 20,000 participants
Annals of General Psychiatry (2023)
-
Bayesian network analysis of antidepressant treatment trajectories
Scientific Reports (2023)
-
A Delphi-method-based consensus guideline for definition of treatment-resistant depression for clinical trials
Molecular Psychiatry (2022)
-
How machine-learning recommendations influence clinician treatment selections: the example of antidepressant selection
Translational Psychiatry (2021)