Abstract
The validity of diagnostic labels of autism spectrum disorder (ASD), attention-deficit/hyperactivity disorder (ADHD), and obsessive compulsive disorder (OCD) is an open question given the mounting evidence that these categories may not correspond to conditions with distinct etiologies, biologies, or phenotypes. The objective of this study was to determine the agreement between existing diagnostic labels and groups discovered based on a data-driven, diagnosis-agnostic approach integrating cortical neuroanatomy and core-domain phenotype features. A machine learning pipeline, called bagged-multiview clustering, was designed to discover homogeneous subgroups by integrating cortical thickness data and measures of core-domain phenotypic features of ASD, ADHD, and OCD. This study was conducted using data from the Province of Ontario Neurodevelopmental Disorders (POND) Network, a multi-center study in Ontario, Canada. Participants (n = 226) included children between the ages of 6 and 18 with a diagnosis of ASD (n = 112, median [IQR] age = 11.7[4.8], 21% female), ADHD (n = 58, median [IQR] age = 10.2[3.3], 14% female), or OCD (n = 34, median [IQR] age = 12.1[4.2], 38% female), as well as typically developing controls (n = 22, median [IQR] age = 11.0[3.8], 55% female). The diagnosis-agnostic groups were significantly different than each other in phenotypic characteristics (SCQ: χ2(9) = 111.21, p < 0.0001; SWAN: χ2(9) = 142.44, p < 0.0001) as well as cortical thickness in 75 regions of the brain. The analyses revealed disagreement between existing diagnostic labels and the diagnosis-agnostic homogeneous groups (normalized mutual information < 0.20). Our results did not support the validity of existing diagnostic labels of ASD, ADHD, and OCD as distinct entities with respect to phenotype and cortical morphology.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD), attention-deficit/hyperactivity disorder (ADHD), and obsessive compulsive disorder (OCD) are complex neurodevelopmental disorders. There is emerging evidence that these diagnostic categories may not correspond to conditions with distinct etiology1,2,3,4,5,6,7,8, biology9, or phenotype10, and that they may not represent distinct underlying mechanisms of dysfunction or predict treatment response11. In this context, several studies have revealed shared characteristics among ASD, ADHD, and OCD across various levels of analysis (e.g., etiology1,2,3,4,5,6,7,8, biology9,12,13, and phenotype10,14,15,16,17,18,19), as well as significant comorbidity among these disorders3,20,21. These studies commonly rely on case-control designs, which use diagnostic labels to define group-level statistics for comparisons. Although these approaches can identify group differences in means when distributions are close to normal, they cannot characterize group overlap in the presence of large within-group variability that may arise from existence of subgroups within each group. This is an important consideration when analyzing complex disorders, such as ASD, ADHD, and OCD, which present with strikingly large within-disorder heterogeneity in etiology21,22,23,24,25,26,27,28,29, neurobiology30,31,32,33,34,35,36, and phenotypic presentation37.
The between-group overlap and the large within-group heterogeneity motivate a shift away from traditional case-control designs to trans-diagnostic analyses based on diagnosis-agnostic and continuous measures. This approach may provide insight into the structure of individual variability in biology and phenotype, including discovery of homogeneous subgroups and/or continua characterized by different biologies. To this end, we propose a data-driven, diagnosis-agnostic approach to derive sub-groups that share biological and phenotypic characteristics. Previous attempts have been made to discover homogeneous subgroups within each disorder37,38,39,40,41,42 or on a single level of analysis16,43, however, to our knowledge cross-disorder, multi-level stratification has not been examined previously.
We examined homogeneity in neuroanatomy, measured by cortical thickness, and core-domain phenotypic characteristics of each disorder. Neuroanatomical similarities can provide an intermediate phenotype that links multiple genetic variants31 given that genetic findings are rare and unknown for the majority of individuals with ASD, OCD, and ADHD. Cortical thickness is a heritable measure of cortical columnar structure, suggested to reflect cellular maturational changes in the cortex (i.e., dendritic arborization and pruning, myelination), as well as cognitive and behavioral differences44,45,46.
Materials and methods
Participants
Participants were recruited through the Province of Ontario (Canada) Neurodevelopmental Disorders Network (POND), a multi-center research network studying neurodevelopmental disorders. Participants who had capacity to consent provided informed consent. For others, consent was obtained from guardians and assent was obtained from the participants. Ethics approval was obtained from the research ethics boards at Holland Bloorview Kids Rehabilitation Hospital and the Hospital for Sick Children.
The included participants were 6–18 years old, had sufficient English comprehension to complete the testing protocols, and did not have contraindications for MRI. For the clinical groups, a primary diagnosis of ASD, ADHD, or OCD was required. Diagnoses for the clinical groups were confirmed using in-depth assessments (ASD: Autism Diagnostic Observation Schedule–2 (ADOS)47 and Autism Diagnostic Interview–Revised (ADI-R)48; ADHD: Parent Interview for Child Symptoms (PICS)49; OCD: K-SADS and the Children’s Yale–Brown Obsessive Compulsive Scale (CY-BOCS)50. The controls did not have a neurodevelopmental, psychiatric and/or neurological diagnosis and were born after 35 weeks gestation.
Behavioral measures
Our analyses focused on primary domains affected in ASD, ADHD, and OCD, quantified using continuous measures of autism features (Social Communication Questionnaire (SCQ)51), inattention (inattentive subscale of the Strengths and Weaknesses of ADHD-symptoms and Normal Behavior (SWAN) rating scale52), and obsessive-compulsive traits (Toronto Obsessive Compulsive rating scale (TOCS)53). Participants also completed the Child-Behaviour Checklist (CBCL)59. Full-scale IQ was estimated using the age-appropriate Wechsler or Stanford-Binet scales.
Imaging data
Structural MRI data was collected on the 3-Tesla Siemens Trio TIM at the Hospital for Sick Children, in Toronto, Ontario for 184 participants. The remaining participants were scanned after a hardware updated to the Siemens Prisma scanner. Cortical thickness measures were extracted from T1-weighted images using the CIVET pipeline (version 2.1.0)54. The pipeline applies a non-uniformity correction on the images54 followed by stereotaxic registration to the Montreal Neurologic Institute (MNI ICBM152) template (non-linear 6th generation target)55,56. Next, brains were masked, extracted, and classified into gray matter, white matter, and cerebrospinal fluid. Tissue classification images were used to generate gray and white matter surfaces57,58,59,60,61. A surface-diffusion kernel was applied62, and regions were registered to the automated anatomical labeling atlas63,64,65. Cortical thickness measurements were taken from the distance between the two smoothed surfaces66. Quality assurance was carried out at the time of the scan for motion artifact, and was analyzed through the CIVET quality control (QC) analysis pipeline. Scans that were flagged on the QC analysis were manually reviewed for quality and excluded if needed.
Cortical thickness measurements from 76 regions of the brain were regressed against age, sex, and scanner type in a sequential manner and the z-scored residuals were used in subsequent analyses.
Analysis
Inspired by the concepts of multi-view clustering67 and bagging68, a machine learning pipeline was designed to analyze the multi-dimensional brain-behavior data. This pipeline (Fig. 1), called bagged-multiview clustering, integrates three features: (1) clustering to discover groups of participants who present with “similar” characteristics in both neuroanatomy and phenotype, (2) bagging to improve cluster stability, and (3) feature weight calculation to determine the cortical regions that contributed most to determining the clusters. The clustering analyses were performed using the Scikit-learn toolbox in Python. Statistical analyses were carried out using R 3.3.3 and Matlab 2017a.
Bagged clustering
The bagged-multiview clustering pipeline consisted of bagging and spectral clustering69. Resampling methods such as bagging68 generate and aggregate decisions based on multiple random subsets of data to improve the accuracy, stability, and generalizability of machine learning algorithms69. For this study, a full run of the bagged-multiview clustering pipeline consisted of 50,000 subsamples, each using a random subset of 63.2% of participants, two (of three) dimensions of phenotypic data, cortical thickness measurements from seven (of 76) cortical regions, and the number of clusters randomly chosen between 2 and 15. The size of the random subsets was determined following seminal works in bagging70,71. The range for the number of clusters was determined based on visual inspection of affinity matrices. Each iteration generated a participant connectivity matrix, with entries of one if two participants were grouped in the same cluster and zero otherwise. To confirm that the clustering result was indicative of true connections between participants, the analyses were run on two sets created by (1) randomly sampling a uniform distribution across the range of the data, and (2) randomly permuting the cortical thickness and phenotypic data. The distribution of connectivity values due to random chance was computed. Two participants were deemed “similar” if they were grouped together more times than the 99th percentile of the connection values for the random data.
For each iteration of clustering, the following steps were performed. First, using the Gaussian similarity function, affinity matrices were computed for the SCQ, SWAN, and TOCS scores as well as cortical thickness measurements for each of the 76 cortical regions (total of 3 + 76 matrices). The parameter for the Gaussian kernel was set as the 75th percentile of pairwise distances in each measure. Second, a subset of cortex features were chosen for fusion with the phenotype data. The selection maximized within-to-between cluster similarity using sequential-feature-forward selection72. Within-cluster similarity was defined as the median of the medians similarities for participants in the same cluster. Between-cluster similarity was the median of the 99th percentile similarity between participants in one clusters and those in all other clusters (99th percentile chosen to deal with the sparsity of the matrix). The overall ratio was computed as the average of the within-to-between ratios, where the ratio for each cluster was weighted by the number of participants in that cluster. Participant similarities were obtained from an affinity matrix resulting from the fusion of cortical thickness and phenotypic matrices. The matrices corresponding to the same data type (cortical measurement or phenotype) were fused using element-wise arithmetic averaging. This type of fusion allows for clusters to match along any of the features combined. This procedure results in two affinity matrices: one for cortical thickness and one for phenotypic measurements. These two matrices were fused using a geometric mean. This requires the final clusters to match across cortical thickness and phenotypic dimensions. To further reduce variability and improve generalizability, the entire pipeline was run 10 times (×50,000 iteration each time) and the median of the participant similarity matrices was used to generate the results reported in the following sections. The 10 iterations were performed further to reduce variability in clustering in a computationally efficient manner.
Feature weight calculation
Feature weights were computed based on an adapted version of the permutation accuracy importance70,71. In particular, each feature’s prediction accuracy was calculated as the difference between the final labels and the labels generated in iterations where that feature was selected. The values of the feature were then permuted and the accuracy was again calculated. The feature weight was defined as the difference in accuracy before and after the feature is permuted.
Agreement between groups
To evaluate the agreement between diagnostic labels and the data-driven cluster assignments, four measures were used:
-
Normalized mutual information73: Roughly, this measure quantifies that amount of information shared between two clustering assignments. This measure takes on values between 0 (independent clusterings) and 1 (identical clusterings).
-
Adjusted Rand score74: This measure is based on counting item pairs who fall in the same or different clusters based on two clusterings. The adjusted Rand score ranges between 0 and 1, with 1 indicating perfect agreement between to clusterings.
-
Homogeneity75: This measure quantifies the extent to which each data-driven cluster contains only participants from a single diagnostic group (0 minimum homogeneity, 1 when each cluster contains only members of a single class).
-
Completeness75: This measures quantifies how well participants in the same diagnostic group are assigned to the same cluster (0 minimum completeness, 1 perfectly complete assignment).
SCQ, SWAN, and TOCS scores, as well as cortical thickness values were compared across clusters using Kruskal–Wallis tests.
Results
Participants
Participant demographic information is shown in Table 1. The diagnostic groups differed significantly in age (χ2(3) = 10.1, p = 0.02), full-scale IQ (χ2(3) = 31.4, p < 0.0001), and measures of core-domain symptomatology namely, SCQ (χ2(3) = 134.8, p < 0.0001), SWAN (χ2(3) = 69.5, p < 0.0001), and TOCS (χ2(3) = 80.2, p < 0.0001). The age difference did not survive correction for multiple comparisons. Post-hoc analyses showed that the proportion of male to female participants was higher in the ASD and ADHD groups compared to the OCD and TD groups. Participants in the ASD group had lower median IQ scores compared to the OCD and TD groups, and the ADHD group had lower median IQ compared to the OCD group. Thirty-eight of the 226 participants were missing IQ data.
The ASD, ADHD, and OCD groups had significantly elevated scores compared to all groups on their respective core-domain measures (SCQ, SWAN, TOCS; p < 0.0001). Interestingly, the ADHD group had significantly higher SCQ scores compared to the TD controls, and the ASD group had significantly elevated SWAN and TOCS scores compared to the TD groups.
In the ASD group, 46 and 40% of the participants met clinical cut-offs on the SWAN and TOCS, respectively. In the ADHD group, 11 and 17% of participants met clinical cut-offs on the SCQ and TOCS. Of the participants in the OCD group, 8 and 24% met the cut-off on the SCQ and SWAN, respectively. None of the TD participants met the cut-offs for SCQ or SWAN, but 2 of the 22 exceeded the cut-off on the TOCS (eTable 1 in the Supplement). The distribution of each of the core-measures scores also evidenced overlap among the diagnostic groups with respect to all three measures (eFig. 1 in the Supplement).
Cluster-diagnosis agreement
The agreement was <0.2 for the normalized mutual information and adjusted rand scores for cluster numbers ranging from 2 to 14 (perfect agreement corresponds to a value of one). Homogeneity and completeness scores were less than 0.3, indicating that data-driven clusters do not represent a single diagnostic category (eFig. 2 in the Supplement).
Clusters
Based on the within-to-between similarity ratio, a 10-cluster solution was chosen for the remaining analyses (eFig. 3 in the Supplement). Figure 2 graphically depicts how clusters emerge as the number of clusters increases. The figure was generated by changing the number of clusters in a spectral clustering algorithm applied to the similarity matrix generated by the bagged-multiview clustering pipeline.
The Kruskal–Wallis test did not show significant cluster differences in age or sex proportions (eFig. 4 in the Supplement). However, the clusters were significantly different in IQ (χ2(9) = 23.6, p = 0.005). Post-hoc testing showed that cluster 1 had significantly higher mean ranks than clusters 5 and 7 (p = 0.03).
Diagnostic labels
Figure 3 shows the percentage of participants from the four diagnostic categories falling into each of the ten clusters. Most clusters contained participants from multiple diagnostic groups. There was also a group of clusters with participants from the neurodevelopmental groups only (referred to as “neurodevelopmental clusters” from here on). There was also a small cluster of participants with ASD only (cluster 10), containing 12% of the participants with an ASD diagnosis.
The Kruskal–Wallis test revealed a significant difference in SCQ and SWAN scores among clusters (SCQ: χ2(9) = 111.21, p < 0.0001; SWAN: χ2(9) = 142.44, p < 0.0001), but the cluster difference in TOCS scores was not significant (eFig. 5 in the Supplement). The clusters were also significantly different in CBCL Social Problems scores (χ2(9) = 56.3, p < 0.0001) and Attention Problems (χ2(9) = 94.8, p < 0.0001), but not OCD Problems. Kruskal–Wallis tests showed a significant effect of cluster on cortical thickness in all regions (Bonferroni corrected p < 0.002), except for the left lingual gyrus.
Figure 4 depicts participant-level SCQ and SWAN scores for the diagnostic groups, as well as the data-driven clusters. This figure highlights the differences between diagnostic classifications and the data-driven solution. The data-driven clusters broadly divide the SCQ-SWAN space into low and high SWAN scores based on a cut-off score of 6; within the low and high SWAN regions, a continuum of SCQ scores can be observed. The neurodevelopmental clusters fall within the high SWAN region, with the exception of the “pure” ASD group.
The participant similarity matrix (eFig. 6 in the Supplement) revealed significant overlap among clusters 1 through 3, and 6 through 9, suggesting a structure more consistent with a continuum rather than distinct clusters within these groups. The matrix also indicates that clusters 5 is poorly defined (low similarity among the participants in the cluster).
Cortical regions
Figure 5 visualizes the contribution of each cortical region to the clustering solution. The top ten highly weighted features are listed in eTable 2 in the Supplement for reference.
The weight distribution among the regions was relatively uniformly decreasing (eFig. 7 in the Supplement), suggesting that no single region is driving the clustering results.
Cluster validity
The distribution of connection values used to derive clustering solutions for participant data showed no overlap with the randomly generated data (eFig. 8 in the Supplement).
Discussion
In this study, we used a data-driven, diagnosis-agnostic approach to examine overlap across three neurodevelopmental disorders (ASD, ADHD, and OCD). Overall, our results suggest that homogeneity in the variables examined in our analyses does not align well with existing diagnostic categories. Instead, we observed that differences in the domains primarily affected in these disorders may exist along a continuum that includes typical development.
Clusters
We started with a grouping of participants categorized into four diagnostic groups, which differed significantly on scores on SCQ, SWAN, and TOCS. Our analyses resulted in a new grouping of these participants into more homogeneous subgroups, which differed significantly in SCQ and SWAN scores as well as cortical thickness, but did not align well with the original diagnostic labels.
The majority of the data-driven clusters contained participants from multiple diagnostic categories, highlighting shared phenotypes and neurobiologies among the diagnostic groups. Social difficulties and inattention are commonly reported as shared features of ASD, ADHD, and OCD10,14,20,76,77. Several studies have also reported shared characteristics in brain structure, function, and connectivity9,12,78,79,80,81,82 in these disorders. Our results support the emerging recognition that the existing behaviorally-defined diagnostic labels may not capture etiologically, biologically, and phenomenologically homogeneous groups29,79,83,84,85,86,87,88,89.
Visually, our results are consistent with the notion that that the ASD-like features, and to some extent inattention traits, exist across a continuum that includes typical development. This model is supported by the substantial etiological overlap between these disorders and typical variation in social communication ability90 and inattention91. This is also consistent with the notion that multiple susceptibility genetic factors may interact with environmental conditions to lead to a continuous dimension of ASD-like and inattention traits, with neurodevelopmental disorders at the extremes of this continuum92,93. This motivates models of neurodevelopmental disorders which focus on continuous variations in traits instead of categorical diagnoses defined based on qualitative cut-offs. Future studies should consider examining other phenotypic characteristics and biological parameters (e.g., metabolic, immune, endocrine markers) to comprehensively describe this continuum.
The data-driven clusters differed significantly in SCQ and SWAN scores, but not TOCS. This pattern was also replicated using the CBCL measures of social, attention, and OCD problems. Moreover, the majority of participants with an OCD diagnosis clustered together with the typical controls. This has been observed in two other studies which examined social perception abilities10 and white matter structure9 using the same cohort. In addition, a study of a community sample found that those with a sibling with ASD showed more ADHD, but not OCD traits compared to those without a sibling with ASD20. Replication on larger samples is needed to further explore shared characteristics and differences across these disorders.
Finally, it is important to note that discovery of the exact clusters/subgroups that can be translated into clinical practice requires replication and integration of findings across a large number of studies and measures. This paper is a first step to accomplish this. Our results motivate a paradigm shift to challenge how ASD, ADHD, and OCD are currently defined, diagnosed, and treated. In particular, this paper adds to the evidence that these diagnoses may not exist as uniquely-defined diagnostic constructs, and highlights the need to discover other groupings that may be more closely aligned with biology and/or response to treatment.
Our results also have implications for the research community. Most existing studies commonly rely on case-control designs, which use diagnostic labels to define group-level statistics for comparisons. These approaches are often not able to characterize group overlap in the presence of large within-group variability that is revealed in our study. In this context, our results highlight the need to move beyond traditional statistical approaches to more advanced computational approaches to examine variability and overlap in/across these disorders. To our knowledge, this is the first examination of cross-disorder, multi-level stratification across ASD, ADHD, and OCD.
Cortical features
Our results add to the emerging evidence that the existing diagnostic categories may not be associated with unique patterns of difference in brain structure, paralleling a recent study showing significant heterogeneity in brain volume across 26 mouse models of ASD31.
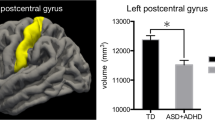
Broadly, the regions contributing most to the data-driven groupings were involved in social function, emotion processing, language, attention, and inhibitory control. Many of these regions have been previously implicated in studies of cortical morphology in ASD (e.g., middle temporal gyrus94,95,96,97,98,99, supramarginal gyrus78,96,97,98,99, angular gyrus100, middle frontal gyrus94,96,99,100, cingulate94,97,99, inferior frontal gyrus96,98,99, postcentral gyrus96,98,99,100, inferior temporal gyrus94,98,99), ADHD (e.g., cingulate101,102,103,104, dorsolateral prefrontal cortex102, inferior frontal cortex102, anterior cingulate cortex13, temporoparietal regions13), and OCD (e.g., inferior frontal gyrus105, anterior cingulate cortex13,105, supramarginal gyrus105, dorsolateral prefrontal cortex13, middle frontal gyrus12).
Our results also overlap with those of the very few studies that have examined similarities and differences in brain structure across pairs of ASD, ADHD, and OCD. For example, disorder-specific differences in the left middle temporal gyrus95, right supramarginal gyrus78, and the prefrontal cortex106 have been reported for ASD and ADHD. Looking at ADHD and OCD, differences have been reported in the cingulate cortex and dorsolateral prefrontal cortex13. Decreased volume in the anterior cingulate cortex has been suggested as a shared finding in ASD and OCD 12.
Limitations
Our analyses were conducted on a single measure of cortical structure and three phenotypic measures as well as a specific age group. These levels of analyses may not fully capture homogeneity across the disorders. Future work should consider running similar types of analyses using multiple measures that can comprehensively characterize the variability across neurodevelopmental disorders. These include brain structure and function and core and comorbid behavioral domains across the life-span, as well as genetic, epigenetic, metabolic, immune, and endocrine markers.
The sample size used for the analyses reported in this paper was limited, with unequal distribution of participants across the diagnostic groups. Replication with larger sample sizes is needed.
To our knowledge, this is the first study of diagnosis-agnostic homogeneity across ASD, ADHD, and OCD using data-driven discovery. Homogeneity in the variables examined in our analyses did not align well with existing diagnostic categories in the sample studied. These results add to the emerging body of literature questioning the validity of existing diagnostic constructs with respect to having distinct biological and phenotype presentation. The results of this study also highlight the need for a shift from case-control models to more complex analyses that can cope with the large between-disorder overlap and within-disorder variability.
References
Ronald, A., Simonoff, E., Kuntsi, J., Asherson, P. & Plomin, R. Evidence for overlapping genetic influences on autistic and ADHD behaviours in a community twin sample. J. Child Psychol. Psychiatry Allied Discip. 49, 535–542 (2008).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Rommelse, N. N. J., Franke, B., Geurts, H. M., Hartman, C. A. & Buitelaar, J. K. Shared heritability of attention-deficit/hyperactivity disorder and autism spectrum disorder. Eur. Child Adolesc. Psychiatry 19, 281–295 (2010).
Lichtenstein, P., Carlström, E., Råstam, M., Gillberg, C. & Anckarsäter, H. The genetics of autism spectrum disorders and related neuropsychiatric disorders in childhood. Am. J. Psychiatry 167, 1357–1363 (2010).
Guo, W. et al. Polygenic risk score and heritability estimates reveals a genetic relationship between ASD and OCD. Eur. Neuropsychopharmacol. 27, 657–666 (2017).
Lionel, A. C. et al. Disruption of the ASTN2/TRIM32 locus at 9q33.1 is a risk factor in males for autism spectrum disorders, ADHD and other neurodevelopmental phenotypes. Hum. Mol. Genet. 23, 2752–2768 (2014).
Lionel, A. C. et al. Rare copy number variation discovery and cross-disorder comparisons identify risk genes for ADHD. Sci. Transl. Med. 3, 95ra75-95ra75 (2011).
Geller, D. et al. Examining the relationship between obsessive-compulsive disorder and attention-deficit/hyperactivity disorder in children and adolescents: a familial risk analysis. Biol. Psychiatry 61, 316–321 (2007).
Ameis, S. H. et al. A diffusion tensor imaging studyin children with ADHD, autism spectrum disorder, OCD, and matched controls: distinct and non-distinct white matter disruption and dimensional brain-behavior relationships. Am. J. Psychiatry 173, 1213–1222 (2016).
Baribeau, D. A. et al. Examining and comparing social perception abilities across childhood-onset neurodevelopmental disorders. J. Am. Acad. Child Adolesc. Psychiatry 54, 479–486.e1 (2015).
Insel, T. et al. Research Domain Criteria (RDoC): toward a new classification framework for research on mental disorders. Am. J. Psychiatry 167, 748–751 (2010).
Carlisi, C. O. et al. Comparative multimodal meta-analysis of structural and functional brain abnormalities in autism spectrum disorder and obsessive-compulsive disorder. Biol. Psychiatry. 82, 83–102 (2017).
Norman, L. J. et al. Structural and functional brain abnormalities in attention-deficit/hyperactivity disorder and obsessive-compulsive disorder: a comparative meta-analysis. JAMA Psychiatry 73, 815–825 (2016).
Anholt, G. E. et al. Autism and adhd symptoms in patients with ocd: Are they associated with specific oc symptom dimensions or oc symptom severity. J. Autism Dev. Disord. 40, 580–589 (2010).
Zandt, F., Prior, M. & Kyrios, M. Repetitive behaviour in children with high functioning autism and obsessive compulsive disorder. J. Autism Dev. Disord. 37, 251–259 (2007).
Van Der Meer, J. M. J. et al. Are autism spectrum disorder and attention-deficit/hyperactivity disorder different manifestations of one overarching disorder? Cognitive and symptom evidence from a clinical and population-based sample. J. Am. Acad. Child Adolesc. Psychiatry. 51, 1160–1172 (2012).
Bora, E. & Pantelis, C. Meta-analysis of social cognition in attention-deficit/hyperactivity disorder (ADHD): Comparison with healthy controls and autistic spectrum disorder. Psychol. Med. 46, 699–716 (2016).
Gargaro, B. A., Rinehart, N. J., Bradshaw, J. L., Tonge, B. J. & Sheppard, D. M. Autism and ADHD: How far have we come in the comorbidity debate? Neurosci. Biobehav. Rev. 35, 1081–1088 (2011).
Ruzzano, L., Borsboom, D. & Geurts, H. M. Repetitive behaviors in autism and obsessive–compulsive disorder: new perspectives from a network analysis. J. Autism Dev. Disord. 45, 192–202 (2014).
van der Plas, E., Dupuis, A., Arnold, P., Crosbie, J. & Schachar, R. Association of autism spectrum disorder with obsessive-compulsive and attention-deficit/hyperactivity traits and response inhibition in a community sample. J. Autism Dev. Disord. 46, 3115–3125 (2016).
Abramovitch, A., Dar, R., Mittelman, A. & Wilhelm, S. Comorbidity between attention deficit/hyperactivity disorder and obsessive-compulsive disorder across the lifespan. Harv. Rev. Psychiatry 23, 245–262 (2015).
Vorstman, J. A. S. et al. Autism genetics: opportunities and challenges for clinical translation. Nat. Rev. Genet. 18, 362–376 (2017).
De La Torre-Ubieta, L., Won, H., Stein, J. L. & Geschwind, D. H. Advancing the understanding of autism disease mechanisms through genetics. Nat. Med. 22, 345–361 (2016).
Gizer, I. R., Ficks, C. & Waldman, I. D. Candidate gene studies of ADHD: a meta-analytic review. Hum. Genet. 126, 51–90 (2009).
Neale, B. M. et al. Meta-analysis of genome-wide association studies of attention-deficit/hyperactivity disorder. J. Am. Acad. Child Adolesc. Psychiatry 49, 884–897 (2010).
Li, Z., Chang, Shua, Zhang, Lyan, Gao, L. & Wang, J. Molecular genetic studies of ADHD and its candidate genes: A review. Psychiatry Res. 219, 10–24 (2014).
Hawi, Z. et al. The molecular genetic architecture of attention deficit hyperactivity disorder. Mol. Psychiatry 20, 289–297 (2015).
Nestadt, G., Grados, M. & Samuels, J. F. Genetics of obsessive-compulsive disorder. Psychiatr. Clin. North Am. 33, 141–158 (2010).
Pauls, D. L., Abramovitch, A., Rauch, S. L. & Geller, D. A. Obsessive-compulsive disorder: An integrative genetic and neurobiological perspective. Nat. Rev. Neurosci. 15, 410–424 (2014).
Anagnostou, E. & Taylor, M. J. Review of neuroimaging in autism spectrum disorders: What have we learned and where we go from here. Mol. Autism. 2, 4 (2011).
Ellegood, J. et al. Clustering autism: using neuroanatomical differences in 26 mouse models to gain insight into the heterogeneity. Mol. Psychiatry 20, 118–125 (2015).
Wang, J. B. et al. Inconsistency in abnormal brain activity across cohorts of ADHD-200 in children with attention deficit hyperactivity disorder. Front Neurosci. 11, 320 (2017).
Lenet, A. E. Shifting focus: from group patterns to individual neurobiological differences in attention-deficit/hyperactivity disorder. Biol. Psychiatry 82, e67 (2017).
Piras, F. et al. Widespread structural brain changes in OCD: a systematic review of voxel-based morphometry studies. Cortex 62, 89–108 (2015).
Fouche, J. P. et al. Cortical thickness in obsessive-compulsive disorder: multisite mega-analysis of 780 brain scans from six centres. Br. J. Psychiatry 210, 67–74 (2017).
Eng, G. K., Sim, K. & Chen, S. H. A. Meta-analytic investigations of structural grey matter, executive domain-related functional activations, and white matter diffusivity in obsessive compulsive disorder: An integrative review. Neurosci. Biobehav. Rev. 52, 233–257 (2015).
Bragdon, L. B. & Coles, M. E. Examining heterogeneity of obsessive-compulsive disorder: Evidence for subgroups based on motivations. J. Anxiety Disord. 45, 64–71 (2017).
Roberts, B. A., Martel, M. M. & Nigg, J. T. Are there executive dysfunction subtypes within ADHD? J. Atten. Disord. 21, 284–293 (2017).
Leung, P. & Chan, F. Neurocognitive deficits underlying attention-deficit/hyperactivity disorder (ADHD): A clustering/subgrouping analysis. Eur. Psychiatry 33, S131 (2016).
Ben-Sasson, A. & Podoly, T. Y. Sensory over responsivity and obsessive compulsive symptoms: A cluster analysis. Compr. Psychiatry 73, 151–159 (2017).
Hasanpour, H. et al. A critical appraisal of heterogeneity in Obsessive-Compulsive Disorder using symptom-based clustering analysis. Asian J. Psychiatr. 28, 89–96 (2017).
Grzadzinski, R., Huerta, M. & Lord, C. DSM-5 and autism spectrum disorders (ASDs): an opportunity for identifying ASD subtypes. Mol. Autism. 4, 12, (2013).
Lim, L. et al. Disorder-specific predictive classification of adolescents with attention deficit hyperactivity disorder (ADHD) relative to autism using structural magnetic resonance imaging. PLoS ONE. 8, e63660, (2013).
Huttenlocher, P. R. Morphometric study of human cerebral cortex development. Neuropsychologia 28, 517–527 (1990).
Sowell, E. R. Longitudinal Mapping of Cortical Thickness and Brain Growth in Normal Children. J. Neurosci. 24, 8223–8231 (2004).
Panizzon, M. S. et al. Distinct genetic influences on cortical surface area and cortical thickness. Cereb. Cortex. 19, 2728–2735 (2009).
Lord, C. et al. Autism Diagnostic Observation Schedule (ADOS). J. Autism Developmental Disord. 30, 205–223 (2000).
Lord, C., Rutter, M. & Le Couteur, A. Autism Diagnostic Interview-Revised: A revised version of a diagnostic interview for caregivers of individuals with possible pervasive developmental disorders. J. Autism Dev. Disord. 24, 659–685 (1994).
Ickowicz, A. et al. The parent interview for child symptoms: a situation-specific clinical research interview for attention-deficit hyperactivity and related disorders. Can. J. Psychiatry 51, 325–328 (2006).
Scahill, L. et al. Children’s Yale-Brown obsessive compulsive scale: reliability and validity. J. Am. Acad. Child Adolesc. Psychiatry 36, 844–852 (1997).
Rutter Bailey, A. & Lord, C. M. Social Communication Questionnaire. Los Angeles, CA. 2003.
Swanson, J. M. et al. Categorical and dimensional definitions and evaluations of symptoms of ADHD: history of the SNAP and the SWAN rating scales. Int. J. Educ. Psychol. Assess. 10, 51–70 (2012).
Park, L. S. et al. The Toronto Obsessive-Compulsive Scale: psychometrics of a dimensional measure of obsessive-compulsive traits. J. Am. Acad. Child Adolesc. Psychiatry 55, 310–318 (2016).
Sled, J. G., ZijdenbosA. P. & EvansA. C. A nonparametric method for automatic correction of intensity nonuniformity in MRI data. IEEE Trans. Med Imaging 17, 87–97 (1998).
Grabner, G. et al. Symmetric atlasing and model based segmentation: an application to the hippocampus in older adults. Med Image Comput Comput Assist Interv. Int Conf. Med Image Comput Comput Assist Inter. 9, 58–66 (2006).
Collins, D. L., Neelin, P., Peters, T. M. & Evans, A. C. Automatic 3d intersubject registration of mr volumetric data in standardized talairach space. J. Comput Assist Tomogr. 18, 192–205 (1994).
Smith, S. M. Fast robust automated brain extraction. Hum. Brain Mapp. 17, 143–155 (2002).
Tohka, J., Zijdenbos, A. & Evans, A. Fast and robust parameter estimation for statistical partial volume models in brain MRI. Neuroimage 23, 84–97 (2004).
Zijdenbos A., Forghani R., Evans A. In Proceedings of the First International Conference on Medical Image Computing and Computer-Assisted Intervention, 439–448, 1998.
June, S. K. et al. Automated 3-D extraction and evaluation of the inner and outer cortical surfaces using a Laplacian map and partial volume effect classification. Neuroimage 27, 210–221 (2005).
MacDonald, D., Kabani, N., Avis, D. & Evans, A. C. Automated 3-D extraction of inner and outer surfaces of cerebral cortex from MRI. Neuroimage 12, 340–356 (2000).
Taylor, J. Chung M. K. 2nd IEEE Int Symp Biomed Imaging Nano to Macro (IEEE Cat No 04EX821). 432–435, 2004.
Robbins, S. M. Anatomical Standardization of the Human Brain in Euclidean 3-Space and on the Cortical 2-Manifold. Thesis. 2003.
Lyttelton, O., Boucher, M., Robbins, S. & Evans, A. An unbiased iterative group registration template for cortical surface analysis. Neuroimage 34, 1535–1544 (2007).
Boucher, M., Whitesides, S. & Evans, A. Depth potential function for folding pattern representation, registration and analysis. Med Image Anal. 13, 203–214 (2009).
Lerch, J. P. & Evans, A. C. Cortical thickness analysis examined through power analysis and a population simulation. Neuroimage 24, 163–173 (2005).
Kumar, A., Rai, P. & Daume, H. Co-regularized Multi-view Spectral Clustering. Adv. Neural Inf. Process Syst. 24, 1413–1421 (2011).
Breiman, L. Bagging predictors. Mach. Learn. 24, 123–140 (1996).
Dudoit, S. & Fridlyand, J. Bagging to improve the accuracy of a clustering procedure. Bioinformatics 19, 1090–1099 (2003).
Strobl, C., Boulesteix, A. L., Zeileis, A. & Hothorn, T. Bias in random forest variable importance measures: Illustrations, sources and a solution. BMC Bioinformatics. 8, 25 (2007).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Guyon, I. & Elisseeff, A. An introduction to variable and feature selection. J. Mach. Learn Res. 3, 1157–1182 (2003).
Vinh, N. X. Information theoretic measures for clusterings comparison: variants, properties, normalization and correction for chance. J. Mach. Learn Res. 11, 2837–2854 (2010).
Hubert, L. & Arabie, P. Comparing partitions. J. Classif. 2, 193–218 (1985).
Rosenberg A., Hirschberg J. V-measure: A conditional entropy-based external cluster evaluation measure. In: Proceedings of the Joint Conference on Empirical Methods in Natural Language Processing and Computational Natural Language (EMNLP-CoNLL’07). 2007. p. 410–420.
Sinzig, J., Walter, D. & Doepfner, M. Attention deficit/hyperactivity disorder in children and adolescents With autism spectrum disorder: Symptom or syndrome? J. Atten. Disord. 13, 117–126 (2009).
Leitner, Y. The co-occurrence of autism and attention deficit hyperactivity disorder in children – what do we know? Front Hum. Neurosci. 8, 268 (2014).
Brieber, S. et al. Structural brain abnormalities in adolescents with autism spectrum disorder and patients with attention deficit/hyperactivity disorder. J. Child Psychol. Psychiatry Allied Discip. 48, 1251–1258 (2007).
Ameis, S. H. Heterogeneity within and between autism spectrum disorder and attention-deficit/hyperactivity disorder - challenge or opportunity? JAMA Psychiatry 47, 1093–1094 (2017).
Bethlehem, R. A. I., Romero-Garcia, R., Mak, E., Bullmore, E. T. & Baron-Cohen, S. Structural covariance networks in children with autism or ADHD. Cereb. Cortex. 27, 4267–4276 (2017).
Di Martino, A. et al. Shared and distinct intrinsic functional network centrality in autism and attention-deficit/hyperactivity disorder. Biol. Psychiatry 74, 623–632 (2013).
Carlisi, C. O. et al. Disorder-specific and shared brain abnormalities during vigilance in autism and obsessive-compulsive disorder. Biol. Psychiatry Cogn. Neurosci. Neuroimaging. 2, 644–654 (2017).
Waterhouse, L., London, E. & Gillberg, C. ASD Validity. Rev. J. Autism Dev. Disord. 3, 302–329 (2016).
Müller, R. A. & Amaral, D. G. Editorial: Time to give up on Autism Spectrum Disorder? Autism Res. 10, 10–14 (2017).
London, E. B. Categorical diagnosis: a fatal flaw for autism research? Trends Neurosci. 37, 683–686 (2014).
Karalunas, S. L. et al. Subtyping attention-deficit/hyperactivity disorder using temperament dimensions: toward biologically based nosologic criteria. JAMA Psychiatry 71, 1015–1024 (2014).
Abramovitch, A., Dar, R., Mittelman, A. & Wilhelm, S. Comorbidity between attention deficit/hyperactivity disorder and obsessive-compulsive disorder across the lifespan: a systematic and critical review. Harv. Rev. Psychiatry 23, 245–262 (2015).
Rommelse, N., Buitelaar, J. K. & Hartman, C. A. Structural brain imaging correlates of ASD and ADHD across the lifespan: a hypothesis-generating review on developmental ASD–ADHD subtypes. J. Neural Transm. 124, 259–271 (2017).
Geurts, H. M., Ridderinkhof, K. R. & Scholte, H. S. The relationship between grey-matter and ASD and ADHD traits in typical adults. J. Autism Dev. Disord. 43, 1630–1641 (2013).
Robinson, E. B. et al. Genetic risk for autism spectrum disorders and neuropsychiatric variation in the general population. Nat. Genet. 48, 552–555 (2016).
Levy, F., Hay, Da, McStephen, M., Wood, C. & Waldman, I. Attention-deficit hyperactivity disorder: a category or a continuum? Genet. Anal. 36, 737–744 (1997).
van der Meer, J. M. J. et al. Homogeneous combinations of ASD–ADHD traits and their cognitive and behavioral correlates in a population-based sample. J. Atten. Disord. 21, 753–763 (2017).
Plomin, R., Haworth, C. M. A. & Davis, O. S. P. Common disorders are quantitative traits. Nat. Rev. Genet. 10, 872–878 (2009).
van Rooij, D. et al. Cortical and subcortical brain morphometry differences between patients with autism spectrum disorder and healthy individuals across the lifespan: results from the ENIGMA ASD Working Group. Am. J. Psychiatry. 175, 359–369 (2018).
Lim, L. et al. Disorder-specific grey matter deficits in attention deficit hyperactivity disorder relative to autism spectrum disorder. Psychol. Med. 45, 965–976 (2015).
Khundrakpam, B. S., Lewis, J. D., Kostopoulos, P., Carbonell, F. & Evans, A. C. Cortical thickness abnormalities in autism spectrum disorders through late childhood, adolescence, and adulthood: a large-scale MRI study. Cereb. Cortex. 27, 1721–1731 (2017).
Mensen, V. T. et al. Development of cortical thickness and surface area in autism spectrum disorder. NeuroImage Clin. 13, 215–222 (2017).
Smith, E. et al. Cortical thickness change in autism during early childhood. Hum. Brain Mapp. 37, 2616–2629 (2016).
Foster, N. E. V. et al. Structural gray matter differences during childhood development in autism spectrum disorder: a multimetric approach. Pediatr. Neurol. 53, 350–359 (2015).
Liu, J. et al. Gray matter abnormalities in pediatric autism spectrum disorder: a meta-analysis with signed differential mapping. Eur. Child Adolesc. Psychiatry 26, 933–945 (2017).
Shaw, P. et al. Longitudinal mapping of cortical thickness and clinical outcome in children and adolescents with attention-deficit/hyperactivity disorder. Arch. Gen. Psychiatry 63, 540–549 (2006).
Rubia, K., Alegria, A. & Brinson, H. Imaging the ADHD brain: disorder-specificity, medication effects and clinical translation. Expert Rev. Neurotherapeutics. 14, 519–538 (2014).
Cubillo, A., Halari, R., Smith, A., Taylor, E. & Rubia, K. A review of fronto-striatal and fronto-cortical brain abnormalities in children and adults with Attention Deficit Hyperactivity Disorder (ADHD) and new evidence for dysfunction in adults with ADHD during motivation and attention. Cortex 48, 194–215 (2012).
Bonath, B., Tegelbeckers, J., Wilke, M., Flechtner, H. H. & Krauel, K. Regional gray matter volume differences between adolescents with ADHD and typically developing controls: further evidence for anterior cingulate involvement. J. Atten. Disord. 22, 627–638 (2018).
Hu, X. et al. Meta-analytic investigations of common and distinct grey matter alterations in youths and adults with obsessive-compulsive disorder. Neurosci. Biobehav. Rev. 78, 91–103 (2017).
Dougherty, C. C., Evans, D. W., Myers, S. M., Moore, G. J. & Michael, A. M. A comparison of structural brain imaging findings in autism spectrum disorder and attention-deficit hyperactivity disorder. Neuropsychol. Rev. 26, 25–43 (2016).
Acknowledgements
This research was conducted with the support of the Natural Sciences and Engineering Research Council of Canada and the Ontario Brain Institute, an independent non-profit corporation, funded partially by the Ontario government. The opinions, results and conclusions are those of the authors and no endorsement by the Ontario Brain Institute is intended or should be inferred.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Dr. Kushki holds a patent for the device “Anxiety Meter”. Dr. Anagnostou has served as a consultant to Roche. She has received grant funding from SanofiCanada and SynapDx. She holds a patent for the device, “Anxiety Meter”. She has received royalties from APPI and Springer. Dr. Schachar has consulted to Highland Therapeutics, Eli Lilly and Co., and Purdue Pharma. He has commercial interest in a cognitive rehabilitation software company, “eHave”. Dr. Arnold holds a patent for ‘SLCIAI Marker for Anxiety Disorder’ granted May 6, 2008. The remaining authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kushki, A., Anagnostou, E., Hammill, C. et al. Examining overlap and homogeneity in ASD, ADHD, and OCD: a data-driven, diagnosis-agnostic approach. Transl Psychiatry 9, 318 (2019). https://doi.org/10.1038/s41398-019-0631-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-019-0631-2
This article is cited by
-
Predictors of health-related quality of life for children with neurodevelopmental conditions
Scientific Reports (2024)
-
Behaviour-correlated profiles of cerebellar-cerebral functional connectivity observed in independent neurodevelopmental disorder cohorts
Translational Psychiatry (2024)
-
Modelling the overlap and divergence of autistic and schizotypal traits on hippocampal subfield volumes and regional cerebral blood flow
Molecular Psychiatry (2024)
-
Transdiagnostic Patterns of Sensory Processing in Autism and ADHD
Journal of Autism and Developmental Disorders (2024)
-
Assessing the Contribution of Measures of Attention and Executive Function to Diagnosis of ADHD or Autism
Journal of Autism and Developmental Disorders (2024)