Abstract
Adolescent alcohol drinking is known to contribute to the development and severity of alcohol use disorders (AUDs) later in adulthood. Recent studies have shown that long non-coding RNAs (lncRNAs) are critical for brain development and synaptic plasticity. One such lncRNA is natural occurring brain-derived neurotrophic factor antisense (BDNF-AS) that has been shown to regulate BDNF expression. The role of BDNF-AS lncRNA in the molecular mechanisms of AUD is unknown. Here, we evaluated the expression and functional role of BDNF-AS in postmortem amygdala of either early onset or late onset alcoholics (individuals who began drinking before or after 21 years of age, respectively) and age-matched control subjects. BDNF-AS expression is increased in early onset but not in late onset AUD amygdala and appears to be regulated epitranscriptomically via decreased N6-methyladenosine on BDNF-AS. Upregulation of BDNF-AS is associated with a significant decrease in BDNF expression and increased recruitment of EZH2, which deposits repressive H3K27 trimethylation (H3K27me3) at regulatory regions in the BDNF gene in the early onset AUD group. Drinking during adolescence also contributed to significant decreases in activity-regulated cytoskeleton-associated protein (ARC) expression which also appeared to be mediated by increased EZH2 deposition of repressive H3K27me3 at the ARC synaptic activity response element. These results suggest an important role for BDNF-AS in the regulation of synaptic plasticity via epigenetic reprogramming in the amygdala of AUD subjects who began drinking during adolescence.
Similar content being viewed by others
Introduction
Alcohol use disorders (AUDs) are chronic, debilitating psychiatric illnesses that contribute to considerable socioeconomic and health burden worldwide1. Age of onset is one of the primary risk factors for the development of AUDs, as individuals who began drinking during adolescence are four times more likely to develop AUDs in adulthood2. Adolescence is a critical period in brain development and adolescent drinking decreases orbitofrontal cortex activity and increases amygdala activity3 leading to less executive control, more emotional impulsivity, alterations in decision-making, and have a higher risk to engage in risky behaviors and develop mental health problems later in life4,5. The amygdala acts as a hub for the processing of sensory signaling from cortical regions, and is involved in regulation of emotional reactivity, decision-making, and negative affective state of alcohol addiction6,7. During adolescence, the amygdala undergoes several key changes in specific inter- and intra-region connectivity8, and this process is thought to be disrupted by alcohol use9. Recent studies have begun to examine the molecular basis of these changes in the amygdala in preclinical models of adolescent alcohol exposure and have identified numerous potential molecular components, such as brain-derived neurotrophic factor (BDNF), that are involved in the development of alcohol dependence in adulthood10,11,12.
The neurotrophin BDNF has emerged as a critical molecular player in central nervous system (CNS) development, psychiatric disorders, and AUDs13,14,15. BDNF is associated with survival and differentiation of neurons, activity-dependent neuroplasticity, long-term potentiation, learning, memory, and anxiety13,16. BDNF is critical for normal brain development, as constitutive knockout mice die during second week after birth13 and is known to be involved in synaptogenesis and maintenance of dendrites during development and adulthood13,17. BDNF regulates synaptic plasticity through interactions with the tropomyosin-related kinase B (TrkB) receptor and induction of several signaling cascades that are involved in the regulation of synaptic plasticity and cAMP-responsive element-binding protein (CREB)-dependent gene expression13. Decreased expression of BDNF in the amygdala of rodent models of adolescent alcohol exposure has recently been implicated in anxiety-like and alcohol-drinking behaviors12. Furthermore, mutant versions of BDNF with decreased TrkB activity are associated with increase in alcohol consumption.14,15,18. A particularly important downstream target of BDNF signaling is activity-regulated cytoskeleton-associated protein (ARC) which is a member of the immediate-early gene family and is important in synaptogenesis and synaptic plasticity as well as anxiety and alcohol intake phenotypes19–21. BDNF activates ARC expression through CREB and other transcription factor binding to the synaptic activity response element (SARE) site located in the ARC promoter approximately 7 kb upstream from the transcription start site22,23.
In addition to complex signaling, the BDNF gene also has a complex regulatory and genetic structure, with 9 different promoters encoding at least 20 different splice variants sharing one common exon that encodes the BDNF protein (known as exon IX)13,24 and expression of these variants are temporal- and region-specific25. Studies in animal models have revealed that different BDNF transcript variants are downregulated in the amygdala by adolescent alcohol exposure and associated with alcohol intake in adulthood12.
Long non-coding RNAs (lncRNAs) are broadly defined as RNAs that are greater than 200 bp and lack a functional open-reading frame. The majority of lncRNAs are found exclusively in the brain and are important in normal brain development, with temporal and regional specificity26–28. Most lncRNAs with described function are involved in regulating gene expression and chromatin structure through a number of different mechanisms including cis-tether, trans-regulation, allosteric modulators of epigenetic complexes (e.g. polycomb repressive complex II, PRC2) or transcription factors, or acting as a decoy either for transcription factors or repressors29–31. LncRNAs have recently been identified as being important in AUDs in adults32. Despite several advances in the understanding of lncRNA function, the function of the vast majority of lncRNAs are still unknown. A recent study characterized the role of the lncRNA BDNF-antisense (BDNF-AS) to be a negative regulator of BDNF exon IX expression and neurogenesis, and also described the overlap region between BDNF and BDNF-AS33. However, changes in BDNF and BDNF-AS expression and associated epigenetic regulatory mechanisms in the amygdala of human AUD subjects are currently unknown. We aimed to determine if there are epigenetic changes mediated by BDNF-AS that are operative in the regulation of BDNF expression in adolescence, and whether these changes persist into adulthood in the amygdala of human postmortem brains diagnosed with AUDs that began drinking before the age of 21. Here we report that the epitranscriptome and epigenetic mechanisms interact to regulate BDNF expression in the amygdala and are possibly involved in the pathophysiology of alcoholism that begins with drinking during adolescence.
Materials and methods
Samples
Human postmortem amygdala tissue was acquired from the New South Wales Brain Tissue Resource Center (Sydney, Australia). Criteria for inclusion of AUD samples were: age >18 years, no history of major psychiatric disorders, and no history of other substance abuse disorders. Demographic characteristics are provided in Supplementary Table 1. We have previously described this cohort34. Detailed methods are provided in Supplementary Information.
Quantitative real-time PCR for mRNA levels
Quantitative real-time PCR (qPCR) was run using standard methods with either Taqman probes or specific primers (Supplementary Table 2). Changes in expression were determined using the ∆∆Ct method and normalized to mean Ct values of ACTB and GAPDH. Data are presented as average fold change of controls.
Enzyme-linked immunosorbent assay for protein levels
BDNF protein was ascertained via the Quantikine ELISA total BDNF Assay (R&D Systems) following the manufacturer’s instructions. The optical density of each sample and standard was measured using the Spectra MR microplate reader (Dynex Technologies) and the amount of BDNF was calculated against the BDNF standard curve and expressed as pg/µg of total protein (Supplementary Methods).
Chromatin immunoprecipitation assay
Chromatin immunoprecipitation (ChIP) was run following previously published protocols35,36. Purified DNA was analyzed by qPCR using primers (Supplementary Table 2). The data were analyzed using the ∆∆Ct method, normalizing to input, and the data are expressed as fold change relative to controls.
RNA methylation immunoprecipitation (M6A RIP) assay
M6A RIP was performed using EpiMark® N6-methyladenosine Enrichment Kit (NEB) following the manufacturer’s instructions with some modifications. Purified RNA was sheared to <350 bp using a Covaris ME220 and then purified using Zymo RNA Clean and Concentrator 5 (Zymo) following the manufacturer’s instructions. Purified RNA was then measured for concentration using the Qubit BR RNA Assay (ThermoFisher Scientific) following the manufacturer’s instructions, and 1 µg was used for each pulldown with 2 µL of M6A antibody. Following 1 h incubation and washes, RNA was eluted and purified with MyOne Silane Beads (ThermoFisher Scientific) then prepared for reverse transcriptase in which 1 µL from pulldown or input was put in reverse transcriptase reaction. Resulting cDNA was then used for qPCR using primers designed to predict M6A sites located on BDNF-AS (Supplementary Table 2). M6A sites were predicted using SRAMP37. The data were analyzed using the ∆∆Ct method, normalizing to input, and the data are expressed as fold change relative to controls.
Statistics
Statistical analysis was performed using SigmaStat (Systat Software, San Jose, CA, USA) and data were visualized using GraphPad Prism (La Jolla, CA, USA). Comparisons between groups were performed with Student’s t-tests. If comparisons failed normality tests then Mann–Whitney rank sum tests were performed. Pearson’s correlations were performed to determine linear correlation between two groups. Significance was set at p < 0.05.
Results
Expression of BDNF and BDNF-AS in the postmortem amygdala
Previous studies in rodents have demonstrated a decrease in BDNF expression after adolescent alcohol exposure in the amygdala12, but similar changes in the human amygdala have not yet been examined. In order to identify potential changes that are due to adolescent drinking we evaluated both a cohort that began drinking before the age of 21 (early onset) and a cohort who began drinking after the age of 21 (late onset). We then analyzed these two cohorts to see if there was differences in drinking or other demographics. We found that our early onset group drank 234 ± 56 g (mean ± SEM) of alcohol per day at time of death versus the late onset group which drank 157 ± 28 g of alcohol per day at time of death, although this observation was not statistically significant between early onset and late onset groups (Supplementary Table 1). We also found that there was no statistically significant differences in other metrics evaluated, including age, postmortem interval (PMI), total drinking years, and cigarette pack years (Supplementary Table 1).
We then evaluated BDNF mRNA expression at the common coding exon (BDNF exon IX) and found a significant decrease in BDNF mRNA expression in early onset AUD compared to controls (p = 0.008) (Fig. 1a). Regulation of BDNF expression is complex25,26; however, a previous study has demonstrated that increased BDNF-AS leads to decreased BDNF mRNA expression33. Therefore, we examined changes in the expression of lncRNA BDNF-AS in the postmortem amygdala of alcoholics who began drinking before the age of 21 to determine if this was a regulatory pathway potentially regulating decreased BDNF mRNA expression. BDNF-AS expression was increased in the early onset group (p = 0.033) (Fig. 1b). We also evaluated changes in both BDNF mRNA and BDNF-AS expression in the postmortem amygdala in the late onset group to determine if these changes were specific to early age onset group. There was no significant change in either BDNF mRNA (Fig. 1c) or BDNF-AS (Fig. 1d) expression in the late onset group.
a There is decreased BDNF exon-IX mRNA expression in the human amygdala of individuals with AUDs who began drinking before age of 21 (early onset). Significance was determined using Mann–Whitney test. b The lncRNA BDNF-AS is upregulated in the human amygdala of AUD patients who began drinking before the age of 21. c There is no change in BDNF exon IX mRNA expression in the human amygdala of individuals with AUDs who began drinking after age of 21 (late onset). d The lncRNA BDNF-AS is not changed in the human amygdala of AUD subjects who began drinking after the age of 21. e There is decreased BDNF protein levels in individuals with AUDs who began drinking in adolescence. Values are presented as mean ± SEM. f There is no change in BDNF protein levels in individuals with AUD who began drinking after adolescence. Values are presented as mean ± SEM. Significance was determined using Student’s t-test unless otherwise noted. *p < 0.05 and **p < 0.01. n = 22 (controls), 11 (early onset AUDs) and 11 (late onset AUDs)
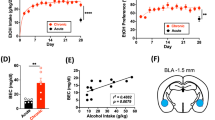
We next analyzed correlations between BDNF and BDNF-AS expression versus changes in several drinking variables (total drinking years, alcohol daily use at time of death (in grams), and standard drinks per week) to determine if there was any relationship. We first analyzed the correlation between BDNF and BDNF-AS expression and found that BDNF and BDNF-AS mRNA levels are negatively correlated (r = −0.0443, p = 0.003). We then evaluated whether changes in BDNF-AS expression was a function of total drinking years, alcohol daily use at the time of death, and standard drinks per week. We found that BDNF-AS expression in total samples (n = 42) was positively correlated with alcohol daily use at time of death (r = 0.523, p = 0.0004) and standard drinks per week (r = 0.466 p = 0.002) but not total drinking years (Supplementary Fig. 1). Examination of AUD subjects who began drinking before the age of 21 revealed that BDNF-AS trended towards being significantly correlated with alcohol use per day at time of death (r = 0.567, p = 0.069), but this was not the case for controls (r = 0.212, p = 0.37) or in the late onset group (r = 0.258, p = 0.45) (Supplementary Fig. 1). Similarly, BDNF mRNA expression in total samples (n = 42) negatively correlated with alcohol per day use at time of death (r = −0.416, p = 0.0062) and standard drinks per week (r = −0.374 and p = 0.015) but not total drinking years (r = −0.253, p = 0.106) (Supplementary Fig. 1). When we determined correlations of BDNF mRNA expression separately versus standard drinks or alcohol daily use at time of death in only early onset drinker and late onset drinker groups we found no significant correlations. However, we did observe a trend towards significant correlation for BDNF and both alcohol use per day at time of death (r = −0.431, p = 0.058) and standard drinks per week (r = −0.43, p = 0.058) (Supplementary Fig. 1) for controls. We also found that there was no statistically significant correlations in other metrics evaluated: age, PMI, and cigarette pack years.
BDNF protein levels in the postmortem amygdala
Concentration of total (free and TrkB bound) BDNF protein in postmortem human amygdala of AUDs and control subjects was determined via enzyme-linked immunosorbent assay. Average (mean ± SEM) BDNF protein (pg/μg of total protein) in controls (0.228 ± 0.011; n = 22) was not significantly different than total alcoholics (0.192 ± 0.019; n = 22). However, further analysis by separation of alcoholics by age of onset of drinking revealed a significant (p = 0.043) decrease in BDNF protein levels in subjects (n = 11) in the early onset group (0.177 ± 0.029) versus controls (0.228 ± 0.011; n = 22) (Fig. 1e). Interestingly, no significant (p = 0.31) differences were observed in the late onset group (0.207 ± 0.025; n = 11) versus controls (0.228 ± 0.011; n = 22) (Fig. 1f). BDNF protein levels in total samples (n = 42) were significantly negatively correlated with both standard drinks per week (r = −0.377, p = 0.014) and alcohol use per day at time of death (r = −0.376, p = 0.0142). When we determined correlation of BDNF protein expression separately in controls, early, and late of onset drinkers with standard drinks per week and alcohol use per day at time of death, we found no significant correlations similar to what was found with BDNF mRNA correlations (Supplementary Fig. 2).
Epigenetic modifications at the BDNF gene in the postmortem amygdala
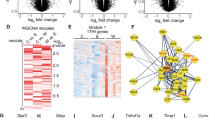
Previous reports in rodent models of adolescent alcohol exposure have demonstrated that changes in BDNF expression is due to changes in epigenetic modifications. Particularly, deficits in histone acetylation (H3K9/14ac) of BDNF exon IV promoter was associated with deficits in BDNF expression12. Further, BDNF-AS is known to contribute to epigenetic changes facilitated by the PRC2 complex at the BDNF exon IX promoter in HEK293 cells33. To evaluate if changes observed in BDNF expression were due to epigenetic changes, we utilized ChIP assay to determine if there were differences in enhancer of zeste homolog 2 (EZH2) occupancy and its catalytic product, repressive H3K27 trimethylation (H3K27me3), at several sites in the BDNF promoter region (Fig. 2a) and at a region where BDNF exon IX and BDNF-AS overlap33. We evaluated EZH2 occupancy at these loci. We observed increases in EZH2 at both the BDNF-IX promoter (p = 0.023) and overlap region (p = 0.027) in the early onset AUD group. The overlap region (Fig. 2a) is the region of overlap between transcribed BDNF and BDNF-AS transcripts (located in the common exon IX of BDNF and exon 5 of BDNF-AS)33. At the BDNF-IV promoter, though there appears to be a trend (p = 0.084) towards an increase in the early onset group (Fig. 2b). We observed no changes in EZH2 occupancy at any of the loci in the late onset group (Fig. 2c). There were significant increases in H3K27me3 associated with the BDNF-IV (p = 0.014) and BDNF-IX (p = 0.049) promoters and also at the overlap region (p = 0.003) (Fig. 2d) in the early onset group. In the late onset group, there was only an increase (p = 0.027) in H3K27me3 at the BDNF-IV promoter (Fig. 2e). We examined for correlations between repressive H3K27me3 at the BDNF-IX, promoter, and BDNF overlap region in total sample (n = 42) with drinking variables. We found no significant correlations with either alcohol use per day at the time of death nor standard drinks per week for either BDNF-IX and BDNF overlap region.
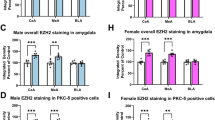
a Schematic demonstrating different BDNF sites evaluated for changes in occupancy of EZH2 and histone H3K27 trimethylation (H3K27me3). b EZH2 occupancy is elevated at the BDNF-IX promoter and the BDNF and BDNF-AS overlap site in the amygdala of individuals who began drinking before the age of 21. Significance was determined by Mann–Whitney test. c EZH2 is not elevated at any of the three BDNF sites measured in individuals who began drinking after the age of 21. Significance was determined by Mann–Whitney test for BDNF-IV, BDNF-IX, and BDNF and BDNF-AS overlap region in later age of onset. d Early onset in the postmortem amygdala of AUD subjects show increases in repressive H3K27me3 associated with several BDNF gene locations while e late onset show only increases in repressive H3K27me3 at the BDNF-IV promoter. Significance was determined by Mann–Whitney test for BDNF-IV for early age of onset and BDNF-IX for both early and late age of onset. Values are presented as mean ± SEM. Significance was determined using Student’s t-test unless otherwise noted. *p < 0.05 and **p < 0.01. AUD alcohol use disorder. n = 22 (controls), 11 (early onset AUDs) and 11 (late onset AUDs)
We next analyzed changes in EZH2 and suppressor of zeste 12 (SUZ12, another member of the PRC2 complex) mRNA levels using qPCR to determine if changes in occupancy of EZH2 could be due to changes in overall levels. We found no change in either EZH2 or SUZ12 mRNA levels in early onset or late onset postmortem human amygdala (Supplementary Fig. 3). SUZ12 is a core component of the PRC2 complex and known binding partner of lncRNAs30. We therefore performed RIP assay to test for an interaction between BDNF-AS and SUZ12 protein and found that SUZ12 binds to BDNF-AS as determined by the RIP assay (Supplementary Fig. 4).
Epigenetic modifications at the ARC SARE site and ARC mRNA levels in the postmortem amygdala
The early immediate gene ARC is a downstream target of BDNF, is involved in synaptic plasticity, and is a critical target of adolescent and adult alcohol exposure in rodent models12,21. Analysis of changes in ARC expression in early onset postmortem amygdala shows decreased ARC (p = 0.016) (Fig. 3a), but not in late onset postmortem amygdala (Fig. 3b). BDNF signaling is known to regulate ARC expression through the ARC SARE site located ~7.7 kb upstream of the transcription start site22,23 and is regulated by enzymes involved in epigenetic modulation22,23, and therefore we examined if there were epigenetic changes at this location (Fig. 3c). H3K27me3 occupancy was increased at the ARC SARE site (p = 0.018) in the early onset AUD group (Fig. 3d). We next evaluated if this change was mediated by increased PRC2 signaling and found increased EZH2 associated with the SARE site in this group as well (p = 0.049) (Fig. 3e). These changes are not present in the late onset AUD group (Supplementary Fig. 5). We also found decreased H3K27ac (a histone post-translational modification associated with increased gene expression) associated with the SARE site in the early onset AUD group (p = 0.009) (Fig. 3f).
a ARC is decreased in the postmortem amygdala of individuals who began drinking during adolescence (early onset). Significance was determined using Mann–Whitney test. b ARC is not decreased in the postmortem amygdala individuals who began drinking in adulthood (late onset). c Representative schematic of the ARC SARE site which responds to BDNF activity and is located up ~7.7 kb upstream from the promoter. d Adolescent drinking increases EZH2 occupancy at the ARC SARE site in human postmortem amygdala. Values are presented as mean ± SEM. e Adolescent drinking increases repressive H3K27me3 associated with the ARC SARE site in human postmortem amygdala. f Adolescent drinking decreases H3K27ac associated with the ARC SARE site in human postmortem amygdala. Significance was determined using Student’s t-test unless otherwise noted. *p < 0.05 and **p < 0.01. n = 22 (controls), 10–11 (early onset AUDs) and 11 (late onset AUDs)
RNA methylation of BDNF-AS in the postmortem amygdala
We next wanted to determine the mechanism for BDNF-AS upregulation in early onset AUD subjects. Our initial experiments focused on H3K4me3 at the BDNF-AS promoter as this mark is known to be involved in active transcription and in the regulation of lncRNAs38,39. However we found no significant increase in H3K4me3 in the early onset group (Supplementary Fig. 6). Recent reports have suggested that post-translational modifications can modulate lncRNA expression40 so we utilized N6-methyladenosine (M6A) RIP assay to determine if there were changes in RNA methylation at predicted M6A sites in BDNF-AS. Our results indicate a decrease (p < 0.001) in BDNF-AS RNA methylation in the early onset AUD group (Fig. 4a), but not in the late onset AUD group (Fig. 4b). We did not observe any change in mRNA expression of several known RNA methyltransferases or the RNA demethylase ALKBH5 (Supplementary Fig. 7a–h).
a Drinking during adolescence decreases RNA methylation at the predicted M6A site in BDNF-AS. b Drinking in adulthood does not change methylation at the M6A site in BDNF-AS. Values are presented as mean ± SEM. Significance was determined using Mann–Whitney rank sum test. ***p < 0.001. n = 21 (controls), 11 (early onset AUDs) and 11 (late onset AUDs)
Discussion
This study demonstrates that early age of onset of alcohol consumption increases BDNF-AS lncRNA expression induced via diminished RNA methylation, which triggers aberrant epigenetic mechanisms to produce deficits in BDNF signaling in the amygdala and possibly associated with pathophysiology of AUD (Fig. 5).
Proposed association of regulation of decreased BDNF expression by BDNF-AS through recruitment of the PRC2 complex in the human amygdala of individuals who began drinking in adolescence. Adolescent drinking appears to increase the expression of BDNF-AS most likely via inhibition of RNA methylation. This is associated with decreases in BDNF expression in the amygdala via recruitment of EZH2 and the associated increase in H3K27 trimethylation (H3K27me3) at the promoter and overlap region of BDNF exon IX. Interestingly, the reduction in BDNF expression is associated with a reduction in activity-regulated cytoskeleton-associated protein (ARC) expression, possibly via an increase in occupancy of EZH2 and H3K27me3 at an important regulatory site of the ARC gene known as synaptic activity response element (SARE). These BDNF-AS-regulated epigenetic mechanisms induced by adolescent drinking in humans appears to be important in the pathophysiology of alcohol use disorders (AUD) later in life
Effects of adolescent drinking and changes in BDNF signaling in adulthood
BDNF signaling is an important modulator of CNS development through mediation of important processes such as neuronal survival, neurogenesis, neurite outgrowth, and neuronal differentiation13,16. Importantly, expression of both BDNF protein and specific BDNF mRNA isoforms are developmentally controlled, with different areas of the amygdala showing different expression patterns during adolescence and adulthood. BDNF expression in the central nucleus of amygdala (CeA) slowly rises to a plateau in adulthood, while the basolateral amygdala has decreased expression in adulthood relative to adolescence41 and disruption to normal BDNF expression and signaling in adolescence is likely to produce detrimental outcomes on normal brain development. Changes in BDNF signaling are likely to be involved in structural changes in the amygdala, which involve synaptic pruning and altered connectivity between the amygdala and other brain regions throughout the adolescent period8. Based on the previous finding that adolescent alcohol exposure in rodents decreases BDNF expression in the CeA12, our present findings confirm the hypothesis that there would be changes in BDNF expression in a cohort of postmortem amygdala from AUD subjects that began drinking during adolescence, demonstrating a conserved pathway that may induce increased drinking and other psychiatric problems in adulthood caused by adolescent alcohol exposure. Previous work has found that individuals with alcohol dependence have decreased circulating BDNF in blood42. Interestingly, transgenic mice harboring a homolog (Val68Met) of a common human polymorphism of BDNF (Val66Met) compulsively consume more alcohol than wild-type mice18. The Val68Met variant leads to decreased intracellular trafficking and activity-dependent BDNF release and function18. Decreasing BDNF or ARC expression in the CeA of rodent models of alcoholism in adulthood increases drinking and anxiety-like behaviors, which are measures of alcohol dependence21,43. These previous findings taken together with the present findings shown here suggest that disruptions to BDNF expression and ARC signaling are likely to increase the risk for alcohol dependence, and the risk is greater if these changes occur at a critical period of brain development44, such as adolescence, due to enduring effects on epigenetic programming. However, the possibility cannot be ruled out that other factors besides drinking during adolescence are involved with changes in BDNF expression that we observed in human postmortem amygdala. These factors could include a longer time period for disruption of normal neural processes by alcohol and a higher lifetime alcohol consumption at the time of death.
Potential role of lncRNAs in epigenetic regulation by adolescent drinking in AUD subjects
LncRNAs have recently emerged as a potential molecular target in AUDs by the use of RNA sequencing32,45. However, evaluation of lncRNAs in a population of individuals with AUDs that began drinking before the age of 21 has not been explored. Previous studies have characterized the lncRNA, BDNF-AS, to be a negative regulator of BDNF expression33. Rodent studies have demonstrated that infusion of BDNF-antisense oligodeoxynucleotides into the CeA increases anxiety and alcohol consumption43. Therefore, we postulated that the naturally occurring BDNF-AS may be involved in changes observed in BDNF expression in our early onset cohort. We found increased BDNF-AS only in individuals who began drinking before the age of 21. Additionally, similar to previous findings demonstrated in vitro33, increased BDNF-AS negatively correlated with BDNF exon IX expression. Intriguingly, we also found that BDNF-AS expression positively correlated with the amount of alcohol consumed using two different metrics of alcohol consumption (grams consumed per day at time of death and standard drinks per week). One possibility is that earlier age of alcohol use may interact with later alcohol consumption to induce BDNF-AS expression. Our results suggest that BDNF-AS may also be important in epigenetic programming during development, and expression of this transcript is dysregulated after drinking in adolescence and associated with AUDs later in life.
Our data indicate that BDNF-AS may be regulated epitranscriptomically by decreased RNA methylation at a M6A site in the amygdala of early onset AUD but not in late onset AUD. Previous reports have found that decreased M6A methylation prevents RNA turnover and decreases targeted degradation by YTH domain family 2, which is recruited to the RNA by M6A methylation46. Other roles for M6A methylation appear to either promote or inhibit recruitment of proteins that bind RNA47, a potential mechanism for the binding of SUZ12 to BDNF-AS and could be explored in future experiments. Nonetheless, the observation that adolescent alcohol exposure decreases BDNF-AS methylation in the amygdala reveals new mechanisms of RNA regulation that add another layer of complexity of epigenetic regulation of BDNF expression in AUD.
Role of PRC2 complex regulating epigenetic changes in early onset AUD
LncRNAs have been demonstrated to interact with SUZ12, a member of the PRC2 complex, via its zinc-finger domain30, and BDNF-AS knockdown disrupts EZH2 recruitment to BDNF promoters in cell culture models33. We tested this hypothesis by examining EZH2 occupancy and its catalytic product, H3K27me3, at regions of the BDNF promoter, including the region of BDNF exon IX that overlaps with BDNF-AS. Our results indicate that there is increased EZH2 recruited to both the BDNF overlap region and the promoter of BDNF exon IX with corresponding increases in H3K27me3 at these locations. Coupled with our data showing that SUZ12 binds directly to BDNF-AS, this suggests that BDNF-AS regulates BDNF-IX expression through direct binding and recruitment of the PRC2 complex and subsequent repression of transcription by deposition of H3K27me3.
In this study, we also found increases of EZH2 and H3K27me3 at the ARC SARE site. This may be due to chromatin remodeling secondary to decreased BDNF signaling and recruitment of class I histone deacetylases (HDACs)22 catalyzing removal of histone acetyl groups, which we have also observed. The recruitment of EZH2 to this site warrants further experimentation to determine if it occurs through decreased BDNF signaling, subsequently decreased occupancy of CREB-binding protein, and increased occupancy of class I HDACs at the SARE site of ARC gene. EZH2 is also heavily implicated in brain development, where it has a role in regulating self-renewal and differentiation in the cerebral cortex48. Since there is significant pruning and differentiation of neurons during adolescence44, EZH2 dysregulation and an increase in repressive H3K27me3 marks at the BDNF and ARC regulatory regions by alcohol drinking during this critical period could contribute to an increased risk of AUDs and higher overall alcohol consumption later in life (Fig. 5).
Conclusions
Our results appear to suggest that BDNF-AS is upregulated most likely via decreased RNA methylation in the amygdala of adult AUD population who began drinking during adolescence, but not in AUD subjects who began drinking in adulthood. However we are unable to determine if these effects are causative without further experimentation, and indeed changes that we observed could be due to other factors and interactions. Future experiments will have to consider these possibilities in larger cohorts of human postmortem brain and through the use of reverse translational approach in animal models. We demonstrate that increased BDNF-AS is associated with decrease in BDNF expression through an epigenetic mechanism involving the recruitment of PRC2 and the repression of gene promoters. Downstream signaling targets of BDNF such as ARC are also decreased through recruitment of EZH2 and the deposition of repressive H3K27me3 marks. Our results provide translational evidence for a mechanism for decreased BDNF in the amygdala previously shown in rodent models of adolescent alcohol exposure: namely, similar decreases in BDNF and ARC that are associated with higher alcohol intake12. These results suggest the possible role for developmentally sensitive lncRNAs in early onset AUD, and that EZH2 inhibitors may prove useful in the treatment of adult psychopathology after adolescent alcohol drinking.
References
Katikireddi, S. V., Whitley, E., Lewsey, J., Gray, L. & Leyland, A. H. Socioeconomic status as an effect modifier of alcohol consumption and harm: analysis of linked cohort data. Lancet Public Health 2, e267–e276 (2017).
Grant, B. F., Stinson, F. S. & Harford, T. C. Age at onset of alcohol use and DSM-IV alcohol abuse and dependence: a 12-year follow-up. J. Subst. Abuse 13, 493–504 (2001).
Xiao, L. et al. Abnormal affective decision making revealed in adolescent binge drinkers using a functional magnetic resonance imaging study. Psychol. Addict. Behav. 27, 443–454 (2013).
Wells, J. E., Horwood, L. J. & Fergusson, D. M. Drinking patterns in mid-adolescence and psychosocial outcomes in late adolescence and early adulthood. Addiction 99, 1529–1541 (2004).
Viner, R. M. & Taylor, B. Adult outcomes of binge drinking in adolescence: findings from a UK national birth cohort. J. Epidemiol. Community Health 61, 902–907 (2007).
Janak, P. H. & Tye, K. M. From circuits to behaviour in the amygdala. Nature 517, 284–292 (2015).
Koob, G. F. & Volkow, N. D. Neurocircuitry of addiction. Neuropsychopharmacology 35, 217–238 (2010).
Qin, S., Young, C. B., Supekar, K., Uddin, L. Q. & Menon, V. Immature integration and segregation of emotion-related brain circuitry in young children. Proc. Natl Acad. Sci. USA 109, 7941–7946 (2012).
O’Daly, O. G. et al. Withdrawal-associated increases and decreases in functional neural connectivity associated with altered emotional regulation in alcoholism. Neuropsychopharmacology 37, 2267–2276 (2012).
Gilpin, N. W. & Roberto, M. Neuropeptide modulation of central amygdala neuroplasticity is a key mediator of alcohol dependence. Neurosci. Biobehav. Rev. 36, 873–888 (2012).
Kyzar, E. J., Zhang, H., Sakharkar, A. J. & Pandey, S. C. Adolescent alcohol exposure alters lysine demethylase 1 (LSD1) expression and histone methylation in the amygdala during adulthood. Addict. Biol. 22, 1191–1204 (2017).
Pandey, S. C., Sakharkar, A. J., Tang, L. & Zhang, H. Potential role of adolescent alcohol exposure-induced amygdaloid histone modifications in anxiety and alcohol intake during adulthood. Neurobiol. Dis. 82, 607–619 (2015).
Autry, A. E. & Monteggia, L. M. Brain-derived neurotrophic factor and neuropsychiatric disorders. Pharmacol. Rev. 64, 238–258 (2012).
Colzato, L. S. et al. BDNF Val66Met polymorphism is associated with higher anticipatory cortisol stress response, anxiety, and alcohol consumption in healthy adults. Psychoneuroendocrinology 36, 1562–1569 (2011).
Pandey, S. C. A critical role of brain-derived neurotrophic factor in alcohol consumption. Biol. Psychiatry 79, 427–429 (2016).
Poo, M. M. Neurotrophins as synaptic modulators. Nat. Rev. Neurosci. 2, 24–32 (2001).
Cohen-Cory, S., Kidane, A. H., Shirkey, N. J. & Marshak, S. Brain-derived neurotrophic factor and the development of structural neuronal connectivity. Dev. Neurobiol. 70, 271–288 (2010).
Warnault, V. et al. The BDNF valine 68 to methionine polymorphism increases compulsive alcohol drinking in mice that Is reversed by tropomyosin receptor kinase B activation. Biol. Psychiatry 79, 463–473 (2016).
Ying, S. W. et al. Brain-derived neurotrophic factor induces long-term potentiation in intact adult hippocampus: requirement for ERK activation coupled to CREB and upregulation of Arc synthesis. J. Neurosci. 22, 1532–1540 (2002).
Moonat, S., Sakharkar, A. J., Zhang, H., Tang, L. & Pandey, S. C. Aberrant histonedeacetylase2–mediated histone modifications and synaptic plasticity in the amygdala predisposes to anxiety and alcoholism. Biol. Psychiatry 73, 763–773 (2013).
Pandey, S. C. et al. Effector immediate-early gene Arc in the amygdala plays a critical role in alcoholism. J. Neurosci. 28, 2589–2600 (2008).
Fukuchi, M. et al. Class I histone deacetylase-mediated repression of the proximal promoter of the activity-regulated cytoskeleton-associated protein gene regulates its response to brain-derived neurotrophic factor. J. Biol. Chem. 290, 6825–6836 (2015).
Kawashima, T. et al. Synaptic activity-responsive element in the Arc/Arg3.1 promoter essential for synapse-to-nucleus signaling in activated neurons. Proc. Natl Acad. Sci. USA 106, 316–321 (2009).
Liu, Q. R. et al. Human brain derived neurotrophic factor (BDNF) genes, splicing patterns, and assessments of associations with substance abuse and Parkinson’s Disease. Am. J. Med. Genet. B Neuropsychiatr. Genet. 134B, 93–103 (2005).
Aid, T., Kazantseva, A., Piirsoo, M., Palm, K. & Timmusk, T. Mouse and rat BDNF gene structure and expression revisited. J. Neurosci. Res. 85, 525–535 (2007).
Rajarajan, P., Gil, S. E., Brennand, K. J. & Akbarian, S. Spatial genome organization and cognition. Nat. Rev. Neurosci. 17, 681–691 (2016).
Mercer, T. R., Dinger, M. E., Sunkin, S. M., Mehler, M. F. & Mattick, J. S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl Acad. Sci. USA 105, 716–721 (2008).
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Guttman, M. & Rinn, J. L. Modular regulatory principles of large non-coding RNAs. Nature 482, 339–346 (2012).
Khalil, A. M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl Acad. Sci. USA 106, 11667–11672 (2009).
Zhao, J. et al. Genome-wide identification of polycomb-associated RNAs by RIP-seq. Mol. Cell 40, 939–953 (2010).
Farris, S. P., Arasappan, D., Hunicke-Smith, S., Harris, R. A. & Mayfield, R. D. Transcriptome organization for chronic alcohol abuse in human brain. Mol. Psychiatry 20, 1438–1447 (2015).
Modarresi, F. et al. Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat. Biotechnol. 30, 453–459 (2012).
Gatta, E. et al. Emerging role of one-carbon metabolism and DNA methylation enrichment on δ-containing GABAA receptor expression in the cerebellum of subjects with alcohol use disorders (AUD). Int. J. Neuropsychopharmacol. 20, 1013–1026 (2017).
Nelson, J. D., Denisenko, O. & Bomsztyk, K. Protocol for the fast chromatin immunoprecipitation (ChIP) method. Nat. Protoc. 1, 179–185 (2006).
Zhang, H. et al. Adolescent alcohol exposure epigenetically regulates CREB signaling in the adult amygdala. Sci. Rep. 8, 10376 (2018).
Zhou, Y., Zeng, P., Li, Y.-H., Zhang, Z. & Cui, Q. SRAMP: prediction of mammalian N6-methyladenosine (m6A) sites based on sequence-derived features. Nucleic Acids Res. 44, e91–e91 (2016).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Kim, J. K., Jeon, S. M., Lee, K. M., Park, E. S. & Cho, H. J. Expression of brain-derived neurotrophic factor in the rat forebrain and upper brain stem during postnatal development: an immunohistochemical study. Neuroscience 146, 1128–1136 (2007).
Joe, K. H. et al. Decreased plasma brain-derived neurotrophic factor levels in patients with alcohol dependence. Alcohol Clin. Exp. Res. 31, 1833–1838 (2007).
Pandey, S. C., Zhang, H., Roy, A. & Misra, K. Central and medial amygdaloid brain-derived neurotrophic factor signaling plays a critical role in alcohol-drinking and anxiety-like behaviors. J. Neurosci. 26, 8320–8331 (2006).
Keshavan, M. S., Giedd, J., Lau, J. Y., Lewis, D. A. & Paus, T. Changes in the adolescent brain and the pathophysiology of psychotic disorders. Lancet Psychiatry 1, 549–558 (2014).
Farris, S. P. & Mayfield, R. D. RNA-Seq reveals novel transcriptional reorganization in human alcoholic brain. Int. Rev. Neurobiol. 116, 275–300 (2014).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Liu, N. & Pan, T. N6-methyladenosine-encoded epitranscriptomics. Nat. Struct. Mol. Biol. 23, 96–102 (2016).
Pereira, J. D. et al. Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc. Natl Acad. Sci. USA 107, 15957–15962 (2010).
Acknowledgements
This work was supported by the National Institute on Alcohol Abuse and Alcoholism (NIAAA) P50AA022538, UO1AA-019971, U24AA-024605 [Neurobiology of Adolescent Drinking in Adulthood (NADIA) project], and RO1AA-010005 grants and the Department of Veterans Affairs (Senior Research Career Scientist award) to S.C.P. We also thank New South Wales Brain Tissue Resource Centre (NSW BTRC supported by NIH-NIAAA R28AA-012725) at the University of Sydney, Australia for providing postmortem human amygdala used in this study. We thank Dr. Eleonora Gatta in the Center for Alcohol Research in Epigenetics (CARE) for technical suggestions and discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bohnsack, J.P., Teppen, T., Kyzar, E.J. et al. The lncRNA BDNF-AS is an epigenetic regulator in the human amygdala in early onset alcohol use disorders. Transl Psychiatry 9, 34 (2019). https://doi.org/10.1038/s41398-019-0367-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-019-0367-z
This article is cited by
-
Advances in brain epitranscriptomics research and translational opportunities
Molecular Psychiatry (2024)
-
EZH2-dependent epigenetic reprogramming in the central nucleus of amygdala regulates adult anxiety in both sexes after adolescent alcohol exposure
Translational Psychiatry (2024)
-
E2F1 Mediates Traumatic Brain Injury and Regulates BDNF-AS to Promote the Progression of Alzheimer’s Disease
Neurotoxicity Research (2024)
-
N6-methyladenosine (m6A) epitranscriptomics in synaptic plasticity and behaviors
Neuropsychopharmacology (2023)
-
Arc controls alcohol cue relapse by a central amygdala mechanism
Molecular Psychiatry (2023)