Abstract
The ε4 allele of Apolipoprotein E (APOE) is the strongest known genetic risk factor of Alzheimer’s disease (AD) but does not account for the entirety of genetic risk. Genetic risk scores (GRSs) incorporating additional genetic variants have been developed to determine the genetic risk for AD, yet there is no systematic review assessing the contribution of GRSs for AD beyond the effect of APOE ε4. The purpose of this systematic PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-analyses)-based review was to summarize original research studies that have developed and validated a GRS for AD utilizing associated single nucleotide polymorphisms (SNPs). The PubMed and Web of Science databases were searched on April 6, 2018 and screening was completed on 2018 citations by two independent reviewers. Eighteen studies published between 2010 and 2018 were included in the review. All GRSs expressed significant associations or discrimination capability of AD when compared to clinically normal controls; however, GRS prediction of MCI to AD conversion was mixed. APOE ε4 status was more predictive of AD than the GRSs, although the GRSs did add to AD prediction accuracy beyond APOE ε4. GRSs might contribute to identifying genetic risk of AD beyond APOE. However, additional studies are warranted to assess the performance of GRSs in independent longitudinal cohorts.
Similar content being viewed by others
Introduction
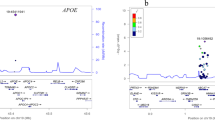
Alzheimer’s disease (AD) is the most common form of dementia and is a critical public health issue across the globe1. The etiology of the disease is thought to be a complex interaction between genes, environmental and lifestyle factors2. The heritability of late-onset AD has been estimated around 74%3. The strongest known genetic risk factor for AD is the ε4 allele of Apolipoprotein E (APOE ε4), but large-scale genome-wide association studies (GWASs) have identified additional genetic loci associated with AD4,5,6,7.
The largest GWAS meta-analysis concerning AD to date (N = 74,046), The International Genomics of Alzheimer’s Project (IGAP), has confirmed at least 20 genetic loci in addition to APOE genotype to be associated with AD8. The IGAP is a large two-stage study based upon GWASs on individuals of European ancestry. In stage 1, IGAP used genotyped and imputed data on 7,055,881 single nucleotide polymorphisms (SNPs) to meta-analyze four previously-published GWAS datasets consisting of 17,008 AD cases and 37,154 controls. In stage 2, 11,632 SNPs were genotyped and tested for association in an independent set of 8572 AD cases and 11,312 controls. Finally, a meta-analysis was performed combining results from stages 1 and 28.
IGAP consortia samples have greatly contributed to advancing genetic risk scores (GRSs) for AD, a strategy developed to deal with the relatively small magnitudes of association of the additional genetic loci for AD. GRSs determine the genetic risk for a disease through the composite consideration of many individual effects of genetic loci, which when considered collectively could account for substantial differences in risk of disease9. Thus, GRSs might present an effective strategy to combine the relatively smaller effects of AD associated loci to assess genetic risk beyond APOE ε4 status. However, the predictive value and methodologies of GRSs vary greatly between studies. For example, Escott-Price et al. analyzed more than 200,000 SNPs, including APOE resulting in an area under the curve (AUC) value of 0.8410, while Tosto et al. used only 21 SNPs excluding APOE and reported an AUC of 0.574.
Assessing the genetic contribution of GRSs to AD is of importance to better identify those with a higher susceptibility to AD and, eventually, enable targeted prevention strategies. To date there is no systematic review assessing GRSs for AD available. The aim of this literature review was to summarize original research studies that have developed and validated a GRS for AD utilizing associated SNPs.
Methods
The literature review was planned and performed using methods specified in the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement for reporting systematic reviews and meta-analyses11. Searches were completed in the PubMed and Web of Science databases (see Supplementary Table 1 for search strategies) on April 6, 2018 following the inclusion criteria: (1) presence of the evaluation of a defined GRS for AD incorporating genetic variants (specifically SNPs); (2) SNPs in GRS directly associated to AD; (3) AD diagnosis as the main outcome; (4) adult population of European descent; and (5) English or German language manuscripts. Specifically excluded were studies with all-cause dementia as an outcome where AD could not be specified as the outcome of interest. Searches were not limited to a specific time period. Based on the eligibility criteria, two reviewers (HS, TM) independently performed the study selection and in case of discrepancy discussion and further review of the issue followed in consultation with a third reviewer (LP).
Data extraction
The reviewers (HS and TM) extracted the following data from the included articles: (1) type of study; (2) validation data set information (study name, sample size, case number, mean age & sex distribution); (3) training data set information (study name, sample size & case number); (4) number of SNPs in the GRS; (5) type of GRS used (weighted or unweighted); (6) association between GRS and AD diagnosis; (7) the covariates considered; and (8) whether APOE was included in GRS. Additionally, information regarding the specific SNPs used in each of the GRSs was extracted including the name, location, gene, and association (odds ratio (OR) or log hazard ratio (HR)).
Quality assessment
The quality of the included studies was assessed independently by two reviewers (HS & TM) through an adapted version of the Newcastle-Ottawa scale (NOS), which assesses the quality of non-randomized studies based on three main categories: (1) the selection of the study groups; (2) the comparability of the groups; and (3) the ascertainment of either the exposure or outcome of interest. This tool was chosen because of the type of studies included12. The assessment tool was adapted to best fit the included studies based upon our inclusion criteria, where the exposure was genetic risk, the outcome of interest was AD diagnosis and the important covariates were age, sex and APOE e4 status. A coding manual was developed to ensure consistency and understanding of assessment. A point was awarded in each of nine categories if the study met the outlined criteria13.
Results
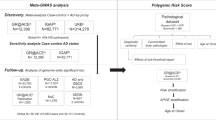
The initial database searches identified 1372 articles from Web of Science and 646 articles from PubMed resulting in a total of 2018 articles. Of the 1638 articles that remained after duplicates were removed (n = 380), 1592 were excluded because of irrelevance to the topic (Fig. 1). Strict inclusion criteria, as outlined above, were applied to the full text of 46 articles. Of these, 18 met the full set of criteria (Table 1)4,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29. All articles were published between 2010 and 2018. PRISMA guidelines were followed throughout the review and reporting process, please see Supplementary Table 2 for a completed PRISMA checklist.
Study characteristics
An overview of the study characteristics can be seen in Table 1. The majority of the studies used a case-control study design comparing AD cases to clinically normal controls4,10,14,17,20,21,22,23,27,28, the remaining studies utilized a longitudinal cohort19,24,25,29,30,31, nested case-control15 or cross-sectional design26 (Table 1). All training samples included individuals of European descent and ranged from 19220 to 74,046 (IGAP meta-analysis)8 individuals. The validation samples were also of European descent and ranged from 20426 to 19,68719 individuals. The majority of studies used IGAP consortia samples for the training and validation sets (Table 1). The selection of SNPs and corresponding magnitudes of associations were derived from a training set while the resulting GRS was assessed in a validation set. Six studies used data sets not associated with IGAP for GRS validation20,23,24,25,27,31. Three studies used a validation sample or group of participants of European descent with a family member afflicted with AD and therefore were not completely representative of a general population of European descent4,14,28. While the majority of included studies compared clinically normal controls to AD participants, two studies examined the ability of a GRS to predict the transition from mild cognitive impairment (MCI) to AD24,25, and one study examined both31.
AD participants within the training sets of the included studies met National Institute of Neurological and Communicative Disorders and Stroke and Alzheimer’s Disease and Related Disorders Association (NINCDS/ADRDA) criteria for probable AD, were autopsy confirmed, or met consensus criteria for AD32. Similarly in all validation sets, AD participants met NINCDS/ADRDA criteria or were confirmed through autopsy with the exception of two studies20,24.
GRS Construction
All included studies developed and validated a defined GRS for AD comprised of varying AD associated SNPs. The number of SNPs included in the GRSs ranged from 522 to 359,50017 (Table 2). SNP inclusion in the GRSs was based on two approaches: (1) selection from genome-wide significant results of previous GWAS (mainly IGAP meta-analysis)4,14,19,20,21,22,24,25,27,28,29,30,31 or (2) following p-value cutoffs including many SNPs10,15,17,23,26 (Table 2). The specific SNPs used in the included studies can be found in Supplementary Table 3 with information regarding location and associated gene. All studies utilized a weighted GRS as outlined by Purcell et al.9. Finally, APOE was either considered as a covariate4,14,19,22,24,25,28,31, included in the GRS10,15,17,20,21,26,27, not included and not considered as a covariate23,29,30.
GRS and AD association
Clinically normal to AD comparison or transition
The GRSs were found to be predictive of AD status or of AD conversion in all included studies, although varied magnitudes of association or discrimination abilities were found. Eight studies measured the disease prediction accuracy of the GRS using the area under the receiver operating characteristic curve4,16,17,20,21,23,27,28. Of which, five GRSs included APOE with an AUC range: 0.62–0.8410,17,20,21,27 and four GRSs excluded APOE with an AUC range: 0.57–0.754,21,23,28. Five studies used time-to-event analysis to evaluate the risk for developing AD15,19,29,30,31. Of those studies, one study included APOE in the GRS and reported a 3.34 fold increased risk of AD in individuals in the 10th decile of the GRS compared to the 1st decile 15 and the remaining four studies did not include APOE in the GRS with HR range: 1.11 (per SD) – 2.36 (84–16 percentile)19,29,30,31. Seven studies expressed statistically significant associations between the GRS and AD with odds ratios, mainly (n = 4) per standard deviation (SD) increase in GRS. Only two GRSs included APOE (OR range: 2.06– 2.32)21,26, while the remaining five GRSs excluded APOE (OR range: 1.14–2.85)4,14,22,23,28. Four studies which reported ORs also reported AUC values and were included in the description above4,21,23,28 (Table 2). For more detailed information including specific covariates considered in each GRS, please see Table 2.
The ability of the GRS in addition to APOE ε4 status to determine AD was investigated in many of the included studies. Possessing one or more APOE ε4 allele expressed greater discrimination ability than the GRSs (which excluded APOE); however, including APOE in the GRS increased AD prediction accuracy (Table 2).
Mild cognitive impairment to AD conversion
One study expressed a statistically significant result in the prediction of AD conversion from MCI, when comparing the 84th to 16th percentile (HR: 1.17, 95%CI: 1.02–1.35)31. The remaining two studies that examined the ability of the GRS to predict MCI conversion to AD did not express a statistically significant result24,25. Rodriguez-Rodriguez et al. reported that the GRS was not significantly associated with risk of AD conversion when comparing the 3rd to the 1st tertile of the GRS (OR: 1.32, 95%CI: 0.57–3.06). The hazard model from Lacour et al. also lacked a significant result (HR: 1.18, 95%CI: 0.37–2.0). Nevertheless, APOE ε4 status was predictive of MCI to AD conversion (Table 2).
Quality assessment
The results of the quality assessment using the adapted NOS are shown in Supplementary Table 4. Included studies were of high quality with a mean score of 7.2 stars (maximum 9) and a range of 5–8 stars. All case-control studies included adequate case and control definitions4,10,17,20,21,22,23,27,28, the vast majority included used representative samples10,15,17,19,21,22,23,24,25,26,28,29,30,31, and controlled for age, sex as well as accounted for APOE ε44,14,15,17,19,21,22,23,24,25,26,28,29,30,31. All included studies attained adequate and appropriate measure of the exposure (genetic risk) and outcome (AD diagnosis).
Discussion
This systematic review outlined and compared the existing GRSs for AD and found that the available GRSs resulted in statistically significant associations or disease prediction accuracy of AD when compared to clinically normal adults. However, results were mixed in predicting MCI to AD conversion and the GRSs were less predictive of AD than APOE ε4 status. Nevertheless they still contributed to disease prediction accuracy beyond APOE ε4.
Evolution of the GRS (clinically normal to AD)
Since 2010 GRSs for AD have advanced to include a higher number of SNPs, longitudinal assessment, pathological diagnosis, and have witnessed an increased rate of development after the publication of the IGAP meta-analysis in 2013. In three of the more recent studies, liberal GRSs (including thousands of SNPs associated to AD) were applied in addition to a conservative GRS (including only the few genome-wide significant SNPs)10,17,23. Conservative GRSs have been the main approach since the development of GRSs for AD, but this may begin to shift. This is evident when comparing the first GRS for AD, which included five SNPs22 to one of the most recent GRSs, which included 205,068 SNPs10. The liberal GRSs have illustrated greater disease prediction accuracy (AUC range: 0.75–0.84)10,17 than the conservative GRSs (AUC range: 0.57–0.72)4,17,20,21,28, suggesting that the conservative approach may be too cautious and that a more liberal method may increase disease prediction accuracy. However, an extremely liberal approach, including all SNPs with p-value < 0.510,17, may also have led to inclusion of many noninformative SNPs, and even better prediction accuracy might be achieved with an intermediate approach (not too restrictive but also not too liberal criteria). This has been demonstrated by two studies that have reported an increase in the ability of a GRS (also based on IGAP data) to differentiate between clinically normal controls and AD cases when including all SNPs p-value < 0.01 or < 0.001 compared to more conservative inclusion, but that after these critical points, discrimination ability plateaued and decreased23,33. It is important to note however that these studies used small validation sets, therefore warranting additional confirmation in larger sample sizes in future studies.
Also, GRSs have evolved to validation within a longitudinal study-design in addition to the previous case-control design. In order to confirm the ability of the GRS to predict AD diagnosis, the use of a longitudinal cohort is superior to a case-control study design due to the progressive nature and age dependence of the disease15. Five of the most recent studies examined the ability of the GRS to predict AD from clinically normal individuals at baseline or as a comparison and were published from 2016–201815,19,29,30,31. All studies reported significant results except one30. The main limitation of these studies is that both training and validation sets were a part of IGAP in all except one31. Additional longitudinal studies investigating the prediction capabilities of a GRS for AD in independent data sets are necessary to assess the plausibility of the GRS in genetic risk assessment.
Only two GRSs to date have been validated in a data set of exclusively pathologically confirmed AD cases10,27. Previous studies mainly utilized NINCDS/ADRDA criteria, which have been shown to have a sensitivity of 81% and specificity of 70% in determining AD34. Although the NINCDS/ADRDA criteria are widely used in research, autopsy confirmation of AD is the gold standard. Escott-Price et al. showed more accuracy in disease prediction in pathologically confirmed cases than in other validation sets without explicit autopsy confirmation, which points to possible AD misdiagnoses in NINCDS/ADRDA confirmed cases10. However this finding needs further replication.
Finally, before the IGAP meta-analysis was published only three studies had been published investigating the use of GRS for AD. Since publication, 15 GRS studies have been published, 11 of which have utilized the IGAP data for the training and validation sets (Supplementary Table 5). Overlap was present in 11 studies, of which only six studies discussed the overlap with five completing additional analysis excluding the overlapping individuals or statistically accounting for overfitting (Supplementary Table 5). The use of overlapping training and validation sets presents a source of possible overfitting. Ideally, completely independent data sets would be used. Although, the IGAP consortia meta-analysis has sparked exponential increase in GRS studies with an unparalleled resource of genetic information, it has also actualized a need for validation of GRSs in independent data sets.
Mild cognitive impairment to AD conversion
The GRS results were mixed in predicting AD conversion in participants with MCI24,25,31. The most recent study, Tan et al., reported a significant association when comparing the 84th to 16th percentile in a larger sample of more than 1650 individuals. Both Lacour et al. and Rodriguez-Rodriguez et al. reported non-significant associations; however, APOE ε4 status did predict AD conversion. Yet, case numbers and power were rather limited in both studies (790 and 118 cases, respectively). More studies are necessary to draw meaningful conclusions regarding the ability of the GRS to predict MCI to AD conversion.
Nonetheless, these results may suggest that other AD susceptibility loci (besides APOE) may not be predictors of AD conversion or have miniscule effects. It is also possible that some bias may exist due to the MCI participants that do not develop AD or develop another form of dementia24. Another viable explanation is the role of cognitive reserve and environmental factors in AD conversion35. Finally, the lack of association may have resulted from chance given the breadth of the confidence intervals.
GRS compared to APOE ε4
The predictive ability of APOE ε4 status to determine AD genetic risk has been well established with one copy and two copies of the APOE ε4 allele resulting in a 3-fold and 15-fold increase in risk respectively36. Although the GRSs in the included studies are significantly associated with AD diagnosis, it is important to investigate whether a GRS adds to genetic risk stratification above and beyond APOE ε4.
The disease prediction accuracy of the GRS (excluding APOE) was worse than APOE ε4 status. However, when the GRS included APOE it did increase the diagnostic accuracy compared to APOE ε4 status alone. The best discrimination ability was seen in GRSs that used a large number of SNPs including those in and around the APOE locus10. It has been estimated that APOE ε4 accounts for only 7% of the 65% total potentially non-modifiable risk factors of AD, suggesting further genetic associations beyond APOE37.
Implications
GRSs for AD are not currently relevant in a clinical setting, but they have the potential for use as a genetic risk stratification tool in clinical trials as well as future therapeutic interventions. Genetic risk stratification has been used in recent years to individualize therapeutic approaches in several diseases including cancer38. In preventable diseases GRSs can help identify those at risk and target preventive strategies accordingly39. In the future, genetic risk assessment through a GRS for AD could be integral in personalized medicine regarding AD.
Recently, the National Institute on Aging and Alzheimer’s Association Research Framework has recommended a shift toward a biological definition and the use of biomarkers for in vivo Alzheimer’s diagnosis40. GRSs have also shown significant associations to Alzheimer biomarkers including beta amyloid, phosphorylated and total tau15,21,41,42, hippocampal and amygdala volume22,23,33,43,44, among others. The results are however mixed with some studies reporting non-significant associations between GRSs and beta amyloid and tau45,46. The relationship between genetic risk and biomarkers of AD can provide deep insights into disease pathology and overall risk. As the definition of Alzheimer’s shifts to a biological basis, the investigation of genetic risk prediction of AD biomarkers may become even more pertinent.
Strengths and limitations
There are several limitations to this review. First, the methods, including choice of SNPs, validation samples, and type of reported measure of association varied across the included studies making it difficult to directly compare results. Furthermore, we focused on GRSs based on and validated within datasets including individuals of European descent, limiting the generalizability of the GRSs described. The populations used in the included studies were also often recruited from clinical settings, which therefore might also limit generalizability. As previously mentioned the largest weakness is the overlap between the training and validation sets, that both used IGAP data (Supplementary Table 5).
The included studies did also exhibit many strengths. All studies used thorough genotyping techniques, clinical diagnoses of AD, and proper control selection (if applicable). Statistical methods and study designs were appropriate and several of the more recent studies utilized a longitudinal cohort design providing deeper insight into the relationship between GRS and AD diagnosis.
The information presented in this systematic review is to our knowledge the first analysis of the existing GRSs for AD, further contributing to the AD literature related to genetic risk. The PRISMA guidelines were followed to ensure a rigorous review, selection, and presentation of the included literature. Furthermore, the topic is very timely with most of the results published recently in a field where the identification of genetic risk will continue to be a critical task.
Conclusion
GRSs including AD associated SNPs seem to be a promising strategy to classify AD genetic risk above and beyond APOE ε4, but the ability to predict MCI to AD conversion remains unclear. However, further validation of the GRSs including liberal approaches (not restricted to SNPs reaching genome-wide significance) and population-based prospective studies are warranted to confirm the results obtained with IGAP data. Finally, risk stratification for AD may be further improved by combining APOE and GRS status with additional data, such as “environmental” risk factors (including lifestyle factors) or other biomarker data known to be associated with AD risk.
References
Van Cauwenberghe, C., Van Broeckhoven, C. & Sleegers, K. The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet. Med. 18, 421 (2016).
Gatz, M. et al. Role of genes and environments for explaining Alzheimer disease. Arch. Gen. Psychiatry 63, 168–174 (2006).
Gatz, M. et al. Heritability for Alzheimer’s disease: the study of dementia in Swedish twins. J. Gerontol. A. Biol. Sci. Med. Sci. 52, M117–M125 (1997).
Tosto, G. et al. Polygenic risk scores in familial Alzheimer disease. Neurology 88, 1180–1186 (2017).
Harold, D. et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat. Genet 41, 1088 (2009).
Naj, A. C. et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat. Genet 43, 436 (2011).
Seshadri, S. et al. Genome-wide analysis of genetic loci associated with Alzheimer disease. JAMA 303, 1832–1840 (2010).
Lambert, J.-C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet 45, 1452–1458 (2013).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Escott-Price, V., Myers, A. J., Huentelman, M., & Hardy, J. Polygenic risk score analysis of pathologically confirmed Alzheimer’s disease. Ann Neurol 82, 311–314 (2017).
Liberati, A. et al. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate health care interventions: explanation and elaboration. PLoS. Med. 6, e1000100 (2009).
Higgins J. P., Green S. Cochrane Handbook for Systematic Reviews of Interventions, Vol. 4 (John Wiley & Sons, New York, 2011).
The Newcastle-Ottawa Scale (NOS) for assessing the quality of nonrandomised studies in meta-analyses. http://www.ohri.ca/programs/clinical_epidemiology/oxford.htm, Accessed 10 Nov 2017.
Tosto, G. et al. The role of cardiovascular risk factors and stroke in familial Alzheimer disease. JAMA Neurol. 73, 1231–1237 (2016).
Desikan, R. S. et al. Genetic assessment of age-associated Alzheimer disease risk: Development and validation of a polygenic hazard score. PLoS. Med. 14, e1002258 (2017).
Escott-Price, V., Shoai, M., Pither, R., Williams, J. & Hardy, J. Polygenic score prediction captures nearly all common genetic risk for Alzheimer’s disease. Neurobiol. Aging 49, 214.e217–214.e211 (2017).
Escott-Price, V. et al. Common polygenic variation enhances risk prediction for Alzheimer’s disease. Brain 138, 3673–3684 (2015).
Limon-Sztencel, A. et al. The algorithm for Alzheimer risk assessment based on APOE promoter polymorphisms. Alzheimer’s Res Ther. 8, 19 (2016).
Chouraki, V. et al. Evaluation of a genetic risk score to improve risk prediction for Alzheimer’s disease. J. Alzheimers Dis: JAD 53, 921–932 (2016).
Yokoyama, J. S. et al. Decision tree analysis of genetic risk for clinically heterogeneous Alzheimer’s disease. Bmc. Neurol. 15, 47 (2015).
Sleegers, K. et al. A 22-single nucleotide polymorphism Alzheimer’s disease risk score correlates with family history, onset age, and cerebrospinal fluid Abeta42. Alzheimers Dement. 11, 1452–1460 (2015).
Biffi, A. et al. Genetic variation and neuroimaging measures in Alzheimer disease. Arch. Neurol. 67, 677–685 (2010).
Lupton, M. K. et al. The effect of increased genetic risk for Alzheimer’s disease on hippocampal and amygdala volume. Neurobiol. Aging 40, 68–77 (2016).
Lacour, A. et al. Genome-wide significant risk factors for Alzheimer’s disease: role in progression to dementia due to Alzheimer’s disease among subjects with mild cognitive impairment. Mol. Psychiatry 22, 153–160 (2017).
Rodriguez-Rodriguez, E. et al. Genetic risk score predicting accelerated progression from mild cognitive impairment to Alzheimer’s disease. J. Neural Transm. (Vienna) 120, 807–812 (2013).
Sabuncu, M. R. et al. The association between a polygenic Alzheimer score and cortical thickness in clinically normal subjects. Cereb. Cortex (New York, NY: 1991) 22, 2653–2661 (2012).
Chaudhury, S. et al. Polygenic risk score in postmortem diagnosed sporadic early-onset Alzheimer’s disease. Neurobiol. Aging 62, 244.e241–244.e248 (2018).
Cruchaga, C. et al. Polygenic risk score of sporadic late-onset Alzheimer’s disease reveals a shared architecture with the familial and early-onset forms. Alzheimers Dement. 14, 205–214 (2018).
Ahmad, S. et al. Disentangling the biological pathways involved in early features of Alzheimer’s disease in the Rotterdam Study. Alzheimers Dement 14, 848–857 (2018).
van der Lee, S. J. et al. The effect of APOE and other common genetic variants on the onset of Alzheimer’s disease and dementia: a community-based cohort study. Lancet Neurol 17, 434–444 (2018).
Tan, C. H. et al. Polygenic hazard score: an enrichment marker for Alzheimer’s associated amyloid and tau deposition. Acta Neuropathol. 135, 85–93 (2018).
Dubois, B. et al. Advancing research diagnostic criteria for Alzheimer’s disease: the IWG-2 criteria. Lancet Neurol. 13, 614–629 (2014).
Mormino, E. C. et al. Polygenic risk of Alzheimer disease is associated with early- and late-life processes. Neurology 87, 481–488 (2016).
McKhann, G. M. et al. The diagnosis of dementia due to Alzheimer’s disease: Recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 7, 263–269 (2011).
Allegri, R. F. et al. Role of cognitive reserve in progression from mild cognitive impairment to dementia. Dement. & Neuropsychol. 4, 28–34 (2010).
Liu, C.-C., Kanekiyo, T., Xu, H. & Bu, G. Apolipoprotein E and Alzheimer disease: risk, mechanisms and therapy. Nat. Rev. Neurol. 9, 106–118 (2013).
Livingston, G. et al. Dementia prevention, intervention, and care. Lancet 390, 2673–2734 (2017).
Nelson, H. D. et al. Risk assessment, genetic counseling, and genetic testing for BRCA-related cancer in women: a systematic review to update the US Preventive Services Task Force recommendation. Ann. Intern. Med. 160, 255–266 (2014).
Khera, A. V. et al. Genetic risk, adherence to a healthy lifestyle, and coronary disease. N. Engl. J. Med. 375, 2349–2358 (2016).
Jack, C. R. et al. NIA-AA Research Framework: Toward a biological definition of Alzheimer’s disease. Alzheimers Dement. 14, 535–562 (2018).
Schultz, S. A. et al. Cardiorespiratory fitness alters the influence of a polygenic risk score on biomarkers of AD. Neurology 88, 1650–1658 (2017).
Ge, T., Sabuncu, M. R., Smoller, J. W., Sperling, R. A., & Mormino, E. C. Dissociable influences of APOE epsilon4 and polygenic risk of AD dementia on amyloid and cognition. Neurology 90, e1605–e1612 (2018).
Foley, S. F. et al. Multimodal brain imaging reveals structural differences in Alzheimer’s disease polygenic risk carriers: a study in healthy young adults. Biol. Psychiatry 81, 154–161 (2017).
Chauhan, G. et al. Association of Alzheimer’s disease GWAS loci with MRI markers of brain aging. Neurobiol. Aging 36, 1765. e1767–1765. e1716 (2015).
Darst, B. F. et al. Pathway-specific polygenic risk scores as predictors of amyloid-β deposition and cognitive function in a sample at increased risk for Alzheimer’s disease. J. Alzheimers Dis. 55, 473–484 (2017).
Martiskainen, H. et al. Effects of Alzheimer’s disease-associated risk loci on cerebrospinal fluid biomarkers and disease progression: a polygenic risk score approach. J. Alzheimers Dis. 43, 565–573 (2015).
Acknowledgements
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors. H.S. is a PhD candidate supported by a scholarship awarded from the Klaus Tschira Foundation. The content of this review is solely the responsibility of the authors and does not necessarily represent the official views of the Klaus Tschira Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stocker, H., Möllers, T., Perna, L. et al. The genetic risk of Alzheimer’s disease beyond APOE ε4: systematic review of Alzheimer’s genetic risk scores. Transl Psychiatry 8, 166 (2018). https://doi.org/10.1038/s41398-018-0221-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-018-0221-8
This article is cited by
-
Death-associated protein kinase 1 as a therapeutic target for Alzheimer's disease
Translational Neurodegeneration (2024)
-
Polygenic effects on the risk of Alzheimer’s disease in the Japanese population
Alzheimer's Research & Therapy (2024)
-
Alzheimer’s polygenic risk scores, APOE, Alzheimer’s disease risk, and dementia-related blood biomarker levels in a population-based cohort study followed over 17 years
Alzheimer's Research & Therapy (2023)
-
Pathophysiological Aspects and Therapeutic Armamentarium of Alzheimer’s Disease: Recent Trends and Future Development
Cellular and Molecular Neurobiology (2023)
-
Capturing additional genetic risk from family history for improved polygenic risk prediction
Communications Biology (2022)