Abstract
Mild-cognitive impairment (MCI) occurs in up to one-fifth of individuals over the age of 65, with approximately a third of MCI individuals converting to dementia in later life. There is a growing necessity for early identification for those at risk of dementia as pathological processes begin decades before onset of symptoms. A cohort of 122 individuals diagnosed with MCI and followed up for a 36-month period for conversion to late-onset Alzheimer’s disease (LOAD) were genotyped on the NeuroChip array along with pathologically confirmed cases of LOAD and cognitively normal controls. Polygenic risk scores (PRS) for each individual were generated using PRSice-2, derived from summary statistics produced from the International Genomics of Alzheimer’s Disease Project (IGAP) genome-wide association study. Predictability models for LOAD were developed incorporating the PRS with APOE SNPs (rs7412 and rs429358), age and gender. This model was subsequently applied to the MCI cohort to determine whether it could be used to predict conversion from MCI to LOAD. The PRS model for LOAD using area under the precision-recall curve (AUPRC) calculated a predictability for LOAD of 82.5%. When applied to the MCI cohort predictability for conversion from MCI to LOAD was 61.0%. Increases in average PRS scores across diagnosis group were observed with one-way ANOVA suggesting significant differences in PRS between the groups (p < 0.0001). This analysis suggests that the PRS model for LOAD can be used to identify individuals with MCI at risk of conversion to LOAD.
Similar content being viewed by others
Introduction
The genetic contribution to late-onset Alzheimer’s disease (LOAD) is now well established, with heritability estimates ranging from 58 to 79%1. The APOE gene located on Chromosome 19 encodes the Apolipoprotein E protein. This gene represents the largest genetic risk factor for LOAD to date, with genetic variation producing three isoforms: ε2 (protective), ε3 (neutral and most common form) and ε4, which is associated with increased risk for LOAD. Further to this a number of both common and rare genetic risk factors have been identified in recent years from genome-wide association studies (GWAS) and next-generation sequencing efforts2,3,4,5. Despite this increase in our knowledge of genetic associations these do not account for the entire heritability of the LOAD phenotype. Although other factors such as epistasis and epigenetics might contribute, it is becoming accepted that far more genes/polymorphisms with much smaller effect sizes are involved in complex diseases than previously envisaged.
Genetic risk calculation studies have typically used only variants identified by GWAS to try and predict LOAD phenotype6,7,8,9,10,11,12, mild- cognitive impairment (MCI) conversion to LOAD13,14, hippocampal cortical thickness15,16, hippocampal volume17, cerebrospinal fluid biomarkers18 and plasma inflammatory biomarkers19.
The development of polygenic risk score (PRS) analysis now allows for the sum of genetic risk from the entire genome to be accounted for, weighted by the effect size estimates attained from established GWAS data, rather than selecting a few specific associated single-nucleotide polymorphism (SNP) markers20. The study by Escott-Price et al.21 demonstrated that the predictability of LOAD from APOE isoform and contribution of the LOAD GWAS SNPs was improved by incorporating further variants from across the genome into their PRS model. The addition of other predictors such as gender and age into the model resulted in a final predictive ability of 78.2%. Several other studies have also demonstrated that this form of analysis can differentiate between controls and LOAD cases with similarly high accuracy22,23.
The pathological features of LOAD have been found to begin decades before the onset of symptoms24,25,26. Therefore, the early detection of those likely to be at risk for LOAD could increase effectiveness of treatments preventing further damage from occurring25,26. MCI is diagnosed in up to one-fifth of individuals over the age of 65 and is considered a prodrome of dementia25. An estimated one-third of those diagnosed with MCI will go on to develop LOAD over time27. Consequently, understanding genetic risk factors within LOAD pathogenic mechanisms can improve detection and promote treatment before the pathogenic state arises24,26,28. A therapeutic intervention at the MCI stage may provide an opportunity to prevent or delay conversion to LOAD13,29. Therefore, there is a need to investigate if MCI to LOAD conversion can be predicted using PRS analysis.
In this study, we developed a PRS model to predict LOAD diagnosis in the brains for dementia research (BDR) cohort30. This model was then applied to a longitudinal sample of individuals with MCI from the Southampton inflammation, cognition and stress (ICOS) study, to see if the model could predict those individuals who converted from MCI to LOAD.
Materials and methods
Samples
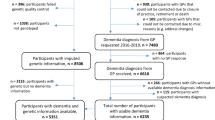
The BDR resource has recruited patients with dementia as well as cognitively healthy controls; post-mortem pathology was used to confirm and classify disease status31. This cohort consists of 302 LOAD cases and 137 controls, with no significant differences in the age at death and percentage of females between the cases and controls. The number of ε4 carriers was significantly higher amongst cases (p < 0.001; Table 1) as expected.
Individuals with MCI were recruited from the ICOS study in Southampton, UK. Diagnoses were made using the Petersen criteria for amnestic MCI32. The study followed-up individuals over a 36-month period (until October 2017) to identify those who converted to a LOAD (‘converters’ n = 49), and those who remained MCI (‘non-converters’ n = 73). Age at recruitment and presence of at least one APOE ε4 allele was not found to be significantly different between the converters and non-converters; although the converter group was found to have a significantly higher proportion of females (p = 0.008; Table 1).
Genotyping
DNA extraction was performed using a standard phenol chloroform method on either 2 ml of blood (MCI) or 100 mg of brain tissue (BDR). DNA quality was assessed using the Agilent 2200 TapeStation DNA integrity number (DIN; average DIN = 8.95) and quantified using Nanodrop 3300 spectrometry. All samples were collected with informed consent as governed by local guidelines at the point of collection; experimental procedures were approved by local ethics committees - Nottingham Research Ethics Committee 2 (REC reference 04/Q2404/130); London City and East NRES (REC reference 08/H0704/128 + 5), and completed in accordance with approved guidelines.
Genotyping was performed on the customised NeuroChip array33. Clustering was completed with the assistance of a cluster file provided by Blauwendraat and colleagues (2017) and the dataset was aligned to the GRCh37/hg19 assembly using files provided by Rayner, W (Personal correspondence, Nov 2017). Quality control of the raw data was completed using GenomeStudio v2 and PLINK v1.934, with samples removed based on a call rate less than 90%, gender mismatch and deviation from European population parameters. SNPs were removed based on a call rate less than 95%, genotype frequencies significantly out of Hardy–Weinberg equilibrium with a Bonferroni corrected p value threshold (p = 1.03 × 10−7) and excess heterozygosity (±3 standard deviations from mean). SNPs where the minor allele was observed in less than 3 individuals were also removed.
The APOE SNPs rs7412 and rs429358 (which determine the isoform) were genotyped with TaqMan assays using standard protocols. At least one APOE ε4 allele was harboured by 70.9% of LOAD cases, 46.9% of MCI converters, 42.9% of MCI non-converters and 40.1% of controls (Table 1).
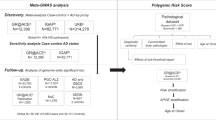
Polygenic-risk score generation and predictability modelling
The APOE gene is the largest known risk factor for LOAD, with the region surrounding the locus displaying levels of strong linkage disequilibrium (LD). Therefore, SNPs which fell within the 500 kb region (chr19:45,160,844-45,660,844; GRCh37/hg19 assembly) surrounding the APOE gene were excluded from the dataset. The genotypes for the APOE ε status SNPs (rs7412 and rs429358) were then reintroduced to the dataset to ensure genetic risk from APOE was captured.
Samples from the 1000 Genomes Project Consortium (1000G; n = 2504) were compared ancestrally to the BDR and MCI samples using common ancestry informative markers. Principal component analysis was carried out in PLINK to verify all NeuroChip-genotyped samples (n = 561) fell within the cluster of European descent (n = 503) to guard against population stratification.
Data from the 1000G samples were also used to calculate LD structure. The clumping algorithm in PRSice-220 was set to identify any SNPs within 250 kb in LD with an r2 threshold greater than 0.1 and ‘clump’ them together to be represented by most significantly associated SNP within each LD block, denoted the index SNP.
PRSice-2 was utilised to generate PRS for LOAD cases and cognitively healthy controls, using summary statistic data from the International Genomics of Alzheimer’s Disease Project (IGAP) genome-wide association study35. Significance values and effect sizes from the IGAP cohort were used as the base dataset to generate the best PRS model which was then applied to the BDR and MCI target datasets. Briefly, the PRS for each individual in the target dataset is generated from a summation of effect sizes from all the SNPs included in the best model. The best model was derived from testing the inclusion of SNPs (19–73,058 SNPs) from a range of p value thresholds in the base dataset (10−6 to 1), to see which threshold gave the largest Nagelkerke’s R2 value. These SNPs were then used to generate PRS for each individual in both the BDR and MCI cohorts.
The significance of differences in mean PRS between the four groups (controls, MCI non-converters, MCI converters and LOAD) was tested using one-way ANOVA with post hoc Tukey in SPSS v24. Spearman’s correlation analysis was also conducted by ranking the diagnosis groups from 0 (control), 1 (non-converters), 2 (converters) to 3 (LOAD).
Predictive ability of APOE alone and APOE with the PRS for identifying LOAD cases and converters in each cohort was calculated using Area Under the precision-recall Curve (AUPRC) in R using the ‘PRROC’ package36 (R Core Team, 2013). AUPRC was calculated to identify predictive ability, values range from 0 to 100%, where 0% is random classification and 100% is perfect classification37.
The effect of non-genetic predictors (age and gender) were estimated in SPSS, and were included with PRS in logistic regression analysis to enable incorporation into the predictive model.
LOAD cases and controls were distributed into deciles based on the range of probability values which accounted for PRS with APOE, gender and age at death. The proportions of LOAD cases which fell into each decile were calculated and are depicted alongside the proportion of controls within the same decile. This was also conducted for the proportions of MCI converters and non-converters within each decile distributed by PRS including APOE, gender and age at recruitment.
Results and discussion
European ancestry of the BDR and MCI samples was confirmed with principal component analysis, as all samples clustered accordingly with European samples from the 1000G dataset. A plot of the first two principal components is presented in Supplementary Fig. 1. With confirmation that the BDR and MCI target datasets were or the same ethnic decent as the IGAP base dataset, analysis could proceed.
PRSice-2 derived Nagelkerke’s R2 values for a range of SNP p value thresholds from 1 × 10−6 to 1 in the IGAP base dataset were used to determine the best significance threshold for inclusion of SNPs required to distinguish between LOAD cases and controls. There was a total of 73,056 SNPs were found in both the IGAP base dataset and NeuroChip target datasets after LD based clumping (r2 > 0.1; kb = 250). The largest Nagelkerke’s R2 value generated was 0.138, suggesting the inclusion of all SNPs at the p value threshold of 1.07 × 10−4 into the PRS model plus the two APOE SNPs (total 167 SNPs; Supplementary Table 1). This model was then applied to both the BDR and MCI cohorts.
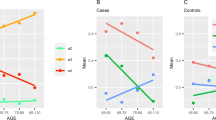
One-way ANOVA analysis of mean PRS scores between diagnosis groups, suggested an overall significance in PRS score (p < 0.0001). Pairwise post hoc Tukey analysis of the groups showed that the mean PRS of the BDR LOAD samples were found to be significantly higher than that of the controls (p < 0.0001 post hoc Tukey), likewise the mean PRS for the MCI converters was found to be higher than that of MCI non-converters, however this was not significant (Fig. 1). Post hoc Tukey also identified significant differences in mean PRS between non-converters and LOAD samples (p < 0.0001).
The range of scores for individuals within each group are described in the figure (grey circles) with the average PRS for each group indicated by the black circle. Significant differences were observed with one-way ANOVA across all four groups (p < 0.0001), with post hoc Tukey indicating significance between pairwise comparisons indicated with ****(p < 0.0001)
Predictability of the LOAD phenotype with PRS, including APOE with gender and age at death incorporated into the model, produced a final AUPRC for predictability for LOAD of 82.5%. The APOE genotypes alone were found to have a predictability of 81.8%. When applied to the MCI cohort for predictability of conversion with age at recruitment in the study instead of age at death, the full model produced a predictability of 61.0% for conversion, with APOE alone showing 43.8% predictability. This demonstrates the utility of the PRS to discern conversion from MCI to LOAD.
Samples for the BDR and MCI cohorts were separately partitioned into deciles of increasing disease risk based on PRS, including APOE genotypes, gender and age; the proportion of LOAD cases and controls which fell into each decile are depicted in Fig. 2. As of December 2017, 49 individuals diagnosed with MCI had converted to LOAD (39.3%); the proportion of converters and non-converters which fell into each decile are also given in Fig. 3. Both figures show the proportion of LOAD/MCI converters increases with increasing PRS.
Individual probabilities were generated using PRSice-2. The polygenic risk scores (PRS) including APOE and covariates for gender and age at death were used to distribute individuals into deciles. The dark bars represent the proportion of LOAD cases which fall into each decile and the light bars represent the proportion of controls. The number of LOAD cases and controls who fall within each decile are indicted in each block
Individual probabilities for the best predictive model were generated from polygenic risk scores (PRS) including APOE and covariates for gender and age at recruitment to distribute individuals into deciles. The dark bars represent the proportion of converting MCIs in each decile and the light bars represent the proportion of non-converting MCIs. The number of converters and non-converters who fall within each decile are listed
PRS is increasingly being used in genetic analyses to determine predictability of complex disease. In this study, cognitively healthy controls and individuals with LOAD or MCI were genotyped on the NeuroChip array33. PRS were generated from IGAP summary statistics and use to calculate risk scores for LOAD cases and controls from the BDR and MCI samples from the Southampton ICOS cohort. The best predictive model (82.5%) to distinguish LOAD cases from controls generated in the BDR cohort was found to be able to predict LOAD converters in the MCI cohort with an accuracy of 61.0%.
The difference between mean PRS of controls, non-converters, converters and LOAD cases was significant with one-way ANOVA, however, pairwise significance (post hoc Tukey) was only observed between LOAD cases and controls and LOAD cases and MCI non-converters. The distribution profile in Fig. 1 and Spearman’s rank order correlation (r = 0.408; p = 0.01) confirms an increased PRS with disease status from control through non-converters, converters to LOAD. The lack of a significant difference in average PRS between the MCI non-converters and converters is mostly like due to the small sample sizes, and it is expected that with increased numbers in each MCI group the mean PRS will move towards controls and LOAD cases, respectively. This hypothesis is supported by several studies suggesting that possession of LOAD-risk alleles are associated with faster cognitive decline, MCI and conversion from MCI to LOAD13,38,39,40, with one study concluding that carrying 6 or more of the non-APOE LOAD risk alleles rapidly increases conversion from MCI to LOAD14.
The resulting risk model, incorporating the APOE SNPs, 165 non-APOE SNPs, gender and age successfully predicted LOAD cases from controls with 82.5% accuracy which is a similar value to previous studies21,22,23,25.
As observed in previous PRS analyses41 there are a number of controls with high risk scores and cases of LOAD individuals with a low-risk scores (Fig. 2). It is possible that in these individuals, lifestyle and environmental factors could be determining the disease presence independent of, or interacting with genetic factors. Controls with higher PRS might represent individuals who would have developed disease had they lived longer as the average age at death of controls in the highest decile was significantly lower (73.5 ± 7.8 years) compared with the lowest decile (86.3 ± 5.0 years; p = 0.032). It is also possible that these individuals harbour unknown protective factors which may reduce disease risk.
Further study highlighted significantly later age at death in LOAD cases with the lowest risk compared to LOAD cases with the greatest risk (average age at death of LOAD in decile 1 = 95.0 ± 11.3; average age at death of LOAD in decile 10 = 81.1 ± 6.4; p = 0.007); this aligns with previous studies identifying later age at onset in cases with low PRS and increased prevalence with age;42,43. This could indicate that genetic load is associated with onset and/or severity of disease.
Predictive ability for LOAD conversion from MCI (61.0%) was less than between controls and LOAD (82.5%). This lower predictive ability is likely due to the smaller sample size of the MCI cohort, furthermore the ICOS cohort is a longitudinal study, and therefore unlike the BDR cohort which have the diagnoses post-mortem verified, the participants are still being clinically and cognitively assessed. It is possible that as the study progresses further individuals in this study will convert and influence the accuracy of the model.
The ability of the LOAD PRS model to predict to some degree the conversion of MCI to LOAD, highlights a possible genetic basis for conversion. Whereas non-converters have a genetic risk similar to that of controls, they may still harbour some genetic variations associated with LOAD which leads to the MCI phenotype though those with the lowest scores may never convert to LOAD. Conversely, the MCI subjects that did convert to LOAD have a PRS that is not significantly different to LOAD cases, which reinforces the idea that MCI can be seen as a prodromal state of LOAD and that by using approaches such as we describe, those at risk of developing LOAD can potentially be identified before the onset of LOAD symptoms and would be the best candidates to evaluate emerging therapeutic approaches.
Change history
11 June 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Gatz, M. et al. Role of genes and environments for explaining Alzheimer disease. Arch. Gen. Psychiatry 63, 168–174 (2006).
Guerreiro, R. et al. TREM2 variants in Alzheimer’s disease. N. Engl. J. Med. 368, 117–127 (2013).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116 (2013).
Lambert, J. C. et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer’s disease. Nat. Genet. 41, 1094–1099 (2009).
Sims, R. et al. Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat. Genet. 49, 1373–1384 (2017).
Chouraki, V. et al. Evaluation of a Genetic Risk Score to Improve Risk Prediction for Alzheimer’s Disease. J. Alzheimers Dis. 53, 921–932 (2016).
Marden, J. R. et al. Using an Alzheimer Disease Polygenic Risk Score to Predict Memory Decline in Black and White Americans Over 14 Years of Follow-up. Alzheimer Dis. Assoc. Disord. 30, 195–202 (2016).
Desikan, R. S. et al. Genetic assessment of age-associated Alzheimer disease risk: Development and validation of a polygenic hazard score. PLoS Med. 14, e1002258 (2017).
Sleegers, K. et al. A 22-single nucleotide polymorphism Alzheimer’s disease risk score correlates with family history, onset age, and cerebrospinal fluid Abeta42. Alzheimers Dement. 11, 1452–1460 (2015).
Verhaaren, B. F. et al. Alzheimer’s disease genes and cognition in the nondemented general population. Biol. Psychiatry 73, 429–434 (2013).
Xiao, Q. et al. Risk prediction for sporadic Alzheimer’s disease using genetic risk score in the Han Chinese population. Oncotarget 6, 36955–36964 (2015).
Yokoyama, J. S. et al. Decision tree analysis of genetic risk for clinically heterogeneous Alzheimer’s disease. BMC Neurol. 15, 47 (2015).
Adams, H. H. et al. Genetic risk of neurodegenerative diseases is associated with mild cognitive impairment and conversion to dementia. Alzheimers Dement. 11, 1277–1285 (2015).
Rodriguez-Rodriguez, E. et al. Genetic risk score predicting accelerated progression from mild cognitive impairment to Alzheimer’s disease. J. Neural Transm. 120, 807–812 (2013).
Harrison, T. M. et al. An Alzheimer’s disease genetic risk score predicts longitudinal thinning of hippocampal complex subregions in healthy older adults. eNeuro 3, 0098–16 (2016).
Sabuncu, M. R. et al. The association between a polygenic Alzheimer score and cortical thickness in clinically normal subjects. Cereb. Cortex. 22, 2653–2661 (2012).
Lupton, M. K. et al. The effect of increased genetic risk for Alzheimer’s disease on hippocampal and amygdala volume. Neurobiol. Aging 40, 68–77 (2016).
Martiskainen, H. et al. Effects of Alzheimer’s disease-associated risk loci on cerebrospinal fluid biomarkers and disease progression: a polygenic risk score approach. J. Alzheimers Dis. 43, 565–573 (2015).
Morgan, A. R. et al. The correlation between inflammatory biomarkers and polygenic risk score in Alzheimer’s disease. J. Alzheimers Dis. 56, 25–36 (2017).
Euesden, J., Lewis, C. M. & O’Reilly, P. F. PRSice: polygenic risk score software. Bioinformatics 31, 1466–1468 (2015).
Escott-Price, V. et al. Common polygenic variation enhances risk prediction for Alzheimer’s disease. Brain 138(Pt 12), 3673–3684 (2015).
Escott-Price, V., Myers, A. J., Huentelman, M. & Hardy, J. Polygenic risk score analysis of pathologically confirmed alzheimer’s disease. Ann. Neurol. 82, 311–314 (2017).
Escott-Price, V., Shoai, M., Pither, R., Williams, J. & Hardy, J. Polygenic score prediction captures nearly all common genetic risk for Alzheimer’s disease. Neurobiol. Aging 49, 214 e7–e11 (2017).
Axelrud, L. K. et al. Polygenic risk score for Alzheimer’s disease: implications for memory performance and hippocampal volumes in early life. Am. J. Psychiatry 175, 555–563 (2018).
Logue, M. W. et al. Use of an Alzheimer’s disease polygenic risk score to identify mild cognitive impairment in adults in their 50s. Mol. Psychiatry 24, 421–430 (2018).
Mormino, E. C. et al. Polygenic risk of Alzheimer disease is associated with early- and late-life processes. Neurology 87, 481–488 (2016).
Mitchell, A. J. & Shiri-Feshki, M. Rate of progression of mild cognitive impairment to dementia—meta-analysis of 41 robust inception cohort studies. Acta Psychiatr. Scand. 119, 252–265 (2009).
Tosto, G. & Reitz, C. Genome-wide association studies in Alzheimer’s disease: a review. Curr. Neurol. Neurosci. Rep. 13, 381 (2013).
Torkamani, A., Wineinger, N. E. & Topol, E. J. The personal and clinical utility of polygenic risk scores. Nat. Rev. Genet. 19, 581–590 (2018).
Francis, P. T., Costello, H. & Hayes, G. M. Brains for dementia research: evolution in a longitudinal brain donation cohort to maximize current and future value. J. Alzheimers Dis. 66(4), 1635–1644, https://doi.org/10.3233/JAD-180699 (2018).
Brookes, K. J. et al. Genotyping of the Alzheimer’s disease genome-wide association study index single nucleotide polymorphisms in the brains for dementia research cohort. J. Alzheimers Dis. 64, 355–362 (2018).
Sussams, R. et al. Systemic inflammatory responses to stress and its impact on cognition in people with mild cognitive impairment. Alzheimer’s Dement. 9, P775 (2013).
Blauwendraat, C. et al. NeuroChip, an updated version of the NeuroX genotyping platform to rapidly screen for variants associated with neurological diseases. Neurobiol. Aging 57, 247.e9–247.e13 (2017).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Lambert, J.-C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Grau, J., Grosse, I. & Keilwagen, J. PRROC: computing and visualizing precision-recall and receiver operating characteristic curves in R. Bioinformatics 31, 2595–2597 (2015).
Saito, T. & Rehmsmeier, M. The precision-recall plot is more informative than the ROC plot when evaluating binary classifiers on imbalanced datasets. PLoS ONE 10, e0118432 (2015).
Carrasquillo, M. M. et al. Late-onset Alzheimer’s risk variants in memory decline, incident mild cognitive impairment, and Alzheimer’s disease. Neurobiol. Aging 36, 60–67 (2015).
Jiang, Y., He, T., Deng, W. & Sun, P. Association between apolipoprotein E gene polymorphism and mild cognitive impairment: a meta-analysis. Clin. Inter. Aging 12, 1941–1949 (2017).
Moreno-Grau, S. & Ruiz, A. Genome research in pre-dementia stages of Alzheimer’s disease. Expert Rev. Mol. Med. 18, e11 (2016).
Chaudhury, S. et al. Polygenic risk score in postmortem diagnosed sporadic early-onset Alzheimer’s disease. Neurobiol. Aging 62, 244 e1–e8 (2018).
Tan, C. H. et al. Polygenic hazard scores in preclinical Alzheimer disease. Ann. Neurol. 82, 484–488 (2017).
Tosto, G. et al. Polygenic risk scores in familial Alzheimer disease. Neurology 88, 1180–1186 (2017).
Acknowledgements
We thank the International Genomics of Alzheimer’s Project (IGAP) for providing summary results data for these analyses. The investigators within IGAP contributed to the design and implementation of IGAP and/or provided data but did not participate in analysis or writing of this report. IGAP was made possible by the generous participation of the control subjects, the patients, and their families. The i-Select chips were funded by the French National Foundation on Alzheimer’s disease and related disorders. EADI was supported by the LABEX (laboratory of excellence programme investment for the future) DISTALZ grant, Inserm, Institut Pasteur de Lille, Université de Lille 2, and the Lille University Hospital. GERAD was supported by the Medical Research Council (Grant no. 503480), Alzheimer’s Research UK (Grant no. 503176), the Wellcome Trust (Grant no. 082604/2/07/Z), and German Federal Ministry of Education and Research (BMBF): Competence Network Dementia (CND) grant nos. 01GI0102, 01GI0711, and 01GI0420. CHARGE was partly supported by the NIH/NIA grant R01 AG033193 and the NIA AG081220 and AGES contract N01-AG-12100, the NHLBI grant R01 HL105756, the Icelandic Heart Association, and the Erasmus Medical Center and Erasmus University. ADGC was supported by the NIH/NIA grants: U01 AG032984, U24 AG021886, and U01 AG016976, and the Alzheimer’s Association grant ADGC-10-196728. We gratefully acknowledge all donors and their families for the tissue provided for this study. Human post-mortem tissue was obtained from the South West Dementia Brain Bank, London Neurodegenerative Diseases Brain Bank, Manchester Brain Bank, Newcastle Brain Tissue Resource and Oxford Brain Bank, members of the Brains for Dementia Research (BDR) Network. The BDR is jointly funded by Alzheimer’s Research UK and the Alzheimer’s Society in association with the Medical Research Council. We also acknowledge the neuropathologists at each centre and BDR Brain Bank staff for the collection and classification of the samples. The South West Dementia Brain Bank is part of the Brains for Dementia Research programme, jointly funded by Alzheimer’s Research UK and Alzheimer’s Society, and is supported by BRACE (Bristol Research into Alzheimer’s and Care of the Elderly) and the Medical Research Council. We thank the donor whose donation of brain tissue to the London Neurodegenerative Diseases Brain Bank allowed this work to take place. The Brain Bank is supported by the Medical Research Council and Brains for Dementia Research (jointly funded by the Alzheimer’s Society and Alzheimer’s Research UK). We acknowledge the support of the Manchester Brain Bank by Alzheimer’s Research UK and Alzheimer’s Society through their funding of the Brains for Dementia Research (BDR) Programme. Manchester Brain Bank also receives Service Support costs from Medical Research Council. Tissue provided by the Newcastle Brain Tissue Resource is funded, in part, by a grant from the UK Medical Research Council (G0400074), by NIHR Newcastle Biomedical Research Centre and Unit awarded to the Newcastle upon Tyne NHS Foundation Trust and Newcastle University, and by a grant from the Alzheimer’s Society and Alzheimer’s Research UK as part of the Brains for Dementia Research Project. Tissue for this study was provided by the Newcastle Brain Tissue Resource, which is funded, in part, by a grant from the UK Medical Research Council (G0400074) and by Brains for Dementia research, a joint venture between Alzheimer’s Society and Alzheimer’s Research UK. We acknowledge the Oxford Brain Bank, supported by the UK MRC, the NIHR Oxford Biomedical Research Centre and the Brains for Dementia Research programme for providing post-mortem specimens. Jose Bras and Rita Guerreiro’s work is funded by Fellowships from the Alzheimer’s Society. The ICOS study was funded by the Alzheimer’s Society (PG 143/150). The Human Genetics Group at the University of Nottingham was supported by Alzheimer’s Research UK (Program Grant 2809) to conduct genotyping and analyses of the BDR cohort and SC was supported by a PhD studentship from the Neurosciences Group at Queen’s Medical Centre (Nottingham) and the School of Life Sciences, University of Nottingham. We thank Dr William Rayner of The Wellcome Trust of Human Genetics for providing the strand files for alignment of the NeuroChip to the GRCh37 assembly.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chaudhury, S., Brookes, K.J., Patel, T. et al. Alzheimer’s disease polygenic risk score as a predictor of conversion from mild-cognitive impairment. Transl Psychiatry 9, 154 (2019). https://doi.org/10.1038/s41398-019-0485-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-019-0485-7
This article is cited by
-
Recent advances in polygenic scores: translation, equitability, methods and FAIR tools
Genome Medicine (2024)
-
A longitudinal study of polygenic score and cognitive function decline considering baseline cognitive function, lifestyle behaviors, and diabetes among middle-aged and older US adults
Alzheimer's Research & Therapy (2023)
-
Genetic loci of beta-aminoisobutyric acid are associated with aging-related mild cognitive impairment
Translational Psychiatry (2023)
-
Polygenic risk of paclitaxel-induced peripheral neuropathy: a genome-wide association study
Journal of Translational Medicine (2022)