Abstract
Schizophrenia (SCZ) is a neuropsychiatric disorder with a complex genetic etiology. The redundancy of the gene networks underlying SCZ indicates that many gene combinations have the potential to cause a system dysfunction that can manifest as SCZ or a related neurodevelopmental disorder. Recent studies show that small non-coding microRNA (miRNA) and long non-coding RNA (lncRNA) are important factors in shaping these networks and are dynamically regulated by neuronal activation. We investigated the genome-wide transcription profiles of 46 human amygdala samples obtained from 22 SCZ patients and 24 healthy controls. Using RNA sequencing (RNA-seq), we determined lncRNA expression levels in all samples and generated miRNA profiles for 27 individuals (13 cases and 14 controls). Previous studies have identified differentially expressed miRNAs in SCZ, including miR-132, miR-212, and miR-34a/miR-34c. Here we report differential expression of a novel miRNA, miR1307, in SCZ. Notably, miR1307 maps to a locus previously associated with SCZ through GWAS. Additionally, one lncRNA that was overexpressed in SCZ, AC005009.2, also maps to a region previously associated with SCZ based on GWAS and overlapped SCZ-related genes. The results were replicated in a large independent data set of 254 dorsolateral prefrontal cortex samples from the CommonMind consortium. Taken together, these results suggest that miRNA and lncRNAs are important contributors to the pathogenesis of SCZ.
Similar content being viewed by others
Introduction
Schizophrenia (SCZ) is a chronic, severe disabling neurodevelopmental disorder that affects people worldwide. While the exact cause of the disorder remains unknown, it is believed to result from an interplay between genetic and environmental factors1, 2. Recent genetic studies have implicated multiple genomic variants in functional gene networks that individually confer modest risk to the disease rather than high-impact mutations in single genes with large effect sizes3,4,5,6. Maintaining optimal network homeostasis requires coordination by an elaborate intracellular network of molecular signal transduction systems, including microRNAs (miRNAs), which are involved in the regulation and expression of genetic networks. Therefore, in addition to gene-specific variants that may affect an individual gene or pathway manifesting as SCZ or a related neurodevelopmental disorder7, 8, miRNAs such as non-coding RNAs that have the potential to dysregulate larger networks of genes have been postulated to be causal factors in the pathogenesis of SCZ9. Two groups of miRNAs, including typical non-coding RNAs and long non-coding RNAs (lncRNAs), have been reported to be associated with SCZ as well as other psychiatric diseases7,10, 11.
The amygdala dysfunction has been implicated in SCZ12,13,14; anatomically it consists of two almond-shaped groups of nuclei located deep and medially within the temporal lobes of the human brain. Functionally, the amygdala is thought to play a primary role in the processing of memory, decision-making, and emotional reactions15. Studies in mouse models involving the amygdala suggest that overexpression of miRNAs can protect the animals against anxiety-associated responses16 and controls their fear response17. Research in human subjects also implicates the amygdala region in emotion processing and recognition of facial expressions18,19,20. To our knowledge, a genome-wide transcription study of non-coding RNAs in the amygdala of SCZ patients has not been previously reported. In this study, we used next-generation sequencing technologies, including RNA sequencing (RNA-seq) and small RNA-seq (SRNA-seq), to assess the expression levels of miRNAs and lncRNAs in postmortem tissue from 46 individuals, including 22 SCZ patients and 24 healthy controls, obtained from the Lieber Institute. Our replication set consisted of 254 dorsolateral prefrontal cortex (DLPFC) tissues obtained from the DLPFC. Data that were generated as part of the CommonMind Consortium21 were of European ancestry.
Materials and methods
RNA extraction and quality assessment
Total RNA including the miRNA portion from postmortem amygdala tissue was extracted from 100 mg of pulverized tissue with all prep DNA/RNA/miRNA Universal Kit (Qiagen). The RNA portion was purified with RNeasy Mini Spin columns and on-column DNase digestion by RNase-Free DNase Set (Qiagen). Total RNA yield was determined by Qubit RNA BR Assay Kit and Qubit Fluorometer (ThermoFisher Scientific). RNA quality was assessed with high-resolution capillary electrophoresis on an Agilent Bioanalyzer 2100 (Agilent Technologies). Approximately 300 ng of total RNA was applied to an RNA 6000 Nano LabChip without prior heating. An RIN, obtained from entire Agilent electrophoretic trace with the RIN software algorithm, was used for the assessment of RNA quality (scale 1–10, with 1 being the lowest and 10 being the highest RNA quality).
Small RNA library preparation and sequencing
The Illumina TruSeq® Small RNA kit was used to generate the small RNA-seq libraries according to the manufacturers’ instructions. Briefly, 3′ (RA3) and 5′ (RA5) adapters were sequentially ligated to Dicer-processed miRNAs. Adaptor-ligated small RNAs were reverse-transcribed and the resultant cDNAs PCR-amplified. The PCR products were size-separated on 2% agarose gels and the 22–30 nucleotide bands excised and purified. Purified products were quantified by qPCR before cluster generation on a cBot (Illumina). Twelve samples were multiplexed per lane and sequenced on an Illumina HiSeq2000at 1 × 36 or 1 × 50 bp.
mRNA library preparation and sequencing
RNA-seq libraries were constructed using illumina TruSeq RNA sample Prep Kit (RS-122-2001) TruSeq RNA Sample Prep Kit V2, Illumina Inc., San Diego, CA, 92122) following the manufacturers’ instructions. The poly-A containing mRNA molecules were purified from 300 to 500 ng DNAse-treated total RNA using oligo (dT) beads. Following the purification, the mRNA was fragmented using divalent cations under elevated temperature (94 degree) for 2 min. Under these conditions, the resultant fragment lengths ranged from 130 to 290 bp with a median length of 185 bp. Reverse transcriptase and random primers were used to generate the first-strand cDNA from the RNA fragments. Second-strand DNA was synthesized using DNA polymerase I and RNaseH. The resulting cDNAs were end-repaired using T4 DNA polymerase, T4 PNK, and Klenow DNA polymerase and a single “A” base added using Klenowexo (3′–5′ exo minus). Illumine PE adapters were ligated using T4 DNA ligase including a barcoding index for multiplexing. These products were then purified and enriched by PCR to create the final cDNA library for high-throughput sequencing on a Highseq2000. The concentration of the RNA-seq libraries was determined by Qubit (Invitrogen, CA) and qPCR, and the quality of the libraries assayed on a Lab ChipGX (Caliper, MA) HT DNA 1 K/12 K/Hi-sensitivity LabChip. The libraries were multiplexed and loaded on a flow cell for cluster generation on a cBot (Illumina). Illumina Real Time Analysis was used for image analysis and base calling. FASTQ files were generated using the BCL Converter (CASAVA v1.8.2).
RNA-seq/small RNA-seq alignment and differential expression test
Following sequencing we obtained an average read depth of 116 M reads per sample, ranging from 34 to 226 M reads across the 46 amygdala samples. Alignments were generated using the Genomic Short-read Nucleotide Alignment Program22 taking common SNPs (dbSNP137) into account during the alignments. The output SAM files were converted to BAM, sorted by index, and unpaired reads were filtered with SAMtools23. Following alignment and filtering an average of 103 M reads were obtained per sample with an average mapping/filtering rate of 89% (Supplementary Table 1). Differential expression tests were performed using the cuffdiff package in cufflink2.2.124 and the GTF file from GENCODE version 1925. The DLPFC RNA-seq data were processed using the same bioinformatics pipeline as previously described for the amygdala data. To reduce algorithm-specific false-positives and to adjust for clinical covariates, we also analyzed the data set using DEseq2 including age, gender, postmortem index (PMI), and manner of death as covariates in the model. Differentially expressed non-coding RNAs were identified from the intersect of the Cuffdiff and DEseq226 data sets.
The small RNA-seq data read depths ranged from 4.7 to 72 M with average 20 M reads per sample. Cutadapt1.8.1 was applied to trim the adaptors27 and the trimmed data were further aligned with Micro Read Fast Alignment Search Tool (MRFast)28. The average mapping rate after trimming was 75.1% with an average read depth of 16 M reads per sample.
Clinical sample selection and sample data QC
A detailed description of the clinical sample is provided in Supplementary Table 6. RNA-seq and small RNA data were generated on all 49 individuals listed in Supplementary Table 6. Following primary analysis of the RNA-seq data, three individuals (10451, 10678, and 10495) were excluded from further analysis because of outlying expression values. The five most highly differentially expressed genes (CCL3, CCL4, CCL8, CXCL10, and CXCL11) in the analysis were driven by these three controls, with extreme fragments per kilobase million (FPKM) values (~×100 higher than the other samples). The outliers may be related to the cause of death of these individuals, two (10451, 10678) died from “complications of smoke inhalation”, and the third (10495) from an accident with multiple head injuries. Of the remaining 46 cases and controls, all were of Caucasian ancestry, 40 were males, and 6 females. Average age at death was 38.4 years. As confounders are an important consideration in postmortem studies, we carried out non-parametric analyses to test for association between age, sex, and PMI with SCZ and gene expression level (FPKM) in the remaining 46 samples. No significant associations were found between FPKM or SCZ and the covariates. Following quality control of the small RNA-seq data, 19 of the 46 individuals with RNA-seq data were excluded from the analysis.
In validation data set DLPFC, RNA was isolated from 50 mg homogenized tissue in Trizol using the RNeasy kit based on the instructional protocol. The mean total RNA yield was 15.3 µg (±5.7). The RNA integrity number (RIN) was determined by fractionating RNA samples on the 6000 Nano chip (Agilent Technologies) on the Agilent 2100 Bioanalyzer. The mean RIN was 7.7 (±0.9), and the mean ratio of 260/280 was 2.0 (±0.02). Processing order was re-randomized prior to ribosomal RNA (rRNA) depletion. Briefly, rRNA was depleted from about 1 µg of total RNA using Ribo-Zero Magnetic Gold kit (Illumina/Epicenter Cat # MRZG12324) to enrich for polyadenylated coding RNA and non-coding RNA. The sequencing library was prepared using the TruSeq RNA Sample Preparation Kit v2 (RS-122–2001-48 reactions) in batches of 24 samples. A pool of 10 barcoded libraries was layered on a random selection of two of the eight lanes of the Illumina flow cell bridge amplified to ~250 million raw clusters. One-hundred base pair paired end reads were obtained on a HiSeq 2500. The sequence data were processed for primary analysis to generate QC values. Samples with a minimum of 50 million mapped reads (~25 million paired end reads) and less than 5% rRNA-aligned reads were retained for downstream analysis21.
Results
Replication of known SCZ-related miRNA from the small RNA-seq data
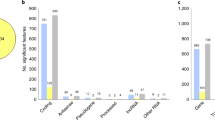
The small RNA-seq data set included 27 individuals (13 cases and 14 controls); differential expression analysis resulted in the identification of 17 statistically significant differentially expressed miRNAs (Table 1). Ten of the seventeen are novel discoveries, the remaining seven have previously been implicated in SCZ and/or other psychiatric phenotypes, including two members of miRNA family 34, mir-34c, -34a (p value = 3e−4 and 0.02, respectively) as well as a number of others including mirs-124,-132, -212, -663, and -144.
miRNA based on RNA-seq
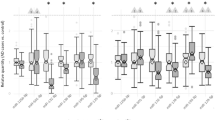
While RNA-seq is not designed to systematically detect expression changes of miRNA, some miRNAs that are affiliated with the RNA class are detectable by RNA-seq. Analysis of the GTF file as described in the Materials and methods section resulted in the identification of differential expression of MIR1307 between the SCZ patients and healthy controls. Expression of MIR1307 was approximately threefold higher in controls (p value = 0.0152). The expression difference and the direction of change were confirmed in the small RNA-seq data set, plotting the expression levels in cases and controls confirms that the association is not driven by outliers (Fig. 1a).
a There are 5 individuals in case has non-zero expression on MIR1307 and 11 individuals in controls. The mean of cases is 5.9 and 11 for controls. b For RP11-724N1.1, the mean gene expression in cases is 3.6 and in controls is 1.4; the overall expression is about three fold higher in SCZ cases. c The mean gene expression of RP11-677M14.2 is 86 in paranoid cases, which is highly driven by one outlier with expression 179; the mean gene expression in controls is 160. d The mean expression of AC005009.2 for undifferentiated cases is 23 and 51 in controls
LncRNAs based on RNA-seq
Differential analysis of the transcriptome data identified 250 lncRNAs from Gencode version 19 that showed significant expression differences between the amygdala regions of cases and controls (Supplementary Table 2). Among those, two lincRNAs RP11-724N1.1 (p value = 0.014) and RP11-677M14.2 (p value = 0.045) are of particular interest as they map to regions previously associated with SCZ based on genome-wide association study (GWAS) (Table 2). RP11-724N1.1 resides within the GWAS locus on chr10, previously shown to be associated with SCZ in multiple studies29, 30. RP11-677M14.2 is located within the SCZ-associated gene, neurogranin (NRGN)30, the human homolog of the neuron-specific rat RC3/neurogranin gene, which is a direct target for thyroid hormone in the human brain, and controls its expression.
Non-coding RNAs for SCZ subtypes
We further categorized the SCZ patients based on their clinical subtypes, including disorganized (seven individuals), paranoid (five individuals), and undifferentiated (nine individuals; Supplementary Table 7). The analysis revealed subtype specificity in non-coding RNA expression levels. MIR1307 showed 10-fold higher expression in controls when compared with the undifferentiated subtype in contrast to a two-fold increase when compared with the paranoid subtype. MIR1282 showed significantly higher expression in the paranoid subtype. We also identified subtype specificity in the expression of miRNAs previously reported to be associated with SCZ or other psychiatric diseases. For instance, the miRNA family 34 were only significant in the undifferentiated subtype and mir-144 was only associated with the disorganized subtype (Supplementary Table 4). For lncRNAs, RP11-677M14.2 was shown to be differentially expressed in the paranoid subtype (p value = 0.0064), whereas RP11-724N1.1 was differentially expressed in undifferentiated SCZ (p value = 0.016; Supplementary Table 5). Finally, some associations were restricted to a particular subtype such as lncRNA AC005009.2, which was not differentially expressed in the combined analysis but is significantly associated with the undifferentiated subtype (p value 0.0016).
Plotting the expression value distributions of the three lncRNAs differentially expressed by subtype (Supplementary Figs. 2a–e) indicates that the results are not driven by outliers. To determine the effects of variables such as age, gender, and manner of death (Supplementary Fig. 3), we used expression values of target lncRNAs as dependent variable (y) and age, gender, and mode of death as independent variable (x). For RP11-724N1.1, the p value of linear regression is 0.312, 0.973, and 0.504 for age of death, gender, and manner of death. For RP11-677M14.2 the p values are 0.2735, 0.3565, and 0.0954, and for AC005009.2 the p values are 0.0429, 0.7158, and 0.7331, respectively. Therefore, the expression level differences of subtypes are not driven by the biological parameters at a significance level of p ≤ 0.01. The entire distribution difference p values of expressed lncRNAs based on Wilcoxon rank-sum between undifferentiated, paranoid, and disorganized are 0.056, 0.15, and 0.89. The statistic results indicate that the lncRNA distribution between undifferentiated and paranoid are close to significant difference, and there are certain level differences between paranoid and disorganized, but the differences between undifferentiated and disorganized are insignificant. As a result, it suggests that paranoid SCZ has unique lncRNA patterns compared to other subtypes.
The DESeq2 results of the three lncRNAs were highly concordant with the cuffdiff outputs, expression fold changes were consistent, and almost identical between two tools (Supplementary Table 8). The boxplot of those three lncRNAs showed that the expression differences are not driven by outliers (Figs. 1b–d).
Replication/validation from independent data set
With limited availability of postmortem amygdala tissue from SCZ patients and no previously reported human amygdala RNA-seq studies available in the public domain, we attempted replication of our findings in DLPFC21, an independent data set from the CommonMind Consortium. RNA-seq was carried out in the DLPFC of totally 254 individuals (120 SCZ and 134 controls) of European ancestry. We applied the same bioinformatics analysis pipeline to the DLPFC data as previously described for the amygdala data to validate our lncRNA/miRNA findings. Adequate coverage was obtained for lncRNA AC005009.2 in the replication set but not for the rest three non-coding RNAs. The read depth of DLPFC data set is about half that of the amygdala data at 60~70 M reads vs. 150 M reads. As shown in Table 3, AC005009.2 replicated with a significant p value, and MIR1307 showed a clear trend toward association, with the same direction of effect, in the DLPFC data (Table 3).
Discussion
SCZ is a complex polygenic disease. Multiple risk genes have already been identified through GWAS and sequencing studies30, 31. To date, no variants, common or rare, conferring large effect sizes have been identified in SCZ. Rather a picture is emerging from the genetic studies implicating the dysregulation of complex gene networks and regulatory mechanisms. Growing evidence indicates that distinct neuronal ncRNAs, particularly miRNAs and lncRNAs, are likely to influence the development of psychiatric diseases, including SCZ. For instance, the lncRNA, Gomafu, is acutely regulated in response to neuronal activation and is involved in SCZ-associated alternative splicing. In this study, we detected altered expression of miRNAs and lncRNAs, well defined in GENCODE annotation, by RNA-seq and SRNA-seq of amygdala tissue samples from 46 subjects including SCZ subjects and controls.
Amygdala dysfunction has been extensively reported in SCZ, yet to our knowledge, gene expression profiling has not been previously reported on human amygdala tissue. Previous studies in mice show that overexpression of miRNA 34 genes in amygdala is associated with anxiety and fear. Overexpression of mir-34c in the amygdala of mice has been shown to induce anxiolytic behavior after challenge16; in our data set, the expression of mir-34c was twofold higher in the SCZ group compared to the control. Mir-34a has been implicated in amygdala-dependent fear memory consolidation in mice through Notch signaling17 and in our data expression of mir-34a was significantly upregulated in cases. Mir-34a has been shown to be differentially expressed in other brain regions, including the prefrontal cortex32, leading to the suggestion that mir-34a levels in blood could serve as a biomarker for SCZ33, 34. In our study, we identify that the 34 family is only differentially expressed in the individuals with “undifferentiated” subtype of SCZ. Another example is mir-132, a well-known miRNA involved with circadian rhythm and previously associated with multiple psychiatric traits in mouse models35 and with network regulation in major depressive disorder in human prefrontal cortex tissue36. Mir-132 is also differentially expressed in human prefrontal cortex tissue in miRNA-profiling data32 and is a potential biomarker in human blood33. In our study, we show that mir-132 is differentially expressed across all of the clinical SCZ subtypes. Another miRNA of interest, mir-663, is a known regulator of neuronal differentiation37. Finally, mir-144 is expressed in the hippocampus of rats and is known to be influenced by both valproate and the mood stabilizer lithium38; mir-144 is found to be only differentially expressed in individuals categorized as “disorganized” SCZ.
In addition to the seven previously reported miRNAs, we further identified 10 miRNAs that will require validation in future human or mouse studies. We also identified over 200 differentially expressed lncRNAs in this study. In order to prioritize the lncRNAs, we focused on differentially expressed lncRNAs that mapped to previously reported GWAS loci. Two lncRNAs, RP11-724N1.1 and RP11-677M14.2, mapped to GWAS loci at chr10q24.3 and chr11q24.2, respectively. RP11-724N1.1 maps to the third most significantly associated region from the PGC2 SCZ meta-analysis (rs11191419, p value = 6.29 × 10−19)39. Similar to miRNAs, RP11-724N1 is also clinical subtype-specific; it is only differentially expressed in undifferentiated SCZ subtype. RP11-724N1.1 overlaps the CNNM2 gene, as shown in Supplementary Fig. 1a. CNNM2 encodes a brain-expressed transmembrane protein that is involved in magnesium transport40. Loss of function mutations in CNNM2 has recently been reported to be causal of hypomagnesemia, seizures, and mental retardation41. RP11-677M14.2 maps to the antisense strand overlapping the NRGN gene (Supplementary Fig. 1b). The NRGN locus has been shown to be associated with SCZ in the PGC2 meta-analysis (rs55661361, p value = 3.68E−12), as well as in multiple other studies that reported association of functional coding variants in NRGN with SCZ39, 42,43,44,45. Post-transcriptional regulation of NRGN through overlapping sense and antisense transcripts has previously been reported during cerebral corticogenesis and synapse function in mice46. We can confirm through our RNA-seq expression data (Supplementary Table 3) that the expression of NRGN is strikingly higher (FPKM = 1251) in the SCZ group compared to controls (FPKM = 0). Subtype analysis further indicated that RP11-677M14.2 under expression was restricted to cases with a clinical diagnosis of paranoid SCZ.
A subtype-specific lncRNA, AC005009.2, has been identified for undifferentiated SCZ individuals; the lncRNA shows twofold higher expressions in control compared to SCZ patients who were diagnosed as the “undifferentiated” subtype. AC005009.2 is also close to GWAS region of PGC2 SCZ meta-analysis (rs12704290, p value = 1.04 × 10−10). Meanwhile, AC005009.2 overlaps the transcript of the metabotropic glutamate receptor 3 gene (GRM3; Supplementary Fig. 5c). GRM3 has been shown to associate with SCZ risks in previous studies47,48,49, and AC005009.2 is only differentially expressed in undifferentiated SCZ subtype with p value = 0.0046 and 1.5-fold higher expression in controls.
MIR1307 also maps to a highly significant SCZ GWAS locus (rs11191419, p value = 6.198e−19) in the PGC2 meta-analysis30. The associated locus contains multiple genes; however, we only detected differential expression in our RNA-seq data set from MIR1307 (Table 4). The expression data raise the possibility that the GWAS association is being driven by MIR1307. Further work will be required to assess the association between the SCZ-associated GWAS variants and MIR1307 expression levels in the brain.
To validate and replicate our findings in the absence of an independent human amygdala RNA-seq data set, we turned to a much larger independent data set of 254 prefrontal cortex tissues. The involvement of the DLPFC in SCZ is well established with multiple reports of association of DLPFC dysfunction correlating with clinical phenotype50, 51. AC005009.2 replicated with a significant p value and MIR1307 showed a clear trend toward association, with the same direction of effect, in the DLPFC data. The two other lncRNAs, RP11-724N1.1 and RP11-677M14.2, did not reach statistical significance in the DLFPC data, which may either be reflective of tissue specificity or simply due to the reduced coverage in the DLPFC data. The average read coverage in the DLPFC was 70 M reads compared with the 150 M reads from the amygdala data.
Finally, we note that several previously reported SCZ-associated miRNAs, including MIR137, 181b, 19, 219, and MIR9, did not replicate in this study. MIR137 and MIR181b were based on GWAS results rather than brain tissues, and MIR181b was reported in Chinese population rather than in Caucasian52,53,54. GWAS studies based on blood showed that MIR137 mediated dysregulation as a previously unknown etiologic mechanism in SCZ52. Our results showed that MIR137 expression is almost same in SCZ and controls, and MIR181b is 1.2-fold higher than SCZ. MIR19 was reported recently to have abnormal expression in neural progenitor cells from SCZ patient-derived induced pluripotent stem cells (iPSCs)55; in our data the expression of MIR19 was 1.2-fold higher in SCZ patients, but it did not reach significance (p value = 0.43). MIR219, a brain-specific miRNA, was reported in both normally developing and SCZ patient iPSC-derived neural stem cells56. In our amygdala data, the expression level of MIR219 was 1.4-fold higher in SCZ; however, it did not reach significance with a p value of 0.19. The association of those two miRNA was initially reported in SCZ NPCs but not in brain tissue, which may explain the lack of replication. MIR-9 was reported to be abundantly expressed in control neural progenitor cells (NPCs) and significantly downregulated in a subset of SCZ NPCs57. In Cuffdiff pipeline, the mir-9 was marked as region with too many fragments, so no p value was generated. In contrast, DESeq2 reports significant q value of mir-9 (0.02).
Conclusion
Differential expression analysis of small RNAs in SCZ brain identified multiple genes that may be contributing to the pathogenesis of the disease. Intersecting the list of differentially expressed small and lncRNAs with loci identified in previously reported GWAS studies has highlighted two genes, miRNA (mir1307) and lncRNAs (AC005009.2), that are differentially expressed in SCZ and as such may underlie or contribute to the observed GWAS signals at these loci. This study demonstrates the utility of transcriptional profiling of relevant disease tissues in identifying genes underlying GWAS signals.
References
Picchioni, M. M. & Murray, R. M. Schizophrenia. Br. Med. J. 335, 91–95 (2007).
van Os, J. & Kapur, S. Schizophrenia. Lancet 374, 635–645 (2009).
Harrison, P. J. & Weinberger, D. R. Schizophrenia genes, gene expression, and neuropathology: on the matter of their convergence. Mol. Psychiatry 10, 40–68 (2005). image 45.
Sun, J. et al. Schizophrenia gene networks and pathways and their applications for novel candidate gene selection. PLoS. ONE 5, e11351 (2010).
Talkowski, M. E. et al. A network of dopaminergic gene variations implicated as risk factors for schizophrenia. Hum. Mol. Genet. 17, 747–758 (2008).
O’Donovan, M. C., Craddock, N. J. & Owen, M. J. Genetics of psychosis; insights from views across the genome. Hum. Genet. 126, 3–12 (2009).
Beveridge, N. J. & Cairns, M. J. MicroRNA dysregulation in schizophrenia. Neurobiol. Dis. 46, 263–271 (2012).
Merico, D. et al. MicroRNA dysregulation, gene networks, and risk for schizophrenia in 22q11.2 deletion syndrome. Front. Neurol. 5, 238 (2014).
Mamdani, F. et al. Coding and noncoding gene expression biomarkers in mood disorders and schizophrenia. Dis. Markers 35, 11–21 (2013).
Kocerha J., Dwivedi Y., Brennand K. J. Noncoding RNAs and neurobehavioral mechanisms in psychiatric disease. Mol. Psychiatry 20, 677–684 (2015).
Barry, G. et al. The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing. Mol. Psychiatry 19, 486–494 (2014).
Lawrie, S. M., Whalley, H. C., Job, D. E. & Johnstone, E. C. Structural and functional abnormalities of the amygdala in schizophrenia. Ann. N. Y. Acad. Sci. 985, 445–460 (2003).
Mahon, P. B. et al. An MRI study of amygdala in schizophrenia and psychotic bipolar disorder. Schizophr. Res. 138, 188–191 (2012).
Rasetti, R. et al. Evidence that altered amygdala activity in schizophrenia is related to clinical state and not genetic risk. Am. J. Psychiatry 166, 216–225 (2009).
Amunts, K. et al. Cytoarchitectonic mapping of the human amygdala, hippocampal region and entorhinal cortex: intersubject variability and probability maps. Anat. Embryol. 210, 343–352 (2005).
Haramati, S. et al. MicroRNA as repressors of stress-induced anxiety: the case of amygdalar miR-34. J. Neurosci. 31, 14191–14203 (2011).
Dias, B. G. et al. Amygdala-dependent fear memory consolidation via miR-34a and Notch signaling. Neuron 83, 906–918 (2014).
Mier, D. et al. Evidence for altered amygdala activation in schizophrenia in an adaptive emotion recognition task. Psychiatry Res. 221, 195–203 (2014).
Pankow, A. et al. Altered amygdala activation in schizophrenia patients during emotion processing. Schizophr. Res. 150, 101–106 (2013).
Suslow, T. et al. Automatic amygdala response to facial expression in schizophrenia: initial hyperresponsivity followed by hyporesponsivity. BMC Neurosci. 14, 140 (2013).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Chen, C., Khaleel, S. S., Huang, H. & Wu, C. H. Software for pre-processing Illumina next-generation sequencing short read sequences. Source Code. Biol. Med. 9, 8 (2014).
Hach, F. et al. mrsFAST: a cache-oblivious algorithm for short-read mapping. Nat. Methods 7, 576–577 (2010).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Schizophrenia Working Group of the Psychiatric Genomics C. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Kim, A. H. et al. MicroRNA expression profiling in the prefrontal cortex of individuals affected with schizophrenia and bipolar disorders. Schizophr. Res. 124, 183–191 (2010).
Sun, X. Y. et al. A preliminary analysis of microRNA as potential clinical biomarker for schizophrenia. Am. J. Med. Genet. B 168, 170–178 (2015).
Lai, C. Y. et al. MicroRNA expression aberration as potential peripheral blood biomarkers for schizophrenia. PLoS. ONE 6, e21635 (2011).
Balu, D. T. et al. Multiple risk pathways for schizophrenia converge in serine racemase knockout mice, a mouse model of NMDA receptor hypofunction. Proc. Natl. Acad. Sci. USA 110, E2400–2409 (2013).
Smalheiser, N. R. et al. MicroRNA expression is down-regulated and reorganized in prefrontal cortex of depressed suicide subjects. PLoS. ONE 7, e33201 (2012).
Shu, R. et al. APP intracellular domain acts as a transcriptional regulator of miR-663 suppressing neuronal differentiation. Cell. Death Dis. 6, e1651 (2015).
Zhou, R. et al. Evidence for selective microRNAs and their effectors as common long-term targets for the actions of mood stabilizers. Neuropsychopharmacology 34, 1395–1405 (2009).
Stefansson, H. et al. Common variants conferring risk of schizophrenia. Nature 460, 744–747 (2009).
Goytain, A. & Quamme, G. A. Functional characterization of ACDP2 (ancient conserved domain protein), a divalent metal transporter. Physiol. Genom. 22, 382–389 (2005).
Arjona, F. J. et al. CNNM2 mutations cause impaired brain development and seizures in patients with hypomagnesemia. PLoS. Genet. 10, e1004267 (2014).
Ruano, D. et al. Association of the gene encoding neurogranin with schizophrenia in males. J. Psychiatr. Res. 42, 125–133 (2008).
Ohi, K. et al. Influence of the NRGN gene on intellectual ability in schizophrenia. J. Hum. Genet. 58, 700–705 (2013).
Walton, E. et al. The impact of genome-wide supported schizophrenia risk variants in the neurogranin gene on brain structure and function. PLoS. ONE 8, e76815 (2013).
Ohi, K. et al. Functional genetic variation at the NRGN gene and schizophrenia: evidence from a gene-based case-control study and gene expression analysis. Am. J. Med. Genet. B 159B, 405–413 (2012).
Ling, K. H. et al. Spatiotemporal regulation of multiple overlapping sense and novel natural antisense transcripts at the Nrgn and Camk2n1 gene loci during mouse cerebral corticogenesis. Cereb. Cortex 21, 683–697 (2011).
Egan, M. F. et al. Variation in GRM3 affects cognition, prefrontal glutamate, and risk for schizophrenia. Proc. Natl. Acad. Sci. USA 101, 12604–12609 (2004).
Mossner, R. et al. Further evidence for a functional role of the glutamate receptor gene GRM3 in schizophrenia. Eur. Neuropsychopharmacol. 18, 768–772 (2008).
Allen, N. C. et al. Systematic meta-analyses and field synopsis of genetic association studies in schizophrenia: the SzGene database. Nat. Genet. 40, 827–834 (2008).
Knable, M. B. & Weinberger, D. R. Dopamine, the prefrontal cortex and schizophrenia. J. Psychopharmacol. 11, 123–131 (1997).
Wible, C. G. et al. Prefrontal cortex, negative symptoms, and schizophrenia: an MRI study. Psychiatry Res. 108, 65–78 (2001).
Schizophrenia Psychiatric Genome-Wide Association Study C. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Silber, J. et al. miR-124 and miR-137 inhibit proliferation of glioblastoma multiforme cells and induce differentiation of brain tumor stem cells. BMC Med. 6, 14 (2008).
Xu, Y. et al. MicroRNAs and target site screening reveals a pre-microRNA-30e variant associated with schizophrenia. Schizophr. Res. 119, 219–227 (2010).
Han, J. et al. Functional implications of miR-19 in the migration of newborn neurons in the adult brain. Neuron 91, 79–89 (2016).
Murai, K. et al. The TLX-miR-219 cascade regulates neural stem cell proliferation in neurodevelopment and schizophrenia iPSC model. Nat. Commun. 7, 10965 (2016).
Topol, A. et al. Dysregulation of miRNA-9 in a subset of schizophrenia patient-derived neural progenitor cells. Cell. Rep. 15, 1024–1036 (2016).
Acknowledgments
We thank Dr. Joel E. Kleinman, Dr. Dan R. Weinberger, and Dr. Thomas M. Hyde from The Lieber Institute for Brain Development | LIBD, Baltimore, MD for contributing the samples. DLPFC data were generated as part of the CommonMind Consortium supported by funding from Takeda Pharmaceuticals Company Limited, F. Hoffman-La Roche Ltd, and from NIH grants R01MH085542, R01MH093725, P50MH066392, P50MH080405, R01MH097276, RO1-MH-075916, P50M096891, P50MH084053S1, R37MH057881, and R37MH057881S1, HHSN271201300031C, AG02219, AG05138, and MH06692. Brain tissue for the study was obtained from the following brain bank collections: the Mount Sinai NIH Brain and Tissue Repository, the University of Pennsylvania Alzheimer’s Disease Core Center, the University of Pittsburgh NeuroBioBank and Brain and Tissue Repositories, and the NIMH Human Brain Collection Core. CMC Leadership: Pamela Sklar, Joseph Buxbaum (Icahn School of Medicine at Mount Sinai), Bernie Devlin, David Lewis (University of Pittsburgh), Raquel Gur, Chang-Gyu Hahn (University of Pennsylvania), Keisuke Hirai, Hiroyoshi Toyoshiba (Takeda Pharmaceuticals Company Limited), Enrico Domenici, Laurent Essioux (F. Hoffman-La Roche Ltd), Lara Mangravite, Mette Peters (Sage Bionetworks), Thomas Lehner, and Barbara Lipska (NIMH).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethical approval
This study had been approved by the Children’s Hospital of Philadelphia with IRB# 4886. All the patients who participated in this project have been consented and they agree to publish the results.
Competing interests
The authors declare that they have no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Funding
This study is funded by an Institutional Development Award to the Center for Applied Genomics from The Children’s Hospital of Philadelphia; Adele and Daniel Kubert donation; MH096891-03S1 (NIMH); RC2 MH089924-02 (NIMH); R01 MH097284-03 (NIMH); and U01-HG008684 (NIH).
Data deposition
The data have been deposited to NCBI SRA site with SRA# 102186.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Y., Chang, X., Hahn, CG. et al. Non-coding RNA dysregulation in the amygdala region of schizophrenia patients contributes to the pathogenesis of the disease. Transl Psychiatry 8, 44 (2018). https://doi.org/10.1038/s41398-017-0030-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-017-0030-5
This article is cited by
-
Neuroinflammation driven by human immunodeficiency virus-1 (HIV-1) directs the expression of long noncoding RNA RP11-677M14.2 resulting in dysregulation of neurogranin in vivo and in vitro
Journal of Neuroinflammation (2024)
-
Significance of an altered lncRNA landscape in schizophrenia and cognition: clues from a case–control association study
European Archives of Psychiatry and Clinical Neuroscience (2023)
-
Dissecting the Relationship Between Neuropsychiatric and Neurodegenerative Disorders
Molecular Neurobiology (2023)
-
Focus on your locus with a massively parallel reporter assay
Journal of Neurodevelopmental Disorders (2022)
-
What genes are differentially expressed in individuals with schizophrenia? A systematic review
Molecular Psychiatry (2022)