Abstract
Neuropsychiatric disorders are diagnosed based on behavioral criteria, which makes the diagnosis challenging. Objective biomarkers such as neuroimaging are needed, and when coupled with machine learning, can assist the diagnostic decision and increase its reliability. Sixty-four schizophrenia, 36 autism spectrum disorder (ASD), and 106 typically developing individuals were analyzed. FreeSurfer was used to obtain the data from the participant’s brain scans. Six classifiers were utilized to classify the subjects. Subsequently, 26 ultra-high risk for psychosis (UHR) and 17 first-episode psychosis (FEP) subjects were run through the trained classifiers. Lastly, the classifiers’ output of the patient groups was correlated with their clinical severity. All six classifiers performed relatively well to distinguish the subject groups, especially support vector machine (SVM) and Logistic regression (LR). Cortical thickness and subcortical volume feature groups were most useful for the classification. LR and SVM were highly consistent with clinical indices of ASD. When UHR and FEP groups were run with the trained classifiers, majority of the cases were classified as schizophrenia, none as ASD. Overall, SVM and LR were the best performing classifiers. Cortical thickness and subcortical volume were most useful for the classification, compared to surface area. LR, SVM, and DT’s output were clinically informative. The trained classifiers were able to help predict the diagnostic category of both UHR and FEP Individuals.
Similar content being viewed by others
Introduction
The current diagnostic model in psychiatry, while the best available, is not highly reliable due to three main factors: patient heterogeneity (i.e., patient’s psychological state, their ability to provide reliable information, and differences in clinical presentation), clinician inconsistency (i.e., different opinions on the same case) and nomenclature inadequacy1,2. As nosology is a key aspect of psychiatry, on which patient assessment and treatment options are based, it would be helpful to have a layer of appraisal centered around objective evaluations to establish a more reliable classification decision. Neuroimaging is one objective measure that might facilitate the diagnostic process, yet it is not currently used in aiding the diagnostic decision in psychiatry, despite much interest.
Machine learning uses statistical methods to find patterns in large amount of data. The learning process starts with the data at hand and improves autonomously over time. Recent advances in machine learning, combined with neuroimaging techniques, are capable of assessing differences in local morphological features of various brain subregions to elucidate novel disorder-related brain patterns3,4,5. Such patterns can be used by computational models to build classifiers for the purpose of aiding the diagnostic decision. Several studies were conducted using similar models to classify patients into their respective diagnostic category for both autism spectrum disorder (ASD)6,7,8,9,10 and schizophrenia11,12,13,14,15,16. However, the majority of these studies focused on distinguishing between typically developing (TD) individuals and those with psychiatric disorders4,11,17. This classification is important for identifying brain patterns that are different from what is considered typical, but does not inform about the variations between different patient groups which is essential for a reliable psychiatric nosology and understanding the overlap between different neuropsychiatric disorders18. Using both approaches, on the other hand, provides a clearer picture about the nature of the disorders and how they differ from one another. Moreover, investigating how the classifiers that are pretrained on distinct clinical phenotypes would perform on “intermediate phenotypes” or early disease states, aids in the quantification of disease progression, predicting outcome, and understanding the intersection between different nosological, phenotypic and neurobiological continua.
Classifying psychiatric disorders, especially schizophrenia and ASD, has been performed using different neuroimaging modalities. In structural magnetic resonance imaging (MRI), several studies classified patients based on a single modality, such as voxel based morphometry19,20, cortical thickness3,4, or surface morphological measures21. Moreover, most of these studies use a single classifier to perform the classification15,22,23,24,25,26,27. Few have conducted the analysis using multiple classifiers, which is important to avoid bias towards a particular classifier28. To our knowledge, there is no study yet that has compared several brain indices, such as cortical thickness, surface area, and subcortical volume using multiple machine learning algorithms between schizophrenia and ASD, and assessed the performance of these classifiers on at-risk and early disease stage patients.
Thus, the current study was aimed to (i) construct and compare classifiers that can distinguish between individuals with schizophrenia, ASD, and TD based on their MRI scans, (ii) uncover the most important brain feature groups contributing to the classification, (iii) assess the consistency of the classifiers with clinical severity, and (iv) predict the diagnostic category of ultra-high risk for psychosis (UHR) and first-episode psychosis (FEP) subjects using the classifiers pretrained on a combination of ASD, TD, and/or schizophrenia data.
Methods
Participants
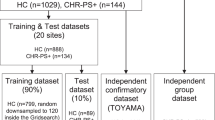
The data of 131 schizophrenia spectrum (26 UHR, 25 FEP, and 80 schizophrenia), 45 high functioning ASD, and 125 TD individuals were included in this study. After assessing the T1-weighted images, 97 schizophrenia spectrum (26 UHR, 17 FEP, and 64 schizophrenia), 36 ASD, and 106 TD scans were analyzed. The ages of the subjects ranged between 14 and 60 years old (y.o.) for schizophrenia (mean ± SD: 29.8 ± 10.1), 20–44 y.o. for ASD (mean ± SD: 30.1 ± 6.7), 16–60 y.o., for TD (mean ± SD: 29.1 ± 6.0), 16–28 y.o. for UHR (mean ± SD: 20.9 ± 3.1), and 17–34 y.o. for FEP (mean ± SD: 23.5 ± 5.2). Individuals with ASD were all males, those with schizophrenia, TD, UHR, and FEP were of mixed sex. The participants were mostly right-handed (ASD, right (R): 28/left (L): 3/mixed (M): 5; TD R: 104/L: 0/M: 2; schizophrenia R: 55/L: 1/M: 8, UHR R: 23/L: 0/M: 3; FEP R: 16/L: 0/M: 1) (Table 1). All participants are ethnically Japanese and were recruited at The University of Tokyo Hospital. The diagnostic criteria for schizophrenia, and ASD and inclusion and exclusion criteria can be found elsewhere29,30,31, additionally, a comprehensive explanation was added to the supplemental materials. The ethical review board of The University of Tokyo Hospital approved this study (Nos. 397 and 2226). All participants gave written informed consent before their participation.
MRI acquisition
The structural MRI images for all of the subjects were acquired using a 3.0-T MRI scanner (GeneralElectric Healthcare, Signa HDxt. v14.0, Milwaukee, Wisconsin), with a standard 8-channel head coil for signal reception. The T1-weighted structural brain images were collected using a three-dimensional Fourier-transform fast-spoiled gradient recalled acquisition with steady state, because it affords excellent contrast between the gray and white matter (repetition time = 6.80 ms, echo time = 1.94 ms, flip angle = 20°, slice thickness = 1.0 mm, field of view = 240 mm, matrix = 256 × 256, number of axial slices = 176). The participant’s head was fixed with foam pads to minimize movement. A trained neuroradiologist (O.A., W.G., or H.T.) checked the scans and found no gross abnormalities in any of the subjects. Magnetic field inhomogeneity in our scanner was monitored with daily quality control. In order to ensure that the images were of appropriate quality, the scans of all subjects were visually examined, slice by slice across all orthogonal directions before any image processing step. The scans were performed between the year 2010 and 2013.
Data processing
Imaging
The structural MRI scans from all subjects were processed with the same procedure, using the FreeSurfer image analysis suite v.6.0 (http://surfer.nmr.mgh.harvard.edu/). This processing step was performed using recon-all pipeline with the default settings. The details of this procedure32,33,34, can be found in the supplemental materials. Even though the FreeSurfer morphometric procedures have been shown to be accurate and reliable35, we implemented additional quality assurance steps. Enhancing NeuroImaging Genetics through Meta-Analysis wrapper script (http://enigma.ini.usc.edu/protocols/imaging-protocols/), was employed after the FreeSurfer processing steps for quality assurance. Subsequently, a visual check was performed on the images to investigate whether there was any sort of abnormality, and manual edits were applied when necessary. When edits were not possible, the scans were discarded from the study. The FreeSurfer output, i.e., cortical thickness (150 regions), surface area (150 regions), and subcortical volume (36 regions) were later used as feature groups in the classification models described in detail in the supplemental materials (Table S1).
Quality control and feature engineering
First, the data were checked for missing values, and subjects with any missing value, were excluded from the analysis. Second, the outliers of each group were detected though the interquartile range method and were removed before the start of the analysis. The total number of excluded subjects throughout the study preprocessing steps were 9 ASD, 16 schizophrenia, 8 FEP, and 19 TD subjects. The features included in each group can be found as a table in the Supplemental material (Table S1).
Before the features were used in the classification process, they were standardized using StandardScalar, part of scikit-learn (SKLearn), by removing the mean and scaling to unit variance. StandardScalar was first applied on the training data set and was then reapplied later with the same transformation on the testing set. These sets were randomly selected based on the train/test split function in SKLearn (See “Classification architecture”).
Classification architecture
All the analyses were implemented using Python v2.7 available at (http://www.python.org) and SkLearn v.0.19.136, a machine learning library for Python. The data was split into training (80%) and testing (20%) sets using the train/test split function in SKLearn. The test set was not used until the very end to assess the performance of the classifiers. StandardScalar was then applied as described in the “Quality control and feature engineering” section. Furthermore, dimensionality reduction was performed using principle component analysis (PCA). PCA utilizes linear dimensionality reduction by using the data’s singular value decomposition to project it to a lower dimensional space37. Moreover, as the features are expected to be collinear, PCA also helps to overcome this multicollinearity problem by producing orthogonal features made from the linear combination of the input vectors, i.e., principle components. PCA was performed inside a pipeline which allows setting different parameters and combines several steps that can be cross validated together. This pipeline was implemented inside GridSearchCV, with a tenfold cross-validation, which performs an exhaustive search over the assigned parameters to construct the best possible classifier using a combination of optimal parameter values. Fine-tuning the classifiers entailed using different parameter combinations inside GridSearchCV. The parameters producing a classification with the best performance were chosen, and the model was fit to the entire training set using those parameter values. All the classifiers utilized in this study were fine-tuned and had the same overall architecture. As the sample size of each group is unbalanced, we used the “class_weight” parameter and set it to “balanced”, to ensure that we had more balanced classes. This parameter option works by weighing classes inversely proportional to their frequency. The classifiers used in our study are logistic regression (LR), support vector machine (SVM), random forest (RF), adaptive boosting (AdaB), decision tree (DT), and k-nearest neighbor (kNN). Several classifiers have been selected to avoid bias toward the use of a particular classifier28, and to compare their performance on our data. The classifiers were run with several subjects and feature group combinations. Only those classifiers that showed relatively high accuracy, with no signs of overfitting, were reported in the manuscript. Four classification runs were performed with each classifier; one multiclass classification (schizophrenia/ASD/TD), and three binary classifications (schizophrenia/ASD, ASD/TD, and schizophrenia/TD). The code is available upon request.
Classifiers
Logistic regression
The logistic function, a core part of the LR, is a sigmoid function that can take a real number and transforms it into a value between 0 and 1, producing a “S”-shaped curve. LR uses the maximum likelihood estimation method to estimate the model coefficients. It is typically used for binary classification problems, but a multiclass classification is also possible, for example, through the one-vs.-rest scheme.
Decision tree
The DT method uses a non-parametric supervised learning approach to solve both regression and classification problems. DT uses a tree representation where each test on an attribute is represented by an internal node, and each leaf denotes a class label. Thus, DT can learn certain decision rules inferred from the features used to build a model that predicts the value of the target variable.
Random forest
RF is an ensemble learning method, consisting of several DTs, that can be used for classification and regression. Ensemble methods are algorithms that incorporate more than one type of algorithm. RF works by constructing a number of DT classifiers which learn and make predictions independently, and outputs a combined single prediction that is the same or better than the output made by the previously constructed DT classifiers.
SVM classifier
SVM is a supervised discriminative classification method that uses the features belonging to several labeled training examples to construct hyperplanes, high-dimensional planes, for optimally separating the data into different groups. The implementation of the C- support vector classification used in our study is based on the library for SVMs (libsvm).
Adaptive boosting
AdaBoost is an ensemble boosting algorithm that combines a set of “weak” classifiers into a weighted sum to create a stronger more accurate “boosted” classifier. AdaBoost starts by fitting a classifier on the dataset, and then fits the same version of that classifier on the same dataset where the weights of the misclassified instances are modified so that the next classifier is improved to work on the more challenging instances.
k-Nearest neighbor
kNN is an instance based learning algorithm, and yet another non-parametric method used for classification and regression. The input consists of a feature space with k closest training examples, where k is assigned by the user, and the output is a class membership. An object is assigned to a class that is most common amongst its nearest neighbors, as the nearest neighbors contribute more to the average than the distant ones.
Classification performance metrics
The chosen indicator of proper classification was not solely based on accuracy. Thus, we further calculated the confusion matrix, recall score (i.e., sensitivity), precision score, and F1/F2 scores. The metrics are described in detail in the supplemental materials.
Classifier consistency with clinical severity
After the classification was complete, the correctly/incorrectly classified (CC)/(IC) instances from both patient groups were extracted, binarized and correlated with their clinical scores. Their clinical scores were assigned accordingly: autism diagnostic interview-revised (ADI-R) sub-scale (social, communication, and restricted and repetitive behavior (RRB)), and autism quotient (AQ) subscale (social skills (SS), attention switching (AS), attention to detail (AD), communication, and imagination) for the ASD group, and the Positive and negative symptom scale (PANSS), which includes positive symptoms, negative symptoms, and general psychopathology for the schizophrenia and FEP groups, the total scores of these scales were also included. The CC and IC classes were coded as “1” and “0”. The classifiers were also coded “1” through “6” representing LR, SVM, RF, AdaB, DT, and kNN. Then, for each classifier a Point-Biserial correlation was run using Statistical Package for Social Sciences (SPSS) v.20. For the ASD group, the ADI-R and AQ sub- and total scores were correlated with the binarized CC and IC instances. The same procedure was conducted for schizophrenia replacing the ADI-R and AQ scores with PANSS sub- and total-scores. The statistical significance threshold for this study was set at P < 0.05. Bonferroni correction was used to correct for multiple comparisons.
UHR and FEP
After training the classifiers, we tested their performance on a group of 26 individuals with UHR and 17 with FEP. We chose the best performing multiclass classifier, and binary classifier (schizophrenia/TD) for that purpose out of all the runs. The recall score was considered here as it represents the ability of the classifier to find the positive samples. The resulting classes were then binarized and a Point-Biserial correlation, evaluating the association of the classification with the PANSS and the global assessment of functioning (GAF) data 1 year or more after the time of the first MRI scan was performed.
To assess whether medication affected the classification, we ran two independent sample t-tests (one sided) on the antipsychotic dose taken by the FEP and UHR subjects that were classified into schizophrenia or TD using the multiclass classifier.
Results
Classifiers
In the multiclass classification, the best results were produced using the cortical thickness feature group, especially using the LR classifier, with an overall accuracy of 69.0% (Table 2).
In the binary classification model between schizophrenia and ASD, the majority of the classifiers performed well with several feature groups. Classification using the whole brain feature group was best in both SVM and kNN with an accuracy of 75% for both, as well as LR with slightly lower accuracy at 70%. Using the subcortical volume feature group, all of the classifiers showed relatively good accuracy; LR, 75%, SVM, 80%, RF 75%, AdaBoost 75%, and kNN 85%. The surface area feature group did the worst overall, although only LR had good results with 70% accuracy. In classifying based on cortical thickness, AdaBoost performed the best, at 85% accuracy, followed by LR (80%), and SVM (75%) (Table 3).
In addition, to further investigate the performance of the classifiers on the patient population versus the TD group, we performed the following classifications between; ASD and TD, and schizophrenia and TD. In the ASD and TD group, only SVM was able to perform well with an accuracy of 75.8% using the whole brain feature group. In the subcortical volume, both LR, accuracy 72.4% and SVM, accuracy 89.6% showed good performance. Lastly, in cortical thickness, DT had the best performance with an accuracy of 75.8% (Table 4).
While in the schizophrenia and TD group, LR performed better than all the other classifiers using the whole brain feature group with 70.5% accuracy. While using subcortical volume, LR accuracy was 64.7%, SVM 67.6%, RF 76.4%, and AdaBoost 73.5%. Lastly, using cortical thickness, only LR, accuracy 67.6% and DT, accuracy 70.5% performed well (Table 5). All of the classifiers’ results, despite accuracy level and overfitting status, in addition to the full metric data such as recall score, F1/F2 scores, and others can be seen in the supplementary material (Fig. S1, Tables S2–S9).
Classifiers and clinical severity
The results of the Point-Biserial correlation showed that LR, SVM, and DT were highly consistent with the clinical severity of the patients with ASD. LR in ASD for example, showed high consistency with ADI-R’s RRB (F(1,46) = 7.91, P corrected = 0.021; CC mean = 5.5, SD = 2.0, IC mean = 3.6, SD = 2.4), and AQ’s AD (F(1,46) = 8.45, P corrected = 0.03; CC mean = 7.2, SD = 2.3, IC mean = 5.2, SD = 1.6). SVM also showed consistency with ADI-R’s communication (F(1,39) = 7.73, P corrected = 0.024; CC mean = 12.9, SD = 3.1, IC mean = 10.4, SD = 2.0), and RRB (F(1,39) = 11.42, P corrected = 0.006; CC mean = 5.6, SD = 2.1, IC mean = 3.4, SD = 1.6), and AQ’s imagination (F(1,39) = 10.41, P corrected = 0.015; CC mean = 6.6, SD = 8.2, IC mean = 8.2, SD = 1.5). Lastly, DT was consistent with ADI-R’s social domain (F(1,14) = 8.23, P = 0.012–0.036; CC mean = 10.9, SD = 5.5, IC mean = 18.5, SD = 4.2). In schizophrenia, no correlation survived after correcting for multiple comparisons.
Classifiers and independent samples
The UHR group that was classified using the multiclass classifier resulted in 15 schizophrenia (57.6%), and 11 TD subjects, but none were classified as ASD. The Point-Biserial correlation showed no relationship with either PANSS or GAF data after correcting for multiple comparison. When run with the schizophrenia/TD subcortical classifier, 96.1% of the sample were classified as schizophrenia.
FEP subjects were also classified using the same procedure as the UHR. 70% of the FEP subjects were classified as schizophrenia by the multiclass classifier, 30% as TD, while none as ASD. No correlation was found with either PANSS or GAF data. When run with the schizophrenia/TD subcortical classifier, 100% of the samples were classified as schizophrenia.
For UHR and FEP, we found no significant difference in antipsychotic dose between the patients classified into schizophrenia and those into TD using the multiclass classifier.
Discussion
To our knowledge, this is the first study to compare cortical thickness, subcortical volume and surface area using multiple machine learning classifiers between schizophrenia, ASD and TD, and investigate how these trained classifiers extrapolate to UHR and FEP. Our findings indicate that, overall, SVM and LR, were the best performing classifiers, producing high accuracy with least overfitting. Second, cortical thickness and subcortical volume were most useful for the classification, compared to surface area. Third, the LR, SVM, and DT’s output were clinically informative as they were consistent with the patients’ clinical severity. Lastly, we showed that a selection of the trained classifiers was able to predict the diagnostic category of UHR and FEP Individuals.
SVM showed a good overall performance, which is consistent with the published literature24,25,38,39,40,41,42,43,44. It is by far the most utilized machine learning classifier in the field of neuroimaging5,45,46. SVM has been used in several studies involving both ASD41,42,43,44 and schizophrenia24,25,38,39,40,47,48. Part of its strength comes from the ability to make inferences at the level of an individual, which is important in a sample of patients with neuropsychiatric disorders having within group heterogeneity5. Moreover, the multivariate nature of SVM allows it to reveal subtle differences in the brain that would otherwise not be detectable through univariate group comparisons5,49, which helps its performance.
LR was the only classifier that showed no overfitting in the multiclass classification model. In binary classification, it showed a similar performance to SVM, with good overall accuracy. LR has been used in many neuroimaging studies as well9,42,50,51. However, we were unable to find structural MRI studies using LR to classify individuals with ASD and TD. Most of the studies that are published use resting-state functional MRI, and show a similar overall classification accuracy as our study42,50. One study classifying individuals with ASD and TD, reported similar results to ours, in which LR and SVM were the best performing classifiers amongst those used (LR, RF, kNN, SVM, linear discriminate analysis, and Naïve Bayes)9. In schizophrenia, on the other hand, Greenstein et al.3 showed that LR was able to classify schizophrenia subjects with a 73.6% accuracy using 74 anatomical brain sub-regions.
Our results also show that RF, DT, kNN, and AdaB did have high performance, at least in specific runs. These classifiers were shown to be useful in several neuroimaging studies of ASD and schizophrenia3,4,7,9,52.
As mentioned previously, cortical thickness and subcortical volume performed better than surface area. Structural volume and cortical thickness features have both shown high accuracy classification in the literature53. A previous study compared their classification performance between individuals with ASD and TD, and found that thickness-based diagnostic models outperformed those that are based on volume in most classifiers28. In our case, the performance of these feature groups was comparable. Another study, by Katuwal et al., showed that surface area performed worse than subcortical volume, which is consistent with our results, though they also showed that surface area performed better than cortical thickness54, which is different from what we report in this study. Cortical thickness’s high overall performance signifies the presence of distinct cortical morphological features that are unique to each diagnostic group. The brain surface area’s stability in adults, where the majority of changes such as neural stem cell proliferation and migration happen during early embryonic development55, compared to that of cortical thickness, where early developmental changes continue into adulthood56, might have contributed to more distinguishable features in cortical thickness than surface area, which was revealed consistently in the performance of all the classifiers used in our study. A previous study comparing different psychiatric disorders including ASD and schizophrenia found that there is more divergence between disorders in cortical thickness than surface area57, which would be another reason why cortical thickness performed better than surface area.
In the same study, by Park et al.57, they demonstrate that ASD shows a trend toward an increase in cortical thickness while schizophrenia towards cortical thinning, this might explain why our results exhibited higher performance distinguishing between ASD/TD, than schizophrenia/TD. Using the multiclass classifier, cortical thickness was the only feature group that was able to distinguish between all three patient groups. This is noteworthy as it suggests that cortical thickness might hold information valuable for distinguishing between schizophrenia and ASD, and that the overlap in their symptoms might be less explained by cortical morphological features. Lastly, the results from the whole brain feature group shows that integrating different modalities doesn’t always improve the overall accuracy58,59, as the combined features, whole brain, did not perform better than the separate features, e.g., cortical thickness.
Most of the classifiers’ output showed an association with the clinical indices of ASD, especially LR and SVM. LR and SVM were highly associated with both ADI-R and AQ, while DT showed an association with ADI-R only. As for schizophrenia, only DT showed an association with the PANSS’s negative symptoms, general symptoms and total score, however this association did not survive Bonferroni correction. To our knowledge, there are no published studies that have assessed this in schizophrenia or ASD.
The early phase of the disease, such as in the UHR as well as FEP, is an important period that can have an outstanding influence on disorder progression60. Early intervention in both has been previously associated with better outcomes60,61. The multiclass classifier was run on 26 individuals with UHR and 17 with FEP. The classifier separated the UHR group into schizophrenia and TD, but not ASD. While when the schizophrenia/TD classifier was used, almost all of the subjects (96%) were classified into the schizophrenic group. The results for FEP were very similar to those in the UHR group. It is interesting that even at these early stages in disease progression, UHR and FEP have such structural brain similarities with schizophrenia. Collectively, this mean that UHR and more strongly FEP, have shared structural neurobiological patterns with schizophrenia, but not with autism. This method was used in a previous study, but showed modest generalization when a classifier, that was trained on schizophrenia and TD, was used to classify individuals with FEP62. It is plausible that the schizophrenia/TD classifier categorized FEP individuals in the schizophrenia group more than the UHR, since the FEP individuals already had their first psychotic episode. These results shed light on the importance of brain structure in understanding disease progression, especially whether or not the patients had their first psychotic episode.
In our sample, the antipsychotic dose that might have been associated with cortical thickness alterations was not predicted by the multiclass classifier. We could not conduct the same analysis using the schizophrenia/TD classifier, as there are not enough samples classified as TD. Given the variability in dose as well as the modest sample size, we are unable to provide a measurably reliable answer on whether, in general, this would influence the classification.
In summary, we found that SVM and LR were the best performing classifiers. Cortical thickness and subcortical volume based classification had better performance across different diagnostic labels and classifiers than surface area. LR, SVM, and DT were consistent with clinical severity of the patients. UHR and FEP show similar neurobiological pattern as schizophrenia. Our findings provide new knowledge about the best performing classifiers between individuals with schizophrenia, ASD, and TD, and reveal the most informative brain feature groups that contribute to the classification. The results also reveal the clinical relevance of these classifiers, in addition to their importance in predicting the diagnostic category. Lastly, they shed light on the structural brain similarities between FEP, UHR and schizophrenia. The knowledge gained from these feature groups can be extrapolated to their use as biomarkers for future targeted therapeutic interventions as well as predicting patients’ disease trajectory.
Parts of the discussion compared our findings to those that are already published, however it should be taken into consideration that comparisons such as accuracies, and class predictions across studies is not ideal as it may be affected by the number of instances, classifiers used, quality and type of images, feature engineering, and other factors45.
Limitations
This study has some limitations. First, the ASD subjects were all males. This, however, did not seem to affect the classification, as we did not see that individuals with ASD were particularly classified better than the other groups. Second, the present study has a modest sample size, nonetheless comparable to other published studies. Third, as few of the ASD subjects, several of the UHR and FEP, and all of the schizophrenia subjects were medicated, this might have affected the results.
References
Aboraya, A., Rankin, E., France, C., El-Missiry, A. & John, C. The reliability of psychiatric diagnosis revisited: the clinician’s guide to improve the reliability of psychiatric diagnosis. Psychiatry 3, 41–50 (2006).
Ward, C. H., Beck, A. T., Mendelson, M., Mock, J. E. & Erbaugh, J. K. The psychiatric nomenclature. Reasons for diagnostic disagreement. Arch. Gen. Psychiatry 7, 198–205 (1962).
Greenstein, D., Malley, J. D., Weisinger, B., Clasen, L. & Gogtay, N. Using multivariate machine learning methods and structural MRI to classify childhood onset schizophrenia and healthy controls. Front. Psychiatry 3, 53 (2012).
Libero, L. E., DeRamus, T. P., Lahti, A. C., Deshpande, G. & Kana, R. K. Multimodal neuroimaging based classification of autism spectrum disorder using anatomical, neurochemical, and white matter correlates. Cortex 66, 46–59 (2015).
Orru, G., Pettersson-Yeo, W., Marquand, A. F., Sartori, G. & Mechelli, A. Using Support Vector Machine to identify imaging biomarkers of neurological and psychiatric disease: a critical review. Neurosci. Biobehav. Rev. 36, 1140–1152 (2012).
Gori, I. et al. Gray matter alterations in young children with autism spectrum disorders: comparing morphometry at the voxel and regional level. J. Neuroimaging 25, 866–874 (2015).
Chen, C. P. et al. Diagnostic classification of intrinsic functional connectivity highlights somatosensory, default mode, and visual regions in autism. NeuroImage Clin. 8, 238–245 (2015).
Iidaka, T. Resting state functional magnetic resonance imaging and neural network classified autism and control. Cortex 63, 55–67 (2015).
Plitt, M., Barnes, K. A. & Martin, A. Functional connectivity classification of autism identifies highly predictive brain features but falls short of biomarker standards. NeuroImage Clin. 7, 359–366 (2015).
Just, M. A., Cherkassky, V. L., Buchweitz, A., Keller, T. A. & Mitchell, T. M. Identifying autism from neural representations of social interactions: neurocognitive markers of autism. PloS ONE 9, e113879 (2014).
Janousova, E., Schwarz, D. & Kasparek, T. Combining various types of classifiers and features extracted from magnetic resonance imaging data in schizophrenia recognition. Psychiatry Res. 232, 237–249 (2015).
Pina-Camacho, L. et al. Predictors of schizophrenia spectrum disorders in early-onset first episodes of psychosis: a support vector machine model. Eur. Child Adolesc. Psychiatry 24, 427–440 (2015).
Radulescu, E. et al. Grey-matter texture abnormalities and reduced hippocampal volume are distinguishing features of schizophrenia. Psychiatry Res. 223, 179–186 (2014).
Zhang, T. & Davatzikos, C. Optimally-Discriminative Voxel-Based Morphometry significantly increases the ability to detect group differences in schizophrenia, mild cognitive impairment, and Alzheimer’s disease. NeuroImage 79, 94–110 (2013).
Zanetti, M. V. et al. Neuroanatomical pattern classification in a population-based sample of first-episode schizophrenia. Prog. Neuropsychopharmacol. Biol. Psychiatry 43, 116–125 (2013).
Takayanagi, Y. et al. Classification of first-episode schizophrenia patients and healthy subjects by automated MRI measures of regional brain volume and cortical thickness. PloS ONE 6, e21047 (2011).
Granziera, C. et al. A multi-contrast MRI study of microstructural brain damage in patients with mild cognitive impairment. NeuroImage Clin. 8, 631–639 (2015).
Krystal, J. H. & State, M. W. Psychiatric disorders: diagnosis to therapy. Cell 1, 201–214 (2014).
Nieuwenhuis, M. et al. Classification of schizophrenia patients and healthy controls from structural MRI scans in two large independent samples. NeuroImage 61, 606–612 (2012).
Calderoni, S. et al. Female children with autism spectrum disorder: an insight from mass-univariate and pattern classification analyses. NeuroImage 59, 1013–1022 (2012).
Bansal, R. et al. Anatomical brain images alone can accurately diagnose chronic neuropsychiatric illnesses. PloS ONE 7, e50698 (2012).
Yu, Y., Shen, H., Zeng, L. L., Ma, Q. & Hu, D. Convergent and divergent functional connectivity patterns in schizophrenia and depression. PloS ONE 8, e68250 (2013).
Fischl, B. et al. Automatically parcellating the human cerebral cortex. Cereb. Cortex 14, 11–22 (2004).
Tang, Y., Wang, L., Cao, F. & Tan, L. Identify schizophrenia using resting-state functional connectivity: an exploratory research and analysis. Biomed. Eng. Online 11, 50 (2012).
Su, L., Wang, L., Shen, H., Feng, G. & Hu, D. Discriminative analysis of non-linear brain connectivity in schizophrenia: an fMRI Study. Front. Hum. Neurosci. 7, 702 (2013).
Bassett, D. S., Nelson, B. G., Mueller, B. A., Camchong, J. & Lim, K. O. Altered resting state complexity in schizophrenia. NeuroImage 59, 2196–2207 (2012).
Anticevic, A. et al. Characterizing thalamo-cortical disturbances in schizophrenia and bipolar illness. Cereb. Cortex 24, 3116–3130 (2014).
Jiao, Y. et al. Predictive models of autism spectrum disorder based on brain regional cortical thickness. NeuroImage 50, 589–599 (2010).
Yassin, W. et al. Paternal age contribution to brain white matter aberrations in autism spectrum disorder. Psychiatry Clin. Neurosci. 73, 649–659 (2019).
Watanabe, T. et al. Diminished medial prefrontal activity behind autistic social judgments of incongruent information. PloS ONE 7, e39561 (2012).
Iwashiro, N. et al. Localized gray matter volume reductions in the pars triangularis of the inferior frontal gyrus in individuals at clinical high-risk for psychosis and first episode for schizophrenia. Schizophrenia Res. 137, 124–131 (2012).
Fischl, B., Liu, A. & Dale, A. M. Automated manifold surgery: constructing geometrically accurate and topologically correct models of the human cerebral cortex. IEEE Trans. Med. Imaging 20, 70–80 (2001).
Fischl, B. et al. Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33, 341–355 (2002).
Fischl, B. et al. Sequence-independent segmentation of magnetic resonance images. NeuroImage 23, S69–S84 (2004).
Reuter, M., Schmansky, N. J., Rosas, H. D. & Fischl, B. Within-subject template estimation for unbiased longitudinal image analysis. NeuroImage 61, 1402–1418 (2012).
Pedregosa, F., Varoquaux, G., Gramfort, A., Michel, V. & Thirion, B. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Halko N, Martinsson PG, Tropp JA. Finding structure with randomness: probabilistic algorithms for constructing approximate matrix decompositions. SIAM Rev 2, 217–288 (2010). https://doi.org/10.1137/090771806?mobileUi=0&.
Costafreda, S. G. et al. Pattern of neural responses to verbal fluency shows diagnostic specificity for schizophrenia and bipolar disorder. BMC Psychiatry 11, 18 (2011).
Fekete, T. et al. Combining classification with fMRI-derived complex network measures for potential neurodiagnostics. PloS ONE 8, e62867 (2013).
Fan, Y., Shen, D., Gur, R. C., Gur, R. E. & Davatzikos, C. COMPARE: classification of morphological patterns using adaptive regional elements. IEEE Trans. Med. Imaging 26, 93–105 (2007).
Deshpande, G., Libero, L. E., Sreenivasan, K. R., Deshpande, H. D. & Kana, R. K. Identification of neural connectivity signatures of autism using machine learning. Front. Hum. Neurosci. 7, 670 (2013).
Uddin, L. Q. et al. Salience network-based classification and prediction of symptom severity in children with autism. JAMA Psychiatry 70, 869–879 (2013).
Ecker, C. et al. Brain anatomy and its relationship to behavior in adults with autism spectrum disorder: a multicenter magnetic resonance imaging study. Arch. Gen. Psychiatry 69, 195–209 (2012).
Segovia, F. et al. Identifying endophenotypes of autism: a multivariate approach. Front. Comput. Neurosci. 8, 60 (2014).
Vapnik, V. & Chapelle, O. Bounds on error expectation for support vector machines. Neural Comput. 12, 2013–2036 (2000).
Arbabshirani, M. R., Plis, S., Sui, J. & Calhoun, V. D. Single subject prediction of brain disorders in neuroimaging: promises and pitfalls. NeuroImage 145, 137–165 (2017).
Gould, I. C. et al. Multivariate neuroanatomical classification of cognitive subtypes in schizophrenia: a support vector machine learning approach. NeuroImage Clin. 6, 229–236 (2014).
Castellani, U. et al. Classification of schizophrenia using feature-based morphometry. J. Neural Transm. 119, 395–404 (2012).
Davatzikos, C. et al. Classifying spatial patterns of brain activity with machine learning methods: application to lie detection. NeuroImage 28, 663–668 (2005).
Murdaugh, D. L. et al. Differential deactivation during mentalizing and classification of autism based on default mode network connectivity. PloS ONE 7, e50064 (2012).
Parikh, M. N., Li, H. & He, L. Enhancing diagnosis of autism with optimized machine learning models and personal characteristic data. Front. Comput. Neurosci. 13, 9 (2019).
Venkataraman, A., Whitford, T. J., Westin, C. F., Golland, P. & Kubicki, M. Whole brain resting state functional connectivity abnormalities in schizophrenia. Schizophrenia Res. 139, 7–12 (2012).
Singh, V., Mukherjee, L. & Chung, M. K. Cortical surface thickness as a classifier: boosting for autism classification. Med. Image Comput. Assist. Interv. 11, 999–1007 (2008).
Katuwal GJ, Baum SA, Michael AM. Early brain imaging can predict autism: application of machine learning to a clinical imaging archive. bioRxiv 471169 (2018). https://www.biorxiv.org/content/10.1101/471169v1.
Rafik, P. Evolution of the neocortex: a perspective from developmental biology. Nat. Rev. Neurosci. 10, 724–735 (2009).
Shaw, P. et al. Neurodevelopmental trajectories of the human cerebral cortex. J. Neurosci. 14, 3586–3594 (2008).
Park, M. T. et al. Neuroanatomical phenotypes in mental illness: identifying convergent and divergent cortical phenotypes across autism, ADHD and schizophrenia. J. Psychiatry Neurosci. 3, 201–212 (2018).
Yang, H., Liu, J., Sui, J., Pearlson, G. & Calhoun, V. D. A hybrid machine learning method for fusing fMRI and genetic data: combining both improves classification of schizophrenia. Front. Hum. Neurosci. 4, 192 (2010).
Vemuri, P. et al. Alzheimer’s disease diagnosis in individual subjects using structural MR images: validation studies. NeuroImage 39, 1186–1197 (2008).
Reading, B. & Birchwood, M. Early intervention in psychosis, rationale and evidence for effectiveness. Dis. Manag. Health Outcomes 13, 53–63 (2005).
Yung, A. R. et al. Psychosis prediction: 12-month follow up of a high-risk (“prodromal”) group. Schizophrenia Res. 60, 21–32 (2003).
Pinaya, W. H. et al. Using deep belief network modelling to characterize differences in brain morphometry in schizophrenia. Sci. Rep. 6, 38897 (2016).
Acknowledgements
We thank Dr. Charles Yokoyama for editing a version of the manuscript and Mr. Dimitris Katsios for checking a version of the code. This work was supported by Grants-in-Aid for Scientific Research (KAKENHI), No. 26670535 to [H.Y.], and No. 19H0357 to [S.K.]; and in part by AMED under Grant Nos. JP19dm0307001, JP19dm0307004, and JP19dm0207069. This study was also supported by UTokyo Center for Integrative Science of Human Behavior (CiSHuB) and the world premier international- International Research Center for Neurointelligence (WPI-IRCN) and from the Japan Society for the Promotion of Science; the Center of Innovation Program and Core Research for Evolutional Science and Technology from Japan Science and Technology Agency (JST); and the Strategic Research Program for Brain Sciences from Japan Agency for Medical Research and Development (JP18dm0107134).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yassin, W., Nakatani, H., Zhu, Y. et al. Machine-learning classification using neuroimaging data in schizophrenia, autism, ultra-high risk and first-episode psychosis. Transl Psychiatry 10, 278 (2020). https://doi.org/10.1038/s41398-020-00965-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-00965-5
This article is cited by
-
Morphometric network-based abnormalities correlate with psychiatric comorbidities and gene expression in PCDH19-related developmental and epileptic encephalopathy
Translational Psychiatry (2024)
-
A primer on the use of machine learning to distil knowledge from data in biological psychiatry
Molecular Psychiatry (2024)
-
Using brain structural neuroimaging measures to predict psychosis onset for individuals at clinical high-risk
Molecular Psychiatry (2024)
-
Sampling inequalities affect generalization of neuroimaging-based diagnostic classifiers in psychiatry
BMC Medicine (2023)
-
Deep learning and machine learning in psychiatry: a survey of current progress in depression detection, diagnosis and treatment
Brain Informatics (2023)