Abstract
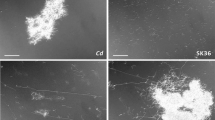
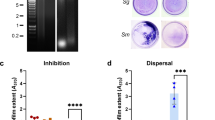
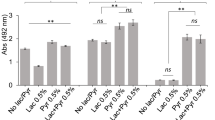
During oral biofilm development, interspecies interactions drive species distribution and biofilm architecture. To understand what molecular mechanisms determine these interactions, we used information gained from recent biogeographical investigations demonstrating an association of corynebacteria with streptococci. We previously reported that Streptococcus sanguinis and Corynebacterium durum have a close relationship through the production of membrane vesicle and fatty acids leading to S. sanguinis chain elongation and overall increased fitness supporting their commensal state. Here we present the molecular mechanisms of this interspecies interaction. Coculture experiments for transcriptomic analysis identified several differentially expressed genes in S. sanguinis. Due to its connection to fatty acid synthesis, we focused on the glycerol-operon. We further explored the differentially expressed type IV pili genes due to their connection to motility and biofilm adhesion. Gene inactivation of the glycerol kinase glpK had a profound impact on the ability of S. sanguinis to metabolize C. durum secreted glycerol and impaired chain elongation important for their interaction. Investigations on the effect of type IV pili revealed a reduction of S. sanguinis twitching motility in the presence of C. durum, which was caused by a decrease in type IV pili abundance on the surface of S. sanguinis as determined by SEM. In conclusion, we identified that the ability to metabolize C. durum produced glycerol is crucial for the interaction of C. durum and S. sanguinis. Reduced twitching motility could lead to a closer interaction of both species, supporting niche development in the oral cavity and potentially shaping symbiotic health-associated biofilm communities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
We are sorry, but there is no personal subscription option available for your country.
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data are available in the main text and the supplementary materials. Raw and processed sequencing data can be found under GSE230560 in the NCBI GEO data repository.
References
Colwell RK, Rangel TF. Hutchinson’s duality: the once and future niche. Proc Natl Acad Sci USA. 2009;106:19651–58.
Kreth J, Merritt J, Qi F, Dong X, Shi W. Antagonistic, synergistic, and counteroffensive strategies for streptococcal interspecies interactions. In: Kolenbrander PE, editor. Oral microbial communities: genomic inquiry and interspecies communication. Washington, DC: ASM Press; 2011. p. 331–44.
Kreth J, Merritt J, Shi W, Qi F. Co-ordinated bacteriocin production and competence development: a possible mechanism for taking up DNA from neighbouring species. Mol Microbiol. 2005;57:392–404.
Roberts AP, Kreth J. The impact of horizontal gene transfer on the adaptive ability of the human oral microbiome. Front Cell Infect Microbiol. 2014;4:124.
Ona L, Giri S, Avermann N, Kreienbaum M, Thormann KM, Kost C. Obligate cross-feeding expands the metabolic niche of bacteria. Nat Ecol Evol. 2021;5:1224–32.
Mark Welch JL, Dewhirst FE, Borisy GG. Biogeography of the oral microbiome: the site-specialist hypothesis. Annu Rev Microbiol. 2019;73:335–58.
Mark Welch JL, Rossetti BJ, Rieken CW, Dewhirst FE, Borisy GG. Biogeography of a human oral microbiome at the micron scale. Proc Natl Acad Sci USA. 2016;113:E791–E800.
Freire M, Nelson KE, Edlund A. The oral host-microbial interactome: an ecological chronometer of health? Trends Microbiol. 2021;29:551–61.
Mira A, Simon-Soro A, Curtis MA. Role of microbial communities in the pathogenesis of periodontal diseases and caries. J Clin Periodontol. 2017;44:S23–S38.
Treerat P, Redanz U, Redanz S, Giacaman RA, Merritt J, Kreth J. Synergism between Corynebacterium and Streptococcus sanguinis reveals new interactions between oral commensals. ISME J. 2020;14:1154–69.
Morillo-Lopez V, Sjaarda A, Islam I, Borisy GG, Mark, Welch JL. Corncob structures in dental plaque reveal microhabitat taxon specificity. Microbiome. 2022;10:145.
Redanz S, Cheng X, Giacaman RA, Pfeifer CS, Merritt J, Kreth J. Live and let die: hydrogen peroxide production by the commensal flora and its role in maintaining a symbiotic microbiome. Mol Oral Microbiol. 2018;33:337–52.
Socransky SS, Dzink JL, Smith CM. Chemically defined medium for oral microorganisms. J Clin Microbiol. 1985;22:303–5.
He X, Wu C, Yarbrough D, Sim L, Niu G, Merritt J, et al. The cia operon of Streptococcus mutans encodes a unique component required for calcium-mediated autoregulation. Mol Microbiol. 2008;70:112–26.
Taciak B, Białasek M, Braniewska A, Sas Z, Sawicka P, Kiraga Ł, et al. Evaluation of phenotypic and functional stability of RAW 264.7 cell line through serial passages. PLoS ONE. 2018;13:e0198943.
Sambrook J, Fritsch, EF, Maniatis, T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989.
Cullin N, Redanz S, Lampi KJ, Merritt J, Kreth J. Murein hydrolase LytF of Streptococcus sanguinis and the ecological consequences of competence development. Appl Environ Microbiol. 2017;83:e01709–e017017.
Zheng L, Chen Z, Itzek A, Ashby M, Kreth J. Catabolite control protein A controls hydrogen peroxide production and cell death in Streptococcus sanguinis. J Bacteriol. 2011;193:516–26.
Zheng X, Zheng H, Lan R, Ye C, Wang Y, Zhang J, et al. Identification of genes and genomic islands correlated with high pathogenicity in Streptococcus suis using whole genome tiling microarrays. PLoS ONE. 2011;6:e17987.
Chen Y-YM, Chiang Y-C, Tseng T-Y, Wu H-Y, Chen Y-Y, Wu C-H, et al. Molecular and functional analysis of the type IV pilus gene cluster in Streptococcus sanguinis SK36. Appl Environ Microbiol. 2019;85:e02788–e027818.
Gurung I, Spielman I, Davies MR, Lala R, Gaustad P, Biais N, et al. Functional analysis of an unusual type IV pilus in the Gram-positive Streptococcus sanguinis. Mol Microbiol. 2016;99:380–92.
Martini AM, Moricz BS, Woods LJ, Jones BD. Type IV pili of Streptococcus sanguinis contribute to pathogenesis in experimental infective endocarditis. Microbiol Spectr. 2021;9:e0175221.
Lasica AM, Ksiazek M, Madej M, Potempa J. The type IX secretion system (T9SS): highlights and recent insights into its structure and function. Front Cell Infect Microbiol. 2017;7:215.
Berry J-L, Gurung I, Anonsen JH, Spielman I, Harper E, Hall AMJ, et al. Global biochemical and structural analysis of the type IV pilus from the Gram-positive bacterium Streptococcus sanguinis. J Biol Chem. 2019;294:6796–808.
Charrier V, Buckley E, Parsonage D, Galinier A, Darbon E, Jaquinod M, et al. Cloning and sequencing of two enterococcal glpK genes and regulation of the encoded glycerol kinases by phosphoenolpyruvate-dependent, phosphotransferase system-catalyzed phosphorylation of a single histidyl residue. J Biol Chem. 1997;272:14166–74.
Liu WZ, Faber R, Feese M, Remington SJ, Pettigrew DW. Escherichia coli glycerol kinase: role of a tetramer interface in regulation by fructose 1,6-bisphosphate and phosphotransferase system regulatory protein IIIglc. Biochemistry. 1994;33:10120–6.
Pettigrew DW, Liu WZ, Holmes C, Meadow ND, Roseman S. A single amino acid change in Escherichia coli glycerol kinase abolishes glucose control of glycerol utilization in vivo. J Bacteriol. 1996;178:2846–52.
Yeh JI, Kettering R, Saxl R, Bourand A, Darbon E, Joly N, et al. Structural characterizations of glycerol kinase: unraveling phosphorylation-induced long-range activation. Biochemistry. 2009;48:346–56.
Weissenborn DL, Wittekindt N, Larson TJ. Structure and regulation of the glpFK operon encoding glycerol diffusion facilitator and glycerol kinase of Escherichia coli K-12. J Biol Chem. 1992;267:6122–31.
López-Lara IM, Geiger O. Bacterial lipid diversity. Biochim Biophys Acta Mol Cell Biol Lipids. 2017;1862:1287–99.
Sohlenkamp C, Geiger O. Bacterial membrane lipids: diversity in structures and pathways. FEMS Microbiol Rev. 2015;40:133–59.
Zhang YM, Rock CO. Membrane lipid homeostasis in bacteria. Nat Rev Microbiol. 2008;6:222–33.
Arora A, Gupta CM. Glycerol backbone conformation in phosphatidylcholines is primarily determined by the intramolecular stacking of the vicinally arranged acyl chains. Biochim Biophys Acta Biomembr. 1997;1324:47–60.
Pettigrew DW, Ma DP, Conrad CA, Johnson JR. Escherichia coli glycerol kinase. Cloning and sequencing of the glpK gene and the primary structure of the enzyme. J Biol Chem. 1988;263:135–9.
Yeh JI, Charrier V, Paulo J, Hou L, Darbon E, Claiborne A, et al. Structures of Enterococcal glycerol kinase in the absence and presence of glycerol: correlation of conformation to substrate binding and a mechanism of activation by phosphorylation. Biochemistry. 2004;43:362–73.
Bizzini A, Zhao C, Budin-Verneuil A, Sauvageot N, Giard JC, Auffray Y, et al. Glycerol is metabolized in a complex and strain-dependent manner in Enterococcus faecalis. J Bacteriol. 2010;192:779–85.
Darbon E, Servant P, Poncet S, Deutscher J. Antitermination by GlpP, catabolite repression via CcpA and inducer exclusion triggered by P-GlpK dephosphorylation control Bacillus subtilis glpFK expression. Mol Microbiol. 2002;43:1039–52.
Deutscher J, Bauer B, Sauerwald H. Regulation of glycerol metabolism in Enterococcus faecalis by phosphoenolpyruvate-dependent phosphorylation of glycerol kinase catalyzed by enzyme I and HPr of the phosphotransferase system. J Bacteriol. 1993;175:3730–3.
Jahreis K, Pimentel-Schmitt EF, Brückner R, Titgemeyer F. Ins and outs of glucose transport systems in eubacteria. FEMS Microbiol Rev. 2008;32:891–907.
Parsons JB, Rock CO. Bacterial lipids: metabolism and membrane homeostasis. Prog Lipid Res. 2013;52:249–76.
Doi Y. Glycerol metabolism and its regulation in lactic acid bacteria. Appl Microbiol Biotechnol. 2019;103:5079–93.
Keevil CW, Williamson MI, Marsh PD, Ellwood DC. Evidence that glucose and sucrose uptake in oral streptococcal bacteria involves independent phosphotransferase and proton-motive force-mediated mechanisms. Arch Oral Biol. 1984;29:871–8.
Redanz S, Masilamani R, Cullin N, Giacaman RA, Merritt J, Kreth J. Distinct regulatory role of carbon catabolite protein A (CcpA) in oral streptococcal spxB expression. J Bacteriol. 2018;200:e00619–e006117.
Craig L, Forest KT, Maier B. Type IV pili: dynamics, biophysics and functional consequences. Nat Rev Microbiol. 2019;17:429–40.
Danne C, Dramsi S. Pili of Gram-positive bacteria: roles in host colonization. Res Microbiol. 2012;163:645–58.
Proft T, Baker EN. Pili in Gram-negative and Gram-positive bacteria - structure, assembly and their role in disease. Cell Mol Life Sci. 2009;66:613–35.
Gurung I, Berry J-L, Hall AMJ, Pelicic V. Cloning-independent markerless gene editing in Streptococcus sanguinis: novel insights in type IV pilus biology. Nucleic Acids Res. 2017;45:e40.
Zähner D, Gandhi AR, Stuchlik O, Reed M, Pohl J, Stephens DS. Pilus backbone protein PitB of Streptococcus pneumoniae contains stabilizing intramolecular isopeptide bonds. Biochem Biophys Res Commun. 2011;409:526–31.
Rozman V, Accetto T, Duncan SH, Flint HJ, Vodovnik M. Type IV pili are widespread among non-pathogenic Gram-positive gut bacteria with diverse carbohydrate utilization patterns. Environ Microbiol. 2021;23:1527–40.
Okahashi N, Nakata M, Terao Y, Isoda R, Sakurai A, Sumitomo T, et al. Pili of oral Streptococcus sanguinis bind to salivary amylase and promote the biofilm formation. Micro Pathog. 2011;50:148–54.
Raynaud C, Sheppard D, Berry J-L, Gurung I, Pelicic V. PilB from Streptococcus sanguinis is a bimodular type IV pilin with a direct role in adhesion. Proc Natl Acad Sci USA. 2021;118:e2102092118.
Piepenbrink KH, Sundberg EJ. Motility and adhesion through type IV pili in Gram-positive bacteria. Biochem Soc Trans. 2016;44:1659–66.
Varga JJ, Nguyen V, O’Brien DK, Rodgers K, Walker RA, Melville SB. Type IV pili-dependent gliding motility in the Gram-positive pathogen Clostridium perfringens and other Clostridia. Mol Microbiol. 2006;62:680–94.
Baker SP, Nulton TJ, Kitten T. Genomic, phenotypic, and virulence analysis of Streptococcus sanguinis oral and infective-endocarditis isolates. Infect Immun. 2018;87:e00703–e007018.
Bishop CJ, Aanensen DM, Jordan GE, Kilian M, Hanage WP, Spratt BG. Assigning strains to bacterial species via the internet. BMC Biol. 2009;7:3.
Kilian M, Mikkelsen L, Henrichsen J. Taxonomic study of viridans streptococci: Description of Streptococcus gordonii sp. nov. and emended descriptions of Streptococcus sanguis (White and Niven 1946), Streptococcus oralis (Bridge and Sneath 1982), and Streptococcus mitis (Andrewes and Horder 1906). Int J Syst Evol Microbiol. 1989;39:471–84.
Kitten T, Munro CL, Zollar NQ, Lee SP, Patel RD. Oral streptococcal bacteremia in hospitalized patients: taxonomic identification and clinical characterization. J Clin Microbiol. 2012;50:1039–42.
Xu P, Alves JM, Kitten T, Brown A, Chen Z, Ozaki LS, et al. Genome of the opportunistic pathogen Streptococcus sanguinis. J Bacteriol. 2007;189:3166–75.
Willis JR, Gabaldón T. The human oral microbiome in health and disease: from sequences to ecosystems. Microorganisms. 2020;8:308.
Peng X, Cheng L, You Y, Tang C, Ren B, Li Y, et al. Oral microbiota in human systematic diseases. Int J Oral Sci. 2022;14:14.
Scannapieco FA. The oral microbiome: Its role in health and in oral and systemic infections. Clin Microbiol Newsl. 2013;35:163–9.
Kelly MS, Plunkett C, Yu Y, Aquino JN, Patel SM, Hurst JH, et al. Non-diphtheriae Corynebacterium species are associated with decreased risk of pneumococcal colonization during infancy. ISME J. 2022;16:655–65.
Barrett SL, Cookson BT, Carlson LC, Bernard KA, Coyle MB. Diversity within reference strains of Corynebacterium matruchotii includes Corynebacterium durum and a novel organism. J Clin Microbiol. 2001;39:943–8.
Esders TW, Michrina CA. Purification and properties of L-alpha-glycerophosphate oxidase from Streptococcus faecium ATCC 12755. J Biol Chem. 1979;254:2710–5.
Mahdi LK, Higgins MA, Day CJ, Tiralongo J, Hartley-Tassell LE, Jennings MP, et al. The pneumococcal alpha-glycerophosphate oxidase enhances nasopharyngeal colonization through binding to host glycoconjugates. EBioMedicine. 2017;18:236–43.
Maurel C, Reizer J, Schroeder JI, Chrispeels MJ, Saier MH. Functional characterization of the Escherichia coli glycerol facilitator, GlpF, in Xenopus oocytes. J Biol Chem. 1994;269:11869–72.
Brisson D, Vohl MC, St-Pierre J, Hudson TJ, Gaudet D. Glycerol: a neglected variable in metabolic processes? Bioessays. 2001;23:534–42.
McCabe ERB, Sadava D, Bullen WW, McKelvey HA, Seltzer WK, Rose CI. Human glycerol kinase deficiency: Enzyme kinetics and fibroblast hybridization. J Inherit Metab Dis. 1982;5:177–82.
Pavlik P, Simon M, Schuster T, Ruis H. The glycerol kinase (GUT1) gene of Saccharomyces cerevisiae: cloning and characterization. Curr Genet. 1993;24:21–25.
Pawlyk AC, Pettigrew DW. Subcloning, expression, purification, and characterization of Haemophilus influenzae glycerol kinase. Protein Expr Purif. 2001;22:52–59.
Seltzer WK, McCabe ER. Human and rat adrenal glycerol kinase: subcellular distribution and bisubstrate kinetics. Mol Cell Biochem. 1984;62:43–50.
Wilk P, Kuśka K, Wątor E, Małecki PH, Woś K, Tokarz P, et al. Structural characterization of glycerol kinase from the thermophilic fungus Chaetomium thermophilum. Int J Mol Sci. 2020;21:9570.
Blötz C, Stülke J. Glycerol metabolism and its implication in virulence in Mycoplasma. FEMS Microbiol Rev. 2017;41:640–52.
Yao J, Rock CO. How bacterial pathogens eat host lipids: Implications for the development of fatty acid synthesis therapeutics. J Biol Chem. 2015;290:5940–6.
Gründling A, Schneewind O. Synthesis of glycerol phosphate lipoteichoic acid in Staphylococcus aureus. Proc Natl Acad Sci USA. 2007;104:8478–83.
Wörmann ME, Corrigan RM, Simpson PJ, Matthews SJ, Gründling A. Enzymatic activities and functional interdependencies of Bacillus subtilis lipoteichoic acid synthesis enzymes. Mol Microbiol. 2011;79:566–83.
Staerck C, Wasselin V, Budin-Verneuil A, Rincé I, Cacaci M, Weigel M, et al. Analysis of glycerol and dihydroxyacetone metabolism in Enterococcus faecium. FEMS Microbiol Lett. 2021;368:fnab043.
Kvalheim SF, Xenaki V, Kvalheim A, Lie S-A, Marthinussen MC, Strand GV, et al. Effect of glycerol on reconstructed human oral mucosa. Eur J Oral Sci. 2019;127:19–26.
Treerat P, McGuire B, Palmer E, Dahl EM, Karstens L, Merritt J, et al. Oral microbiome diversity: The curious case of Corynebacterium sp. isolation. Mol Oral Microbiol. 2022;37:167–79.
Tsuzukibashi O, Uchibori S, Shinozaki-Kuwahara N, Kobayashi T, Takada K, Hirasawa M. A selective medium for the isolation of Corynebacterium species in oral cavities. J Microbiol Methods. 2014;104:67–71.
Forslund A-L, Salomonsson EN, Golovliov I, Kuoppa K, Michell S, Titball R, et al. The type IV pilin, PilA, is required for full virulence of Francisella tularensis subspecies tularensis. BMC Microbiol. 2010;10:227.
Källström H, Islam MS, Berggren P-O, Jonsson A-B. Cell signaling by the type IV pili of pathogenic Neisseria. J Biol Chem. 1998;273:21777–82.
Acknowledgements
This work was supported by an NIH-NIDCR grant DE021726, DE029492, and DE029612 to JK and NIH-NIDCR grant DE028252 to JM. We thank Prof. Todd Kitten for kindly providing S. sanguinis SK strains, and Dr. Claudia S. López (Multiscale Microscopy Core, OHSU) for expert assistance in performing SEM with technical support from Center for Spatial Systems Biomedicine (OCSSB).
Author information
Authors and Affiliations
Contributions
PT, DA, JM and JK designed and conceptualized the research project. PT and DA performed experimental work and PT, DA, RAG and JK performed data analysis. All authors were involved in drafting, revising and finalizing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Treerat, P., Anderson, D., Giacaman, R.A. et al. Glycerol metabolism supports oral commensal interactions. ISME J 17, 1116–1127 (2023). https://doi.org/10.1038/s41396-023-01426-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01426-9