Abstract
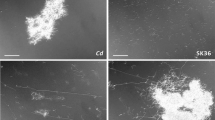
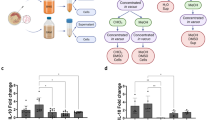
Membrane vesicles are produced by Gram-negative and Gram-positive bacteria. While membrane vesicles are potent elicitors of eukaryotic cells and involved in cell-cell communication, information is scarce about their general biology in the context of community members and the environment. Streptococcus sanguinis, a Gram-positive oral commensal, is prevalent in the oral cavity and well-characterized for its ability to antagonize oral pathobionts. We have found that production and dissemination of membrane vesicles by S. sanguinis is dependent on environmental and community factors. Co-culture with interacting commensal Corynebacterium durum, as well as with the periodontal pathobiont Filifactor alocis had no effect on S. sanguinis vesicle number and size, whereas the periodontal pathobiont Porphyromonas gingivalis abolished S. sanguinis vesicle production. Using both correlation and differential expression analyses to examine the transcriptomic changes underlying vesicle production, we found that differential expression of genes encoding proteins related to the cytoplasmic membrane and peptidoglycan correlate with the abundance of membrane vesicles. Proteomic characterizations of the vesicle cargo identified a variety of proteins, including those predicted to influence host interactions or host immune responses. Cell culture studies of gingival epithelial cells demonstrated that both crude and highly purified membrane vesicles could induce the expression of IL-8, TNF-α, IL-1β, and Gro-α within 6 hours of inoculation at levels comparable to whole cells. Our findings suggest that production of membrane vesicles by S. sanguinis is heavily influenced by community and environmental factors and plays an important role in communication with host cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
We are sorry, but there is no personal subscription option available for your country.
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All raw and normalized RNA sequence data and associated metadata are available on NCBI Gene Expression Omnibus (GEO), under accession GSE225861. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE [1] partner repository with the dataset identifier PXD041791.
References
Caruana JC, Walper SA. Bacterial membrane vesicles as mediators of microbe – microbe and microbe – host community interactions. Front Microbiol. 2020;11:432.
Work E, Knox KW, Vesk M. The chemistry and electron microscopy of an extracellular lipopolysaccharide from Escherichia coli. Ann N. Y Acad Sci. 1966;133:438–49.
Toyofuku M, Nomura N, Eberl L. Types and origins of bacterial membrane vesicles. Nat Rev Microbiol. 2019;17:13–24.
Cecil JD, O’Brien-Simpson NM, Lenzo JC, Holden JA, Chen YY, Singleton W, et al. Differential responses of pattern recognition receptors to outer membrane vesicles of three periodontal pathogens. PLoS ONE. 2016;11:e0151967.
Haurat MF, Aduse-Opoku J, Rangarajan M, Dorobantu L, Gray MR, Curtis MA, et al. Selective sorting of cargo proteins into bacterial membrane vesicles. J Biol Chem. 2011;286:1269–76.
Kim HY, Lim Y, An SJ, Choi BK. Characterization and immunostimulatory activity of extracellular vesicles from Filifactor alocis. Molecular Oral. Microbiology 2020;35:1–9.
Cecil JD, Sirisaengtaksin N, O’Brien-Simpson NM, Krachler AM. Outer membrane vesicle-host cell interactions. Microbiol Spectrum. 2019;7. https://doi.org/10.1128/microbiolspec.PSIB-0001-2018
Farrugia C, Stafford GP, Murdoch C. Porphyromonas gingivalis outer membrane vesicles increase vascular permeability. J Dent Res. 2020;99:1494–501.
Treerat P, Redanz U, Redanz S, Giacaman RA, Merritt J, Kreth J. Synergism between Corynebacterium and Streptococcus sanguinis reveals new interactions between oral commensals. ISME J. 2020;14:1154–69.
Roier S, Zingl FG, Cakar F, Durakovic S, Kohl P, Eichmann TO, et al. A novel mechanism for the biogenesis of outer membrane vesicles in Gram-negative bacteria. Nat Commun. 2016;7:10515.
Lee E-Y, Choi D-Y, Kim D-K, Kim J-W, Park JO, Kim S, et al. Gram-positive bacteria produce membrane vesicles: Proteomics-based characterization of Staphylococcus aureus-derived membrane vesicles. Proteomics. 2009;9:5425–36. https://doi.org/10.1002/pmic.200900338
Díaz-Garrido N, Badia J, Baldomà L. Microbiota-derived extracellular vesicles in interkingdom communication in the gut. J Extracell Vesicles. 2021;10:e12161.
Schlatterer K, Beck C, Hanzelmann D, Lebtig M, Fehrenbacher B, Schaller M, et al. The mechanism behind bacterial lipoprotein release: Phenol-soluble modulins mediate toll-like receptor 2 activation via extracellular vesicle release from Staphylococcus aureus. mBio 2018;9:1–13.
Andreoni F, Toyofuku M, Menzi C, Kalawong R, Shambat SM, François P, et al. Antibiotics Stimulate Formation of Vesicles in Staphylococcus aureus in both Phage-Dependent and-Independent Fashions and via Different Routes. 2019. https://doi.org/10.1128/AAC
Liu H, Zhang Q, Wang S, Weng W, Jing Y, Su J. Bacterial extracellular vesicles as bioactive nanocarriers for drug delivery: Advances and perspectives. Bioactive Materials. 2022;14:169–81. https://doi.org/10.1016/j.bioactmat.2021.12.006
Toyofuku M, Cárcamo-Oyarce G, Yamamoto T, Eisenstein F, Hsiao CC, Kurosawa M, et al. Prophage-triggered membrane vesicle formation through peptidoglycan damage in Bacillus subtilis. Nat Commun 2017;8:481.
Lee JH, Choi CW, Lee T, Kim SI, Lee JC, Shin JH. Transcription factor σB plays an important role in the production of extracellular membrane-derived vesicles in Listeria monocytogenes. PLoS ONE. 2013;8:e73196.
Resch U, Tsatsaronis JA, Le Rhun A, Stübiger G, Rohde M, Kasvandik S, et al. A two-component regulatory system impacts extracellular membrane-derived vesicle production in group a streptococcus. mBio 2016;7:e00207–16.
Welch JLM, Rossetti BJ, Rieken CW, Dewhirst FE, Borisy GG. Biogeography of a human oral microbiome at the micron scale. Proc Natl Acad Sci USA. 2016;113:E791–800.
Kreth J, Zhang Y, Herzberg MC. Streptococcal antagonism in oral biofilms: Streptococcus sanguinis and Streptococcus gordonii interference with Streptococcus mutans. J Bacteriol. 2008;190:4632–40.
Redanz S, Treerat P, Mu R, Redanz U, Zou Z, Koley D, et al. Pyruvate secretion by oral streptococci modulates hydrogen peroxide dependent antagonism. ISME J. 2020;14:1074–88.
Redanz U, Redanz S, Treerat P, Prakasam S, Lin L-J, Merritt J, et al. Differential response of oral mucosal and gingival cells to Corynebacterium durum, Streptococcus sanguinis, and Porphyromonas gingivalis Multispecies Biofilms. Front Cell Infect Microbiol 2021;11:686479 https://doi.org/10.3389/fcimb.2021.686479
Callahan JE, Munro CL, Kitten T. The Streptococcus sanguinis competence regulon is not required for infective endocarditis virulence in a rabbit model. PLoS ONE. 2011;6:e26403.
Crump KE, Bainbridge B, Brusko S, Turner LS, Ge X, Stone V, et al. The relationship of the lipoprotein SsaB, manganese and superoxide dismutase in Streptococcus sanguinis virulence for endocarditis. Mol Microbiol. 2014;92:1243–59.
Ge X, Kitten T, Chen Z, Lee SP, Munro CL, Xu P. Identification of Streptococcus sanguinis genes required for biofilm formation and examination of their role in endocarditis virulence. Infect Immun. 2008;76:2551–9.
Martini AM, Moricz BS, Ripperger AK, Tran PM, Sharp ME, Forsythe AN, et al. Association of novel Streptococcus sanguinis virulence factors with pathogenesis in a native valve infective endocarditis model. Front Microbiol. 2020;11:10.
Van De Rijn I, Kessler RE. Growth characteristics of group a streptococci in a new chemically defined medium. Infection Immunity. 1980;27:444–8. https://journals.asm.org/journal/iai
Standar K, Kreikemeyer B, Redanz S, Münter WL, Laue M, Podbielski A. Setup of an in vitro test system for basic studies on biofilm behavior of mixed-species cultures with dental and periodontal pathogens. PLoS ONE. 2010;5:e13135.
Dos Santos KCG, Desgagné-Penix I, Germain H. Custom selected reference genes outperform pre-defined reference genes in transcriptomic analysis. BMC Genomics. 2020;21:35.
Darwiche R, Struhl K. Pheno-RNA, a method to associate genes with a specific phenotype, identifies genes linked to cellular transformation. Proc Natl Acad Sci USA. 2020;117:28925–9. www.pnas.org/cgi/doi/10.1073/pnas.2014165117
Liu J, Sun L, Liu W, Guo L, Liu Z, Wei X, et al. A nuclease from Streptococcus mutans facilitates biofilm dispersal and escape from killing by neutrophil extracellular traps. Front Cell Infect Microbiol 2017;7:97.
Rahman MM, Hunter HN, Prova S, Verma V, Qamar A, Golemi-Kotra D. The Staphylococcus aureus methicillin resistance factor FmtA is a D-amino esterase that acts on teichoic acids. mBio 2016;7:02070–15.
Garg A, Gupta D. VirulentPred: A SVM based prediction method for virulent proteins in bacterial pathogens. BMC Bioinforma. 2008;9:62.
Jeffery CJ. Protein moonlighting: What is it, and why is it important? Philos Trans R Soc B: Biol Sci. 2018;373:20160523.
C Chen, H Liu, S Zabad, N Rivera, E Rowin, M Hassan et al. MoonProt 3.0: an update of the moonlighting proteins database, Nucleic Acids Research, 49, 8 January 2021, D368-D372, https://doi.org/10.1093/nar/gkaa1101
Papatheodorou I, Oellrich A, Smedley D. Linking gene expression to phenotypes via pathway information. J Biomed Semant. 2015;6:17.
Balibar CJ, Shen X, Tao J. The mevalonate pathway of Staphylococcus aureus. J Bacteriol. 2009;191:851–61.
Matsumoto Y, Yasukawa J, Ishii M, Hayashi Y, Miyazaki S, Sekimizu K. A critical role of mevalonate for peptidoglycan synthesis in Staphylococcus aureus. Sci Rep. 2016;6:22894.
Mehanny M, Koch M, Lehr CM, Fuhrmann G. Streptococcal extracellular membrane vesicles are rapidly internalized by immune cells and alter their cytokine release. Front Immunol. 2020;11:80.
Kent Brown C, Gu ZY, Matsuka YV, Purushothaman SS, Winter LA, Patrick Cleary P, et al. Structure of the streptococcal cell wall C5a peptidase. 2005. https://www.pnas.org
Blue CE, Paterson GK, Kerr AR, Bergé M, Claverys JP, Mitchell TJ. ZmpB, a novel virulence factor of Streptococcus pneumoniae that induces tumor necrosis factor alpha production in the respiratory tract. Infect Immun. 2003;71:4925–35.
Jones MN, Holt RG. Cloning and characterization of an alpha-enolase of the oral pathogen Streptococcus mutans that binds human plasminogen. Biochem Biophys Res Commun. 2007;364:924–9. https://doi.org/10.1016/j.bbrc.2007.10.098
Candela M, Bergmann S, Vici M, Vitali B, Turroni S, Eikmanns BJ, et al. Binding of human plasminogen to Bifidobacterium. J Bacteriol. 2007;189:5929–36.
Kinnby B, Booth NA, Svensäter G. Plasminogen binding by oral streptococci from dental plaque and inflammatory lesions. Microbiology 2008;154:924–31.
Boone TJ, Burnham CAD, Tyrrell GJ. Binding of group B streptococcal phosphoglycerate kinase to plasminogen and actin. Microb Pathogenesis. 2011;51:255–61.
Katakura Y, Sano R, Hashimoto T, Ninomiya K, Shioya S. Lactic acid bacteria display on the cell surface cytosolic proteins that recognize yeast mannan. Appl Microbiol Biotechnol. 2010;86:319–26.
Alkandari SA, Bhardwaj RG, Ellepola A, Karched M. Proteomics of extracellular vesicles produced by Granulicatella adiacens, which causes infective endocarditis. PLoSONE. 2020;15:e0227657 https://doi.org/10.1371/journal.pone.0227657
Bickel M. The role of interleukin-8 in inflammation and mechanisms of regulation. J Periodontol. 1993;64:456–60.
Lan C, Chen S, Jiang S, Lei H, Cai Z, Huang X, et al. Different expression patterns of inflammatory cytokines induced by lipopolysaccharides from Escherichia coli or Porphyromonas gingivalis in human dental pulp stem cells. BMC Oral Health. 2022;22:121 https://doi.org/10.1186/s12903-022-02161-x
Liew FY, Xu D, Brint EK, O’Neill LA. Negative regulation of toll-like receptor-mediated immune responses. Nat Rev Immunol. 2005;5:446–58.
Takashima K, Matsunaga N, Yoshimatsu M, Hazeki K, Kaisho T, Uekata M, et al. Analysis of binding site for the novel small-molecule TLR4 signal transduction inhibitor TAK-242 and its therapeutic effect on mouse sepsis model. Br J Pharm. 2009;157:1250–62.
la Rosa GRM, Gattuso G, Pedullà E, Rapisarda E, Nicolosi D, Salmeri M. Association of oral dysbiosis with oral cancer development (Review). Oncol Lett. 2020;19:3045–58.
Shen X, Zhang B, Hu X, Li J, Wu M, Yan C, et al. Neisseria sicca and Corynebacterium matruchotii inhibited oral squamous cell carcinomas by regulating genome stability. Bioengineered 2022;13:14094–106.
Yerneni SS, Werner S, Azambuja JH, Ludwig N, Eutsey R, Lucas PC, et al. Pneumococcal extracellular vesicles modulate host immunity. mBio 2021;12:0165721.
Bik E, Long C, Armitage G, et al. Bacterial diversity in the oral cavity of 10 healthy individuals. ISME J. 2010;4:962–74. https://doi.org/10.1038/ismej.2010.30
van Niel G, D’Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol. 2018;19:213–28. https://doi.org/10.1038/nrm.2017.125
Moffatt-Jauregui CE, Robinson B, de Moya AV, Brockman RD, Roman AV, Cash MN, et al. Establishment and characterization of a telomerase immortalized human gingival epithelial cell line. J Periodontal Res. 2013;48:713–21.
Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–5. https://doi.org/10.1038/nmeth.2089
Wickham H (2016). ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York. ISBN 978-3-319-24277-4, https://ggplot2.tidyverse.org
Pohlert T (2022). _PMCMRplus: Calculate Pairwise Multiple Comparisons of Mean Rank Sums Extended_.
MARTIN, Marcel. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal, [S.l.], 17, 10-2, may 2011. ISSN 2226-6089. https://doi.org/10.14806/ej.17.1.200.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–60.
Anders S, Pyl PT, Huber W. HTSeq-A Python framework to work with high-throughput sequencing data. Bioinformatics 2015;31:166–9.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: An open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Acknowledgements
We would like to thank the following: OHSU Massive Parallel Sequencing Core (Robert Searles, Amy Carlos), OHSU Microscropy Facility (Claudia Lopez, Steven Adamou), OHSU Imaging Facility (Stefanie Kaech Petrie) and Oregon State University Proteomics Core Facility (Stanislau Stanisheuski). We are grateful to Özlem Yilmaz (Medical University of South Carolina) for sharing F. alocis, Richard J. Lamont (University of Louisville) for sharing P. gingivalis gingipain mutants, and Delaney Shea for assistance with nanoparticle tracking analysis. This work was supported by an NIH-NIDCR grant DE021726, DE029492 and DE029612 to JK and NIH-NIDCR grant DE028252 to J.M.
Author information
Authors and Affiliations
Contributions
E.H., J.M., and J.K .developed and designed the research. EH performed the experiments. E.H., D.C., and JK analyzed data. E.H. drafted the manuscript. E.H., D.C., J.M. and J.K. edited and revised the manuscript. All authors reviewed and approved the results and the revisions made to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Helliwell, E., Choi, D., Merritt, J. et al. Environmental influences on Streptococcus sanguinis membrane vesicle biogenesis. ISME J 17, 1430–1444 (2023). https://doi.org/10.1038/s41396-023-01456-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01456-3