Abstract
The dopaminergic hypothesis of schizophrenia (SZ) postulates that positive symptoms of SZ, in particular psychosis, are due to disturbed neurotransmission via the dopamine (DA) receptor D2 (DRD2). However, DA is a reactive molecule that yields various oxidative species, and thus has important non-receptor-mediated effects, with empirical evidence of cellular toxicity and neurodegeneration. Here we examine non-receptor-mediated effects of DA on gene co-expression networks and its potential role in SZ pathology. Transcriptomic profiles were measured by RNA-seq in B-cell transformed lymphoblastoid cell lines from 514 SZ cases and 690 controls, both before and after exposure to DA ex vivo (100 μM). Gene co-expression modules were identified using Weighted Gene Co-expression Network Analysis for both baseline and DA-stimulated conditions, with each module characterized for biological function and tested for association with SZ status and SNPs from a genome-wide panel. We identified seven co-expression modules under baseline, of which six were preserved in DA-stimulated data. One module shows significantly increased association with SZ after DA perturbation (baseline: P = 0.023; DA-stimulated: P = 7.8 × 10-5; ΔAIC = −10.5) and is highly enriched for genes related to ribosomal proteins and translation (FDR = 4 × 10−141), mitochondrial oxidative phosphorylation, and neurodegeneration. SNP association testing revealed tentative QTLs underlying module co-expression, notably at FASTKD2 (top P = 2.8 × 10−6), a gene involved in mitochondrial translation. These results substantiate the role of translational machinery in SZ pathogenesis, providing insights into a possible dopaminergic mechanism disrupting mitochondrial function, and demonstrates the utility of disease-relevant functional perturbation in the study of complex genetic etiologies.
Similar content being viewed by others
Introduction
Schizophrenia (SZ) is a disabling mental disorder characterized by severe disturbances in thought, behavior, and emotion, including psychotic symptoms and cognitive impairment1. Affecting approximately 1% of individuals globally2, SZ is heritable and highly polygenic3, with a number of neurobiological pathways tentatively implicated in its etiology. The dopaminergic hypothesis is a longstanding SZ model, attributing positive symptoms of the disorder, in particular psychosis, to dysregulation in dopaminergic neurotransmission via the dopamine (DA) receptor D2 (DRD2). This is supported by effects of psychotogenic stimulants (e.g., amphetamines) that activate DA receptors, as in vivo brain imaging studies have shown that amphetamine-induced increases in DA response are correlated with positive symptoms of SZ4,5. Moreover, it is well documented that antipsychotic drugs (e.g., chlorpromazine, haloperidol) block DRD2, with clinical response linked to receptor occupancy6. Meta-analysis of brain imaging data have shown increased post-synaptic DRD2 density in the striatum of SZ patients, but the relationship is complicated by the absence of significant differences between drug-naïve patients and controls, suggesting that the DRD2 upregulation may be due to antipsychotic treatment7. Results from genome-wide association studies (GWAS) do show an association between common variants at the DRD2 locus and SZ8, supporting the contention that D2 receptor variants affect SZ risk.
The action of DA on post-synaptic receptors represents one of the final steps of dopaminergic neurotransmission. A number of studies, however, have identified the most evident dopaminergic abnormality in SZ as being pre-synaptic and likely non-receptor-mediated, related to DA synthesis capacity, baseline synaptic DA levels, and/or DA release. Elevated pre-synaptic striatal DA levels have emerged as a fairly robust feature of SZ7,9,10,11, with increases also observed in the prodromal phase of the disorder that is linked to symptom severity and onset of psychosis12. In contrast, recent topographic analyses of extra-striatal brain regions have revealed DA deficits in the dorsolateral prefrontal cortex (DLPFC), with hypofunction associated with the activation of working memory in this region13. With current drug treatments primarily acting upon the same mechanism, namely D2/D3 receptor blockade, future research and drug development for SZ is needed to better target pre-synaptic DA abnormalities.
The molecular pathways linking aberrant DA levels to SZ etiology are not well understood. In addition to synaptic transmission, DA is known to have oxidative mechanisms that lead to apoptosis, a process that contributes to DA neuron loss in Parkinson’s disease (PD) and other neurodegenerative disorders14,15. Experimental studies provide a potential means of identifying such pathogenic pathways, without confounding due to drug treatment and other factors. Towards this end, we recently investigated non-receptor-mediated effects of DA on cellular gene expression in B-cell transformed lymphoblastoid cell lines (LCLs) and SZ risk. Using a cell perturbation approach ex vivo, we measured transcriptomic profiles by RNA sequencing (RNAseq) before and after exposure to DA in LCLs from SZ cases and controls, revealing differentially expressed genes enriched for brain expression and for functions related to immunity and apoptosis16. In this paper, we expand upon this work by examining differences in co-expression patterns (i.e., joint changes in gene behavior) due to DA stimulation. Such network-based approaches offer a means of clustering correlations in the transcriptome that tend to be biologically meaningful and can reveal insights into the larger genetic architecture of complex disorders. Using weighted gene co-expression network analysis, we successfully identified a co-expression module present at both baseline and in the DA-stimulated data, and whose association with SZ varies as a function of DA, with heightened disease correlation upon DA perturbation. This module is highly enriched for ribosomal proteins, as well as for genes implicated in neurodegenerative disorders, providing potential insights into non-receptor-mediated effects of DA in SZ pathogenesis.
Materials and methods
Samples
The RNAseq sample consists of 514 SZ cases and 690 controls after quality control (QC) processing (total sample = 1204), as previously described16. The subjects are of European ancestry and represent a subset of the Molecular Genetics of Schizophrenia (MGS) collection selected for GWAS and analyses of CNVs and transcriptomics16,17,18,19,20. There are 639 males (263 cases and 376 controls) and 565 females (251 cases and 314 controls) in this sample, with study enrollment ages ranging from 15 to 84 years, with detailed phenotypic data available17,21. Cases are severely affected on average, with most (~98%) exhibiting positive, psychotic symptoms (i.e., delusions, hallucinations). The NorthShore University HealthSystem Institutional Review Board approved this study, with informed consent obtained from all subjects.
Cell culture and DA perturbation
LCLs for the study sample were obtained from the Rutgers University Cell and DNA Repository22, for which we measured EBV load, viable cell count, and ATP level at cell harvest, all known to have an effect on gene expression23. For the design of the DA perturbation model, different DA concentrations were tested in a pilot study of four LCLs from control subjects16. At 100 μM, significant changes were observed for gene expression throughout the genome, affecting approximately 13% of genes, with only limited effects on LCL growth (reduction by ~20%), and thus 100 μM was selected for DA perturbation of the larger study sample. Cells were grown in independent wells in the presence or absence of DA, with DA exposure lasting 24 h. For comparison, in vivo DA concentrations in the human striatum, a brain region associated with dopaminergic abnormalities in SZ, is highly varied, with DA levels ranging from nM to μM concentrations, with significant differences between tonic DA release into extra-synaptic spaces (i.e., background activity) and the more transient and intense phasic activation that occurs in the synapses, with interactions between the two mechanisms [Grace, 1991]. Our ex vivo model represents a steady-state of DA exposure, albeit an extreme one, with an intended focus on pre-synaptic and non-receptor-mediated DA effects.
RNAseq and data processing
RNA sequencing was performed for baseline and DA-stimulated samples at the University of Minnesota Genomics Center on an Illumina HiSeq2000 at approximately 10 million reads per sample, with RNA quality scores indicative of high quality. Resulting RNAseq data were processed as previously described16,20,24. Alignment of 50-bp single reads to the Gencode v. 20 (GRCh38) human genome reference assembly was achieved with Tophat v. 2.0.5. Gene expression levels were calculated as reads per kilobase (kb) of transcript, per million mapped reads (RPKM) for the exon model of the longest transcript of a gene, with each quantile-normalized to account for batch biases and log2 transformed for variation stabilization. QC of gene RPKM levels, as previously described in detail16, involved several steps, including analysis of technical and biological replicates (i.e., same RNA and independent cell cultures of same LCL sample), consistency of sex chromosome gene expressions versus reported sex, comparison of RNAseq-called genotypes with previous GWAS SNP genotypes17,18, comparison of sample completion rate between cases and controls for each gene at baseline or DA-stimulation conditions, and identification of expression outliers based on PCA. QC processing yielded gene expression data for 21,043 genes for both baseline and DA-stimulated conditions. For downstream analyses, log2 transformed RPKM expression levels were residualized for the following relevant covariates: sex, age, cell counts and ATP levels at cell harvest under the two respective conditions, genotypic ancestry PCs 1–5, Epstein-Barr virus load, and sequencing batch. The variance in gene expression explained by these biological and technical confounds are presented in Supplementary Figures 1 and 2, with non-significant differences observed in the overall variance estimates for the baseline and DA-stimulated data sets (s2baseline = 4.20; s2DA = 4.18; P = 0.48).
Weighted gene co-expression network analysis (WGCNA)
To identify correlation patterns in the expression data for baseline and DA-stimulated conditions, WGCNA was performed using the R package of the same name (WGCNA v. 3.3.3)25. The WGCNA pipeline is as follows: (1) Using the command “pickSoftThreshold”, we assessed whether gene expression data have “scale-free topology” (i.e., frequency distribution of k, which is the summation of pairwise correlation coefficients for each gene, follows a power law) and identified values of the exponential parameter β for achieving it. A transformation such as this down-weights weaker correlations between genes, resulting in more cohesive co-expression networks that are centered on and stabilized by highly connected “hub genes”, which the authors of this approach argue are robust to random changes in connection patterns and more closely resemble true biology26. (2) Adjacency matrices were computed for each data set (i.e., baseline and DA-stimulation), representing pairwise correlation coefficients (i.e., Pearson’s r) transformed by the aforementioned β to ensure a scale-free correlation structure. We employed the unsigned method, in which absolute values of the coefficients were transformed, thus avoiding ablation of any strongly negative co-expression relationships (note: we also performed WGCNA based on signed adjacency matrices, in order to investigate the robustness of our identified modules and their associations with SZ). (3) From the adjacency matrices, topological overlap matrices (TOMs) were computed, representing the “interconnectedness” between pairs of genes, both directly, as well as indirectly, with connection strengths mediated by shared gene neighbors that are one-step away, which reportedly achieves more cohesive and biologically meaningful modules than ones determined solely from direct correlations27. These values were then used to calculate a dissimilarity distance measure, DistTOM, equating to 1—TOM. (4) We constructed dendrograms for the 21,043 genes with available gene expression data based on hierarchical clustering of DistTOM scores. This was achieved using the well-established UPGMA method via the R command “hclust”. Modules of co-expressed genes were then determined from branches in the resulting dendrograms using the command “cutreeDynamic”, which performs adaptive branch pruning based on various criteria, including a minimum cluster size of 50 genes and a conservative branch cut height of 0.99. The Partitioning Around Medoids option, which is a greedy algorithm for identifying outlying genes for module inclusion, was not utilized in order to maintain cohesive modules within the dendrograms. With these particular settings, modules of highly correlated genes will be preferentially detected, with the larger number of genes exhibiting weaker network connectivities left unassigned, including potential risk genes. (5) Module eigengenes (ME) were computed for each subject, which simply represent the first principal component of expression levels of genes assigned to a particular module28. ME scores were then used to calculate pairwise correlations between modules, as well as with SZ status, which in turn were used to generate dissimilarity distance matrices (DistME) by subtracting the coefficients from one and construct dendrograms via UPGMA. A distance threshold of 0.25 was used to identify closely related modules, which were then merged, necessitating the recalculation of MEs for final module assignments.
WGCNA results were visualized through a combination of dendrograms and heatmaps (R commands “TOMplot” and “plotEigengeneNetworks”), as well as network topologies comprised of gene nodes and edges (i.e., gene-gene connection strengths from the adjacency matrices; threshold = 0.01) that were constructed in Cytoscape v. 3.6.029. Significance of gene overlap between modules across the two conditions was determined through pairwise hypergeometric testing of 2 × 2 contingency tables (i.e., Fisher’s exact test). Modules were originally assigned random color names, which were subsequently changed such that modules with significant overlap have matching color names, in an effort to simplify for the reader the downstream analyses that compare WGCNA results across the two conditions.
Association between WGCNA modules and SZ
The relationships of the baseline modules to SZ status (i.e., case or control) were examined through logistic regression analysis of the respective ME scores in R v. 3.3.2 (note: no other covariates were included since gene expression data were residualized; see above), as well as for recomputed MEs based on the DA-stimulated data for the same set of module genes. Association results for the two sets of MEs were compared, with goodness-of-fit assessed by differences in Akaike Information Criteria (ΔAIC) for the DA-stimulated and baseline models (note: due to multicollinearity, multiple regression models that include both MEs were not examined). Moreover, we calculated the proportion of genes in each module that have been linked to SZ risk in the large GWAS conducted by the SZ Working Group of the Psychiatric Genomics Consortium (PGC)8. The 108 SNPs and indels that displayed genome-wide significance in the PGC study were assigned to 1308 genes, including non-protein coding RNAs, using a 250 K bp window around the loci in the GRCh38 reference assembly, with the significance of the enrichments determined via permutations (10,000×) weighted by total gene connectivity scores (k) from the adjacency matrices.
Preservation of WGCNA modules
How well the baseline modules were preserved under DA-stimulation (and vice versa) was assessed in the WGCNA program through various statistics related to module density and connectivity in the two data sets30, including correlations of: intramodular connectivity (kIM), which is the summation of adjacency matrix values, or connection strengths, of a module gene with other module genes; and eigengene-based connectivity (kME), representing the correlation of gene expression profiles with MEs of a given module. To summarize the different preservation metrics, a composite statistic based on median rank was computed for each module. The kIM scores were also used to screen for so-called “hub genes”, highly connected nodes that may represent key drivers in the co-expression modules.
Gene set enrichment analysis
To identify classes of genes or biological features that are over-represented in our WGCNA modules, we performed enrichment analyses for gene lists defined by GO-terms (molecular functions and biological processes) and KEGG pathways using the online tool DAVID v. 6.831, relative to the total set of genes comprising our expression data. Significance of gene enrichment was evaluated using a modified, and more conservative, Fisher’s Exact P-value, as well as false discovery rate (FDR). In addition, we employed the functional annotation clustering algorithm available in DAVID, which accounts for the redundant nature of gene annotations, and reports groups of related annotation terms from across various bioinformatics databases, yielding enrichment scores representing the geometric means of Fisher’s Exact P-values for individual gene sets tested in the primary analyses.
Genome-wide association testing
We screened the genome for quantitative trait loci (QTL) influencing co-expression patterns of select WGCNA modules. Genotype association testing was performed on ME scores in PLINK v. 1.07 via linear regression32, which included the first five principal components as covariates to control for potential population substructure. The genotype data were generated on the Affymetrix 6.0 array, with 671,422 SNPs passing QC filtering, as previously described17. In addition, we tested for SNP*SZ interaction in the linear model in an effort to identify QTL effects that are modified by case-control status (i.e., comparing the SNP regression coefficients of the two groups).
Results
WGCNA modules
While we have previously identified a large number of genes whose expression level response to DA perturbation varies between LCLs from SZ cases compared to ones from unaffected individuals16, the interpretation of these transcriptome-wide results are complex. We therefore employed an alternative analytical approach, one that focuses on gene co-expression networks—and thus presumably biology—to potentially yield additional insights. The first step in WGCNA was to evaluate the scale-free topology of our gene expression data sets. This is characterized by a correlation structure in the data that is comprised of highly connected hub genes, with relatively weaker pairwise correlations outside the hubs. Neither the baseline nor the DA-stimulated data are scale-free according to the WGCNA standard, which is the norm for gene expression studies. Thus, the baseline and DA data sets needed to be transformed for the adjacency matrices by down-weighting weaker correlations, with the β parameter set to 8 and 10, respectively, to achieve an r2 ≥ 0.9 with the scale-free model. Given our interest in the changes in gene expression in response to DA-stimulation (and whether it varies between SZ cases and controls), we also evaluated differential expression levels between the two conditions (i.e., response to DA perturbation), which were the target phenotype of a previous paper reporting on transcriptomic signatures of SZ by our research group16. However, the response data are not scale-free, and we failed to achieve robust fits to the scale-free model when transformed under a series of β powers (ranging from 1 to 20) for the total sample, as well as for SZ cases and controls separately. The authors of the WGCNA approach recommend against using such differential data, as it alters gene-gene co-expression structure and leads to single, highly correlated networks, invalidating the scale-free assumption. Thus, we performed WGCNA on the baseline and DA-stimulated data sets separately, and then compared the resulting modules.
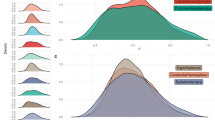
For the baseline data, 13 co-expression modules were initially identified, each randomly assigned to a color (Supplementary Figure 3), which were reduced to seven after merging modules with similar expression profiles (DistME < 0.25). The sizes of the seven modules range from 74 to 521 genes, with mean intramodular k values ranging from 1.17 to 8.61 (overall mean of 5.44), indicating that the identified modules comprise of highly intercorrelated genes. Most of the genes in the dataset are unassigned (n = 19,366), labeled as “grey”. As is evident from the deep branching in the dendrogram and the corresponding red highlights in the heatmap in Fig. 1a, the brown and blue modules exhibit the strongest gene-gene interconnectedness based on TOM dissimilarity distances. When comparing modules based on their eigengene profiles (Fig. 1b), the black and green modules show the strongest connections between one another (DistME = 0.39), sharing a clade with the green-yellow and magenta modules. To better visualize the gene co-expressions beyond what are represented in these binary-tree hierarchies, we constructed a two-dimensional network topology in Cytoscape v. 3.6.0 (Fig. 1c). With a minimum threshold of 0.01 for gene-gene connection strength, the purple module emerges as an outlier, with only a small number of network edges paired with black nodes. Overall, the network shows where connections are most concentrated (e.g., between the brown and black modules) and where they are not (e.g., between the brown and blue modules).
a Heatmap of pairwise TOM scores, aligned with the relevant dendrogram branches, of genes assigned to the seven baseline modules. b Correlation heatmap and dendrogram of eigengene profiles for baseline modules (including SZ status). c Two-dimensional network of gene-gene connection strengths (adjacency matrix values; minimum of 0.01) created in Cytoscape v. 3.6.0. Node colors correspond to baseline modules to which the genes were assigned. Top results from gene enrichment analyses of GO-terms and KEGG pathways are also shown, with Fisher’s Exact P-values in parentheses
For the DA-stimulated data, seven co-expression modules were initially identified (Supplementary Figure 4), which were reduced to six after merging of two similar modules. The six modules range from 62 to 198 genes. Relative to the baseline results, the number of unassigned genes (n = 20,401) increased by 1,035 genes, perhaps indicating transcriptome-wide disruption in co-expression patterns upon DA exposure of the LCLs. To understand how these modules relate to the ones generated in the baseline data, we performed pairwise hypergeometric testing across the two sets, thus identifying modules that show significant overlap in gene membership (Table 1). All six DA-stimulated modules were found to have substantial and exclusive overlap with a baseline module, ranging from 86.9% to all of the genes being shared. Similar to the baseline, the brown DA-stimulated module displays the strongest connectivity in the gene heatmap, harboring the deepest branches in the corresponding dendrogram (Fig. 2a). The black and green modules again were closest in their co-expression levels (Fig. 2b; DistME = 0.46), which is evident in the Cytoscape network (Fig. 2c), with most of the edges positioned between these two modules.
a Heatmap of pairwise TOM scores, aligned with the relevant dendrogram branches, of genes assigned to the six DA-stimulated modules. b Correlation heatmap and dendrogram of eigengene profiles for DA-stimulated modules (including SZ status). c Two-dimensional network of gene-gene connection strengths (adjacency matrix values; minimum of 0.01) created in Cytoscape v. 3.6.0. Node colors correspond to DA-stimulated modules to which the genes were assigned. Top results from gene enrichment analyses of GO-terms and KEGG pathways are also shown, with Fisher’s Exact P-values in parentheses
Given the significant overlap in the genes comprising the baseline and DA-stimulated modules, it is perhaps not surprising that the gene-gene interconnectedness of the baseline modules is well preserved in the DA-stimulated expression data (Supplementary Table 1), with strong correlations in both kIM, ranging from 0.78 to 0.99, and kME, ranging from 0.82 to 0.99 (excluding the purple module outlier). Moreover, for three of the six matching modules, the same hub genes are identified based on the top kIM scores (Supplementary Table 2): RPL11 for the brown modules; HNF1b for the black modules; and NFKB2 for the green modules. The median ranks of the various preservation statistics from the WGCNA program indicate that the brown module is the best preserved of the seven baseline modules in the DA-stimulated data, and vice versa.
Gene set enrichment analyses
To investigate the potential biological relevance of the WGCNA modules, we performed gene set enrichment analyses for GO-terms and KEGG pathways in DAVID (Table 2). For both the baseline and DA-stimulated results, highly significant, replicated enrichments were observed in the pair of matching brown modules: GO-term “SRP-dependent co-translational protein targeting to membrane”, representing 42.3% and 70.6% of the genes, respectively (FDR = 4.0 × 10−141 and 2.7 × 10−161); and KEGG pathway “Ribosome”, representing 40.7% and 68.8% of the same genes (FDR = 2.3 × 10−110 and 1.1 × 10−125). This is also reflected in the functional annotation clustering of the entire enrichment results (Supplementary Table 3), as the brown modules produced the highest overall scores in each data set for terms related to ribosomal translation. Some of the other top enrichments that were observed in both matching modules include: KEGG pathway “Protein processing in endoplasmic reticulum” for the blue modules; GO-term “Inflammatory response” for the green modules; KEGG pathway “Cell cycle” and GO-term “Cell division” for the green-yellow modules; and GO-term “Poly(A) RNA binding” for the magenta modules. Among the top enrichment results that are not shared, most remain highly enriched in the alternate module but are simply not the top hit (e.g., TNF signaling is the second highest enriched KEGG pathway in the DA-stimulated green module; FDR = 4.0 × 10−8). Although DA-stimulation does appear to have broadly weakened the top enrichment signals observed in the baseline modules, some dramatically (e.g., GO-term “Defense response to virus” in the black modules, with the baseline FDR of 5.2 × 10−17 becoming a non-significant 0.59), with the exceptions being the GO-term “Cell division” for the green-yellow modules and the top results from the brown modules.
Associations with SZ
The gene network modules were initially identified and subsequently characterized using the entire gene expression data sets, without taking SZ status into account. Thus, to determine whether the co-expression levels of a module differ between cases and controls, ME scores for the baseline modules (proportion of the variation explained ranging from 0.48 to 0.57) were tested for association with SZ (Table 3). With the exception of the green-yellow (P = 0.11) and brown modules (P = 0.023), the gene co-expressions of the baseline modules show highly significant associations with SZ (P < 10−5), with the green module, which is enriched for genes related to inflammatory response (see above), having the most significant result (P < 2.0 × 10−16). This is in general agreement with our previous report which found a vast number of genes to be differentially expressed between cases and controls at baseline16. To investigate the impact of DA-stimulation on these findings, we recomputed the MEs using the DA-stimulated expression data for the exact same sets of module genes (accounting for 0.42–0.52 of the variation), and then retested the associations. In other words, we compared our two conditions by generating respective pairs of co-expression profiles for each baseline module and thus fixed for gene membership. For six of the modules, less significant associations were observed, reflected by decreases in the estimated regression coefficients. The lone exception is the brown module (enriched for translational machinery), for which the P-value decreased from 0.023 to 7.8 × 10−5, corresponding to an increase in absolute effect size from −1.12 ± 0.96 to −1.95 ± 0.96 (Z = 1.19; P = 0.11). This module is also the only one exhibiting greater goodness-of-fit between SZ and its DA-stimulated MEs (ΔAIC = −10.5), representing approximately 190-fold increase in model likelihood33. As one would expect, similar association results were observed for MEs from the matching DA-stimulated modules (Supplementary Table 4). Notably, 4.1% of the genes in the brown module are located within 250 K bp of the 108 SZ risk loci reported by the PGC8, which is the highest percentage among the seven modules, and represents near significant enrichment relative to the 2.5% observed for unassigned genes in the baseline data set (empirical P = 0.079).
Signed WGCNA networks
The modules reported here are based on unsigned networks, which incorporate negative correlations between genes. To determine how this potentially impacted our network construction (i.e., TOM calculations) and association test results, we re-ran WGCNA using a signed adjacency matrix for the baseline data. All seven of the original baseline modules were found to have matching modules from the signed network, with highly significant overlaps in gene memberships (P < 1.9 × 10−163), and each corresponding to the same top enrichments in GO-terms and KEGG pathways (e.g., brown module: SRP-dependent co-translational protein targeting, FDR = 3.6 × 10−117; and ribosome, FDR = 5.5 × 10−105), except for the purple module. Moreover, the matching green module from the signed network is again the most strongly associated with SZ (P < 2.0 × 10−16), and similarly the brown module is the only one exhibiting a marked increase in association upon DA-stimulation, producing an identical ΔAIC of -10.5, which is again a stark outlier among all the other identified modules (ΔAIC ranges from −0.4 to 21). All of this underscores the robustness of our key findings.
QTLs for the brown module
Given that the brown module is the only module that exhibits stronger association with SZ upon DA-stimulation, with preserved connectivity patterns and gene enrichment profile from the baseline condition, we scanned its co-expression levels for QTLs, representing genetic “drivers” of regulatory systems that may underlie the architecture of transcriptional patterns (e.g., transcription factors, non-coding RNAs (ncRNAs), feedback mechanisms), by testing its ME scores for association with genome-wide SNP data (n = 671,423) available for the study sample. The top-5 association results for the brown module under DA-stimulation are presented in Table 4, with the MEs accounting for 64% of the variation. None achieved genome-wide significance (FDR = 0.35; genomic inflation factor λ = 1.0), with the top SNP, rs6504934 (P = 2.7 × 10−6), located upstream (~18 kb) of a long non-coding RNA (lncRNA), RN7SKP14. For the other four SNPs listed in the table, all are in close proximity and exhibit strong LD (pairwise r2 > 0.87), located within or near FASTKD2. This gene encodes a pro-apoptotic protein that is required for mitochondrial ribosome assembly34,35, and has been associated with memory performance and hippocampal structure36.
We also tested for interaction between SNP genotypes and case-control status on co-expression scores in order to assess whether QTL effects vary as a function of SZ under DA-stimulation. The top-5 SNP*SZ interaction effects are given in Table 5 (note: results for the baseline data are available in Supplementary Table 5). Again, none achieved genome-wide significance (FDR = 0.18; λ = 0.99). The top SNP, rs10497316 (P = 3.8 × 10−7), is located in the gene XIRP2, which encodes an actin-binding protein, and does not show evidence of association with SZ in the PGC GWAS results, although strong enrichment of rare loss-of-function (LoF) mutations in XIRP2 were observed among SZ cases in a recent whole-exome sequence study37. For the other SNPs listed in the table, they are located either within or closest to lncRNA genes, with three in near-perfect LD (pairwise r2 > 0.99), and all four showing nominal evidence of association with SZ in the PGC data (P-values ranging from 0.050 to 0.0043).
Discussion
With the rapid expansion of transcriptome sequencing over the past decade, researchers have increasingly focused on the expression of genes relative to each other (i.e., co-expression) as a means of assigning putative functions to genes and ncRNAs based on the annotations of wider co-expression networks (so-called “guilt-by-association”), as well as gaining insight into potential regulatory relationships that underlie biological processes. Such network-based approaches reduce the inherent complexity and dimensionality of genome-wide expression data, as most genes are weakly correlated with other genes and thus disregarded, depending on chosen thresholds. Moreover, co-expression analyses have been used to identify novel risk genes for various human diseases, including psychiatric disorders, as coordinated gene expression is critical for brain development and function. This includes SZ, for which studies of postmortem brain specimens and peripheral blood have found dysregulated gene networks, including modules enriched for genes involved in synaptic transmission38, immune function39,40, oxidative stress and mitochondria41,42, and neurogenesis and neuron differentiation42,43.
In our network analysis of the genome-wide effects of DA perturbation on gene expression in LCLs, we examined the total sample, comprising both SZ cases and controls. Prior studies of the brain transcriptome have found gene co-expression patterns to be organized into distinct cellular and functional categories44, with significant overlap observed in gene membership of case- and control-only modules, suggesting that module composition and gene-gene connectivity per se are not likely to be key determinants in the pathogenesis of SZ42, although this does not preclude the possibility of certain regulatory relationships between smaller numbers of genes. Thus, our aim was to identify changes in co-expression levels within our networks that are associated with SZ status, to which one module, related to ribosomal translation, has revealed a marked increase in disease risk upon DA-stimulation in our sample.
Based on WGCNA, we identified six co-expression modules under baseline conditions that are preserved under DA-stimulation, both in terms of gene-gene network connectivities and significant overlap in gene membership with corresponding modules generated from the DA-stimulated data. When tested for their association with SZ status, the co-expression profiles (i.e., eigengenes) of the green baseline module (n = 127 genes) show the strongest effect, which is reflected by its median FDR of 1.1 × 10−9 for the genes within the module when analyzed individually in prior gene-based association results16. This module has a significant enrichment of genes related to inflammatory response, in particular TNF signaling and NF-kappa B signaling—cytokine pathways that are involved in systemic inflammation, apoptosis, immune response, and synaptic plasticity, with substantial evidence implicating them in the pathophysiology of SZ45,46,47,48. Under DA-stimulation, the corresponding module (n = 62 genes) also produced the most significant association with SZ, with the same pattern of primary enrichment for genes involved in inflammation and NF-kappa B and TNF signaling. These results are consistent with our previous studies on these expression data16,20,22, which found immune-related genes enriched among both baseline and DA-stimulated transcripts that are differentially expressed by affection status, as well as transcriptome studies by other groups49, thus further supporting the immune and cytokine hypothesis for SZ.
Overall, five of the seven baseline modules are strongly associated with SZ (P < 10−5), both for the baseline and DA-stimulated data, with the co-expression profiles of the unassigned genes also exhibiting a highly significant association (P = 4.3 × 10−11). This is consistent with our gene-based results, which showed a large number of SZ-associated gene expressions under both conditions (31 and 21%, respectively; FDR < 0.05). As previously discussed16, we did not find any evidence that these results are a consequence of any technical artifacts, and thus assume that they reflect real biology in our well-controlled LCL study. Intriguingly, this may reflect the recently proposed “omnigenic” model50, where most, if not all, genes outside core disease-related pathways indirectly contribute to disease risk, especially for transcriptomics and regulatory networks that involve higher-order, interconnected structures.
DA-stimulated effects related to translational machinery
Of the six co-expression modules that correspond across the two conditions, only the brown module (n = 194 genes at baseline) shows stronger evidence of association with SZ after DA perturbation, with the estimated effect changing from −1.12 ± 0.96 (P = 0.023) to a highly significant −1.95 ± 0.96 (P = 7.8 × 10−5), with the latter representing a substantial 190-fold increase in model likelihood. Moreover, eight of the baseline module genes (4.1%) are in proximity to the 108 significant GWAS loci reported by the PGC for SZ, the highest proportion observed among our modules (empirical P = 0.079), seven of which are downregulated and exhibit more significant associations with SZ after DA-stimulation (top FDR = 1.7 × 10−4): MRPS21, NDUFB3, NFATC3, RBX1, RPL13A, RPS11, SRP14, and ZFAS1. Despite cellular stress and transcriptome-wide changes caused by the DA perturbation of the LCLs (~91% of genes responsive to DA at FDR < 0.05)16, gene-gene connectivity patterns in the brown module are strongly preserved in the DA-stimulated expression data (see Supplementary Table 1).
For the brown module under both conditions, but especially DA-stimulation, highly significant enrichments were observed for genes involved in signal recognition particle (SRP)-dependent targeting for endoplasmic reticulum (ER) translation (~134-fold; FDR = 2.7 × 10−161) and ribosomal translation (~43-fold; FDR = 1.1 × 10−125), accounting for 67.6 and 69.4% of the module genes, respectively. Coupled with the increased association with SZ under DA conditions, these results for the brown module are compelling, as ribosomal proteins and the broader translational machinery have been implicated in neurodevelopment and SZ in a number of recent papers. A gene knockdown study in the rat forebrain found neuronal maturation to be associated with a considerable expansion of ribosomal proteins, with translational insufficiency impairing dendritic growth and neuronal connectivity51. In another study involving human stem cell-derived neural progenitor cells from SZ patients and controls52, increased levels of global protein synthesis and translational machinery, including ribosomal proteins, were observed in the SZ cells. In contrast, for human olfactory neurosphere-derived cells, discovery-based proteomics and functional analyses revealed significant reductions in particular ribosomal proteins among SZ patients, including total ribosomal signal intensity53. And, perhaps most intriguingly, Zhou et al54. reveal that the interactome of ZNF804A, a SZ risk gene robustly replicated in different populations55,56,57,58, is highly represented by ribosomal and mitochondrial proteins, with ZNF804A modulating translational efficiency. Moreover, the ribosomal protein RPSA interacts with ZNF804A, and rescues neuronal migration and translational defects caused by knockdown of the ZNF804A homolog in mice, linking the SZ risk gene to neurodevelopment and translational control. Although ZNF804A was not assigned to our brown modules, RPSA was for both baseline and DA-stimulated data, and interestingly, the DA-stimulated expression levels of RPSA show a significant negative correlation with SZ (FDR = 3.9 × 10−6), with its differential expression between the two conditions (i.e., DA response) yielding the top association result among the 110 overlapping module genes (FDR = 0.027)16.
Links to mitochondrial function?
Although we were unable to identify genome-wide significant QTLs for the brown module under DA-stimulation, the top results are nonetheless of interest. In particular, FASTKD2 harbors a linkage disequilibrium block of strongly associated SNPs (minimum P = 2.8 × 10−6), with the gene playing a critical role in the biogenesis of mitochondrial ribosomes and translation34,35. The encoded RNA-binding protein modulates apoptosis, and is involved in the assembly of mitochondrial RNA granules, often induced under conditions of cellular stress, with abnormal accumulation of such granules linked to some neurodegenerative disorders59,60. Interestingly, the top enrichment result for the brown module, SRP-dependent targeting for ER translation, serves as a mechanism for mRNA to escape stress granule sequestration through localization to the ER61, which has obvious implications for neuronal cell survival. Furthermore, a SNP within FASTKD2, rs7594645, has been previously linked to memory performance in older adults, with carriers exhibiting neuroprotective effects, including increased hippocampal volume and gray matter density, and decreased cerebrospinal fluid levels of apoptotic mediators, all of which are features of Alzheimer’s disease36. And in a pair of interactome studies based on affinity capture-mass spectrometry62,63, FASTKD2 was identified as a protein-protein interaction partner of NGRN, a SZ risk gene with downstream effects on ZNF804A in the aforementioned work by Zhou et al.54, as part of a larger oxidative phosphorylation (OXPHOS) network regulating mitochondrial translation.
For SNP*SZ interactions on DA-stimulated co-expression, rs10497316 in XIRP2 was our top finding for the brown module (P = 3.8 × 10−7), reflecting a marked difference in the magnitude and direction of its main effect among SZ cases (β = 0.013 ± 0.0030; P = 6.77 × 10−6) versus controls (β = −0.0063 ± 0.0050; P = 0.014). The gene encodes a protein that protects actin filaments from depolymerization and is involved in neuronal integrity. In a large whole-exome sequencing study by the UK10K Consortium37, a disproportionately high number of LoF mutations was reported for XIRP2 among SZ cases, corresponding to its top gene-based association for this class of variants (P = 3.5 × 10−5). Moreover, XIRP is a known target of MEF2A64,65, a DNA-binding transcription factor that mediates neuronal differentiation and survival, as it is cleaved by mitochondrial apoptotic caspases during excitotoxic neuronal stress66, and XIRP2 has previously shown decreased expression in brain samples of PD patients67. For the other top interaction effects listed in Table 5, the SNPs all show nominal association with SZ in the PGC data (P < 0.05), and are positioned within or closest to lncRNAs, an abundant class of RNA molecules that regulate gene expression and are believed to play a critical role in neuronal development68, with prior co-expression network analyses implicating them in early onset SZ69.
Taken together, these SNP association results for the brown module suggest potential links with mitochondrial processes and cell survival/apoptosis, in particular neurodegeneration, as impaired ribosome production has been documented in neurodegenerative diseases70,71,72. This is borne out in our enrichment analyses for the KEGG lists in the baseline module (Fig. 1c; see Supplementary Table 6), as the next most enriched pathways after ribosomes are genes related to: OXPHOS in mitochondria (FDR = 2.4 × 10−15); Huntington’s disease (FDR = 9.0 × 10−14); Parkinson’s disease (FDR = 1.2 × 10−12); and Alzheimer’s disease (FDR = 3.9 × 10−12). For the DA-stimulated module, however, all four of these failed to show significant enrichment (minimum FDR = 0.20), stemming from the non-inclusion of 21 genes belonging to these pathways, 16 of which are shared. Taking a closer look at this splinter group, we found the association of its eigengenes with SZ status to change in a manner similar to the larger brown module (ΔAIC = −6.6), going from a non-significant association at baseline (β = −0.70 ± 0.94; P = 0.16) to a significant one (β = −1.44 ± 0.94; P = 3.50 × 10−3), with DA-stimulation weakening the co-expression between genes from this subgroup and the rest of the brown module, as evidenced by pairwise correlations in their eigengene scores (baseline: r = 0.81 ± 0.018; DA-stimulation: r = 0.69 ± 0.029).
Mitochondrial dysfunction characterizes various neurodegenerative diseases, reflected by the gene enrichments of the aforementioned KEGG pathways. Mitochondria play key roles in ATP generation, reactive oxygen species formation, and apoptosis, with neurons particularly dependent on mitochondria because of their high energy demands, and thus susceptible to cellular damage due to oxidative stress73. What is more, impaired mitochondrial function and variants in related genes have been repeatedly implicated in SZ74,75,76, with prior co-expression network analyses of SZ reporting similar modular enrichments39,41,69. SZ cases have been shown to exhibit abnormal mitochondrial respiration77, increased oxidative stress78, and altered gene expression of mitochondrialcomplexes79. Remarkably, in a recent study of pyramidal neurons from the DLPFC of postmortem SZ subjects, significant transcriptome alterations were identified in molecular pathways related to mitochondrial energy production and the regulation of protein translation, which corroborates our main findings80. These alterations of nuclear-encoded genes involved in energy production are consistent with the poor activation of DLPFC circuitry during increased cognitive demands that has been long observed in SZ81. Moreover, subjects responsive to antipsychotic treatment were identified in another study as having approximately a 40% decrease in the number of mitochondria per synapse in dorsal striatum at postmortem, with treatment-resistant cases having normal levels82, thus suggesting that there may be a biological distinction between treatment response and treatment resistance in SZ, for which DA may represent a key trigger. DA has been shown to have non-receptor-mediated effects on mitochondrial function, along with antipsychotics, as it inhibits Complex I activity and ATP production of the respiratory chain through its uptake into the organelle83,84,85,86. This potential link between DA and mitochondrial dysfunction in SZ risk is compelling and, given our findings, warrants further investigation.
Conclusion
The results of our study identify a co-expression network that exhibits increased risk for SZ upon DA-stimulation of LCLs, highly enriched for ribosomal proteins and translational machinery. This suggests that relevant but cryptic pathological mechanisms underlying SZ can become detectable by functional perturbation, which makes replication of these findings more involved given our unique study design. Moreover, the use of LCLs as a cellular model versus brain presents some clear limitations for our study of a presumably brain-related disorder, as expression of some genes in LCLs will substantially differ from that in brain, although most expression signatures are shared between different tissues87,88,89,90,91. However, LCLs also have distinct advantages, as they are derived from the most accessible tissue, yielding sizeable samples, and allow for experimental manipulations such as DA perturbation, as we have discussed previously in greater detail16. Nonetheless, our findings provide key insights into the long-standing dopaminergic hypothesis for SZ based on gene co-expression changes, revealing non-receptor-mediated DA disturbances in translational machinery and mitochondrial function, including genes involved in neurodegeneration, with potential treatment targets of pre-synaptic dopaminergic features commonly observed in the disorder.
References
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders. 5th Edition, (American Psychiatric Publishing, Washington, DC, 2013).
van Os, J. & Kapur, S. Schizophrenia. Lancet 374, 635–645 (2009).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Abi-Dargham, A. et al. Increased striatal dopamine transmission in schizophrenia: confirmation in a second cohort. Am. J. Psychiatry 155, 761–767 (1998).
Breier, A. et al. Schizophrenia is associated with elevated amphetamine-induced synaptic dopamine concentrations: evidence from a novel positron emission tomography method. Proc. Natl Acad. Sci. USA 94, 2569–2574 (1997).
Kapur, S. et al. Relationship between dopamine D(2) occupancy, clinical response, and side effects: a double-blind PET study of first-episode schizophrenia. Am. J. Psychiatry 157, 514–520 (2000).
Howes, O. D. et al. The nature of dopamine dysfunction in schizophrenia and what this means for treatment. Arch. Gen. Psychiatry 69, 776–786 (2012).
Ripke, S. et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Howes, O., Williams, M., Ibrahim, K. & Leung, G. Midbrain dopamine function in schizophrenia and depression: a post-mortem and positron emission tomographic imaging study. Brain 136, 3242–3251 (2013).
Abi-dargham, A. et al. Increased baseline occupancy of D2 receptors by dopamine in schizophrenia. Proc. Natl Acad. Sci. USA 97, 8104–8109 (2000).
Kegeles, L. S. et al. Increased synaptic dopamine function in associative regions of the striatum in schizophrenia. Arch. Gen. Psychiatry 67, 231–239 (2010).
Howes, O. D., McCutcheon, R., Owen, M. J. & Murray, R. The role of genes, stress and dopamine in the development of schizophrenia. Biol. Psychiatry 81, 9–20 (2017).
Slifstein, M. et al. Deficits in prefrontal cortical and extrastriatal dopamine release in schizophrenia. JAMA Psychiatry 10032, 1–9 (2015).
Segura-Aguilar, J. et al. Protective and toxic roles of dopamine in Parkinson’s disease. J. Neurochem. 129, 898–915 (2014).
Meiser, J., Weindl, D. & Hiller, K. Complexity of dopamine metabolism. Cell Commun. Signal. 11, 34 (2013).
Duan, J., Transcriptomic signatures of schizophrenia revealed by dopamine perturbation in an ex vivo model. Transl. Psychiatry. 8, 158 (2018).
Shi, J. et al. Common variants on chromosome 6p22.1 are associated with schizophrenia. Nature 460, 753–757 (2009).
Schizophrenia Psychiatric Genome-Wide Association Study Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Levinson, D. F. et al. Copy number variants in schizophrenia: confirmation of five previous findings and new evidence for 3q29 microdeletions and VIPR2 duplications. Am. J. Psychiatry 168, 302–316 (2011).
Sanders, A. R. et al. Transcriptome sequencing study implicates immune-related genes differentially expressed in schizophrenia: new data and a meta-analysis. Transl. Psychiatry 7, e1093 (2017).
Sanders, A. R. et al. The Internet-based MGS2 control sample: self report of mental illness. Am. J. Psychiatry 167, 854–865 (2010).
Sanders, A. R. et al. Transcriptome study of differential expression in schizophrenia. Hum. Mol. Genet. 22, 5001–5014 (2013).
Choy, E. et al. Genetic analysis of human traits in vitro: drug response and gene expression in lymphoblastoid cell lines. PLoS. Genet. 4, e1000287 (2008).
Duan, J. et al. Transcriptome outlier analysis implicates schizophrenia susceptibility genes and enriches putatively functional rare genetic variants. Hum. Mol. Genet. 24, 4674–4685 (2015).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 9, 559 (2008).
Zhang B., Horvath S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, 17 (2005).
Yip, A. & Horvath, S. Gene network interconnectedness and the generalized topological overlap measure. BMC Bioinform. 8, 22 (2007).
Langfelder, P. & Horvath, S. Eigengene networks for studying the relationships between co-expression modules. BMC. Syst. Biol. 1, 54 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Langfelder, P., Luo, R., Oldham, M. C. & Horvath, S. Is my network module preserved and reproducible? PLoS. Comput. Biol. 7, e1001057 (2011).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Purcell, S. et al. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am. J. Hum. Genet. 81, 559–575 (2007).
Burnham, K. P. & Anderson, D. R. Model Selection and Multimodel Inference: A Practical Information—Theoretic Approach. 2nd Edition, (Springer-Verlag, New York, NY, 2002).
Antonicka, H. & Shoubridge, E. A. Mitochondria RNA granules are centers for posttranscriptional RNA processing and ribosome biogenesis. Cell Rep. 10, 920–932 (2015).
Popow, J. et al. FASTKD2 is an RNA-binding protein required for mitochondrial RNA processing and translation. RNA 11, 1873–1884 (2015).
Ramanan, V. K. et al. FASTKD2 is associated with memory and hippocampal structure in older adults. Mol. Psychiatry 20, 1197–1204 (2015).
Singh, T. et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat. Neurosci. 19, 571–577 (2016).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Hess, J. L. et al. Transcriptome-wide mega-analyses reveal joint dysregulation of immunologic genes and transcription regulators in brain and blood in schizophrenia. Schizophr. Res. 176, 114–124 (2016).
de Jong, S. et al. A gene co-expression network in whole blood of schizophrenia patients is independent of antipsychotic-use and enriched for brain-expressed genes. PLoS. ONE. 7, e39498 (2012).
Maschietto, M. et al. Co-expression network of neural-differentiation genes shows specific pattern in schizophrenia. BMC Med. Genom. 8, 23 (2015).
Torkamani, A., Dean, B., Schork, N. J. & Thomas, E. A. Coexpression network analysis of neural tissue reveals perturbations in developmental processes in schizophrenia. Genome Res. 20, 403–412 (2010).
Chen, C. et al. Two gene co-expression modules differentiate psychotics and controls. Mol. Psychiatry 18, 1308–1314 (2013).
Oldham, M. C. et al. Functional organization of the transcriptome in human brain. Nat. Neurosci. 11, 1271–1282 (2008).
Hoseth, E. Z. et al. A study of TNF pathway activation in schizophrenia and bipolar disorder in plasma and brain tissue. Schizophr. Bull. 43, 881–890 (2017).
Goldsmith, D. R., Rapaport, M. H. & Miller, B. J. A meta-analysis of blood cytokine network alterations in psychiatric patients: comparisons between schizophrenia, bipolar disorder and depression. Mol. Psychiatry 21, 1696–1709 (2016).
O’Brien, S. M., Scully, P. & Dinan, T. G. Increased tumor necrosis factor-alpha concentrations with interleukin-4 concentrations in exacerbations of schizophrenia. Psychiatry Res. 160, 256–262 (2008).
Song, X.-Q., Lv, L.-X., Li, W.-Q., Hao, Y.-H. & Zhao, J.-P. The interaction of nuclear factor-kappa B and cytokines is associated with schizophrenia. Biol. Psychiatry 65, 481–488 (2009).
Fillman, S. G. et al. Increased inflammatory markers identified in the dorsolateral prefrontal cortex of individuals with schizophrenia. Mol. Psychiatry 18, 206–214 (2013).
Boyle, E. A., Li, Y. I. & Pritchard, J. K. An expanded view of complex traits: from polygenic to omnigenic. Cell 169, 1177–1186 (2017).
Slomnicki, L. P. et al. Requirement of neuronal ribosomes synthesis for growth and maintenance of the dendritic tree. J. Biol. Chem. 291, 5721–5739 (2016).
Topol, A. et al. Increased abundance of translation machinery in stem cell-derived neural progenitor cells from four schizophrenia patients. Transl. Psychiatry 5, e662 (2015).
English, J. A. et al. Reduced protein synthesis in schizophrenia patient-derived olfactory cells. Transl. Psychiatry 5, e663 (2015).
Zhou, Y. et al. Interactome analysis reveals ZNF804A, a schizophrenia risk gene, as a novel component of protein translational machinery critical for embryonic neurodevelopment. Mol. Psychiatry 23, 952–962 (2017).
O’Donovan, M. C. et al. Identification of loci associated with schizophrenia by genome-wide association and follow-up. Nat. Genet. 40, 1053–1055 (2008).
Zhang, R. et al. Population-based and family-based association studies of ZNF804A locus and schizophrenia. Mol. Psychiatry 16, 360–361 (2011).
Williams, H. J. et al. Fine mapping of ZNF804A and genome-wide significant evidence for its involvement in schizophrenia and bipolar disorder. Mol. Psychiatry 16, 59–66 (2011).
Riley, B. et al. Replication of association between schizophrenia and ZNF804A in the Irish case-control study of schizophrenia sample. Mol. Psychiatry 15, 29–37 (2010).
Li, Y. R., King, O. D., Shorter, J. & Gitler, A. D. Stress granules as crucibles of ALS pathogenesis. J. Cell Biol. 201, 361 (2013).
Ramaswami, M., Taylor, J. P. & Parker, R. Altered ribostasis: RNA-protein granules in degenerative disorders. Cell 154, 727–736 (2013).
Unsworth, H., Raquz, S., Edwards, H. J., Higgins, C. F. & Yagüe, E. mRNA escape from stress granule sequestration is dictated by localization to the endoplasmic reticulum. FASEB J. 24, 3370–3380 (2010).
Arroyo, J. D. et al. A genome-wide CRISPR death screen identifies genes essential for oxidative phosphorylation. Cell. Metab. 24, 875–885 (2016).
Huttlin, E. L. et al. Architecture of the human interactome defines protein communities and disease networks. Nature 545, 505–509 (2017).
Huang, H.-T. et al. Myomaxin is a novel transcriptional target of MEF2A that encodes a xin-related α-actinin-interacting protein. J. Biol. Chem. 281, 39370–39379 (2006).
McCalmon, S. A. et al. Modulation of angiotensin II-mediated cardiac remodeling by the MEF2A target gene Xirp2. Circ. Res. 106, 952–960 (2010).
Okamoto, S. et al. Dominant-interfering forms of MEF2 generated by caspase cleavage contribute to NMDA-induced neuronal apoptosis. Proc. Natl Acad. Sci. USA 99, 3974–3979 (2002).
Tatura, R. et al. Parkinson’s disease: SNCA-, PARK2-, and LRRK2-targeting microRNAs elevated in cingulate gyrus. Park. Relat. Disord. 33, 115–121 (2016).
D’haene, E. et al. Identification of long non-coding RNAs involved in neuronal development and intellectual disability. Sci. Rep. 6, 28396 (2016).
Ren, Y. et al. A co-expression network analysis reveals lncRNA abnormalities in peripheral blood in early-onset schizophrenia. Pro Neuropsychopharmacol. Biol. Psychiatry 63, 1–5 (2015).
Honda, K. et al. Ribosomal RNA in Alzheimer disease is oxidized by bound redox-active iron. J. Biol. Chem. 280, 20978–20986 (2005).
Vilotti, S. et al. Parkinson’s disease DJ-1 L166P alters rRNA biogenesis by exclusion of TTRAP from the nucleolus and sequestration into cytoplasmic aggregates via TRAF6. PLoS. ONE. 7, e35051 (2012).
Lee, J. et al. Dysregulation of upstream binding factor-1 acetylation at K352 is linked to impaired ribosomal DNA transcription in Huntington’s disease. Cell Death Differ. 18, 1726–1735 (2011).
Bhat, A. H. et al. Oxidative stress, mitochondrial dysfunction and neurodegenerative diseases; a mechanistic insight. Biomed. Pharmacother. 74, 101–110 (2015).
Anglin, R. E., Mazurek, M. F., Tarnopolsky, M. A. & Rosebush, P. I. The mitochondrial genome and psychiatric illness. Am. J. Med Genet. Part B 159B, 749–759 (2012).
Manji, H. et al. Impaired mitochondrial function in psychiatric disorders. Nat. Rev. Neurosci. 13, 293–307 (2012).
Gonçalves, V. F. et al. A comprehensive analysis of nuclear-encoded mitochondrial genes in schizophrenia. Biol. Psychiatry 83, 780–789 (2018).
Rosenfeld, M., Brenner-Lavie, H., Ari, S. G., Kavushansky, A. & Ben-Shachar, D. Perturbation in mitochondrial network dynamics and in complex I dependent cellular respiration in schizophrenia. Biol. Psychiatry 69, 980–988 (2011).
Wang, J. F., Shao, L., Sun, X. & Young, L. T. Increased oxidative stress in the anterior cingulate cortex of subjects with bipolar disorder and schizophrenia. Bipolar Disord. 11, 523–529 (2009).
Karry, R., Klein, E. & Ben-Shachar, D. Mitochondrial complex I subunits expression is altered in schizophrenia: a postmortem study. Biol. Psychiatry 55, 676–684 (2004).
Arion, D. et al. Transcriptome alterations in prefrontal pyramidal cells distinguish schizophrenia from bipolar and major depressive disorders. Biol. Psychiatry 82, 594–600 (2017).
Weinberger, D. R., Berman, K. F. & Zec, R. F. Physiologic dysfunction of dorsolateral prefrontal cortex in schizophrenia. I. Regional cerebral blood flow evidence. Arch. Gen. Psychiatry 43, 114–124 (1986).
Roberts, R. C. Postmortem studies on mitochondria in schizophrenia. Schizophr. Res. 187, 17–25 (2017).
Burkhardt, C., Kelly, J. P., Lim, Y. H., Filley, C. M. & Parker, W. D. Jr. Neuroleptic medications inhibit complex I of the electron transport chain. Ann. Neurol. 33, 512–517 (1993).
Whatley, S. A. et al. Superoxide, neuroleptics and the ubiquinone and cytochrome b5 reductases in brain and lymphocytes from normal and schizophrenic patients. Mol. Psychiatry 3, 227–-237 (1998).
Casademont, J. et al. Neuroleptic treatment effect on mitochondrial electron transport chain: peripheral blood mononuclear cells analysis in psychotic patients. J. Clin. Psychopharmacol. 27, 284–288 (2007).
Bergman, O. & Ben-Shachar, D. Mitochondrial oxidative phosphorylation system (OXPHOS) deficits in schizophrenia: possible interactions with cellular processes. Can. J. Psychiatry 61, 457–469 (2016).
Cheung, V. G. et al. Natural variation in human gene expression assessed in lymphoblastoid cells. Nat. Genet. 33, 422–425 (2003).
Dixon, A. L. et al. A genome-wide association study of global gene expression. Nat. Genet. 39, 1202–1207 (2007).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008).
Ioannidis, J. P. et al. Repeatability of published microarray gene expression analyses. Nat. Genet. 41, 149–155 (2009).
van Heerden, J. H. et al. Parallel changes in gene expression in peripheral blood mononuclear cells and the brain after maternal separation in the mouse. BMC Res. Notes 2, 195 (2009).
Acknowledgements
We thank Winton Moy for RNA-seq preparation, Deli He and Jessica Freda for technical support on cell culture. We thank the study participants of MGS. MGS was mainly supported by R01MH059571, R01MH081800, and U01MH079469 (to P.V.G.); and other NIH grants for other MGS sites (R01MH067257 to N.G.B., R01MH059588 to B.J.M., R01MH059565 to R.F., R01MH059587 to F.A., R01MH060870 to W.F.B., R01MH059566 to D.W.B., R01MH059586 to J.M.S., R01MH061675 to D.F.L., R01MH060879 to C.R.C., U01MH046276 to C.R.C., and U01MH079470 to D.F.L). The transcriptomic work was primarily supported by RC2MH090030 (to A.R.S.), R01MH094116 (to H.H.H.G.), and R01MH094091 (to P.V.G.). This work was also supported by NorthShore University HealthSystem Research Career Development Award (to J.D.) and R01MH106575 (to J.D.) and conducted in part in facilities constructed under the support of NIH grant 1C06RR020547.
MGS
P. V. Gejman6, A. R. Sanders6, J. Duan6, D. F. Levinson7, J. Shi8, N. G. Buccola9, B. J. Mowry10, R. Freedman11, A. Olincy11, F. Amin12, D. W. Black13, J. M. Silverman14, W. F. Byerley15, C. R. Cloninger16, D. M. Svrakic16
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Members of the Molecular Genetics of Schizophrenia (MGS) Collaboration are listed below Acknowledgements.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kos, M.Z., Duan, J., Sanders, A.R. et al. Dopamine perturbation of gene co-expression networks reveals differential response in schizophrenia for translational machinery. Transl Psychiatry 8, 278 (2018). https://doi.org/10.1038/s41398-018-0325-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-018-0325-1