Abstract
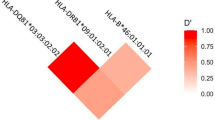
Antithyroid drug (ATD) is a mainstay of Graves’ disease (GD). About 0.1–0.5% of patients with GD treated with ATD exhibit ATD-induced agranulocytosis, which is characterized by severe reduction of circulating neutrophils. Immune-mediated responses have been proposed as a possible mechanism for the pathogenesis of ATD-induced agranulocytosis. Although it has been reported that the HLA class II allele (HLA-DRB1*08:03) was associated with ATD-induced agranulocytosis in multiple populations, the entire HLA region have not been explored in Japanese. Therefore, we performed HLA sequencing for 10 class I and 11 class II genes in 87 patients with ATD-induced agranulocytosis and 384 patients with GD who did not show ATD-induced agranulocytosis. By conducting case-control association studies at the HLA allele and haplotype levels, we replicated the association between HLA-DRB1*08:03:02 and ATD-induced agranulocytosis (P = 5.2 × 10−7, odds ratio = 2.80), and identified HLA-B*39:01:01 as an independent risk factor (P = 1.4 × 10−3, odds ratio = 3.35). To verify reproducibility of the novel association of HLA-B*39:01:01, we reanalyzed allele frequency data for HLA-B*39:01:01 from previous case-control association studies. The association of HLA-B*39:01:01 was significantly replicated in Chinese (P = 9.0 × 10−3), Taiwanese (P = 1.1 × 10−3), and European populations (P = 5.2 × 10−4). A meta-analysis combining results from the previous and current studies reinforced evidence of the association between HLA-B*39:01:01 and ATD-induced agranulocytosis (Pmeta = 1.2 × 10−9, pooled OR = 3.66, 95% CI; 2.41–5.57). The results of this study will provide a better understanding of the pathogenesis of ATD-induced agranulocytosis in the context of HLA-mediated hypersensitivity reactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Weetman AP. Graves’ disease. N Engl J Med. 2000;343:1236–48.
Cooper DS. Antithyroid drugs. N Engl J Med. 1984;311:1353–62.
Nakamura H, Miyauchi A, Miyawaki N, Imagawa J. Analysis of 754 cases of antithyroid drug-induced agranulocytosis over 30 years in Japan. J Clin Endocrinol Metab. 2013;98:4776–83.
Vicente N, Cardoso L, Barros L, Carrilho F. Antithyroid drug-induced agranulocytosis: state of the art on diagnosis and management. Drugs R D 2017;17:91–6.
Chen PL, Shih SR, Wang PW, Lin YC, Chu CC, Lin JH, et al. Genetic determinants of antithyroid drug-induced agranulocytosis by human leukocyte antigen genotyping and genome-wide association study. Nat Commun. 2015;6:7633.
Cheung CL, Sing CW, Tang CS, Cheng VK, Pirmohamed M, Choi CH, et al. HLA-B*38:02:01 predicts carbimazole/methimazole-induced agranulocytosis. Clin Pharm Ther. 2016;99:555–61.
He Y, Zheng J, Zhang Q, Hou P, Zhu F, Yang J, et al. Association of HLA-B and HLA-DRB1 polymorphisms with antithyroid drug-induced agranulocytosis in a Han population from northern China. Sci Rep. 2017;7:11950.
Thao MP, Tuan PVA, Linh LGH, Van Hoang L, Hen PH, Hoa LT, et al. Association of HLA-B∗38:02 with antithyroid drug-induced agranulocytosis in Kinh Vietnamese patients. Int J Endocrinol. 2018;2018:7965346.
Hallberg P, Eriksson N, Ibañez L, Bondon-Guitton E, Kreutz R, Carvajal A, et al. Genetic variants associated with antithyroid drug-induced agranulocytosis: a genome-wide association study in a European population. Lancet Diabetes Endocrinol. 2016;4:507–16.
Chen WT, Chi CC. Associations of HLA genotypes with antithyroid drug-induced agranulocytosis: a systematic review and meta-analysis of pharmacogenomics studies. Br J Clin Pharmacol. 2019;85:1878–87.
Tamai H, Sudo T, Kimura A, Mukuta T, Matsubayashi S, Kuma K, et al. Association between the DRB1*08032 histocompatibility antigen and methimazole-induced agranulocytosis in Japanese patients with Graves disease. Ann Intern Med. 1996;124:490–4.
Hosomichi K, Shiina T, Tajima A, Inoue I. The impact of next-generation sequencing technologies on HLA research. J Hum Genet. 2015;60:665–73.
Hirata J, Hosomichi K, Sakaue S, Kanai M, Nakaoka H, Ishigaki K, et al. Genetic and phenotypic landscape of the major histocompatibilty complex region in the Japanese population. Nat Genet. 2019;51:470–80.
Robinson J, Soormally AR, Hayhurst JD, Marsh SGE. The IPD-IMGT/HLA database—new developments in reporting HLA variation. Hum Immunol. 2016;77:233–7.
Hosomichi K, Jinam TA, Mitsunaga S, Nakaoka H, Inoue I. Phase-defined complete sequencing of the HLA genes by next-generation sequencing. BMC Genom. 2013;14:355.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler Transform. Bioinformatics. 2009;25:1754–60.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Browning BL, Browning SR. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am J Hum Genet. 2009;84:210–23.
Nakaoka H, Inoue I. Meta-analysis of genetic association studies: methodologies, between-study heterogeneity and winner’s curse. J Hum Genet. 2009;54:615–23.
Cochran WG. The combination of estimates from different experiments. Biometrics. 1954;10:101–29.
Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002;21:1539–58.
Balduzzi S, Rücker G, Schwarzer G. How to perform a meta-analysis with R: a practical tutorial. Evid Based Ment Health. 2019;22:153–60.
González-Galarza FF, Takeshita LY, Santos EJ, Kempson F, Maia MH, da Silva AL, et al. Allele frequency net 2015 update: new features for HLA epitopes, KIR and disease and HLA adverse drug reaction associations. Nucleic Acids Res. 2015;43:D784–8.
Ueda S, Oryoji D, Yamamoto K, Noh JY, Okamura K, Noda M, et al. Identification of independent susceptible and protective HLA alleles in Japanese autoimmune thyroid disease and their epistasis. J Clin Endocrinol Metab. 2014;99:E379–83.
Shin DH, Baek IC, Kim HJ, Choi EJ, Ahn M, Jung MH, et al. HLA alleles, especially amino-acid signatures of HLA-DPB1, might contribute to the molecular pathogenesis of early-onset autoimmune thyroid disease. PLoS ONE 2019;14:e0216941.
Nguyen DV, Vidal C, Chi HC, Do NTQ, Fulton R, Li J, et al. A novel multiplex polymerase chain reaction assay for detection of both HLA-A*31:01/HLA-B*15:02 alleles, which confer susceptibility to carbamazepine-induced severe cutaneous adverse reactions. HLA. 2017;90:335–42.
Wang Q, Sun S, Xie M, Zhao K, Li X, Zhao Z. Association between the HLA-B alleles and carbamazepine-induced SJS/TEN: A meta-analysis. Epilepsy Res. 2017;135:19–28.
Kim YJ, Kim HY, Lee JH, Yu SJ, Yoon JH, Lee HS, et al. A genome-wide association study identified new variants associated with the risk of chronic hepatitis B. Hum Mol Genet. 2013;22:4233–8.
Jiang DK, Ma XP, Yu H, Cao G, Ding DL, Chen H, et al. Genetic variants in five novel loci including CFB and CD40 predispose to chronic hepatitis B. Hepatology. 2015;62:118–28.
Shin JG, Cheong HS, Kim JY, Lee JH, Yu SJ, Yoon JH, et al. Identification of additional EHMT2 variant associated with the risk of chronic hepatitis B by GWAS follow-up study. Genes Immun. 2019;20:1–9.
Mallal S, Phillips E, Carosi G, Molina JM, Workman C, Tomazic J, et al. HLA-B*5701 screening for hypersensitivity to abacavir. N Engl J Med. 2008;358:568–79.
Redwood AJ, Pavlos RK, White KD, Phillips EJ. HLAs: key regulators of T-cell-mediated drug hypersensitivity. HLA. 2018;91:3–16.
Illing PT, Vivian JP, Dudek NL, Kostenko L, Chen Z, Bharadwaj M, et al. Immune self-reactivity triggered by drug-modified HLA-peptide repertoire. Nature. 2012;486:554–8.
Ostrov DA, Grant BJ, Pompeu YA, Sidney J, Harndahl M, Southwood S, et al. Drug hypersensitivity caused by alteration of the MHC-presented self-peptide repertoire. Proc Natl Acad Sci USA. 2012;109:9959–64.
Wei CY, Chung WH, Huang HW, Chen YT, Hung SI. Direct interaction between HLA-B and carbamazepine activates T cells in patients with Stevens-Johnson syndrome. J Allergy Clin Immunol. 2012;129:1562–9.e5.
Pan RY, Chu MT, Wang CW, Lee YS, Lemonnier F, Michels AW, et al. Identification of drug-specific public TCR driving severe cutaneous adverse reactions. Nat Commun. 2019;10:3569.
Yun J, Mattsson J, Schnyder K, Fontana S, Largiadèr CR, Pichler WJ, et al. Allopurinol hypersensitivity is primarily mediated by dose-dependent oxypurinol-specific T cell response. Clin Exp Allergy. 2013;43:1246–55.
Acknowledgements
We are grateful to Shigeru Shuto, Yoko Watanabe, and Shigeko Wakiya for their contributions to sample collection. We thank Junko Kajiwara, Junko Kitayama, and Yumiko Sato for their technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nakakura, S., Hosomichi, K., Uchino, S. et al. HLA-B*39:01:01 is a novel risk factor for antithyroid drug-induced agranulocytosis in Japanese population. Pharmacogenomics J 21, 94–101 (2021). https://doi.org/10.1038/s41397-020-00187-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-020-00187-4