Abstract
Bacterial growth often alters the environment, which in turn can impact interspecies interactions among bacteria. Here, we used an in vitro batch system containing mucin beads to emulate the dynamic host environment and to study its impact on the interactions between two abundant and prevalent human gut bacteria, the primary fermenter Bacteroides thetaiotaomicron and the butyrate producer Roseburia intestinalis. By combining machine learning and flow cytometry, we found that the number of viable B. thetaiotaomicron cells decreases with glucose consumption due to acid production, while R. intestinalis survives post-glucose depletion by entering a slow growth mode. Both species attach to mucin beads, but only viable cell counts of B. thetaiotaomicron increase significantly. The number of viable co-culture cells varies significantly over time compared to those of monocultures. A combination of targeted metabolomics and RNA-seq showed that the slow growth mode of R. intestinalis represents a diauxic shift towards acetate and lactate consumption, whereas B. thetaiotaomicron survives glucose depletion and low pH by foraging on mucin sugars. In addition, most of the mucin monosaccharides we tested inhibited the growth of R. intestinalis but not B. thetaiotaomicron. We encoded these causal relationships in a kinetic model, which reproduced the observed dynamics. In summary, we explored how R. intestinalis and B. thetaiotaomicron respond to nutrient scarcity and how this affects their dynamics. We highlight the importance of understanding bacterial metabolic strategies to effectively modulate microbial dynamics in changing conditions.
Similar content being viewed by others
Introduction
The human gut is populated by hundreds of microbial species interacting with each other and with their host. Gut bacterial species differ in their distribution along the human digestive tract and favor different microhabitats, such as the lumen of the large intestine, colonic crypts and mucus layers [1, 2]. Diverse growth rates and attachment properties of gut bacteria help establish spatially structured communities. Also, the spatially heterogeneous environment influences the colonization success of different members of the gut microbiota and their competitive or cooperative relationships with one another [3,4,5,6,7,8].
An efficient way to acquire nutrients is essential for microbial persistence in the human gut. In addition to dietary fiber in the lumen, host-derived mucin glycans provide an alternative source of nutrients for members of the gut microbiota [9]. The colonic mucus layer, located at the interface between the epithelium and the gut microbiota, is key to maintain intestinal homeostasis and to enhance the resilience to perturbations of the gut microbiome [10, 11]. Colon mucus is continuously secreted by goblet cells and is organized into two distinct layers: an inner dense layer tightly attached to the epithelium, which appears to be essentially sterile when healthy; and an outer loose layer with an expanded volume, which supports a metabolically diverse community. The outer layer provides mucin polysaccharides and serves as attachment sites for the gut species that co-evolved to utilize these substrates, giving rise to complex microbial interactions [9, 10, 12, 13]. Bacteroides is one of the most abundant but also most variable genera across human fecal samples [14]. Its most studied species Bacteroides thetaiotaomicron (B. thetaiotaomicron) holds a large repertoire of genes involved in sensing and hydrolyzing many different polysaccharides, and B. thetaiotaomicron was reported to strongly express genes involved in the degradation of dietary plant glycans [15, 16]. As a nutritional generalist, B. thetaiotaomicron can alter its metabolic response to mucin glycans in the absence of dietary nutrients, to colonize the mucus layer and maintain its persistence in the human gut [17].
Short-chain fatty acids (SCFAs) including acetate, propionate and butyrate, are the major end products of the anaerobic fermentation of polysaccharides by the human gut microbiota. All these SCFAs exert multiple health benefits, in particular butyrate, which is used preferentially as an energy source to fuel colonic enterocytes and may trigger signals regulating mucin synthesis and secretion, as well as to maintain the integrity of epithelial barriers [18, 19]. Metabolic cross-feeding of the human gut microbiota impacts SCFA production [20]. For example, members of the most abundant genera, including Faecalibacterium, Eubacterium and Roseburia, are able to convert acetate and/or lactate into butyrate using the prevalent butyrate-producing pathway via butyryl-CoA:acetate CoA-transferase [21, 22]. Lactate is also an important end product of anaerobic fermentation in the gut [23]. In addition to acetate as a key intermediate that promotes cross-feeding interactions of the gut microbiota, the role of lactate as substrate for butyrate producers or for propionate producers via the acrylate pathway may explain its low levels of accumulation in the human colon of healthy adults [24, 25]. Upon exhaustion of the primary carbon sources such as carbohydrates in the gut, microbes have to hunt for another carbon source to thrive. It was reported that acetate could promote the growth of a common intestinal butyrate-producing species Roseburia intestinalis (R. intestinalis) during growth on glucose [26,27,28].
Several studies employing in vitro fermentation systems have shown mucin as a driving force in forming a microbial community that is phylogenetically and metabolically distinct from the luminal one [29, 30]. 16S rRNA gene-based results of complex communities from an in vitro gut model showed that R. intestinalis colonized mucin-covered carriers [31]. For unclear reasons, Roseburia but not Bacteroides decreased in relative abundance with mucin supplementation [32]. We thus require a mechanistic understanding of how mucin formed microhabitats impact the abundance of different gut species. Here, we built an in vitro batch system containing mucin beads, to systematically investigate how a spatially structured environment influences the growth and interactions of the model gut species B. thetaiotaomicron and R. intestinalis (Fig. 1). The growth dynamics of monocultures and co-cultures in the liquid medium and attached to the beads were monitored using live/dead cell staining followed by flow cytometry in combination with machine learning techniques to quantify their proportions in co-culture. Subsequently, we characterized the effect of mucin monosaccharides on the abundance of B. thetaiotaomicron and R. intestinalis, and found that it depended on the sugar level being considered. Transcriptomic analyses of monoculture and co-culture in medium with and without mucin beads at several time points revealed different nutrient harvesting strategies employed by B. thetaiotaomicron and R. intestinalis, which resulted in their different growth modes, and also changed their interaction mechanisms depending on nutrient availability. We further developed a kinetic model that accounts for pH conditions, metabolites, and growth profiles, to mechanistically understand how gel-entrapped mucin influences the proliferation of B. thetaiotaomicron and R. intestinalis and the dynamics of their relationship. Our methodology could potentially be applied to other ecosystems where microbial growth alters environmental conditions, which in turn affect microbial interactions.
We used a bottle system containing mucin beads in WC medium to mimic the spatially structured environment of the human gut. The growth dynamics of monoculture and co-culture in the liquid medium and attached to the beads were monitored using flow cytometry with live/dead cell staining. Live cells of B. thetaiotaomicron and R. intestinalis were differentiated from each other and from debris using a UMAP-based classification strategy (details in the “Methods” section). Fermentation products, including short-chain fatty acids and mucin-derived monomeric sugars, were measured by high-performance liquid chromatography and gas chromatography. The presence and quantity of mRNA in samples collected at different time points were assessed by RNA-seq. Further, a kinetic model with ordinary differential equations was developed that summarizes our understanding of the system. O2 oxygen, WC Wilkins–Chalgren medium, BT Bacteroides thetaiotaomicron, RI Roseburia intestinalis.
Materials and methods
Human gut bacterial strains
Bacterial strains of B. thetaiotaomicron VPI-5482 (DSM 2079T), DSM 24846, DSM 108160, DSM 108161 and R. intestinalis L1-82 (DSM 14610T) were obtained from the Deutsche Sammlung von Mikroorganismen und Zellkulturen (DSMZ, Germany). The strains were frozen in Wilkins–Chalgren Anaerobe Broth (WC; Oxoid Ltd, Basingstoke, UK) plus 20% glycerol and maintained at −80 °C until use.
Mucin bead preparation
Mucin beads were generated based on a protocol previously described [33]. Briefly, a polysaccharide gel was prepared that contained (in g/100 ml) Gelrite gellan gum (2.5, Sigma), xanthan gum (0.25, Sigma) and porcine mucin type II (5, Sigma; pretests of the strains grown in WC medium showed no difference between the beads supplemented with type II and type III mucins). The dried powders were suspended in 100 ml of 90 °C pre-heated distilled water containing 0.2 g sodium citrate (Sigma) and 350 μl 10 M NaOH. This polymer solution was mixed until dissolved and then autoclaved for 20 min at 121 °C. The bead generation was performed under sterile conditions and was based on a two-phase dispersion process [34]. The autoclaved solution was cooled down to 50 °C on a heater and slowly poured with a distance of 40 cm into 45 °C sunflower oil while keeping the stirring speed at 300 rpm, to obtain a suspension of aqueous gel droplets in the hydrophobic phase. Gel beads were formed when cooling the suspension down to 15 °C on ice. The oil was carefully washed off and beads were subsequently hardened by soaking in a 0.1 M CaCl2 solution (adjusted to pH 5.2) for 20 min at room temperature. Beads with a diameter of 500 μm were selected by wet sieving and stored at 4 °C in aseptic conditions until use.
Cultivation and sample collection
Serum bottles (120 ml) containing 60 ml WC medium and around 8000 mucin beads were used as a spatially structured system to grow B. thetaiotaomicron and R. intestinalis. Briefly, each bottle was first filled up with 50 ml WC medium, and 3 g wet mass of 500 μm mucin beads in 10 ml WC were pipetted into each bottle under sterile conditions. The bottles were transferred to the anaerobic workstation (Don Whitley A35 with HEPA filter; 10% H2, 10% CO2, 80% N2, 55% humidity) to be deoxygenated for 120 h. Oxygen-free conditions in liquid and in the beads were confirmed through the color change of redox indicator resazurin. The bottles were closed with rubber stoppers and sealed with aluminum caps that were UV sterilized for 1 h in a laminar flow chamber before use. For bottle experiments without mucin beads, bottles were filled with 60 ml WC medium that was prepared in the same way as described above.
To prepare the inoculum, 100 μl of B. thetaiotaomicron or R. intestinalis from frozen stocks was first inoculated in a 10-ml serum vial (Wheaton) containing 5 ml WC medium as preculture. These serum vials were prepared by adding autoclaved medium and were deoxygenated in the anaerobic workstation for 24 h. The vials were closed with rubber stoppers and sealed with aluminum caps before use. After 16 h, each species’ preculture was diluted to an OD600 of 0.1 determined by a portable spectrometer (DR1900; Hach Lange GmbH, Germany) and used as the inoculum. The serum bottles were inoculated with 1 ml of the diluted preculture and were incubated at a constant stirring rate of 170 rpm at 37 °C in a shaker (KS 4000 i; IKA, Staufen, Germany). Cultivations of the monoculture and co-culture were followed for 120 h after inoculation, with samples taken from both the liquid and beads every four hours for the first 48 h and every 12 h afterwards. For further investigation of the B. thetaiotaomicron and R. intestinalis growth and their interactions in the long term, an experiment was designed for the co-culture to follow for 288 h with samples taken every 24 h since 120 h after inoculation. These batch fermentation experiments were performed in closed bottles, i.e., metabolites and gas composition were not kept constant. All experiments were performed at least in three biological replicates and always had a negative control bottle without inoculation but with sampling for each time, to verify its sterility.
For each time point, 2 ml of the fermentation broth containing the suspended mucin beads were sampled to 2-ml tubes (Eppendorf). The serum bottles were opened and closed carefully each time to avoid contamination in the anaerobic workstation. During sampling the beads settled downward to the bottom within seconds. The supernatant was carefully pipetted into new 2-ml tubes in the anaerobic workstation, and subsequently was used to measure pH (PH-922, ISFET, Japan) and OD600, and to profile metabolites as well as quantify bacterial cell counts by live/dead staining followed by flow cytometry.
Spent media and pH preference experiments
Several spent media assays were conducted in serum bottles to clarify how B. thetaiotaomicron and R. intestinalis interact with each other, and how abiotic factors influenced their growth. First, we grew a culture of B. thetaiotaomicron in WC for 48 h without mucin beads and sterilized the spent medium through a filtration with 0.2-μm sterile syringe filters (PES; VWR International). In addition, the filtered medium was autoclaved and deoxygenated for 24 h in the anaerobic workstation. Four conditions were prepared to cultivate B. thetaiotaomicron in the spent medium: (i) the original spent medium without adjusting pH (pH 5) and without adding carbon source, (ii) spent medium with pH adjusted to 6.5, (iii) spent medium with glucose and pyruvate added in the same concentrations as in WC at pH 5 and (iv) spent medium with added glucose and pyruvate plus pH adjusted to 6.5. We grew B. thetaiotaomicron in three biological replicates in the serum bottles for a period of 48 h and sampled every 8 h for measuring pH, OD600, metabolites and live/dead cell counts. Second, we grew B. thetaiotaomicron in WC plus mucin beads in different periods of 24, 48 and 72 h, as well as in WC without mucin beads for 48 h. The cell- and oxygen-free spent media were prepared in the same way as described above, and the pH was adjusted to 6.5 before autoclaving.
We further adjusted the pH of WC medium to the specified values of 4.5, 5.0, 5.5 and 6.0 by using 1 M HCl solution, and also included the original WC of pH 6.7 for testing in serum bottles. The growth of B. thetaiotaomicron was followed for 48 h, with samples taken at 0, 4, 8, 12, 24, 36 and 48 h and measured as described above.
Detachment experiment
To estimate the rate at which bacterial cells detach from particles, we followed the population change of planktonic cells in a fresh WC medium, into which mucin beads with attached B. thetaiotaomicron cells were added. We first grew B. thetaiotaomicron culture in a serum bottle containing 60 ml WC plus mucin beads, with a same inoculum size as described: 1 ml of OD600 0.1. After 24 h cultivation, 2 ml of the fermentation broth containing the suspended mucin beads were sampled to each 2-ml tube. 0.5 ml of phosphate buffer (PBS, pH 7.3; Invitrogen, Thermo Fisher Scientific, USA) was added to each tube after washing with 1 ml PBS. The beads in PBS were transferred to a 12-well plate (Sarstedt, Germany), with 1.5 ml sterile WC medium in each well. For comparison, we included wells of fresh WC medium and WC medium supplemented with sterile mucin beads, both of which were inoculated with B. thetaiotaomicron in a volume of 25 µl of OD600 0.1. The experiment was performed in four replicate wells for each group. The plates were incubated at a constant stirring rate of 170 rpm at 37 °C, with liquid samples taken at 0, 30, 60, 90, 150 and 240 min for live/dead cell staining followed by flow cytometry.
Mucin monomeric sugar assays
The degradation products of mucin including N-acetylneuraminic acid (Neu5Ac), N-acetylgalactosamine, N-acetylglucosamine, fucose, mannose and galactose were tested for their effects on the growth of B. thetaiotaomicron and R. intestinalis. The tests for each species were performed by using 24-well plates (Sarstedt, Germany) filled with two types of media: the standard WC and WC without carbon sources glucose and pyruvate. Each medium was split into eight aliquots, each of which was supplemented with either one of those mucin sugars in 1 mM or a mixture of all six sugars together in 1 mM or without any sugar added as the control. We first inoculated 0.5 ml preculture with OD600 0.1 to 20 ml of each aliquot, and then transferred 1.2 ml to each well in three replicates. The plates of B. thetaiotaomicron or R. intestinalis were incubated at a constant stirring rate of 120 rpm at 37 °C for 24 or 48 h, respectively, with samples taken every 12 h for pH and live/dead staining followed by flow cytometry measurements.
Quantification of bacterial cell counts using flow cytometry
DNA-based staining using SYBR Green I (SG; Invitrogen) in combination with propidium iodide (PI; Invitrogen) was applied to quantify and differentiate bacterial cells with intact and damaged cytoplasmic membranes following the protocols described before [35]. Briefly, the stock staining solution contained per ml: 10 μl of SG (10,000 concentrate), 20 μl of 20 mM PI and 970 μl of the filtered dimethylsulfoxide. Cells of the fresh culture in liquid from different time points were diluted in a filter-sterilized PBS buffer for the live/dead cell staining. Dilutions in 1:10 and 1:200 were performed for the first two time points (0 and 4 h) and the later time points (after 8 h), respectively. Under anoxic conditions, cell samples were diluted in 1-ml 96-well plates (Eppendorf, Germany) and stained with 3 μl of the SG/PI staining solution, and subsequently were incubated for 20 min in the dark at 37 °C right before flow cytometric measurements. To quantify the attached cell counts, the mucin beads sampled from different time points were first washed with 1 ml filter-sterilized PBS in 2-ml tubes (Eppendorf). After carefully removing the supernatant, we added another 0.5 ml PBS to the tubes that later were sonicated in the water bath of an ultrasonic cleaner (USC200TH, VWR International) for 5 min at 45 kHz. The fresh detached cells in PBS were diluted 1:10 in 96-well plates for all time points and were used for the live/dead cell staining as described above. To confirm method accuracy, we also used a green fluorescent protein (GFP)-tagged B. thetaiotaomicron containing the modified plasmid pWW3452 (CTAATG in the start codon of GFP was replaced with CATATG), which can stably fluoresce when grown under anoxic conditions with subsequent exposure to oxygen for 60 min as described [36].
Flow cytometric counts of the fresh cells were obtained by using a benchtop CytoFLEX S flow cytometer (Beckman Coulter, Brea, USA), equipped with the identical lasers and detectors as described previously [37]. Cell events of each sample were recorded for 1 min at a sample flow rate of 10 μl/min, using the CytExpert software (version 2.4.0.28). Threshold values of 3000 and 2000 were applied to the channels of forward scatter and side scatter, respectively. These thresholds were selected based on our tests of B. thetaiotaomicron and R. intestinalis cells from different growth phases in serum bottles containing WC medium without and with mucin beads to exclude most of the background events and to obtain accurate cell counts. In addition to the instrumental calibration with CytoFLEX Daily QC Fluorospheres, 0.5 μm and 1 μm green fluorescent beads (Thermo Fisher Scientific, USA) were applied to each batch of experiments as internal standards, to monitor the instrument stability and to make cytometric data from different batches more comparable.
Flow cytometry data processing
We used an in-house pipeline to quantify the absolute abundance of B. thetaiotaomicron and R. intestinalis cells by clustering populations of flow cytometric events in a UMAP space [38]. The detailed protocol and scripts to reproduce our cell population counts are available at: bit.ly/3WNrslL. Briefly, we normalized and scaled raw flow cytometry data, transforming the values with the arcsin function for uncorrelated unit variances and zero means. We then projected this data into three-dimensional UMAP space using Euclidean distance, with parameters of 25 and 0.1 for the number of neighbors and minimum distance. Flow cytometry runs containing cells and runs containing only the media (blank) allowed classification into four categories: (1) “live” (SG positive, PI negative, and not in the blanks); (2) “inviable” (not in the blanks, but PI positive) and; (3) “debris” (PI and SG negative, but not in the blanks); (4) “blank” (overlap with the blank files). PI and SG positive events were determined based on a threshold signal in their respective channels, which were empirically estimated from the distribution of all the data samples. The overlap with blank events from blank control files was determined by drawing a circle around the points in the blank files, whose radius is proportional to the density of blank points found in its proximity and classifying the area spanned by this circle as blanks in a combined UMAP projection of samples and blank files. This approach that mimics the process of drawing a circle around the events in the blank file was sufficient because the UMAP projection showed a clear separation between the blank and cell events. Using the density as a basis for the circles allowed us to automatically account for the cases where there is a small carryover from cell events to the blank runs, preventing the automatic detection of cell events as blanks. Only the events classified as “live” were selected for the next steps. We then used the monoculture samples of each respective time point to classify cells in the co-culture samples. For this we first built a training space consisting of a labeled UMAP projection of 5000 random events selected from each monoculture biological replicate. Next, we used supervised UMAP to reproject the monoculture and co-culture live-cell events into the training space. We then used the K-nearest neighbors vote classifier (with parameters: n_neighbors = 50, weights = distance, and metric = Mahalanobis) to identify the species labels of the co-culture events based on their proximity to the monoculture events in the supervised UMAP space. Before filtering out the blanks, each sample was overlayed with their corresponding blank runs and manually inspected using clickable three-dimensional scatter plots, where we could visually assure that the populations classified as blank in our samples overlap with the populations in the blank files and not with the cell populations. Examples of these scatter plots are available at: bit.ly/3WNrslL. Similarly, we manually verified that each classified species population overlapped with their monoculture runs for similar time points. The manual inspection also showed that our parameter choices for UMAP and the K-nearest vote classifier were robust.
Contamination check and confirmation of abundance profiles with 16S rRNA genes
Sanger sequencing of 16S rRNA genes was performed regularly to check each strain identity. In addition, we selected four biological replicates for each time point from the co-cultivation experiments of B. thetaiotaomicron and R. intestinalis in WC with and without mucin beads, to confirm the abundance profile of B. thetaiotaomicron and R. intestinalis in co-culture with 16S rRNA gene amplicon sequencing (Fig. S1). Sequencing was performed using the MiSeq platform (Illumina) to generate paired-end reads of 250 base pairs. The taxonomy was assigned by using the RDP classifier v2.13 with a confidence threshold of 80% [39].
Metabolite profiling
Liquid samples in 1 ml were collected and centrifuged for 20 min at 21,130 × g at 4 °C (Centrifuge 5424R; Eppendorf, Hamburg, Germany). The supernatants were used for chemical analyses by applying external standards for calibration and quality control. For the calibration curves, all compounds (Merck) were in high purity and acidic form (except sugars), and were dissolved or diluted in ultrapure water for calibration. The concentrations of trehalose, glucose, pyruvate, succinate, formate, acetate, lactate, n-butyrate and Neu5Ac were determined by high-performance liquid chromatography (HPLC). Propionate was also measured but not detected in our tests since B. thetaiotaomicron requires vitamin B12 (absent in our medium) to produce propionate via the succinate pathway [40, 41]. HPLC analysis was carried out on an Agilent 1200 series HPLC system (Agilent Technologies, USA) equipped with a Bio-Rad Aminex HPX-87H column (Bio-Rad, USA) and a refractive index detector at 40 °C. Samples in 20 μl were injected and eluted for 60 min using 5 mM H2SO4 as the mobile phase at a flow rate of 0.6 ml/min.
Concentrations of mucin monomeric sugars galactose, fucose and mannose were relatively quantified by using gas chromatography, carried out by the Metabolomics Core of KU Leuven. The supernatant in 100 μl of each sample was diluted to 1 ml in 80% methanol, and left overnight to allow the precipitation of lipids and proteins. After centrifugation, 100 μl of the supernatant was transferred and evaporated to dryness on a speedvac. A 2% methoxyamine solution was prepared by dissolving methoxyamine hydrochloride (Supelco, ref. 226904-5g) in pyridine (Sigma-Aldrich, ref. 270970-1 L). 25 μl of the methoxyamine solution was added to each sample and left to react for 1.5 h at 37 °C for methoximation of the hexoses. Afterwards, 80 μl N,O-bis(trimethylsilyl)trifluoroacetamide with 1% trimethylchlorosilane (Supelco, ref. 15238) was added and left to react for 30 min at 60 °C for trimethylsilylation of the methoximated hexoses. One microliter of each methoximated and trimethylsilylated sample was injected by 2:1 split injection by an Agilent Technologies 7693 autosampler (Agilent Technologies, Santa Clara; California) into an Agilent 7890 A gas chromatograph equipped with a fused silica capillary column (0.25 mm × 30 m) with a chemically bonded 0.25 μm HP-5MS stationary phase (Agilent J&W GC columns, Folsom; California, ref 19091S-433). The injector temperature was 250 °C, and the septum purge flow rate was 3 ml/min. The helium flow rate through the column was 0.9 ml/min. The column temperature was held at 160 °C for 23.5 min and then increased by 100 °C/min to 300 °C, with 1 min holding there. The column effluent then entered an Agilent 7000 triple quadrupole mass spectrometer. Temperatures of the transfer line and electron impact ion source were 250 °C and 230 °C, respectively. Ions were generated by a 70 eV electron beam. Spectra were acquired in a selected ion monitoring mode, with fragment masses based on the analyses of standards.
RNA extraction and sequencing
We included a total of 69 samples representing the different growth phases of B. thetaiotaomicron and R. intestinalis in three biological replicates for RNA sequencing, which were selected from the following six independent experiments: monoculture B. thetaiotaomicron in WC (4, 12 and 36 h), monoculture R. intestinalis in WC (4, 12 and 48 h), co-culture B. thetaiotaomicron and R. intestinalis in WC (12, 24 and 36 h), monoculture B. thetaiotaomicron in WC plus mucin beads (4, 12, 24, 48 and 120 h), monoculture R. intestinalis in WC plus mucin beads (16, 28, 40 and 48 h), and co-culture B. thetaiotaomicron and R. intestinalis in WC plus mucin beads (6, 12, 24, 48 and 120 h).
Total RNA was extracted and purified from RNAprotect-treated (Qiagen, Hilden, Germany) frozen liquid samples using the phenol-free RNeasy Plus Mini Kit (Qiagen) according to the instructions of the manufacturer. The cell pellets from −80 °C were first lysed by adding the lysis buffer RLT from the kit with 10 μl/ml β-mercaptoethanol. The content of PowerBead tube (glass, 0.1 mm; Qiagen) was further added to the lysate prior to using a bead beater (TissueLyser II, Qiagen) for homogenizing the mixture for 5 min at a speed of 30 beats/s. Other steps were performed by following the instructions. The eluted RNA samples in nuclease free water were stored at −80 °C until use. RNA concentrations were determined with a Nanodrop ND 1000 spectrophotometer (Thermo Fisher Scientific, USA) and a Qubit 2.0 fluorometer using the Qubit RNA High Sensitivity Assay Kit (Invitrogen). Samples with a 260/230 absorbance ratio below 0.5 were purified using the MinElute PCR Purification kit (Qiagen) and re-measured afterwards, with all samples above 0.5 after purification. RNA integrity and yield were evaluated using RNA Nano and Pico 6000 LapChips (Agilent Technologies, Santa Clara, CA, USA) run on an Agilent 2100 Bioanalyzer (Agilent Technologies).
Library preparation and sequencing of RNA-seq libraries were carried out by the Genomics Core of KU Leuven. All samples were prepared using the NEBNext Ultra II Directional RNA Library Prep Kit for Illumina (New England BioLabs, #E7760L), with an initial amount of 100 ng total RNA in 12 μl. Bacterial rRNA depletion was proceeded using the rRNA Depletion Kit (Bacteria) (New England BioLabs, #E7850L), and the residual mRNA was quality controlled and converted to the libraries that were sequenced on a NovaSeq 6000 (Illumina, San Diego, CA, USA) on a v1.5 S4_300 kit in XP mode. The resulting data of paired-end reads in fastq file format were obtained and demultiplexed before further analysis.
RNA-seq data processing
The analysis of the raw sequencing reads was performed as follows: low quality reads and adapters were trimmed with fastp [42]. High quality RNA reads were mapped to their respective reference transcripts with Salmon run in a selective alignment mode and with a decoy-aware index constructed from each organism’s genome [43]. The latest reference genomes and transcripts for B. thetaiotaomicron and R. intestinalis were downloaded from the ensemble bacteria [44] and BV-BRC databases [45], respectively:
GCA_000011065.ASM1106v1.dna.toplevel.fa (B. thetaiotaomicron VPI-5482), GCA_000011065.ASM1106v1.cds.all.fa (B. thetaiotaomicron VPI-5482), GCA_900537995.1_Roseburia_intestinalis_strain_L1-82_genomic.fna (R. intestinalis L1-82), GCA_900537995.1_Roseburia_intestinalis_strain_L1-82_cds_from_genomic.fna (R. intestinalis L1-82). Raw transcript counts were extracted from salmon’s output and used for differential gene expression analysis. COG categories, COG numbers and KO numbers were extracted from NCBI COG and KEGG database [46, 47], for B. thetaiotaomicron. To obtain annotations for R. intestinalis, we mapped the respective transcripts to the eggNOG database v5.0 with the eggNOG-mapper 2.1.9 [48, 49].
Statistics
To measure gene expression levels, we used methods of transcripts per million and DESeq2’s median of ratios to normalize RNA read counts [50]. A strict cutoff of at least 1.5-fold change and a Benjamini-Hochberg adjusted p value less than or equal to 0.01 were applied for differential gene expression calculated by DESeq2. Statistical analyses were performed using R (version 4.2.0).
Modeling
Details of model definition and parameterization can be found in the Supplementary file 2.
Results
Different responses to sugar depletion
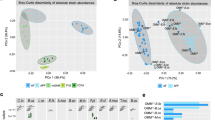
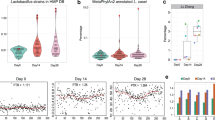
Glucose and pyruvate were provided as the main carbon sources in WC medium (Fig. 2a). When grown in monoculture, B. thetaiotaomicron produced succinate, lactate, formate, and acetate during fermentation, which decreased the pH from 6.7 to 5.0. At 20 h, B. thetaiotaomicron reached its maximal density of 1.2 × 106 viable cells/µl and hereafter its density decreased. After the depletion of glucose and pyruvate, the acidic conditions led to a loss of viable B. thetaiotaomicron cells but did not affect the total cell count. Loss of viability at low pH (5.0) was further confirmed by spent media and pH experiments. We re-grew B. thetaiotaomicron in its spent medium (collected at 48 h cultivation when glucose is depleted), and found that an initial pH of 6.5 is required for growth (on trehalose; Fig. S2). Supplementing the spent medium with glucose and pyruvate only enhanced growth when the pH was also increased (Fig. S2a). In addition, B. thetaiotaomicron did not grow on glucose and pyruvate when the pH of WC medium was lowered to 4.5 and 5.0, respectively, and its growth rate was decreased at pH 5.5 and 6.0 compared to pH 6.7 (Fig. S2b). R. intestinalis in monoculture produced lactate, formate, acetate and butyrate from glucose and pyruvate, corresponding to a pH decrease from 6.7 to 5.8 at 24 h and reached a maximal density of 7.8 × 105 viable cells/µl (Fig. 2b). After the depletion of glucose and pyruvate in 24 h, R. intestinalis gradually entered a slow growth mode, seen as a drop in viable cells to 1.8–3.8 × 105 cells/µl from 40 to 120 h in two independent experiments. In addition, we observed a further significant (p < 0.001 by Student’s t test) increase of butyrate concentration concurrent with decreasing acetate and, in some cases, lactate (Fig. 2b and Fig. S3 for lactate consumption in R. intestinalis monoculture), resulting in a pH increase from 5.7 to 6.0. In brief, viable B. thetaiotaomicron cells decline primarily due to the lowered pH, whereas R. intestinalis maintains higher cell densities by entering a slow growth mode.
Cell density, concentrations of glucose, pyruvate and fermentation products, and pH values for B. thetaiotaomicron monoculture (a), R. intestinalis monoculture (b) and their co-culture (c) in WC medium during 120 h cultivation; Cell density, concentrations of glucose, pyruvate and fermentation products, pH values as well as concentrations of mucin monomeric sugars for B. thetaiotaomicron monoculture (d), R. intestinalis monoculture (e) and their co-culture (f) in WC with mucin beads during 120 h cultivation. Circles represent the mean values of biological replicates. Error bars indicate the standard deviation of biological replicates (n = 5 for B. thetaiotaomicron and R. intestinalis monocultures in WC experiment; n = 6 for co-culture in WC; n = 3 and n = 5 for B. thetaiotaomicron and R. intestinalis monocultures in WC with mucin beads, respectively; n = 5 for co-culture in WC with mucin beads). Concentrations of mucin sugars in the barplots are represented by the integrated area under the peak corresponding to the metabolite. To show the same range on the y axis of barplots, we divided these values by factors of 1,000,000, 5,000,000 and 10,000 for galactose, mannose and fucose, respectively. WC Wilkins–Chalgren medium, BT Bacteroides thetaiotaomicron, RI Roseburia intestinalis, mono monoculture; co co-culture, Neu5Ac N-acetylneuraminic acid.
Given the observed diauxic shift from glucose/pyruvate to lactate/acetate, we performed RNA-seq of R. intestinalis grown in monoculture for several time points and replicates to investigate its metabolic mechanism. During the exponential growth period, genes associated with glycolytic and pyruvate-utilizing pathways were more highly expressed at mid-log phase of 12 h compared to at 4 h (Fig. S4b). Although glucose ran out at 48 h, we found that R. intestinalis kept expressing glycolysis genes and also non-essential genes such as those involved in cell motility despite growing in a well-mixed environment (Fig. S5). We further compared the effect of different normalization methods (transcripts per million and DESeq2’s median of ratios [50]), which showed consistent results except for some glycolysis genes (Fig. S6). When comparing the transcript levels of R. intestinalis in monoculture between 12 and 48 h, the significant up-regulation of genes encoding lactate dehydrogenase (RIL182_00335 and 01719) and butyryl-CoA:acetate CoA-transferase (RIL182_00413) corroborates the finding that R. intestinalis converts lactate/acetate to butyrate after glucose depletion. The energy yielded by butyrate production from lactate/acetate (ΔfG0 = −46 kJ/mol) is much lower than that from glucose (ΔfG0 = −250 kJ/mol) [51]. Therefore, it is not surprising that R. intestinalis grows more slowly on lactate/acetate. In summary, R. intestinalis maintains its slow growth in the absence of glucose by converting acetate and lactate to butyrate.
Effects of mucin on monoculture
B. thetaiotaomicron survives longer in the presence of mucin beads than without them. After depleting glucose and pyruvate at 24 h, planktonic B. thetaiotaomicron retained a viable cell count higher than 1.0 × 106 cells/µl (Fig. 2d), which was also maintained when further extending the cultivation period from 120 h to 288 h (Fig. S7a, b for three non-type strains). In contrast, the growth of R. intestinalis on glucose/pyruvate seems to be impeded by mucin, demonstrated by significantly lower cell densities at the same time points and lower maximum of 4.5 × 105 viable cells/µl at 36 h when compared with R. intestinalis grown without mucin beads (Fig. 2e; Fig. S8). After nutrient depletion, R. intestinalis still exhibited the slow growth mode characterized by a further increase of butyrate and decrease of acetate, but with a much lower density of 0.3–0.5 × 105 viable cells/µl than in the absence of mucin. In addition to the planktonic cells in liquid, we also quantified the cells of B. thetaiotaomicron and R. intestinalis attached to the mucin beads at different time points. Attached B. thetaiotaomicron and R. intestinalis in monoculture reached maximal densities of 6.0 × 105 viable cells/µl at 24 h and 1.1 × 105 viable cells/µl at 32 h, respectively (Fig. 2d, e). Thus, in the presence of mucin, B. thetaiotaomicron can maintain high densities in the long term, whereas the density of R. intestinalis declines.
Regarding the mucin degradation products, the concentration of Neu5Ac showed a significant (p < 0.001 by Student’s t test) increase of 0.12 mM in B. thetaiotaomicron monoculture after 120 h and no significant change in R. intestinalis monoculture (Fig. 2d, e). We also observed higher levels of fucose and galactose for B. thetaiotaomicron grown with mucin beads than without it. The dynamics of galactose in R. intestinalis monoculture indicates that R. intestinalis also hydrolyzed mucin and used resulting degradation products such as galactose. To investigate the effects of mucin degradation products on the growth of B. thetaiotaomicron and R. intestinalis at different glucose levels, monosaccharides including Neu5Ac, N-acetylgalactosamine, N-acetylglucosamine, fucose, galactose, mannose, and their mixture were added to the standard WC medium and WC medium without glucose and pyruvate in 24-well plates. In the presence of glucose/pyruvate, none of the supplemented sugars influenced B. thetaiotaomicron growth significantly (p > 0.05 by Student’s t test) compared to the WC control (Fig. S9a). In contrast, Neu5Ac increased R. intestinalis growth by 43% while other sugars had negative effects (14–46%; Fig. S9c). In the absence of glucose/pyruvate, all sugars except Neu5Ac had positive effects on B. thetaiotaomicron growth (2–35%; Fig. S9b), and all sugars, in particular galactose, promoted R. intestinalis growth (13%–197%; Fig. S9d). In conclusion, mucin-derived monosaccharides have species-specific effects that in addition depend on the level of glucose.
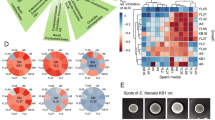
The RNA-seq data further elucidated the different responses of B. thetaiotaomicron and R. intestinalis to mucin. For B. thetaiotaomicron, we first collected a list of genes/operons associated with the polysaccharide utilization loci of mucin O-glycans, which were possibly activated by two types of transcriptional regulators, namely extracytoplasmic function sigma/anti-sigma pair (ECF-σ) and hybrid two-component system (HTCS) [52]. The genes linked to HTCS and ECF-σ showed significantly high expression in the first 4 h (Fig. 3a; p < 0.001 for both by Student’s t test), and genes of ECF-σ-linked loci were also expressed after the glucose/pyruvate depletion (48 and 120 h), indicating BT utilization of mucin O-glycan components. Specifically, after glucose/pyruvate ran out at 24 h, we observed a significantly (p < 0.001 in all cases; Student’s t test) increased expression of genes encoding the extracellular galactosidase (BT_3340, BT_4684) and fucosidase (BT_1842), as well as genes involved in the utilization of galactose (BT_0370–71, BT_0623) and fucose (BT_3606, BT_1273-75). These results are consistent with B. thetaiotaomicron metabolizing mucin O-glycans including galactose and fucose (Fig. 2d and S9b). In addition, R. intestinalis consistently increased the expression of genes involved in glycolysis, pyruvate utilization, mucin hydrolysis, galactose and mannose utilization, and the butyrate-producing process after depletion of glucose/pyruvate, with or without mucin beads present in the medium (Figs. 3b and S4b). At a high glucose/pyruvate level (Fig. S10a), the presence of mucin significantly reduced the expression of genes associated with the transport of glucose (RIL182_03793, RIL182_03794, RIL182_03796), pyruvate (RIL182_01689), galactose (RIL182_01067-69) and N-acetylglucosamine (RIL182_02688-89) across the outer membrane. At a low glucose/pyruvate level, genes of lactate dehydrogenase (RIL182_00335) and butyryl-CoA:acetate CoA-transferase (RIL182_00413) were significantly down-regulated (Fig. S10b). Thus, mucin supports the long-term survival of B. thetaiotaomicron, while it undermines R. intestinalis’ growth on glucose/pyruvate and possibly also on lactate/acetate.
Gene expression of the key metabolic routes of glucose and pyruvate fermentation, as well as mucin hydrolysis and mucin sugar utilization in B. thetaiotaomicron (a) and R. intestinalis (b) monocultures in WC medium with mucin beads; differential gene expression between co-culture and monocultures of B. thetaiotaomicron (c) and R. intestinalis (d). In each barplot of a and b, gene expression levels (x axis) were measured in transcripts per million of RNA samples at time (y axis) 4, 12, 24, 48 and 120 h for B. thetaiotaomicron, and at time 16, 28, 40 and 48 h for R. intestinalis. In each barplot of c and d, differential gene expression was calculated with DESeq2 for four categories: high levels of glucose/pyruvate in WC (I), low levels of glucose/pyruvate in WC (II), high levels of glucose/pyruvate in WC with mucin beads (III) and low levels of glucose/pyruvate in WC with mucin beads (IV); positive and negative values of log2fold-change are shown in red and blue on the x axis, respectively. Numbers in each barplot stand for the Gene ID in the genomes of B. thetaiotaomicron and R. intestinalis. The solid and dashed lines represent single- and multi-enzyme reactions between the two indicated molecules, respectively. Error bars indicate the standard deviation of three biological replicates. Metabolic maps with gene expression in B. thetaiotaomicron and R. intestinalis monocultures in WC without mucin beads are shown in Fig. S4 and the Supplementary files 3, 4. ECF extracytoplasmic function sigma/anti-sigma pair, HTCS hybrid two-component system, GlcR HTH-type transcriptional repressor of glucose, ppGpp alarmone guanosine-5′-3′-bispyrophosphate.
Impact of mucin on interaction dynamics
For the co-culture grown in WC, the co-utilization of glucose and pyruvate resulted in two separate density peaks of 7.4 and 4.6 × 105 viable cells/µl for B. thetaiotaomicron and R. intestinalis, respectively (Fig. 2c). The pH decreased from 6.7 to 5.3, which was not as acidic as in the B. thetaiotaomicron monoculture (5.0) but lower than in R. intestinalis monoculture (5.7). After depletion of glucose and pyruvate, B. thetaiotaomicron’s viable cell numbers gradually declined, as in monoculture. R. intestinalis presented a second peak of 4.1 × 105 viable cells/µl at 38 h. In addition, the net concentration of lactate in co-culture was significantly lower than in each monoculture (p < 0.001 in both cases; Student’s t test). For the co-culture grown in the presence of mucin beads, B. thetaiotaomicron reached a two-fold higher maximal density than without beads (1.5 × 106 viable cells/µl at 16 h). In contrast, R. intestinalis in the mucin co-culture exhibited significantly lower cell densities than in WC co-culture (p < 0.01 by Student’s t test), accompanied by a lower maximal density of 1.8 × 105 viable cells/µl. The pH of 5.1 was nearly as acidic as in the B. thetaiotaomicron monoculture with or without mucin. A second peak of R. intestinalis density in mucin co-culture was observed at 120 h, corresponding to a roughly ten-fold increase in density from 96 h to 120 h (Fig. S7a). The significant (p < 0.001 by Student’s t test) increase of galactose in the co-culture compared to the B. thetaiotaomicron monoculture suggests that both species hydrolyzed mucin (Fig. 2f).
To investigate the interaction dynamics of B. thetaiotaomicron and R. intestinalis and mucin’s impact on it, we compared viable cell counts in co-culture and monoculture over time. We computed the interaction strength, here defined as the impact of one organism on the other’s abundance, as the log ratio of viable cell counts in liquid in co- versus monocultures. The interaction strength and in some cases its sign (i.e., positive or negative impact) changed over time for both organisms with and without mucin beads (Fig. S11). In the presence of glucose/pyruvate, the competition for these nutrients resulted in negative interaction strengths for both organisms. After depletion of glucose/pyruvate and in the absence of mucin, R. intestinalis impacts B. thetaiotaomicron positively, likely by increasing the pH in the co-culture compared to the B. thetaiotaomicron monoculture. In the presence of mucin, B. thetaiotaomicron’s impact on R. intestinalis during the initial growth on glucose/pyruvate (20–48 h) changed from negative to positive, likely by reducing the concentration of mucin degradation products that affect R. intestinalis’ growth negatively in the presence of glucose. In summary, changing conditions alter interaction mechanisms between the two species, which lead to varying interaction signs and strengths.
To further elucidate their changing interaction mechanisms across growth phases, we compared the differential gene expression between co-culture and monoculture (Fig. 3c, d). In the presence of R. intestinalis, B. thetaiotaomicron genes associated with the transport of glucose (BT_0371, BT_1758) were significantly down-regulated in WC without mucin (Fig. 3c; I, II). In addition, the expression of six out of ten glycolysis genes as well as genes involved in succinate, acetate and formate production was significantly lowered in WC at low glucose/pyruvate levels (Fig. 3c; II). With mucin, B. thetaiotaomicron up-regulated glycolysis genes and genes involved in producing formate, acetate, succinate and lactate (Fig. 3c; III, IV). At low nutrient levels, it also up-regulated genes encoding the exocellular galactosidase (BT_3340, BT_4684), fucosidase (BT_1842), mannosidase (BT_0458) and sialidase (BT_0455) in response to mucin hydrolysis, and genes associated with the utilization of galactose and N-acetylgalactosamine (Fig. 3c; IV). B. thetaiotaomicron genes (BT_0617-22) belonging to the membrane-associated, proton-translocating ferredoxin:NAD+ oxidoreductase (Rnf complex) were also more expressed with mucin (Fig. 3c; III, IV). In conclusion, B. thetaiotaomicron down-regulates the genes of glycolysis and pyruvate oxidation when R. intestinalis is present in WC but does the opposite when mucin is also present.
In the presence of B. thetaiotaomicron, R. intestinalis genes associated with glucose transport (RIL182_03793-94, RIL182_03796, RIL182_01587-88) and the ATP generation via proton pump (RIL182_00203-10) were down-regulated at high and low glucose/pyruvate levels, respectively (Fig. 3d; I, II, respectively). A candidate gene of the alarmone ppGpp biosynthesis was significantly up-regulated (RIL182_00174). It is part of R. intestinalis’ response to stress such as carbon starvation and may be involved in triggering its diauxic shift. R. intestinalis genes associated with the import of glucose (RIL182_03794, RIL182_03796), pyruvate (RIL182_01689), galactose (RIL182_01067-69) and N-acetylglucosamine (RIL182_02688-89), as well as seven out of ten glycolysis genes were systematically down-regulated at high glucose/pyruvate levels with mucin (Fig. 3d; III). In addition, R. intestinalis genes associated with the butyrate-producing process via butyryl-CoA:acetate CoA-transferase pathway were more expressed when glucose/pyruvate was depleted, with or without mucin (Fig. 3d; II, IV). Thus, B. thetaiotaomicron reduces R. intestinalis’ import of glucose and may assist its production of butyrate from lactate/acetate, regardless of mucin supplementation.
Kinetic model reproduces the observed dynamics
To reproduce physiological observables such as total and viable cell counts as well as concentrations of key nutrients and metabolites, we developed a model with ordinary differential equations that summarizes our understanding of the interaction mechanisms (Fig. 4a). Since we see a roughly linear relationship between counts of viable attached and free-swimming cells (Fig. S12), the model assumes that a fixed fraction of the cells attaches to mucin beads and that only these cells can degrade mucin. Mucin is treated as an inexhaustible resource. We further assume that changes in SCFAs fully determine changes in the pH, since we observed a strong linear relationship between pH and the sum of the concentrations of SCFAs (Fig. S13). Based on the literature, we also assume that a low pH affects the growth of B. thetaiotaomicron more strongly than that of R. intestinalis (i.e., B. thetaiotaomicron is more sensitive to acidity [53]). In addition, B. thetaiotaomicron and R. intestinalis can both grow on glucose and mucin sugars, which was well-supported by our fermentation data and also confirmed at the transcriptional level. Given our microtiter plate data, we further assume that mucin degradation products inhibit R. intestinalis’ growth on glucose (Fig. S9c). R. intestinalis only switches to its lactate/acetate-consuming slow growth mode when the concentrations of glucose and mucin degradation products are low (Fig. S10b). Finally, we hypothesize that glucose inhibits B. thetaiotaomicron’s growth on mucin sugars, which is supported by the significant overexpression of the master regulator of carbohydrate utilization in the presence of glucose (log2fold-change of BT_4338 in mucin monoculture: 1.8 for 4 h vs 12 h, 3.5 for 4 h vs 24 h and 1.6 for 12 h vs 24 h). We simplified the model by treating glucose and pyruvate, acetate and lactate as well as mucin sugars as single entities (see model scheme in Fig. 4a). The slow growth mode of R. intestinalis and the behavior of B. thetaiotaomicron on mucin after depletion of sugar are represented by separate states (in both cases referred to as “slow growth mode”). Parameter values including metabolite production and consumption as well as state switching rates were determined manually. The equations, parameter values and initial conditions are given in the Supplementary file 2.
a Scheme of the kinetic model. b Dynamics generated by the model without mucin (six upper plots) and with mucin (six lower plots). The equations, parameter values and initial conditions are given in Supplementary file 2.
The model qualitatively reproduces our observations, including the second peak observed for B. thetaiotaomicron at 32 h in WC monoculture, the long-term survival of R. intestinalis in WC and of B. thetaiotaomicron in the presence of mucin beads (Fig. 4b). However, the model fails to reproduce the second peak for R. intestinalis in WC in co-culture, implying that additional mechanisms are at play that are not yet fully understood.
Discussion
Here, we investigated the molecular mechanisms underlying the ecological interactions between two representative gut bacteria in the presence of simple and complex carbon sources within a spatially structured environment (Fig. 5). The greater affinity of B. thetaiotaomicron for the mucin beads (with a mean percentage of 33% attached cells out of total cells, compared to 25% for R. intestinalis), coupled with the more beneficial effect of mucin on the growth of B. thetaiotaomicron than R. intestinalis, agrees with their known niches in the human gut: B. thetaiotaomicron thrives as a mucus resident, while R. intestinalis tends to inhabit the lumen [17, 32].
Signs of positive, negative and absence of an interaction are shown as +, − and 0, respectively. The thickness of the arrow represents the interaction strength quantified as the logarithm of viable cell density ratios in co- versus monocultures (detailed in Figs. S8 and S11). Main metabolic pathways including glycolysis, converting lactate/acetate to butyrate, mucin hydrolysis and mucin sugar utilization are colored in teal, pink and black for B. thetaiotaomicron, R. intestinalis and both, respectively. Data shown in the figure represent measurements except for the metabolic pathways, which were taken from the literature or inferred from RNA-seq data. Neu5Ac N-acetylneuraminic acid, GlcNAc N-acetylglucosamine, GalNAc N-acetylgalactosamine.
We found that even when feeding on similar resources, B. thetaiotaomicron and R. intestinalis used alternative strategies to cope with and anticipate nutrient scarcity. Growth data combined with temporal gene expression patterns suggest that B. thetaiotaomicron acts as a “funnel” with its central carbon/energy metabolism capable of tapping several monosaccharides. Independent entry points are funneled into glycolysis until phosphoenolpyruvate is produced, which is converted into oxaloacetate by fixing a CO2 molecule with the generation of an extra ATP [54]. CO2 could be supplied from either the external environment or the conversion of pyruvate to acetyl-CoA. Although this strategy results in accelerated growth, it comes at the cost of significantly inferior performance when actively consumed nutrients are depleted. In contrast, R. intestinalis uses carbon chain elongation to generate additional energy and maintain a redox balance (Fig. 3b), resulting in butyrate production. Although its growth is slower than that of B. thetaiotaomicron, it can switch its energy source to acetate and lactate, which are also produced by B. thetaiotaomicron. This diauxic shift resembles the nutrient utilization strategies adopted by other bacterial species [55], mostly well characterized in Escherichia coli [56,57,58,59]. Some butyrate producers such as R. intestinalis can overcome the energetic barrier to convert lactate to pyruvate by employing the flavin-based electron confurcation: driving electron flow from lactate to NAD+ at the cost of exergonic electron flow from reduced ferredoxin to NAD+. However, producing butyrate from lactate and acetate is a much lower energy-yielding process, compared to glucose [51]. Slow growth is also a known strategy to deal with starvation in other nonsporulating bacteria [60, 61].
B. thetaiotaomicron’s growth strategy results in a dramatic reduction in the environmental pH, which significantly inhibits its own growth. This self-inhibition resembles the hypothesis of ecological suicide suggested previously [62]. At low pH, B. thetaiotaomicron cells remain viable for an extended period of time when mucin beads are present. Without mucin, when glucose and pyruvate are depleted, a loss of viability is observed (Fig. 2a). When the pH was restored in a spent medium assay, B. thetaiotaomicron metabolized an alternative carbon source and restored growth (Fig. S2). Thus, the depletion of nutrients per se is probably not the cause of this viability loss. Taken together, these results suggest that similar to other bacteria [63], the metabolic activity of B. thetaiotaomicron requires an energy source in an acidic environment, possibly channeling its ATP from growth to maintaining its intracellular pH within a physiologically viable range. When this source of energy is no longer available, cells quickly lose viability and can no longer exploit new nutrients. Although the two species compete for glucose and pyruvate, and thus initially sustain a negative interaction, the growth of R. intestinalis results in a smaller decrease in the environmental pH than the growth of B. thetaiotaomicron, leading to an indirect positive effect on B. thetaiotaomicron at later stages (Fig. 5).
B. thetaiotaomicron and R. intestinalis respond differently to mucin, depending also on the presence or absence of glucose. In the absence of glucose, most of the mucin sugars tested boost B. thetaiotaomicron’s growth, except for Neu5Ac. Analysis of its genome reveals the absence of a Neu5Ac lyase (nanL) gene. B. thetaiotaomicron is unable to catabolize Neu5Ac and did not grow on it in a minimal medium [64, 65]. As mucin beads and the tested mucin sugars did not significantly affect B. thetaiotaomicron’s growth during glucose fermentation (Fig. S9a), we assume that glucose prevents B. thetaiotaomicron from simultaneously consuming other carbon sources such as mucin-derived galactose and fucose. This assumption was also necessary to reproduce B. thetaiotaomicron’s behavior with our model. In E. coli, further uptake of carbon sources is suppressed by the transcriptional regulator cAMP-Crp when nutrient uptake surpasses anabolism [55]. B. thetaiotaomicron uses another master regulator of carbohydrate utilization (BT_4338) particularly for mucin utilization [66], which we found to be significantly over-expressed when B. thetaiotaomicron consumed glucose. As B. thetaiotaomicron was reported to easily switch its metabolism from dietary input to mucin glycans [17], we assume that it relies on BT_4338 to cope with excessive carbon flux. In contrast, R. intestinalis does not grow well when glucose and mucin are supplied together (Fig. 2e). Adding mucin beads also significantly up-regulated the gene of HTH-type transcriptional repressor GlcR (RIL182_03930), which indicated the repression of glucose catabolism by mucin sugars. Combined with slower growth on mucin sugars, this may explain the negative effects of mucin sugars on its growth on glucose. This is also supported by its significantly increased abundance in the mucin co-culture (Fig. 2f), which likely resulted from a decreased concentration of mucin sugars such as mannose due to uptake by B. thetaiotaomicron. However, a mechanism of R. intestinalis’ carbon uptake strategy has to be explored further.
Glycan-mediated attachment was reported as an important strategy for gut bacteria to access nutrients [8]. Since both B. thetaiotaomicron and R. intestinalis attached to the mucin beads, the physical proximity could influence their interactions on the beads. The beads were also not saturated by cells, illustrated by a roughly linear relationship between density on beads and in the liquid of co-culture (Figs. S12 and S14). In addition, the observed fast attachment and detachment implies that the dynamics between attached and free-swimming cells may be tightly coupled. Spatial transcriptomics at a single-cell resolution may further reveal the interaction mechanisms of human gut bacteria in a spatial context [67].
In the presence of B. thetaiotaomicron, R. intestinalis exhibited additional density peaks at 38 h without mucin and at 120 h with mucin, which were not predicted by our model. In most of the cases, the classification technique we used showed high accuracy in distinguishing B. thetaiotaomicron and R. intestinalis on a basis of their optical characteristics in flow cytometry, and the method accuracy was confirmed with the data of a GFP labeled B. thetaiotaomicron (Fig. S15). However, when the cells are losing viability (Fig. 2c, 24–38 h), the similarity between live and inviable cells causes difficulties in species separation. Thus, the transition from the first peak of viable R. intestinalis cells to the second in the WC co-culture may be smoother than observed. There are also alternative explanations for the peak. Since B. thetaiotaomicron can produce a higher concentration of lactate than R. intestinalis in monoculture, a transiently sufficient amount of lactate in the presence of B. thetaiotaomicron may reverse the flow from lactate to pyruvate in R. intestinalis so that lactate is consumed instead of produced. In addition, under the stress of an acidic pH B. thetaiotaomicron may release toxins that contribute to the loss of viability of R. intestinalis [68]. Some viable cells of R. intestinalis may also grow by necrophagy on accumulated dead cells [69].
We found that interaction strength significantly varies in value and sign along the growth curve, both with and without mucin (Fig. 5), due to the change in pH and nutrient levels caused by bacterial growth. Such an interplay between microbial growth and the environment is not only limited to the human gut, but also commonly described for other natural or engineered ecosystems, such as soil [70] or anaerobic digesters [71]. Understanding it requires new experimental and analytical approaches that track ecosystems in time and reveal how and when interactions change with the environment. However, widely used population models such as the generalized Lotka-Volterra assume interaction strengths to be constant. Thus, the generalized Lotka-Volterra is only applicable in systems where environmental conditions (including pH and nutrient levels) are kept constant, i.e., chemostats. The intestine is not well described by a chemostat, since nutrient levels, moisture and pH fluctuate with the daily intake of meals and the circadian rhythm [72, 73]. As confirmed by in situ measurements with capsules, the pH is also known to be variable across colon segments and across colons [74]. To cope with this variability, models can either account for changing interaction strengths as done here, which requires knowledge of interaction mechanisms and parameter values, or average over interaction strengths encountered in typical scenarios in the environment of interest. Such an ensemble approach may better represent the interaction between two organisms than the usual “snapshot” strategy of quantifying interactions in one particular condition but may not lend itself to predictive modeling. Thus, changing interaction strengths are a challenge for predictive models of the human gut microbiota and other ecosystems that sustain dynamic environmental conditions.
While our experiments were designed to mimic some aspects of the human colon, they neglected others. Not only is the chemical environment in the colon more complex than WC medium, but also the commercially available pig gastric mucins and subsequent treatment (e.g., autoclavation) alters the composition and structural properties compared to the colonic mucins, which could potentially alter the attachment and further utilization abilities of gut bacteria. In addition, nutrient availability and pH may vary less in the colon than in our experiments since sorption across the epithelium reduces the accumulation of fermentation products. Lastly, we studied interaction mechanisms in a specific context using specific strains. It remains uncertain how these findings would apply to different strains of these two species or in varied environmental settings (e.g. different medium). Although context and strain-specific dynamics are widely expected in microbial ecosystems—with strains differentiated by as few as 100 bp SNPs possibly having markedly different biotic interactions [75]—we observed a similar impact of pH on the growth of other B. thetaiotaomicron strains (Fig. S7b) and some genes involved in the key processes that we describe (e.g. mucus degradation and butyrate formation) are encoded in the genomes of other Roseburia strains as reported previously [76].
A deeper understanding of alternative metabolic strategies and changing species interactions in gut microbiota is necessary to rationally design next-generation probiotics and to predict the outcome of fecal microbiota transplantation. By deciphering the interaction mechanisms between two common gut bacteria across changing conditions, we have taken another step towards this goal.
Data availability
Raw amplicon sequencing data and RNA-seq data have been deposited to the Sequence Read Archive database under accession numbers of SAMN32182740-827 and SAMN32321133-55, respectively (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA911421/ and https://www.ncbi.nlm.nih.gov/bioproject/PRJNA914119/). Raw flow cytometry data have been deposited to flowrepository.org, under Repository IDs of FR-FCM-Z6YM, FR-FCM-Z6YN, FR-FCM-Z6YP, FR-FCM-Z6YQ, FR-FCM-Z6YR, FR-FCM-Z6YS, FR-FCM-Z6YT, FR-FCM-Z6YU, FR-FCM-Z6YV and FR-FCM-Z6YX. The processed RNA-seq data, flow cytometry data and fermentation data are available at: https://github.com/msysbio/BTRI_mucin_study_data.
Code availability
Flow cytometry data processing: bit.ly/3WNrslL (also available at Zenodo: https://zenodo.org/record/8251714), MATLAB file for the kinetic model and RNA-seq processing scripts: https://github.com/msysbio/BTRI_mucin_study_scripts.
References
Donaldson GP, Lee SM, Mazmanian SK. Gut biogeography of the bacterial microbiota. Nat Rev Microbiol. 2015;14:20–32.
Tropini C, Earle KA, Huang KC, Sonnenburg JL. The gut microbiome: connecting spatial organization to function. Cell Host Microbe. 2017;21:433–42.
Nadell CD, Drescher K, Foster KR. Spatial structure, cooperation and competition in biofilms. Nat Rev Microbiol. 2016;14:589–600.
Welch JLM, Hasegawa Y, McNulty NP, Gordon JI, Borisy GG. Spatial organization of a model 15-member human gut microbiota established in gnotobiotic mice. Proc Natl Acad Sci USA. 2017;114:E9105–14.
Van Hoek MJA, Merks RMH. Emergence of microbial diversity due to cross-feeding interactions in a spatial model of gut microbial metabolism. BMC Syst Biol. 2017;11:56.
Lowery NV, Ursell T. Structured environments fundamentally alter dynamics and stability of ecological communities. Proc Natl Acad Sci USA. 2019;116:379–88.
Geng J, Ji B, Li G, López-Isunza F, Nielsen J. CODY enables quantitatively spatiotemporal predictions on in vivo gut microbial variability induced by diet intervention. Proc Natl Acad Sci USA. 2021;118:e2019336118.
Patnode ML, Guruge JL, Castillo JJ, Couture GA, Lombard V, Terrapon N, et al. Strain-level functional variation in the human gut microbiota based on bacterial binding to artificial food particles. Cell Host Microbe. 2021;29:664–.e5.
Desai MS, Seekatz AM, Koropatkin NM, Kamada N, Hickey CA, Wolter M, et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell. 2016;167:1339–53.
Johansson MEV, Phillipson M, Petersson J, Velcich A, Holm L, Hansson GC. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc Natl Acad Sci USA. 2008;105:15064–9.
Duncan K, Carey-Ewend K, Vaishnava S. Spatial analysis of gut microbiome reveals a distinct ecological niche associated with the mucus layer. Gut Microbes. 2021;13:1874815.
Johansson MEV, Holmén Larsson JM, Hansson GC. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host-microbial interactions. Proc Natl Acad Sci USA. 2011;108:4659–65.
Salyers AA, West SEH, Vercellotti JR, Wilkins TD. Fermentation of mucins and plant polysaccharides by anaerobic bacteria from the human colon. Appl Environ Microbiol. 1977;34:529–33.
Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR, et al. Enterotypes of the human gut microbiome. Nature. 2011;473:174–80.
Xu J, Bjursell MK, Himrod J, Deng S, Carmichael LK, Chiang HC, et al. A genomic view of the human-Bacteroides thetaiotaomicron symbiosis. Science. 2003;299:2074–6.
Sonnenburg JL, Xu J, Leip DD, Chen CH, Westover BP, Weatherford J, et al. Glycan foraging in vivo by an intestine-adapted bacterial symbiont. Science. 2005;307:1955–9.
Bjursell MK, Martens EC, Gordon JI. Functional genomic and metabolic studies of the adaptations of a prominent adult human gut symbiont, Bacteroides thetaiotaomicron, to the suckling period. J Biol Chem. 2006;281:36269–79.
Morrison DJ, Preston T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes. 2016;7:189–200.
Wrzosek L, Miquel S, Noordine M-L, Bouet S, Chevalier-Curt MJ, Robert V, et al. Bacteroides thetaiotaomicron and Faecalibacterium prausnitzii influence the production of mucus glycans and the development of goblet cells in the colonic epithelium of a gnotobiotic model rodent. BMC Biol. 2013;11:61.
Ríos-Covián D, Ruas-Madiedo P, Margolles A, Gueimonde M, De los Reyes-Gavilán CG, Salazar N. Intestinal short chain fatty acids and their link with diet and human health. Front Microbiol. 2016;7:185.
Louis P, Flint HJ. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol Lett. 2009;294:1–8.
Falony G, Verschaeren A, De Bruycker F, De Preter V, Verbeke K, Leroy F, et al. In vitro kinetics of prebiotic inulin-type fructan fermentation by butyrate-producing colon bacteria: implementation of online gas chromatography for quantitative analysis of carbon dioxide and hydrogen gas production. Appl Environ Microbiol. 2009;75:5884–92.
Louis P, Duncan SH, Sheridan PO, Walker AW, Flint HJ. Microbial lactate utilisation and the stability of the gut microbiome. Gut Microbiome. 2022;3:e3.
Louis P, Flint HJ. Formation of propionate and butyrate by the human colonic microbiota. Environ Microbiol. 2017;19:29–41.
Duncan SH, Louis P, Flint HJ. Lactate-utilizing bacteria, isolated from human feces, that produce butyrate as a major fermentation product. Appl Environ Microbiol. 2004;70:5810–7.
Duncan SH, Hold GL, Barcenilla A, Stewart CS, Flint HJ. Roseburia intestinalis sp. nov., a novel saccharolytic, butyrate-producing bacterium from human faeces. Int J Syst Evol Microbiol. 2002;52:1615–20.
Duncan SH, Barcenilla A, Stewart CS, Pryde SE, Flint HJ. Acetate utilization and butyryl coenzyme A (CoA):acetate-CoA transferase in butyrate-producing bacteria from the human large intestine. Appl Environ Microbiol. 2002;68:5186–90.
D’hoe K, Vet S, Faust K, Moens F, Falony G, Gonze D, et al. Integrated culturing, modeling and transcriptomics uncovers complex interactions and emergent behavior in a three-species synthetic gut community. eLife. 2018;7:e37090.
Macfarlane S, Woodmansey EJ, Macfarlane GT. Colonization of mucin by human intestinal bacteria and establishment of biofilm communities in a two-stage continuous culture system. Appl Environ Microbiol. 2005;71:7483–92.
Van den Abbeele P, Roos S, Eeckhaut V, Mackenzie DA, Derde M, Verstraete W, et al. Incorporating a mucosal environment in a dynamic gut model results in a more representative colonization by lactobacilli. Micro Biotechnol. 2012;5:106–15.
Van den Abbeele P, Belzer C, Goossens M, Kleerebezem M, De Vos WM, Thas O, et al. Butyrate-producing Clostridium cluster XIVa species specifically colonize mucins in an in vitro gut model. ISME J. 2013;7:949–61.
Van Herreweghen F, De Paepe K, Roume H, Kerckhof FM, Van de Wiele T. Mucin degradation niche as a driver of microbiome composition and Akkermansia muciniphila abundance in a dynamic gut model is donor independent. FEMS Microbiol Ecol. 2018;94:fiy186.
Cinquin C, Le Blay G, Fliss I, Lacroix C. Immobilization of infant fecal microbiota and utilization in an in vitro colonic fermentation model. Microb Ecol. 2004;48:128–38.
Lamboley L, Lacroix C, Champagne CP, Vuillemard JC. Continuous mixed strain mesophilic lactic starter production in supplemented whey permeate medium using immobilized cell technology. Biotechnol Bioeng. 1997;56:502–16.
Hernandez-Sanabria E, Slomka V, Herrero ER, Kerckhof FM, Zaidel L, Teughels W, et al. In vitro increased respiratory activity of selected oral bacteria may explain competitive and collaborative interactions in the oral microbiome. Front Cell Infect Microbiol. 2017;7:235.
Whitaker WR, Shepherd ES, Sonnenburg JL. Tunable expression tools enable single-cell strain distinction in the gut microbiome. Cell. 2017;169:538–.e12.
Van de Velde CC, Joseph C, Biclot A, Huys GRB, Pinheiro VB, Bernaerts K, et al. Fast quantification of gut bacterial species in cocultures using flow cytometry and supervised classification. ISME Commun. 2022;2:40.
McInnes L, Healy J, Saul N, Großberger L. UMAP: uniform manifold approximation and projection. J Open Source Softw. 2018;3:861.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–7.
Reichardt N, Duncan SH, Young P, Belenguer A, Leitch CM, Scott KP, et al. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 2014;8:1323–35.
Shelton AN, Seth EC, Mok KC, Han AW, Jackson SN, Haft DR, et al. Uneven distribution of cobamide biosynthesis and dependence in bacteria predicted by comparative genomics. ISME J. 2019;13:789–804.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890.
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods. 2017;14:417–9.
Yates AD, Allen J, Amode RM, Azov AG, Barba M, Becerra A, et al. Ensembl Genomes 2022: an expanding genome resource for non-vertebrates. Nucleic Acids Res. 2022;50:D996–D1003.
Olson RD, Assaf R, Brettin T, Conrad N, Cucinell C, Davis JJ, et al. Introducing the Bacterial and Viral Bioinformatics Resource Center (BV-BRC): a resource combining PATRIC, IRD and ViPR. Nucleic Acids Res. 2023;51:D678–D689.
Galperin MY, Wolf YI, Makarova KS, Alvarez RV, Landsman D, Koonin EV. COG database update: focus on microbial diversity, model organisms, and widespread pathogens. Nucleic Acids Res. 2021;49:D274–D281.
Kanehisa M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 2019;28:1947–51.
Huerta-Cepas J, Szklarczyk D, Heller D, Hernández-Plaza A, Forslund SK, Cook H, et al. EggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 2019;47:D309–14.
Cantalapiedra CP, Hernandez-Plaza A, Letunic I, Bork P, Huerta-Cepas J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol Biol Evol. 2021;38:5825–9.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Shetty SA, Boeren S, Bui TPN, Smidt H, de Vos WM. Unravelling lactate-acetate and sugar conversion into butyrate by intestinal Anaerobutyricum and Anaerostipes species by comparative proteogenomics. Environ Microbiol. 2020;22:4863–75.
Martens EC, Chiang HC, Gordon JI. Mucosal glycan foraging enhances fitness and transmission of a saccharolytic human gut bacterial symbiont. Cell Host Microbe. 2008;4:447–57.
Duncan SH, Louis P, Thomson JM, Flint HJ. The role of pH in determining the species composition of the human colonic microbiota. Environ Microbiol. 2009;11:2112–22.
Fischbach MA, Sonnenburg JL. Eating for two: how metabolism establishes interspecies interactions in the gut. Cell Host Microbe. 2011;10:336–47.
Okano H, Hermsen R, Hwa T. Hierarchical and simultaneous utilization of carbon substrates: mechanistic insights, physiological roles, and ecological consequences. Curr Opin Microbiol. 2021;63:172–8.
Monod J. The growth of bacteria cultures. Annu Rev Microbiol. 1949;3:371–94.
Monod J. From enzymatic adaptation to allosteric transitions. Science. 1966;154:475–83.
Basan M, Honda T, Christodoulou D, Hörl M, Chang YF, Leoncini E, et al. A universal trade-off between growth and lag in fluctuating environments. Nature. 2020;584:470–4.
Balakrishnan R, de Silva RT, Hwa T, Cremer J. Suboptimal resource allocation in changing environments constrains response and growth in bacteria. Mol Syst Biol. 2021;17:e10597.
Gray DA, Dugar G, Gamba P, Strahl H, Jonker MJ, Hamoen LW. Extreme slow growth as alternative strategy to survive deep starvation in bacteria. Nat Commun. 2019;10:890.
Schofield WB, Zimmermann-Kogadeeva M, Zimmermann M, Barry NA, Goodman AL. The stringent response determines the ability of a commensal bacterium to survive starvation and to persist in the gut. Cell Host Microbe. 2018;24:120–.e6.
Ratzke C, Denk J, Gore J. Ecological suicide in microbes. Nat Ecol Evol. 2018;2:867–72.
Krulwich TA, Sachs G, Padan E. Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol. 2011;9:330–43.
Marcobal A, Barboza M, Sonnenburg ED, Pudlo N, Martens EC, Desai P, et al. Bacteroides in the infant gut consume milk oligosaccharides via mucus-utilization pathways. Cell Host Microbe. 2011;10:507–14.
Brigham C, Caughlan R, Gallegos R, Dallas MB, Godoy VG, Malamy MH. Sialic acid (N-acetyl neuraminic acid) utilization by Bacteroides fragilis requires a novel N-acetyl mannosamine epimerase. J Bacteriol. 2009;191:3629–38.
Townsend GE II, Han W, Schwalm ND III, Hong X, Bencivenga-Barry NA, Goodman AL, et al. A master regulator of Bacteroides thetaiotaomicron gut colonization controls carbohydrate utilization and an alternative protein synthesis factor. mBio. 2020;11:e03221–19.
Dar D, Dar N, Cai L, Newman DK. Spatial transcriptomics of planktonic and sessile bacterial populations at single-cell resolution. Science. 2021;373:758.
Coyne MJ, Béchon N, Matano LM, McEneany VL, Chatzidaki-Livanis M, Comstock LE. A family of anti-Bacteroidales peptide toxins wide-spread in the human gut microbiota. Nat Commun. 2019;10:3460.
Finkel SE. Long-term survival during stationary phase: evolution and the GASP phenotype. Nat Rev Microbiol. 2006;4:113–20.
Ratzke C, Gore J. Modifying and reacting to the environmental pH can drive bacterial interactions. PLoS Biol. 2018;16:e2004248.
Liu B, Sträuber H, Centler F, Harms H, da Rocha UN, Kleinsteuber S. Functional redundancy secures resilience of chain elongation communities upon pH shifts in closed bioreactor ecosystems. Environ Sci Technol. 2023. https://doi.org/10.1021/acs.est.2c09573.
Zmora N, Suez J, Elinav E. You are what you eat: diet, health and the gut microbiota. Nat Rev Gastroenterol Hepatol. 2018;16:35–56.
Zarrinpar A, Chaix A, Yooseph S, Panda S. Diet and feeding pattern affect the diurnal dynamics of the gut microbiome. Cell Metab. 2014;20:1006–17.
Shalon D, Culver RN, Grembi JA, Folz J, Treit PV, Shi H, et al. Profiling the human intestinal environment under physiological conditions. Nature. 2023;617:581–91.
Goyal A, Bittleston LS, Leventhal GE, Lu L, Cordero OX. Interactions between strains govern the eco-evolutionary dynamics of microbial communities. eLife. 2022;11:e74987.
Pichler MJ, Yamada C, Shuoker B, Alvarez-Silva C, Gotoh A, Leth ML, et al. Butyrate producing colonic Clostridiales metabolise human milk oligosaccharides and cross feed on mucin via conserved pathways. Nat Commun. 2020;11:3285.
Acknowledgements
We would like to thank Emma Hernandez-Sanabria, Veronica Lloréns-Rico and Jelle Matthijnssens for input and support concerning bacterial transcriptomics. In addition, we thank Vitor Pinheiro, Lieve Vanmellaert and Charlotte van de Velde for their help in constructing the GFP tagged B. thetaiotaomicron strain as well as Jeroen Raes for access to the CytoFLEX. We are also grateful to Thi Thuy Duyen Nguyen, Leen Rymenans and Geert Huys of the Raes lab for assistance in flow cytometry, 16S rRNA gene library preparation and anaerobic cultivation, as well as to Raul Yhossef Tito Tadeo for processing raw sequence reads. We also thank Tobie Martens from MULTIPHOTON of KU Leuven for his technical assistance in microscopy and the team of Genomics Core and Metabolomics Core of KU Leuven for data acquisition. This work was supported by funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program under grant agreement number 801747 (EcoBox) and by KU Leuven Internal funding for Interdisciplinary Networks (ID-N) PlasticDaphnia.
Author information
Authors and Affiliations
Contributions
BL, DRG and KF designed the experiments and BL and DRG carried them out. AK processed raw RNA-seq data. DG developed the kinetic model and performed simulations. KS measured metabolite concentrations. AG and KB gave advice on experimental design and interpretation. BL, DRG, KF and DG analyzed the results and BL, DRG, and KF wrote the manuscript. All authors read the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, B., Garza, D.R., Gonze, D. et al. Starvation responses impact interaction dynamics of human gut bacteria Bacteroides thetaiotaomicron and Roseburia intestinalis. ISME J 17, 1940–1952 (2023). https://doi.org/10.1038/s41396-023-01501-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01501-1