Abstract
Genetic factors are strongly implicated in the susceptibility to develop externalizing syndromes such as attention-deficit/hyperactivity disorder (ADHD), oppositional defiant disorder, conduct disorder, and substance use disorder (SUD). Variants in the ADGRL3 (LPHN3) gene predispose to ADHD and predict ADHD severity, disruptive behaviors comorbidity, long-term outcome, and response to treatment. In this study, we investigated whether variants within ADGRL3 are associated with SUD, a disorder that is frequently co-morbid with ADHD. Using family-based, case-control, and longitudinal samples from disparate regions of the world (n = 2698), recruited either for clinical, genetic epidemiological or pharmacogenomic studies of ADHD, we assembled recursive-partitioning frameworks (classification tree analyses) with clinical, demographic, and ADGRL3 genetic information to predict SUD susceptibility. Our results indicate that SUD can be efficiently and robustly predicted in ADHD participants. The genetic models used remained highly efficient in predicting SUD in a large sample of individuals with severe SUD from a psychiatric institution that were not ascertained on the basis of ADHD diagnosis, thus identifying ADGRL3 as a risk gene for SUD. Recursive-partitioning analyses revealed that rs4860437 was the predominant predictive variant. This new methodological approach offers novel insights into higher order predictive interactions and offers a unique opportunity for translational application in the clinical assessment of patients at high risk for SUD.
Similar content being viewed by others
Introduction
Substance use disorders (SUD) and addiction represent a global public health problem of substantial socioeconomic implications1,2. In 2010, 147.5 million cases of alcohol and drug abuse were reported (Whiteford et al., 2015), and SUD prevalence is expected to increase over time. Genetic factors have been implicated in SUD etiology, with genes involved in the regulation of several neurobiological systems (including dopaminergic and glutamatergic) found to be important (for a review see Prom-Wormley et al., 20173). However, limitations intrinsic to most genetic epidemiological studies support the search for additional risk genes.
Attention-deficit/hyperactivity disorder (ADHD), the most common neurodevelopmental behavioral disorder4,5, is frequently co-morbid with disruptive behaviors such as oppositional defiant disorder (ODD), conduct disorder (CD), and SUD6,7. The close association between ADHD and disruptive behaviors is summarized by longitudinal observations in ADHD cohorts6,8,9. Children diagnosed with ADHD monitored during the transition into adolescence exhibit higher rates of alcohol, tobacco, and psychoactive drug use than control groups of children without ADHD10,11. It has been estimated that the lifetime risk for SUD is ~50% in subjects with childhood ADHD persisting into adulthood12,13. Reciprocally, the prevalence of ADHD is high in adolescents with SUD9,14,15 and the presence of an ADHD diagnosis affects SUD prognosis, with ADHD being associated with both earlier and more frequent alcohol-related relapses16 and lower likelihood of cannabis-dependence treatment completion17.
Strong evidence from family, twin, and genome-wide linkage and association studies suggests that genetic factors play a crucial role in shaping the susceptibility to both ADHD and SUD18,19,20,21. During the last 15 years, we have collected families clustering individuals affected with ADHD and disruptive behaviors from disparate regions around the world6,18,22,23. Although the prevalence of ADHD co-morbid with disruptive behaviors is variable across populations, we found a higher frequency of CD, ODD, and SUD (mainly nicotine dependence and alcohol abuse) in ADHD individuals than in unaffected relatives6,22,24. Using genome-wide data from extended multigenerational families, we found evidence of linkage of ADHD to markers in chromosomes 4q13.2, 5q33.3, 8q11.23, 11q22, and 17p1125, and co-segregation of ADHD and disruptive behaviors with loci at 2p21-22.3, 4q13.2, 5p13.1-p13.3, 8q24, 8q15, 11q22, 12p11.23-13.3, and 14q21.1-22.28. Fine mapping of the 4q13.2 region identified variants in the adhesion G-protein-coupled receptor L3 gene (ADGRL3, also known as latrophilin 3 or LPHN3) that predispose to ADHD22,24,26,27,28,29,30.
Characterization of the association between ADHD and ADGRL3 has provided key information to better predict the severity of ADHD, the long-term outcome, the patterns of brain metabolism, and the response to stimulant medication24,27,29,31,32,33,34. To the best of our knowledge, ADGRL3 linkage and association results represent some of the most robustly replicated genetic and pharmacogenetic findings in ADHD genetic research. While ADGRL3 has also shown association with disruptive behaviors in the context of ADHD18,24,35, a direct link to SUD has not been systematically investigated. In this manuscript we tested the hypothesis that ADHD risk variants harbored at the ADGRL3 locus interact with clinical, demographic, and environmental variables associated with SUD.
Subjects and methods
Subjects
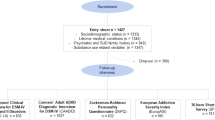
We used independent populations from disparate regions of the world (n = 2698) ascertained through patients affected with ADHD co-morbid with disruptive behaviors (Paisa, Spanish and MTA samples) or SUD (Spanish and Kentucky samples).
Paisa sample
This population isolate is unique in that it was used to identify ADHD susceptibility genes by linkage and association strategies. Detailed clinical and demographic information on this sample has been published elsewhere23,25,29. The sample consists of 1176 people (adults, adolescents, and children), mean age 28 ± 17 years, ascertained from 18 extended multigenerational and 136 nuclear Paisa families inhabiting the Medellin metropolitan area in the State of Antioquia, Colombia. Initial coded pedigrees were obtained through a fixed sampling scheme from a parent or grandparent of an index proband after having collected written informed consent from all subjects or their parent/guardian, as approved by the University of Antioquia and the NIH Ethics Committees, and in accordance with the Helsinki Declaration. Patients were recruited under NHGRI protocol 00-HG-0058 (NCT00046059).
Exclusion criteria for ADHD participants were IQ < 80, or any autistic or psychotic disorders. Parents underwent a full psychiatric structured interview regarding their offspring (Diagnostic Interview for Children and Adolescents—Revised—Parents version (DICA-IV-P, Spanish version translated with permission from Dr. Wendy Reich (Washington University, St. Louis). All adult participants were assessed using the Composite International Diagnostic Interview (CIDI), as well as the Disruptive Behavior Disorders module from the DICA-IV-P modified for retrospective use. The interview was conducted by a “blind” rater (either a psychologist, a neuropsychologist, or a psychiatrist) at the Neurosciences Clinic of the University of Antioquia, or during home visits. ADHD status was defined by the best estimate method. Specific information regarding clinical diagnoses and co-morbid disruptive disorders, affective disorders, anxiety, and substance use has been published elsewhere3.
From the 1176 individuals in this cohort, only founder members were included in analyses (n = 472). This was done to avoid kinship relatedness bias and to exclude children and adolescents, as they may have not been exposed to substances of abuse yet. Of these 472 individuals, 17% (n = 79) fulfilled criteria for ADHD, 17% (n = 78) for ODD, 18% (n = 84) for CD, 22% nicotine dependence (n = 102), 27% alcohol dependence (n = 124), 3% drug dependence (n = 12), 37% social/simple phobia (n = 156), 13% any other anxiety disorder (n = 58), and 25% major depressive disorder (n = 117) (Table 1).
Spanish sample
The ADHD sample consisted of 670 adult ADHD patients, mean age 33 ± 10 years, 69% males (n = 461), recruited and evaluated at the Psychiatry Department of the Hospital Universitari Vall d’Hebron (Barcelona, Spain) according to DSM-IV TR criteria. ADHD diagnosis was based on the Spanish version of the Conners Adult ADHD Diagnostic Interview for DSM-IV (CAADID)36. Comorbidity was assessed by Structured Clinical Interview for DSM-IV Axis I and Axis II Disorders (SCID-I and SCID-II). ODD during childhood and adolescence was retrospectively evaluated with the Schedule for Affective Disorders and Schizophrenia (SADS) for School-Age Children, Present and Lifetime Version (K-SADS). Thirty-nine percent of ADHD patients (n = 263) fulfilled diagnostic criteria for SUD, 21% for disruptive behavior disorders (CD and/or ODD; n = 142), 21% for depression (n = 143), 13% for anxiety (n = 89), and 8% for phobias (n = 50). The level of impairment was measured with the Clinical Global Impression (CGI) included in the CAADID Part II and the Sheehan Disability Inventory. Exclusion criteria for ADHD patients were IQ < 80; pervasive developmental disorders; schizophrenia or other psychotic disorders; presence of mood, anxiety or personality disorders that might explain ADHD symptoms; birth weight ≤ 1.5 kg; and other neurological or systemic disorders that might explain ADHD symptoms.
The SUD sample consisted of 494 adults (mean age 37 ± 9 years and 76% males, n = 376) recruited and evaluated at the Addiction and Dual Diagnosis Unit of the Psychiatry Department at the Hospital Universitari Vall d’Hebron with the Structured Clinical Interview for DSM-IV Axis I Disorders (SCID-I). All patients fulfilled DSM-IV criteria for drug dependence beyond nicotine dependence. None were evaluated for ADHD.
The control sample consisted of 483 blood donors (mean age 42 ± 20 years, 74% males) in which DSM-IV lifetime ADHD symptomatology was excluded under the following criteria: (1) not having been diagnosed with ADHD and (2) answering negatively to the lifetime presence of the following DSM-IV ADHD symptoms: (a) often has trouble keeping attention on tasks, (b) often loses things needed for tasks, (c) often fidgets with hands or feet or squirms in seat, and (d) often gets up from seat when remaining in seat is expected. Individuals affected with SUD were excluded from this sample. None of them had self-administered drugs intravenously. It is important to mention that the exposure criterion was not applied; therefore, this set cannot be classified as “pure” controls.
All patients and controls were Spanish of Caucasian descent. This study was approved by the ethics committee of the Hospital Universitari Vall d’Hebron and informed consent was obtained from all subjects in accordance with the Helsinki Declaration.
MTA sample
The Multimodal Treatment Study of Children with ADHD (MTA) was designed to evaluate the relative efficacy of treatments for childhood ADHD, combined subtype, in a 14-month randomized controlled trial of 579 children assigned to four treatment groups: medication management, behavior modification, their combination, and treatment as usual in community care. After the 14-month treatment-by-protocol phase, the MTA continued as a naturalistic follow-up in which self-selected use of psychoactive medication was monitored. A local normative comparison group of 289 randomly selected classmates group-matched for grade and sex was added when the ADHD participants were between 9–12 years of age. The outcomes in childhood (14, 24, and 36 months after baseline), and adolescence (6 and 8 years after baseline) and into adulthood (12, 14, and 16 years after baseline) have been reported10,11,37,38,39,40,41,42,43. Substance use was assessed with a child/adolescent-reported questionnaire adapted for the MTA11,43. The measure included items for lifetime and current (previous 6 months) use of alcohol, cigarettes, tobacco, cannabis, and other recreational drugs. Also included were items for non-prescribed use or misuse of psychoactive medications, including stimulants. The measure was modeled after similar substance use measures in longitudinal or national survey studies of alcohol and other drug use that also rely on confidential youth self-report as the best source of data44,45. A National Institutes of Health (NIH) Certificate of Confidentiality further strengthened the assurance of privacy. Substance use was coded positive if any of the following behaviors, selected after examining distributions, were endorsed as occurring in the participant’s lifetime up to 8 years post-baseline: (1) alcohol consumption (more than just a sip) more than five times or drunk at least once; (2) cigarette smoking or tobacco chewing more than a few times; (3) cannabis use more than once; or (4) use of inhalants, hallucinogens, cocaine, or any of amphetamines/stimulants, barbiturates/sedatives, and opioids/narcotics without a prescription or misused a prescription (used in greater quantity or more often than prescribed). Each of the four types of substances, as well as daily use of tobacco and the number of substance use classes endorsed (0, 1, 2, or more), were explored in secondary analyses.
DSM-IV abuse or dependence was based on a positive parent or child report with the Diagnostic Interview Schedule for Children version 2.3/3.0 (DISC)46 at the 6- and 8-year follow-up assessments. The DISC includes both lifetime and past year diagnoses. The Diagnostic Interview Schedule-IV47 was used at the 8-year follow-up for 18 + year-olds (n = 111). SUD was defined as the lifetime presence of any abuse or dependence (excluding tobacco dependence, due to differences in the meaning of abuse/dependence for tobacco versus other substances).
Additional analyses explored SUD for alcohol, tobacco, and cannabis/other drugs (recreational or misused prescription medications) separately10. All patients in this study provided informed written consent as approved by the NIH Ethics Committee.
Kentucky sample
A sample of 560 inpatients and outpatients with severe SUD from Central Kentucky psychiatric facilities was collected during a pharmacogenetics investigation48. Patient interviews and medical record information (including urine drug screens and substance abuse counselor notes) were used by the research nurse to assess the Clinician Rating of Alcohol (CRAUD) and Drug Use Disorder (CRDUD)49,50 that provides a score from 1 = abstinence (not used in the assessed period) to 5 = severe dependence. Scores of 3 and higher are pathological and were considered positive in our analyses. All drugs were combined into one rating48. Descriptions of the training provided to research nurses to assess the CRAUD and CRDUD were published elsewhere48,51.
DNA was available from 533 of 560 study subjects. Of the 533 subjects with available DNA, 53% (n = 285) were male, 82% (n = 436) were Caucasian, 16% (n = 87) were African American, and 2% (n = 10) were from other ethnicities. Additional clinical information for this sample has been described elsewhere48,51 and included: (1) clinical diagnosis obtained from medical records, (2) prior psychiatric history, (3) history of daily smoking, (4) reviews of current and psychiatric medication use, and (5) body mass index (Supplemental Table 2). All participants in the Kentucky study provided informed written consent as approved by the University of Kentucky IRB.
Genotyping
DNA was extracted from whole blood (Paisa, Spanish and MTA sample) or buccal swabs (Kentucky sample) using standard protocols. The Paisa sample was genotyped using the service provided by Illumina (San Diego, CA). The Spanish, MTA, and Kentucky samples were genotyped for select variants using pre-designed TaqMan® SNP genotyping assays (Thermo Fisher Scientific, Waltham, MA). Allelic discrimination real-time PCR reactions were performed in a 384-well plate format for each individual sample according to the manufacturer’s instructions. Briefly, 20 ng of genomic DNA were mixed with 2.5 μL of 2X TaqMan Universal PCR Master Mix and 0.25 μL of 20X SNP Genotyping Assay in a total volume of 5 μL per reaction. Assays were run in an ABI 7900HT Fast Real-Time PCR System (Thermo Fisher Scientific). Allele calling was made by end-point fluorescent signal analysis using the ABI’s SDS2.3 software. In addition, we had previously collected exome genotype data from the MTA sample26 using the Infinium® HumanExome-12 v1.2 BeadChip kit (Illumina), which covers putative functional exonic variants selected from over 12,000 individual exome and whole-genome sequences. Processed and raw intensity signals for the array data can be accessed at GEO (GSE112652). SNP markers harbored at the ADGRL3 gene were filtered in from this dataset and added to those genotyped using TaqMan® assays.
Dataset quality control and preparation for analysis
Genotype data were imported into Golden Helix® SVS 8.3.1 (Golden Helix, Bozeman, MT) for quality control analysis. Markers with a minor allele frequency (MAF) < 0.01 (rare variants), significant deviation from Hardy–Weinberg equilibrium (P-values < 0.0001), and a genotyping success rate < 90%, were excluded. For the Paisa and Spanish samples, a subset of variants in the ADGRL3 minimal critical region (MCR), 5′UTR and 3′UTR were selected based on a previous ADHD association study30. Because the Paisa sample is a family-based cohort and recursive-partitioning analysis does not correct for kinship relatedness, only founder members from the pedigrees were included in the analyses. For the MTA sample, a total of 8568 markers with a MAF ≥ 1.0 % from the 244,414 markers genotyped with the exome chip were filtered out using linkage disequilibrium (LD) pruning, and variants within ADGRL3 were selected for analyses. For the Kentucky sample, only four ADGRL3 variants were selected for analyses after LD pruning of a list of markers located within the ADGRL3 5′UTR and MCR regions that was available to us. Variants rs7659636 and rs5010235 had been imputed from ADHD genome-wide association data funded through the Genetics Analysis Information Network (GAIN) initiative, a public-private partnership between the NIH and the private sector (https://www.genome.gov/19518664/genetic-association-information-network-gain/#al-4). ADGRL3 variants used in this study for each cohort are presented in Supplemental Table 3.
Advanced recursive-partitioning (tree-based) approach (ARPA)
Association studies of ADGRL3 variants with ADHD, ODD, CD, response to stimulant treatment and severity outcome have been published elsewhere for the Paisa and Spanish populations24,29,32,52. We used ARPA to build a predictive framework to forecast the behavioral outcome of children with ADHD, suitable for translational applications. Our goal was to test the hypothesis that ADGRL3 variants predisposing to ADHD also increase the risk of co-morbid disruptive symptoms, including SUD.
ARPA is a tree-based method widely used in predictive analyses because it accounts for non-linear and interaction effects, offers fast solutions to reveal hidden complex substructures and provides truly non-biased statistically significant analyses of high-dimension, seemingly unrelated data53. In a visionary manuscript, D.C. Rao suggested that recursive-partitioning techniques could be useful for genetic dissection of complex traits54. ARPA accounts for the effect of hidden interactions better than alternative methods, and is independent of the type of data (i.e., categorical, continuous, ordinal, etc.) and of the type of data distribution (i.e., fitting or not fitting normality)54. Furthermore, results supplied by tree-based analytics are easy to interpret visually and logically53. Therefore, to generate the most comprehensive and parsimonious classificatory model to predict the susceptibility to disruptive behaviors, we applied ARPA using a set of different modules implemented in the Salford Predictive Modeler® (SPM) software, namely, Classification and Regression Trees (CART), Random Forest, and TreeNet (http://www.salford-systems.com). One important advantage of SPM when compared to other available data mining software is its ability to use raw data with sparse or empty cells, a problem frequently encountered in genetic data.
Briefly, CART is a non-parametric approach whereby a series of recursive subdivisions separate the data by dichotomization55. The aim is to identify, at each partition step, the best predictive variable and its best corresponding splitting value while optimizing a splitting statistical criterion, so that the dataset can be successfully split into increasingly homogeneous subgroups55. We used a battery of different statistical criteria as splitting rules (e.g., GINI Index, Entropy, and Twoing) to determine the splitting rule, maximally decreasing the relative cost of the tree while increasing the prediction accuracy of target variable categories55. The best split at each dichotomous node was chosen by either a measure of between-node dissimilarity or iterative hypothesis testing of all possible splits to find the most homogeneous split (lowest impurity). Similarly, we used a wide range of empirical probabilities (priors) to model numerous scenarios recreating the distribution of the targeted variable categories in the population55. Following this iterative process, each terminal node was assigned to a class outcome. To avoid finishing with an over-fitted CART predictive model (a common problem in CART analyses), and to ensure that the final splits were well substantiated, we applied tree pruning. During the procedure, predictor variables that were close competitors (surrogate predictors with comparable overall classification error to the optimal predictors) were pruned to eliminate redundant commonalities among variables, so the most parsimonious tree would have the lowest misclassification rate for an individual not included in the original data55.
Additionally, we applied the Random Forest (RF) methodology using a bagging strategy to exactly identify the most important set of variables predicting disruptive behaviors56. The RF strategy differs from CART in the use of a limited number of variables to derive each node while creating hundreds to thousands of trees. This strategy has proved to be immune to the over fitting generated by CART56. In RF, variables that appeared repeatedly as predictors in the trees were identified. The misclassification rate was recorded for each approach.
The TreeNet strategy was used as a complement to the CART and RF strategies because it reaches a level of accuracy that is usually not attainable by single models such as CART or by ensembles such as bagging (i.e., RF)57. The TreeNet algorithm generates thousands of small decision trees built in a sequential error-correcting process converging on an accurate model57. The number of variables considered to derive each node with RF was \(\sqrt n\), where n is the number of independent variables (either 3 or 4).
To derive honest assessments of the derived models and have a better view of their performance on future unseen data, we applied a cross-validation strategy where both training with all the data and then indirectly testing with all the data were performed. To do so, we randomly divided the data into separate partitions (folds) of different sizes. This strategy allowed us to review the stability of results across multiple replications55. We used a 10-fold cross-validation as implemented in the SPM software.
A fixed-effects meta-analysis of the overall fraction of correctly classified individuals (accuracy) using the derived models from each of the four samples was applied to derive a general perspective of the SUD predictive capacity of this demographic-clinical-genetic framework.
Results
A series of predictive models were built on our data using combinations of the following criteria: (i) the rules of splitting (GINI index, twoing, order twoing, and entropy); (ii) the priors; (iii) the size of the terminal nodes; (iv) the costs; (v) the depth of branching; and (vi) the size of the folds for cross-validation, to maximize the accuracy of the derived classification tree while considering class assignment, tree pruning, testing and cross-validation.
A parsimonious and informative reconstructed predictive tree derived from CART for the Paisa sample revealed demographic (age), clinical (CD), and genetic variables (rs5010235 and rs4860437) (Fig. 1a). The importance of these variables was corroborated, and their potential over fitting discarded by the TreeNet analyses that revealed a set of predictors for SUD containing those derived by CART (Fig. 1b). This predictive model displays good sensitivity and specificity as shown by areas under the receiver-operating characteristic (ROC) curve (0.954 and 0.87 for the learning and the test data, respectively) during TreeNet cross-validation using folding (Fig. 1c). The proportions of misclassification for SUD cases in the cross-validation experiment for the learning and testing data were 0.124 and 0.177, respectively (Fig. 1d).
a Derived Classification and Regression Tree (CART) for SUD status as categorical target variable (disjunctive affection status, i.e., substance use of either alcohol, or nicotine, or other drugs). Only founder individuals were included in the analysis to avoid kinship relatedness bias. Class 0 (unaffected) is indicated in red and class 1 (affected) in blue. This derived tree for the Paisa sample included demographic (age), clinical (conduct disorder (CD)), and genetic variables (markers rs5010235 and rs4860437). The T allele of the rs4860437 variant (node 4) generates a highly discriminant split in combination with age (45.5 years) to terminal node 3 of ADHD individuals without CD (see root node 1). b Variable importance scores derived by Random Forest and TreeNet analysis were compatible with the variables included in the tree derived by CART. c, d TreeNet analysis to maximize the ROC area and minimize the classification error using 200 trees. The areas under the ROC curve (AUC) were 0.954 and 0.87 for learning and testing samples (blue and red curves and values, respectively), while the proportions of misclassification for SUD cases in the cross-validation experiment were 0.124 and 0.177 for learning and testing data sets, respectively
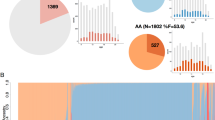
In the case of the Spanish sample, a parsimonious and informative tree was reconstructed with CART revealing demographic (sex), clinical (CD, ODD, depression, and ADHD), and genetic variables (rs4860437 and rs1868790) (Fig. 2a). The TreeNet analysis revealed a set of predictors for SUD containing those derived by CART (Fig. 2b). This predictive model displayed good sensitivity and specificity as shown by areas under the ROC curve (AUC) of 0.911 and 0.897 for learning and testing samples, respectively, during TreeNet cross-validation using folding (Fig. 2c). The proportions of misclassification for SUD cases obtained by TreeNet analysis for learning and testing data were 0.151 and 0.175, respectively (Fig. 2d).
a Derived tree by CART for SUD status as categorical target variable (disjunctive affection status, i.e., substance use of either alcohol, or nicotine, or other drugs). This derived tree for the Spanish sample included demographic (sex), clinical (CD, ODD, depression, and ADHD), and genetic variables (markers rs677642, rs4860437, rs1868790). b Variable importance scores derived by Random Forest and TreeNet analysis were compatible with the variables included in the tree derived by CART. c, d TreeNet analysis to maximize the AUC and minimize the classification error using 200 trees. The AUC were 0.911 and 0.897 for learning and testing samples while the proportions of misclassification for SUD cases in the cross-validation experiment were 0.151 and 0.175 for learning and testing data sets, respectively. Conventions as in Fig. 1
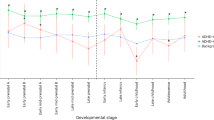
As in the previous cohorts, for the MTA sample we derived a parsimonious and informative predictive tree with CART depicting demographic (site of ascertainment), and genetic variables (rs2172802, rs61747658, rs12509110, and rs6856328) (Fig. 3a). The TreeNet analyses revealed a set of predictors for SUD containing those derived by CART (Fig. 3b). This predictive model displays good sensitivity and specificity as showed by AUC of 0.808 and 0.643 for learning and testing samples, respectively, during TreeNet cross-validation using folding (Fig. 3c). The proportions of misclassification for SUD cases obtained by TreeNet analysis for learning and testing data were 0.314 and 0.358, respectively (Fig. 3d).
a Derived tree by CART for the SUD status as categorical target variable (disjunctive affection status, i.e., substance use of either alcohol, or nicotine, cannabis, or other drugs). As the MTA is a longitudinal study, we used SUD status at 96 and 120 month follow-ups and applied a lag analysis of SUD emergence. The derived tree included demographic (site of ascertainment), and genetic variables (markers rs2172802, rs61747658, rs12509110, and rs6856328). The combination of variants rs61747658 and rs2172802 generated an important discriminant splitting of SUD affected and unaffected classes. b Variable importance scores derived by Random Forest and TreeNet analysis were compatible with the variables included in the tree derived by CART. c, d TreeNet analysis to maximize ROC area and minimize classification error using 200 trees. The AUC were 0.808 and 0.643 for learning and testing samples, while the proportions of misclassification for SUD cases in the cross-validation experiment, for learning and testing data were 0.314 and 0.358, respectively. Conventions as in Fig. 1
Finally, for the Kentucky sample, we derived a parsimonious and informative predictive tree with CART involving demographic (sex), clinical (high body mass index (HBMI) and schizophrenia diagnosis), and genetic variables (rs4860437 and rs7659636) (Fig. 4a). The TreeNet analyses revealed a set of predictors for SUD containing those derived by CART (Fig. 4b). This predictive model displays good sensitivity and specificity as showed by AUC of 0.811 and 0.744 for learning and testing samples, respectively, during TreeNet cross-validation using folding (Fig. 4c). The proportions of misclassification for SUD cases obtained by TreeNet analysis for learning and testing data were 0.285 and 0.252, respectively (Fig. 4d). The results from the RF analysis were consistent with those produced by TreeNet cross-validation using folding.
a Derived CART tree for SUD status as categorical target variable (disjunctive affection status, i.e., substance use of either alcohol, or nicotine, or other drugs). This derived tree for the Kentucky sample included demographic (sex), clinical (high Body Mass index [HBMI], and schizophrenia diagnosis), and genetic variables (markers rs4860437 and rs7659636). Notably, the T allele of the rs4860437 variant generated a split in the same direction as occurred for the derived tree in the Paisa and in the Spain samples. b Variable importance scores derived by Random Forest and TreeNet analysis were compatible with the variables included in the tree derived by CART. c, d TreeNet analysis to maximize ROC area and minimize classification error using 200 trees. The AUC were 0.811 and 0.744 for learning and testing samples, respectively, while the proportions of misclassification for SUD cases in the cross-validation experiment, for learning and testing data were 0.285 and 0.252, respectively. Conventions as in Fig. 1
A fixed-effects meta-analysis for overall accuracy returned a value of 0.727 (95% CI = 0.710–0.744) (Fig. 5), suggesting potential eventual clinical utility of predictive values. Overall, ADGRL3 marker rs4860437 was the most important variant predicting susceptibility to SUD, a commonality suggesting that these networks may be accurate in predicting the development of SUD based on ADGRL3 genotypes.
We conducted independent analyses for alcohol or nicotine dependence and compared these results with those of our composite SUD phenotype, as defined by the disjunctive presence of substance use phenotypes and explained by likely common neuropathophysiological mechanisms. In general, across cohorts, we found significant alcohol and nicotine risk variants, some of which have reasonably high odd ratios (OR). For instance, in the Spain sample, marker rs2271339 conferred significant risk to nicotine use: the heterozygote genotype A/G confers 43% increased risk of being diagnosed with nicotine use (OR = 1.43, 95% CI = 1.12–1.82). In the same vein, we found in the Paisa sample that the heterozygote A/T genotype for rs1456862 confers 83% increased risk to nicotine use (corrected OR = 1.84, 95 CI% = 1.03–3.38) than the A/A genotype. Regarding alcohol use, we found in the Paisas that the heterozygote C/T genotype for rs2159140 confers susceptibility, whereas the C/C genotype does not (corrected OR = 1.64, 95 CI% = 1.01–2.72). Supplemental Fig. 1 shows the ROC curves of nicotine and alcohol use prediction in the Paisa sample. Note that the AUC is greater than 0.7 in both cases, which suggests a straight performance of markers rs1456862 and rs2159140 in predicting nicotine and alcohol use, respectively.
To determine the significance of improvement of prediction when genetic markers are introduced in the ARPA-based predictive model for SUD, we compared the performance measures (i.e., sensitivity, specificity, classification rate, and lift) across all cohorts under two disjunctive scenarios: inclusion of genetic markers or not. We found that including genetic markers improved the performance measures of the resulting ARPA-based predictive model of SUD, regardless of cohort (Supplemental Fig. 2 and Supplemental Table 1). For instance, the AUC for the Spain sample was 81.6% (95% CI = 79.8–83.4) when genetic information was included, and 77.5 (95% CI = 75.9–79.1) when it was excluded. A bootstrap-based test with 10,000 replicates revealed that the former AUC was statistically greater than the latter (P < 0.0001, Supplemental Table 1). Similar results were obtained for the Paisa sample: the AUC was 90% (95% CI = 86.6–93.0) when genetic information was included versus 78.8% (95% CI = 75.8–81.7) when it was not (P < 0.0001, Supplemental Table 1). Improvements were also observed in the correct classification rate for the Spanish and Paisa samples, the sensitivity values in all samples, the specificity in the Spanish and Paisa samples, and the lift in the Paisa sample (Supplemental Table 1). Similar results were observed for the MTA and Kentucky samples, where including genetic information in the predictive model for SUD drastically improved these performance measures (Supplemental Table 1).
Discussion
SUD genetic epidemiological studies across multiple substances have been plagued with inconsistency in the replication of genetic association results. This may be due to reasons such as: (i) small effect size of variants expected to influence the SUD phenotype, as with any complex disease;58 (ii) insufficient power to detect significant associations due to small sample size;59 (iii) phenotypic heterogeneity of SUD across samples that may reflect different disease stages or multiple subtypes (i.e., single-drug versus poly-drug dependence/use); (iv) genetic heterogeneity arising from distinct risk genes sets; (v) ethnicity inconsistencies between discovery and replication samples;60,61 and (vi) comorbidity with other psychiatric conditions (e.g., ADHD) with shared genetic and environmental architecture62,63. Consequently, additional studies are required to identify new SUD candidate genes and to help dissect genetic contributions in the context of complex interactions with co-morbid conditions.
In this study, we present a demographic, clinical and genetic framework generated using ARPA that is able to predict the risk of developing SUD. Interestingly, marker rs4860437 showed a differential splitting pattern in the Paisa, Spain, and Kentucky cohorts. For instance, in Fig. 1a, rs4860437 splits into (G/G, G/T) and T/T; in Fig. 2a, the same variable splits into (G/G, T/T) and G/T; and in Fig. 4a, it splits into (G/T, T/T) and G/G. The most parsimonious and plausible explanation of this splitting pattern is the presence of genomic variability surrounding this proxy marker, reflecting ancestral composition. Future studies of genomic regions surrounding rs4860437 might reveal a cryptic mechanism. It is particularly compelling that ADGRL3 marker rs4860437, which is a major predictor variable component in the trees for SUD, is in complete LD with ADHD susceptibility markers rs6551665 and rs1947274 in Caucasians28,30,52, suggesting that the phenotype underpinning SUD is under the pleiotropic effect of ADGRL3 variants. Unfortunately, rs4860437 was not included in the exome chip used to genotype the MTA sample and, therefore, could not be included in the analyses for this sample. Given the limited overlap of markers across datasets and possible stratification differences among study populations, a gene- rather than a marker-level approach has been advocated64.
Adopting such a perspective, our results suggest that genetic variants harbored in the ADGRL3 locus confer susceptibility to SUD in populations from disparate regions of the world. These populations are from three different countries and involve different investigators, diverse inclusion criteria, and different clinical assessments, which suggests that our results may replicate in other settings and are likely to be clinically relevant. Of particular interest is the generalization of our findings to a longitudinal study (the MTA sample), where adding genetic information to baseline data predicted the development of SUD at later ages, as determined from information gathered over a period of more than 10 years. Additionally, our results generalized to a sample of patients with severe SUD from Kentucky (U.S.) that were not ascertained on the basis of ADHD diagnosis.
The first genome-wide significant ADHD risk loci were published recently65. Marker rs4860437 is not represented in this dataset; however, this study was not aimed at identifying loci shared between ADHD and SUD. In any case, while genome-wide association studies are a useful tool for discovering novel risk variants—as it involves a hypothesis-free interrogation of the entire genome—the lack of genetic association may be a reflection of the polygenic, multifactorial nature of ADHD, with both common and rare variants likely contributing small effects to its etiology66,67,68. In addition, an important factor may be the genetic heterogeneity of ADHD subtypes, which may have different underlying genetic mechanisms. Therefore, genome-wide significance may identify loci with larger genetic effects, while others with smaller effects remain undetected for a given population size.
Variation in ADGRL3 has been implicated in ADHD in diverse populations24,27,28,29,31,32,33,34,69. ADGRL3 is a member of the latrophilin subfamily of G-protein-coupled receptors (GPCR)70 and is most strongly expressed in brain regions implicated in the neurophysiological basis of ADHD32,52,71. Mouse and zebrafish knockout models also support ADGRL3 implication in ADHD pathophysiology72,73. More recently, Martinez et al35. identified a brain-specific transcriptional enhancer within ADGRL3 that contains an ADHD risk haplotype associated with reduced ADGRL3 mRNA expression in the thalamus. This haplotype was associated not only with ADHD, but also with disruptive behaviors, including SUD35. A member of the family of leucine-rich repeat transmembrane (FLRT) proteins has been identified as an endogenous postsynaptic ligand for latrophilins74. Interference with this interaction reduces excitatory synapse density in cultured neurons and decreases afferent input strength and dendritic spine number in dentate granule cells, which implicates ADGRL3 and FLRT3 in glutamatergic synapse development74. Similarly, convergent evidence from a network analysis of a gene set significantly associated and/or linked to ADHD and SUD revealed pathways involved in axon guidance, regulation of synaptic transmission, and regulation of transmission of nerve impulse18. These data altogether suggest that ADGRL3 may be an important SUD susceptibility gene.
Strong evidence from clinical and genetic association studies suggests that genetic factors play a crucial role in shaping the susceptibility to both ADHD and SUD75,76,77,78,79,80,81. More strikingly, ADHD treatment has been shown to reduce the risk of SUD82,83,84. Though the neurobiological basis for this association remains unclear, a variety of causal pathways from ADHD to SUD have been proposed that involve conduct problems77,79. Clinical studies have suggested that the link between SUD and ADHD disappears after controlling for co-morbid CD84,85,86. In agreement with these studies, the presence of CD was a major predictor of SUD in the ARPA-based predictive models for SUD in the Paisa and Spanish cohorts (both assessed first for ADHD) (Figs. 1 and 2).
Some researchers implicate genetically mediated personality traits, such as impulsivity and lack of inhibitory control (common to ADHD and disruptive behaviors) as a link between ADHD and SUD resulting from common neurological substrates87. Some investigators have proposed that patients with ADHD use addictive substances to self-medicate88 and that the differential response to drugs of abuse and atypical behavioral regulation in response to social cues (e.g., social modeling and peer selection) may fuel substance use77,89. Others suggest that the poor judgment and impulsivity associated with ADHD contribute to the development of substance dependence79. Clinical variables from childhood have also been associated with SUD in patients with ADHD, such ADHD subtype, temper characteristics (fear, accident propensity), sexual abuse, suspension from school, and a family history of ADHD13.
In summary, our results support a possible functional role for ADGRL3 in modulating drug seeking behavior. Regardless of the type of abused substance, longitudinal studies generally find that the onset of ADHD precedes that of SUD, suggesting that the psychopathology of ADHD is not secondary to SUD in most patients79. Accordingly, it is reasonable to consider that timely diagnosis and treatment of ADHD with stimulant medication may reduce the occurrence and/or severity of SUD. Based on the relationship with medication response30, we speculate that ADGRL3 variants may underlie a differential genetic susceptibility not only to SUD, but also to the long-term protective effects of medication treatment. Confirmation of such hypothesis would have substantial public health implications. Inasmuch as ADGRL3 participates in synaptic formation and function, its involvement in SUD could be mediated by either influencing brain development or moderating drug-induced changes in synaptic strength. Molecular studies are required to elucidate the pathogenic mechanism(s) associated with ADGRL3 dysfunction in SUD.
References
Whiteford, H. A., Ferrari, A. J., Degenhardt, L., Feigin, V. & Vos, T. The global burden of mental, neurological and substance use disorders: an analysis from the Global Burden of Disease Study 2010. PLoS ONE 10, e0116820 (2015).
Heslin K. C., Elixhauser A., Steiner C. A. Hospitalizations Involving Mental and Substance Use Disorders Among Adults, 2012: Statistical Brief #191 Healthcare Cost and Utilization Project (HCUP) Statistical Briefs. (Agency for Healthcare Research and Quality, Rockville, MD, 2010).
Prom-Wormley, E. C., Ebejer, J., Dick, D. M. & Bowers, M. S. The genetic epidemiology of substance use disorder: a review. Drug Alcohol. Depend. 180, 241–259 (2017).
Centers for Disease C, Prevention. Increasing prevalence of parent-reported attention-deficit/hyperactivity disorder among children—United States, 2003 and 2007. Mmwr. Morb. Mortal. Wkly. Rep. 59, 1439–1443 (2010).
Visser, S. N. et al. Trends in the parent-report of health care provider-diagnosed and medicated attention-deficit/hyperactivity disorder: United States, 2003–2011. J. Am. Acad. Child Adolesc. Psychiatry 53, 34–46 e2 (2014).
Palacio, J. D. et al. Attention-deficit/hyperactivity disorder and comorbidities in 18 Paisa Colombian multigenerational families. J. Am. Acad. Child Adolesc. Psychiatry 43, 1506–1515 (2004).
Sibley, M. H. et al. The delinquency outcomes of boys with ADHD with and without comorbidity. J. Abnorm. Child Psychol. 39, 21–32 (2011).
Jain, M. et al. Attention-deficit/hyperactivity disorder and comorbid disruptive behavior disorders: evidence of pleiotropy and new susceptibility loci. Biol. Psychiatry 61, 1329–1339 (2007).
Kuperman, S. et al. Developmental sequence from disruptive behavior diagnosis to adolescent alcohol dependence. Am. J. Psychiatry 158, 2022–2026 (2001).
Molina, B. S. et al. Adolescent substance use in the multimodal treatment study of attention-deficit/hyperactivity disorder (ADHD) (MTA) as a function of childhood ADHD, random assignment to childhood treatments, and subsequent medication. J. Am. Acad. Child Adolesc. Psychiatry 52, 250–263 (2013).
Molina, B. S. G. & Pelham, W. E. Childhood predictors of adolescent substance use in a longitudinal study of children with ADHD. J. Abnorm. Psychol. 112, 497–507 (2003).
Biederman, J. et al. Psychoactive substance use disorders in adults with attention-deficit hyperactivity Disorder (ADHD)—effects of ADHD and psychiatric comorbidity. Am. J. Psychiatry 152, 1652–1658 (1995).
Nogueira, M. et al. Early-age clinical and developmental features associated to substance use disorders in attention-deficit/hyperactivity disorder in adults. Compr. Psychiatry 55, 639–649 (2014).
DeMilio, L. Psychiatric syndromes in adolescent substance abusers. Am. J. Psychiatry 146, 1212–1214 (1989).
Horner, B. R. & Scheibe, K. E. Prevalence and implications of attention-deficit hyperactivity disorder among adolescents in treatment for substance abuse. J. Am. Acad. Child Adolesc. Psychiatry 36, 30–36 (1997).
Ercan, E. S., Coskunol, H., Varan, A. & Toksoz, K. Childhood attention deficit/hyperactivity disorder and alcohol dependence: a 1-year follow-up. Alcohol. Alcohol. 38, 352–356 (2003).
White, A. M. et al. Predictors of relapse during treatment and treatment completion among marijuana-dependent adolescents in an intensive outpatient substance abuse program. Subst. Abus. 25, 53–59 (2004).
Arcos-Burgos, M., Velez, J. I., Solomon, B. D. & Muenke, M. A common genetic network underlies substance use disorders and disruptive or externalizing disorders. Hum. Genet. 131, 917–929 (2012).
Gorwood, P. et al. Genetics of dopamine receptors and drug addiction. Hum. Genet. 131, 803–822 (2012).
Hart, A. B. et al. Genome-wide association study of d-amphetamine response in healthy volunteers identifies putative associations, including cadherin 13 (CDH13). PLoS ONE 7, e42646 (2012).
Li, M. D. & Burmeister, M. New insights into the genetics of addiction. Nat. Rev. Genet. 10, 225–231 (2009).
Acosta, M. T. et al. Latent class subtyping of attention-deficit/hyperactivity disorder and comorbid conditions. J. Am. Acad. Child Adolesc. Psychiatry 47, 797–807 (2008).
Arcos-Burgos, M. et al. Attention-deficit/hyperactivity disorder (ADHD): feasibility of linkage analysis in a genetic isolate using extended and multigenerational pedigrees. Clin. Genet. 61, 335–343 (2002).
Acosta, M. T. et al. A two-locus genetic interaction between LPHN3 and 11q predicts ADHD severity and long-term outcome. Transl. Psychiatry 1, e17 (2011).
Arcos-Burgos, M. et al. Attention-deficit/hyperactivity disorder in a population isolate: linkage to loci at 4q13.2, 5q33.3, 11q22, and 17p11. Am. J. Hum. Genet. 75, 998–1014 (2004).
Acosta, M. T. et al. ADGRL3 (LPHN3) variants are associated with a refined phenotype of ADHD in the MTA study. Mol. Genet. Genom. Med. 4, 540–547 (2016).
Bruxel, E. M. et al. LPHN3 and attention-deficit/hyperactivity disorder: a susceptibility and pharmacogenetic study. Genes. Brain. Behav. 14, 419–427 (2015).
Hwang, I. W., Lim, M. H., Kwon, H. J. & Jin, H. J. Association of LPHN3rs6551665 A/G polymorphism with attention deficit and hyperactivity disorder in Korean children. Gene 566, 68–73 (2015).
Jain, M. et al. A cooperative interaction between LPHN3 and 11q doubles the risk for ADHD. Mol. Psychiatry 17, 741–747 (2012).
Arcos-Burgos, M. et al. A common variant of the latrophilin 3 gene, LPHN3, confers susceptibility to ADHD and predicts effectiveness of stimulant medication. Mol. Psychiatry 15, 1053–1066 (2010).
Labbe, A. et al. Refining psychiatric phenotypes for response to treatment: contribution of LPHN3 in ADHD. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 159B, 776–785 (2012).
Ribases, M. et al. Contribution of LPHN3 to the genetic susceptibility to ADHD in adulthood: a replication study. Genes. Brain. Behav. 10, 149–157 (2011).
Song, J. et al. Association of SNAP-25, SLC6A2, and LPHN3 with OROS methylphenidate treatment response in attention-deficit/hyperactivity disorder. Clin. Neuropharmacol. 37, 136–141 (2014).
Gomez-Sanchez, C. I. et al. Attention deficit hyperactivity disorder: genetic association study in a cohort of Spanish children. Behav. Brain. Funct. 12, 2 (2016).
Martinez, A. F. et al. An ultraconserved brain-specific enhancer within ADGRL3 (LPHN3) underpins attention-deficit/hyperactivity disorder susceptibility. Biol. Psychiatry 80, 943–954 (2016).
Ramos-Quiroga, J. A. et al. Criterion and concurrent validity of Conners Adult ADHD Diagnostic Interview for DSM-IV (CAADID) Spanish version. Rev. Psiquiatr Salud Ment. 5, 229–235 (2012).
Group, T. M. C. A 14-month randomized clinical trial of treatment strategies for attention-deficit/hyperactivity disorder. The MTA Cooperative Group. Multimodal treatment study of children with ADHD. Arch. Gen. Psychiatry 56, 1073–1086 (1999).
Jensen, P. S. et al. 3-year follow-up of the NIMH MTA study. J. Am. Acad. Child Adolesc. Psychiatry 46, 989–1002 (2007).
Molina, B. S. et al. The MTA at 8 years: prospective follow-up of children treated for combined-type ADHD in a multisite study. J. Am. Acad. Child Adolesc. Psychiatry 48, 484–500 (2009).
Hechtman, L. et al. Functional adult outcomes 16 years after childhood diagnosis of attention-deficit/hyperactivity disorder: MTA results. J. Am. Acad. Child Adolesc. Psychiatry 55, 945–52 e2 (2016).
Swanson, J. M. et al. Young adult outcomes in the follow-up of the multimodal treatment study of attention-deficit/hyperactivity disorder: symptom persistence, source discrepancy, and height suppression. J. Child Psychol. Psychiatry 58, 663–678 (2017).
Roy, A. et al. Childhood predictors of adult functional outcomes in the multimodal treatment study of attention-deficit/hyperactivity disorder (MTA). J. Am. Acad. Child Adolesc. Psychiatry 56, 687–95 e7 (2017).
Molina, B. S. et al. Delinquent behavior and emerging substance use in the MTA at 36 months: prevalence, course, and treatment effects. J. Am. Acad. Child Adolesc. Psychiatry 46, 1028–1040 (2007).
Johnson, C. et al. Pooled association genome scanning for alcohol dependence using 104,268 SNPs: validation and use to identify alcoholism vulnerability loci in unrelated individuals from the collaborative study on the genetics of alcoholism. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 141B, 844–853 (2006).
Uhl GR. Molecular genetics of substance abuse vulnerability: remarkable recent convergence of genome scan results. Ann. N. Y. Acad. Sci. 1025, 1–13 (2004).
Shaffer, D., Fisher, P., Lucas, C. P., Dulcan, M. K. & Schwab-Stone, M. E. NIMH Diagnostic Interview Schedule for Children Version IV (NIMH DISC-IV): description, differences from previous versions, and reliability of some common diagnoses. J. Am. Acad. Child Adolesc. Psychiatry 39, 28–38 (2000).
Robins L. N., Cottler L., Bucholz K., Compton W. The Diagnostic Interview Schedule for DSM-IV (DIS-IV). (Washington University School of Medicine, St. Louis, MO, 2000).
de Leon, J., Susce, M. T., Diaz, F. J., Rendon, D. M. & Velasquez, D. M. Variables associated with alcohol, drug, and daily smoking cessation in patients with severe mental illnesses. J. Clin. Psychiatry 66, 1447–1455 (2005).
Drake, R. E., Mchugo, G. J. & Biesanz, J. C. The test-retest reliability of standardized instruments among homeless persons with substance use disorders. J. Stud. Alcohol. 56, 161–167 (1995).
Drake, R. E., Rosenberg, S. D. & Mueser, K. T. Assessing substance use disorder in persons with severe mental illness. New Dir. Ment. Health Serv. 1996, 3–17 (1996).
de Leon, J., Armstrong, S. C. & Cozza, K. L. Clinical guidelines for psychiatrists for the use of pharmacogenetic testing for CYP450 2D6 and CYP450 2C19. Psychosomatics 47, 75–85 (2006).
Arcos-Burgos, M. & Muenke, M. Toward a better understanding of ADHD: LPHN3 gene variants and the susceptibility to develop ADHD. Atten. Defic. Hyperact Disord. 2, 139–147 (2010).
Wong, M. L., Dong, C., Andreev, V., Arcos-Burgos, M. & Licinio, J. Prediction of susceptibility to major depression by a model of interactions of multiple functional genetic variants and environmental factors. Mol. Psychiatry 17, 624–633 (2012).
Rao, D. C. CAT scans, PET scans, and genomic scans. Genet. Epidemiol. 15, 1–18 (1998).
Ryzin, J. V., Breiman, L., Friedman, J. H., Olshen, R. A. & Stone, C. J. Classification and regression trees. J. Am. Stat. Assoc. 81, 253 (1986).
Breiman L. Random Forests. In Machine Learning, Vol. 45.(ed. Schapire, R. E.) (Kluwer Academic Publishers, The Netherlands, 2001).
Friedman J. H. Greedy Function Approximation: a Gradient Boosting Machine. (University of Stanford, Stanford, CA, 1999).
Marjoram, P., Zubair, A. & Nuzhdin, S. V. Post-GWAS: where next? More samples, more SNPs or more biology? Hered. (Edinb.) 112, 79–88 (2014).
Visscher, P. M., Brown, M. A., McCarthy, M. I. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Melroy-Greif, W. E. et al. Examination of the involvement of cholinergic-associated genes in nicotine behaviors in European and African Americans. Nicotine. Tob. Res. 19, 417–425 (2017).
Polimanti, R., Yang, C., Zhao, H. & Gelernter, J. Dissecting ancestry genomic background in substance dependence genome-wide association studies. Pharmacogenomics 16, 1487–1498 (2015).
Bi, J., Gelernter, J., Sun, J. & Kranzler, H. R. Comparing the utility of homogeneous subtypes of cocaine use and related behaviors with DSM-IV cocaine dependence as traits for genetic association analysis. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 165B, 148–156 (2014).
Palmer, R. H. et al. Examining the role of common genetic variants on alcohol, tobacco, cannabis and illicit drug dependence: genetics of vulnerability to drug dependence. Addiction 110, 530–537 (2015).
Neale, B. M. & Sham, P. C. The future of association studies: gene-based analysis and replication. Am. J. Hum. Genet. 75, 353–362 (2004).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Stergiakouli, E. et al. Investigating the contribution of common genetic variants to the risk and pathogenesis of ADHD. Am. J. Psychiatry 169, 186–194 (2012).
Poelmans, G., Pauls, D. L., Buitelaar, J. K. & Franke, B. Integrated genome-wide association study findings: identification of a neurodevelopmental network for attention deficit hyperactivity disorder. Am. J. Psychiatry 168, 365–377 (2011).
Williams, N. M. et al. Genome-wide analysis of copy number variants in attention deficit hyperactivity disorder: the role of rare variants and duplications at 15q13.3. Am. J. Psychiatry 169, 195–204 (2012).
Choudhry, Z. et al. LPHN3 and attention-deficit/hyperactivity disorder: interaction with maternal stress during pregnancy. J. Child Psychol. Psychiatry 53, 892–902 (2012).
Martinez, A. F., Muenke, M. & Arcos-Burgos, M. From the black widow spider to human behavior: Latrophilins, a relatively unknown class of G protein-coupled receptors, are implicated in psychiatric disorders. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 156B, 1–10 (2011).
Koob, G. F. & Volkow, N. D. Neurocircuitry of addiction. Neuropsychopharmacology 35, 217–238 (2010).
Lange, M. et al. The ADHD-susceptibility genelphn3.1 modulates dopaminergic neuron formation and locomotor activity during zebrafish development. Mol. Psychiatry 17, 946–954 (2012).
Wallis, D. et al. Initial characterization of mice null for Lphn3, a gene implicated in ADHD and addiction. Brain Res. 1463, 85–92 (2012).
O'Sullivan, M. L. et al. FLRT proteins are endogenous latrophilin ligands and regulate excitatory synapse development. Neuron 73, 903–910 (2012).
Chassin L., Colder C. R., Hussong A., Sher K. J. Substance Use and Substance Use Disorders. (ed. Cicchetti, D.) Developmental Psychopathology, 2nd edn, Vol. 3. (John Wiley & Sons, Inc., Hoboken, NJ, 2016) pp 833-897.
Brook, J. S., Balka, E. B., Zhang, C. & Brook, D. W. ADHD, conduct disorder, substance use disorder, and nonprescription stimulant use. J. Atten. Disord. 21, 776–782 (2017).
Molina, B. S. & Pelham, W. E. Jr. Attention-deficit/hyperactivity disorder and risk of substance use disorder: developmental considerations, potential pathways, and opportunities for research. Annu. Rev. Clin. Psychol. 10, 607–639 (2014).
Zulauf, C. A., Sprich, S. E., Safren, S. A. & Wilens, T. E. The complicated relationship between attention deficit/hyperactivity disorder and substance use disorders. Curr. Psychiatry Rep. 16, 436 (2014).
Wilens, T. E. & Biederman, J. Alcohol, drugs, and attention-deficit/ hyperactivity disorder: a model for the study of addictions in youth. J. Psychopharmacol. 20, 580–588 (2006).
Estevez-Lamorte, N. et al. Adult attention-deficit/hyperactivity disorder, risky substance use and substance use disorders: a follow-up study among young men. Eur. Arch. Psychiatry Clin. Neurosci. (2018). https://doi.org/10.1007/s00406-018-0958-3.
Chen, Q. et al. Common psychiatric and metabolic comorbidity of adult attention-deficit/hyperactivity disorder: a population-based cross-sectional study. PLoS ONE 13, e0204516 (2018).
Quinn, P. D. et al. ADHD medication and substance-related problems. Am. J. Psychiatry 174, 877–885 (2017).
Chang, Z. et al. Stimulant ADHD medication and risk for substance abuse. J. Child Psychol. Psychiatry 55, 878–885 (2014).
Groenman, A. P. et al. Substance use disorders in adolescents with attention deficit hyperactivity disorder: a 4-year follow-up study. Addiction 108, 1503–1511 (2013).
Flory, K. & Lynam, D. R. The relation between attention deficit hyperactivity disorder and substance abuse: what role does conduct disorder play? Clin. Child Fam. Psychol. Rev. 6, 1–16 (2003).
Lee, S. S., Humphreys, K. L., Flory, K., Liu, R. & Glass, K. Prospective association of childhood attention-deficit/hyperactivity disorder (ADHD) and substance use and abuse/dependence: a meta-analytic review. Clin. Psychol. Rev. 31, 328–341 (2011).
Valero, S. et al. Personality profile of adult ADHD: the alternative five factor model. Psychiatry Res. 198, 130–134 (2012).
Khantzian, E. J. The self-medication hypothesis of substance use disorders: a reconsideration and recent applications. Harv. Rev. Psychiatry 4, 231–244 (1997).
Pedersen, S. L., Harty, S. C., Pelham, W. E., Gnagy, E. M. & Molina, B. S. G. Differential associations between alcohol expectancies and adolescent alcohol use as a function of childhood ADHD. J. Stud. Alcohol. Drugs 75, 145–152 (2014).
Acknowledgements
First and foremost, we thank all the patients who participated in this study. This study was supported by the National Human Genome Research Institute intramural funds (to M.M.) for the ascertainment of patients from the Paisa population. The Kentucky sample was ascertained using funds from the Eli Lilly Research Foundation (to J.d.L.), a National Alliance for Research on Schizophrenia and Depression (NARSAD) Independent Award (to J.d.L.), University of Kentucky internal funding (to J.d.L.), and Roche Molecular Systems, Inc., who provided free genotyping and laboratory supplies (to J.d.L.). J.I.V. is partially supported by research grant FOFICO 32101-511035-PE0031 from Universidad del Norte, Barranquilla, Colombia. The Multimodal Treatment Study of Children with ADHD (MTA) was a National Institute of Mental Health (NIMH) cooperative agreement randomized clinical trial, continued under an NIMH contract as a follow-up study and finally under a National Institute on Drug Abuse (NIDA) contract followed by a data analysis grant (DA039881). The Spanish sample was recruited, assessed and genotyped using funds from the Instituto de Salud Carlos III, Spain (PI12/01139, PI14/01700, PI15/01789, PI16/01505), and co-financed by the European Regional Development Fund (ERDF); Agència de Gestió d’Ajuts Universitaris i de Recerca-AGAUR, Generalitat de Catalunya (2014SGR1357, 2014SGR0932); Departament de Salut, Generalitat de Catalunya; Ministerio de Economía, Industria y Competitividad, Spain (SAF2015-68341-R); the European College of Neuropsychopharmacology (ECNP network: ‘ADHD across the lifespan’) and a NARSAD Young Investigator Grant from the Brain & Behavior Research Foundation. This project has also received funding from the European Union’s Horizon 2020 research and innovation programme (CoCA under grant agreement 667302). C.S.-M. is recipient of a Sara Borrell contract from the Instituto de Salud Carlos III, Ministerio de Economía, Industria y Competitividad, Spain (CD15/00199). M.R. is a recipient of a Miguel de Servet contract from the Instituto de Salud Carlos III, Ministerio de Economía, Industria y Competitividad, Spain (CP09/00119 and CPII15/00023). N.F.-C. was supported by a contract of the Centro de Investigación Biomédica en Red de Enfermedades Raras (CIBERER).
Collaborators from NIMH
Benedetto Vitiello, M.D. (Child & Adolescent Treatment and Preventive Interventions Research Branch), Joanne B. Severe, M.S. (Clinical Trials Operations and Biostatistics Unit, Division of Services and Intervention Research), Peter S. Jensen, M.D. (currently at The REACH Institute), L. Eugene Arnold, M.D., M.Ed. (currently at Ohio State University), and Kimberly Hoagwood, Ph.D. (currently at NYU School of Medicine); previous contributors from NIMH to the early phases: John Richters, Ph.D. (currently at National Institute of Nursing Research) and Donald Vereen, M.D. (currently at NIDA). Principal investigators and co-investigators from the sites are: University of California, Berkeley/San Francisco: Stephen P. Hinshaw, Ph.D. (Berkeley), Glen R. Elliott, Ph.D., M.D. (San Francisco); Duke University: Karen C. Wells, Ph.D., Jeffery N. Epstein, Ph.D. (currently at Cincinnati Children's Hospital Medical Center), and Desiree W. Murray, Ph.D.; previous Duke contributors to early phases: C. Keith Conners, Ph.D. (former PI; deceased) and John March, M.D., M.P.H.; University of California, Irvine: James Swanson, Ph.D. and Timothy Wigal, Ph.D.; previous contributor from UCLA to the early phases: Dennis P. Cantwell, M.D.(deceased); New York University School of Medicine: Howard B. Abikoff, Ph.D.; Montreal Children's Hospital/ McGill University: Lily Hechtman, M.D.; and New York State Psychiatric Institute/Columbia University/Mount Sinai Medical Center: Laurence L. Greenhill, M.D. (Columbia) and Jeffrey H. Newcorn, M.D. (Mount Sinai School of Medicine). University of Pittsburgh: Brooke Molina, Ph.D., Betsy Hoza, Ph.D. (currently at University of Vermont), and William E. Pelham, Ph.D. (PI for early phases, currently at Florida International University). Follow-up phase statistical collaborators: Robert D. Gibbons, Ph.D. (University of Illinois, Chicago); Sue Marcus, Ph.D. (Icahn School of Medicine at Mt. Sinai) and Kwan Hur, Ph.D. (University of Illinois, Chicago). Original study statistical and design consultant: Helena C. Kraemer, Ph.D. (Stanford University). Collaborator from the Office of Special Education Programs/US Department of Education: Thomas Hanley, Ed.D. Collaborator from Office of Juvenile Justice and Delinquency Prevention/Department of Justice: Karen Stern, Ph.D.
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Conflict of interest
J.A.R.-Q has served on the speakers’ bureau and acted as consultant for Eli Lilly and Co., Janssen-Cilag, Novartis, Lundbeck, Shire, Ferrer, and Laboratorios Rubió. He has received travel awards from Eli Lilly and Co., Janssen-Cilag, and Shire for participating in psychiatric meetings. The ADHD Program chaired by J.A.R.-Q. has received unrestricted educational and research support from Eli Lilly and Co., Janssen-Cilag, Shire, Rovi, Ferrer and Laboratorios Rubió in the past two years. The remaining authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arcos-Burgos, M., Vélez, J.I., Martinez, A.F. et al. ADGRL3 (LPHN3) variants predict substance use disorder. Transl Psychiatry 9, 42 (2019). https://doi.org/10.1038/s41398-019-0396-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-019-0396-7
This article is cited by
-
Structure, function and drug discovery of GPCR signaling
Molecular Biomedicine (2023)
-
Novel non-stimulants rescue hyperactive phenotype in an adgrl3.1 mutant zebrafish model of ADHD
Neuropsychopharmacology (2023)
-
adgrl3.1-deficient zebrafish show noradrenaline-mediated externalizing behaviors, and altered expression of externalizing disorder-candidate genes, suggesting functional targets for treatment
Translational Psychiatry (2023)
-
Relation between latrophilin 3 (LPHN3) gene polymorphism (rs2345039) and attention deficit hyperactivity disorder in children
Egyptian Pediatric Association Gazette (2022)
-
ADGRL3 genomic variation implicated in neurogenesis and ADHD links functional effects to the incretin polypeptide GIP
Scientific Reports (2022)