Abstract
In-depth understanding of metabolite-mediated plant-nematode interactions can guide us towards novel nematode management strategies. To improve our understanding of the effects of secondary metabolites on soil nematode communities, we grew Arabidopsis thaliana genetically altered in glucosinolate, camalexin, or flavonoid synthesis pathways, and analyzed their root-associated nematode communities using metabarcoding. To test for any modulating effects of the associated microbiota on the nematode responses, we characterized the bacterial and fungal communities. Finally, as a proxy of microbiome-modulating effects on nematode invasion, we isolated the root-associated microbiomes from the mutants and tested their effect on the ability of the plant parasitic nematode Meloidogyne incognita to penetrate tomato roots. Most mutants had altered relative abundances of several nematode taxa with stronger effects on the plant parasitic Meloidogyne hapla than on other root feeding taxa. This probably reflects that M. hapla invades and remains embedded within root tissues and is thus intimately associated with the host. When transferred to tomato, microbiomes from the flavonoid over-producing pap1-D enhanced M. incognita root-invasion, whereas microbiomes from flavonoid-deficient mutants reduced invasion. This suggests microbiome-mediated effect of flavonoids on Meloidogyne infectivity plausibly mediated by the alteration of the abundances of specific microbial taxa in the transferred microbiomes, although we could not conclusively pinpoint such causative microbial taxa.
Similar content being viewed by others
Background
Nematodes are the most abundant metazoans on Earth [1]. They are widely distributed and are broadly classified as root-feeding plant parasitic nematodes (PPNs), free-living bacterial or fungal feeders, predators, or omnivores [2]. It is estimated that PPNs cause more than 10% yield losses in crops worldwide [3]. PPNs are categorized as (i) ectoparasites feeding on root cells from outside the root; (ii) semi-endoparasites penetrating the roots with the anterior part of the body; (iii) migratory endoparasites moving destructively through root tissues while feeding on root cells; or (iv) sedentary endoparasites (i.e. root-knot nematodes (Meloidogyne spp.) and cyst nematode) in which the infective second-stage juvenile (J2) invades the root and migrates towards vascular cells forming permanent feeding sites [4]. Plant-PPN interactions are dependent on sophisticated chemical signaling, and a number of plant-synthesized metabolites such as glucosinolates, camalexins, and flavonoids released into the rhizosphere may attract [5, 6] or repel nematodes, or affect nematode development, reproduction, and survival [7, 8]. Glucosinolates are anionic thioglucosides containing N-hydroxysulfate, and they are important phytoanticipins in Brassicaceae [9]. Upon cellular disruption, e.g. wounding by nematodes orpathogen intrusion, the activity of the endogenous enzyme myrosinase hydrolyses the thioglucoside linkage, resulting in the formation of nitrogenous compounds including nitrile, isothiocyanate, thiocyanate, and epithionitrile [8, 9] that are active against pathogens and herbivores. For instance, it was shown that the transgenic Arabidopsis thaliana line CYP79A1 with an altered exogenous glucosinolate profile significantly affected root and rhizosphere associated microbial communities [10]. Camalexin (3-thiazol-2′-yl-indole) is a sulfur-containing phytoalexin in Arabidopsis thaliana (hereafter Arabidopsis) that is produced from tryptophan via indole-3 acetaldoxime from the glucosinolate pathway [11]. It is known to accumulate in response to biotic stressors such as herbivorous insects, fungal, and bacterial pathogens [12,13,14]. Root gall numbers of the root-knot nematode Meloidogyne incognita were higher in the Arabidopsis mutant myb34myb51 having reduced glucosinolate and camalexin concentrations, and pad3 mutant having negligible camalexin contents [15]. Similarly, numbers of the cyst nematode Heterodera schachtii were significantly higher in the two mutants pgip1-1 and cyp79B2cyp79B3, which do not produce glucosinolates or camalexin [16], while the pad3 mutant had more H. schachtii females, bigger syncytia and cysts [17]. Several studies have suggested that glucosinolates and camalexin affects PPNs negatively in planta, while other studies indicated that it is myrosinase-catalyzed glucosinolate breakdown-products that are the defense compounds acting against PPNs [18, 19].
Flavonoids are a large group of secondary metabolites [20] that are known to accumulate as aglycones and glycosides in root tips and root cap cells [21], and are actively exuded into the rhizosphere [22, 23]. They play important roles in plant root development, pigmentation, UV-light protection, pollinator attraction, the symbiosis between rhizobia and legumes, and signaling between plants and root-associated microbes [20]. The bitter taste of flavonoids renders plants with high flavonoid content unattractive to some herbivores [24]. Moreover, flavonoids are involved in the development of feeding sites of sedentary nematodes and in plant defense responses to PPNs [25]. For instance, they act as defense compounds against M. incognita in soybean [26] and chinaberry plants [27], H. glycines in soybean [28], Radopholus similis in banana roots [29], and Ditylenchus dipsaci in white clover [30]. In vitro assays similarly showed that flavonoids were nematicidal against M. incognita and R. similis [31, 32]. In addition, flavonoids have been reported to affect the growth rates of rhizosphere bacteria and fungi [33,34,35].
Most current understanding of plant metabolite-nematode interactions derives from studies involving single nematode species and plants grown in synthetic media or sand, or from in vitro bioassays. However, to elucidate mechanisms behind metabolite-mediated plant-nematode interactions in natural soils, we need both in vivo and in vitro studies [8]. Additionally, little is known about the effects of root metabolites on overall nematode communities, particularly free-living nematodes, despite their importance in soil ecosystems [36, 37]. Finally, root-associated microbial communities could modulate the complex interactions between roots, metabolites, and nematodes [38, 39], and thus analyses of the associated microbiomes in plant-nematode studies are highly relevant.
We hypothesized that genetic disruption of secondary metabolites synthesis would affect particularly endoparasitic nematodes forming intimate host associations. We also predicted that the associated microbial communities would modulate such interactions. To test these hypotheses, we characterized root-associated nematode communities from Arabidopsis mutants altered in glucosinolate, camalexin, and flavonoid biosynthesis, and their parental lines in a system using arable soil. Further, to elucidate the potential role of microbiomes in metabolite-nematode interactions, we analysed co-occurrence patterns between nematode and microbial taxa, and we tested the effects of microbiomes isolated from Arabidopsis mutants on M. incognita invasion capacity in its natural host tomato.
Materials and methods
Plant material
We used 14 well-characterized Arabidopsis lines carrying mutations in glucosinolate, camalexin, or flavonoid pathways, and their parental background lines Col-0 or Ler-0 as controls (Table 1). Arabidopsis parental lines and the mutants gsm1-1, gsm2-1, myb51, tgg1tgg2, pad2-1, pad3-1, tt3-1tt5-1, ttg1ttg2, and pap1-D were supplied by Nottingham Arabidopsis Stock Centre, UK. The cyp79B2, cyp79B3, and cyp79B2cyp79B3 mutants were kindly provided by Professor Judith Bender (Brown University, USA), while the tt3-1 mutant was supplied by Professor Wendy Ann Peer (University of Maryland, USA). The glucosinolate pathway, including the pathway leading to camalexin, and the flavonoid pathway are outlined in Supplementary Figs. 1, 2.
Experiment 1: Nematode communities and microbiomes associated with metabolite mutants
In this experiment, we collected root- and rhizosphere samples from Arabidopsis mutants and their respective parental lines to analyze the composition of nematode and microbial root/rhizosphere communities of the different genotypes.
Seeds of lines Col-0, Ler-0, cyp79B2, cyp79B3, cyp79B2cyp79B3, gsm1-1, gsm2-1, tgg2-1, tgg1tgg2, pad2-1, pad3-1, tt3-1, tt3-1tt5-1, and pap1-D (Table 1) were sown in pots with potting soil and vernalized at 4 °C for 4 days, before transfer to a glasshouse (AU Flakkebjerg, Slagelse, Denmark; 55.173912°N,11.25624°E) under 12/12 h light-dark and 22-24 °C. On day 16, plants of each line were transferred into four pots (five plants per pot) containing 400 g hand-mixed soil (sandy-clayey soil, pH 5.9) collected from a field previously cultivated with Faba bean (Skælskør, Denmark; 55.2521101°N,11.2942011°E). The field was infested with the root-knot nematode Meloidogyne hapla (Emma Skov, personal communication). After 40 days, we gently uprooted the plants and shook the roots to remove loosely adhering soil. Roots with remaining rhizosphere soil were placed in 2 ml Eppendorf tubes and flash-frozen in liquid nitrogen. We pooled roots from the five plants from each of four replicate pots. All samples were freeze-dried for 3 days, homogenized, and ground in a Geno/Grinder 2010 (RAMCON, Birkerød, Denmark) at 1500 rpm for 6 × 1 min. To analyze the composition of nematode and microbial communities, samples were stored at −20 °C until DNA extraction, followed by amplicon sequencing as described below (Supplementary Fig. 3).
Experiment 2: Analysis of the effects of root microbiomes from Arabidopsis lines on M. incognita root invasion
In Experiment 2, we assessed the effects of root microbiomes isolated from Arabidopsis lines selected from Experiment 1 (cyp79B2, cyp79B2cyp79B3, gsm1-1, tgg1tgg2, pad3-1, tt3-1tt5-1, pap1-D, and their parental lines), and an additional flavonoid mutant, ttg1ttg2 (Table 1) on M. incognita invasion of tomato roots. In overview, we collected root microbiomes from the different genotypes and transferred them to tomato roots before inoculating infective M. incognita juveniles (stage J2) to the tomato roots to assess their invasion success. Microorganisms, including microorganisms attaching to the surface of J2 juveniles, affect J2 mobility and vigour before the juveniles encounter and penetrate host roots [40]. To allow pre-infection interactions between microbiomes and J2s, we pre-exposed the J2s to the microbiomes from the different genotypes before inoculating them to tomato roots. In the following, we describe the details of the experimental procedures.
A set-up like the one in Experiment 1 was used to grow the Arabidopsis lines in 6 replicate pots, each containing 400 g of a batch of 80 kg hand-mixed soil collected from the same field as in Experiment 1 (Supplementary Fig. 4). After 40 days, we pooled the roots of five plants from each pot, including the attached rhizosphere soil. To isolate root microbiomes, we transferred roots to 30 ml of sterile 0.085% NaCl before shaking at 350 rpm for 50 min, followed by centrifugation at 500 × g for 5 min. The supernatant was passed through a 500 µm sieve to remove larger particles. The filtrate was centrifuged at 5000 x g for 10 min, and the pellet was resuspended in 15 ml 0.085% NaCl. We used these microbiome suspensions to evaluate their effect on M. incognita J2 invasion of tomato roots. To characterize the microbiomes by metabarcoding, we centrifuged 2 ml of each microbiome suspension at 10,000 x g for 10 min and stored the pellet at −80 °C until DNA extraction.
A culture of M. incognita was maintained on tomato plants in a glasshouse at Aarhus University, Flakkebjerg (55.173912°N,11.25624°E). Nematode eggs were harvested using a 1.5% NaOCl solution and purified by sucrose gradient centrifugation [41]. To hatch juveniles (stage J2), eggs were transferred to a Baermann tray, and freshly hatched J2s were collected every day for one week [42] and surface sterilized in a mixture of 200 mg/l streptomycin sulfate, 25 mg/l rifampicin, and 1× CellGuard (PanReac, AppliChem, Darmstadt, Germany) for 4 h [43]. After sterilization, the J2s were washed in sterile 0.085% NaCl.
We pre-incubated the J2s with the microbiomes isolated from different Arabidopsis mutants and their parental lines the day after the microbiomes were isolated. Approximately two thousand surface-sterilized J2s were transferred to individual wells of 24-well plates containing 2 ml of microbiome suspension from the different Arabidopsis lines. To sustain microbial activity during the pre-incubation, we added tryptic soy broth (TSB, VWR International, Søborg, Denmark) (1:300) to the wells. Plates were sealed with parafilm and incubated at 21 °C for 2 weeks with frequent aeration. Each treatment was replicated 6 times.
Tomato seeds were surface sterilized using 70% ethanol for 1 min and 2.5% NaOCl for 3 min, washed with sterile distilled water, and germinated on ½ MS media (Murashige & Skoog, Duchefa Biochemie, Haarlem, Netherlands) for 4 days. Seedlings were transplanted into pots containing 400 g of sand which had been autoclaved at 121 °C for 90 min. At the same time, seedlings were drenched with 5 ml of microbiome suspension of each Arabidopsis line. We included a control treatment without microbiome addition. We used six replications of each treatment. Eleven days after transfer to pots, tomato seedlings were inoculated with approximately one thousand pre-incubated J2s by transferring the nematodes to three holes around the shoot base using a pipette. In addition, we added 5 ml of microbiome suspension around the roots of each plant. After one week, we removed plants from the pots and stained the whole root system of each plant with acid fuchsin to count the total numbers of invaded J2s using a stereomicroscope [42]. Data were checked for normality and homogeneity of variance in R [44]. One-way ANOVA was performed to identify significant effects of treatments, followed by pairwise comparisons between treatments using the Tukey test in R.
DNA extraction, library preparation, and amplicon sequencing (Experiments 1 and 2)
We extracted DNA from 250 mg of each of the freeze-dried root samples (Experiment 1), and from the pellets of 2 ml of microbiome suspension (Experiment 2) using a PowerLyzer soil DNA extraction kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions, except that samples were initially homogenized in a Geno/Grinder 2010 at 1500 rpm for 3 × 30 seconds to improve DNA extraction efficiency. The DNA concentration of each sample was measured using a Qubit fluorometer (Thermo Fisher Scientific, Waltham, Massachusetts, USA). Equal amounts of DNA from each sample were used to profile nematode, bacterial, and fungal communities.
The first PCR was performed in a reaction mixture of 25 μl consisting of 5 µl PCR buffer (Promega 5X, Promega, Madison, USA), 1.5 µl MgCl2 (25 mM), 2 µl dNTPs (2.5 mM), 0.5 μl of each primer (10 µM), 0.1 µl GoTaq Flexi polymerase (5U, Promega), 2 µl DNA template, and 13.4 µl water. We amplified the V6-V8 region of the 18 S rRNA gene using the nematode primers Nemf, 5′-GGGGAAGTATGGTTGCAAA-3′ and 18Sr2b, 5′-TACAAAGGGCAGGGACGTAAT-3′ [45, 46]. This primer pair efficiently amplifies the diversity of root-associated soil nematodes [47]. We amplified the fungal ITS2 region using fITS7, 5´-GTGARTCATCGAATCTTTG-3´ and ITS4, 5´-TCCTCCGCTTATTGATATGC-3´ [48], and the V3-V4 regions of the 16 S rRNA gene using S-D-Bact-0341-b-S-17, 5´-CCTACGGGNGGCWGCAG-3´ and S-D-Bact-0785-a-A-21, 5´-GACTACHVGGGTATCTAATCC-3´ [49]. The PCR conditions were 94 °C for 5 min followed by 35 cycles of 94 °C for 1 min, 53 °C for 30 s, 72 °C for 1 min for nematodes, 25 cycles of 94 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min for bacteria, 33 cycles of 94 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min for fungi, followed by a final extension step at 72 °C for 10 min, and 4 °C on hold.
The master mix of the second PCR was identical to that of the first PCR, except that 2 μl of the different combinations of index primers were used. Each index primer consisted of a unique 8 bp multiplex identifier and the Illumina adapter overhang sequence. The second PCR was performed at 94 °C for 5 min, followed by 13, 12, or 10 cycles of 94 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min for nematodes, fungi, and bacteria, respectively, and 72 °C for 10 min. Amplicons were visualized by gel electrophoresis, pooled equimolarly, precipitated, and the pellet dissolved in water. After gel electrophoresis, amplicons of the expected size (approximately 500 bp for nematodes and bacteria, 300-450 bp for fungi) were excised and purified using the QIAquick Gel Extraction kit (Qiagen) according to the manufacturer’s instruction. Finally, the DNA concentrations of amplicons were measured using a Qubit fluorometer and sequenced on a MiSeq sequencer with PE300 (Illumina, Eurofins Genomics, Konstanz, Germany).
qPCR for quantification of Meloidogyne hapla in Arabidopsis roots (Experiment 1)
In Experiment 1, we quantified Meloidogyne hapla in Arabidopsis roots by quantitative PCR (qPCR) using an Applied Biosystems ViiA7 Real-Time PCR system (Thermo Fischer Scientific). We specifically targeted M. hapla, as this species was previously diagnosed in the field soil used in the experiment, and M. hapla is the only Meloidogyne species known to be present in the country. A reaction mixture of 13 µl was prepared to contain 1.35 µl of each of the forward (5ʹ-TGGTTCAGGGTCATTTTTCTATAAAGT-3ʹ) and reverse (5ʹ- CAAATCGCTGCGTACCAACA-3ʹ) primers (10 µM), 0.75 µl probe (5ʹ-FAM-CCATTGGCACTATAAC-MGB-3ʹ) (5 µM), 7.5 µl qPCRBio Probe Mix LoROX (PCR Biosystems, London, United Kingdom), 2.05 µl H2O, and 2 µl DNA template [50]. As a control, we extracted M. hapla from carrot root galls and extracted DNA from recovered eggs and J2s using DNeasy Blood and Tissue kit (Qiagen). The identity of M. hapla eggs and J2s was confirmed by 18 S rDNA sequencing [50]. A 1:10 dilution series was prepared from this M. hapla DNA (0.83 ng/µl) to generate a standard curve. The thermal cycles in the real-time PCR were 95 °C for 10 min, followed by 40 cycles of 95 °C for 60 s and 60 °C for 1 min, and three technical replicates were used for each reaction. We checked data for normality by visualizing Normal Q-Q plots and homogeneity of variance by inspecting residuals vs. fitted value plots and fitted linear models using the generalized least squares function in R [44]. The effect of Arabidopsis genotype was tested with one-way ANOVA, followed by pairwise comparisons between individual genotypes using the Tukey test in R.
Sequence analysis (Experiments 1 and 2)
Sequence reads obtained from the MiSeq were demultiplexed using the python command line “demultiplexer.py”. Single-ended sequence reads were merged to paired-end reads using VSEARCH of QIIME2 [51], using a default overlap of minimum 10 bp (nematodes and fungi) or 20 bp (bacteria), and reads with quality Phred scores <30 were removed. Internal barcodes, forward and reverse primers, and reads with less than 250 bp were also excluded. Following this, sequences were screened for chimeras and clustered at 99% (nematode) or 97% (fungi and bacteria) similarity using VSEARCH of QIIME2 [51]. ITS2 reads were extracted using ITSx extractor v1.0.6 [52] before clustering fungal datasets. Taxonomy assignments for the clustered operational taxonomic units (OTUs) were done using the SILVA 132 reference database for nematodes (18 S) and bacteria (16 S) [53, 54] and the UNITE database version 04.02.2020 for fungi [55] using assign_taxonomy.py [56]. Moreover, all nematode OTUs were searched against the NCBI Genbank to confirm their taxonomic assignment due to the low specificity and coverage of the SILVA 132 reference database. Top hits with coverage of 100% in the BLAST search and sequence similarities of 100 % at species and ≥ 99 % at genus rank were considered.
Statistical analysis of sequence read abundances (Experiments 1 and 2)
We analyzed nematode diversity using the “phyloseq” [57] packages in R [44]. The OTU table was transformed to relative abundances before the calculation of beta diversity. For beta diversity and partitioning of variance, Bray-Curtis dissimilarity matrices were subjected to permutational analysis of variance (PERMANOVA) using the Adonis test with 1000 permutations from the “vegan” package [58]. Dissimilarity matrices were visualized using unconstrained principal coordinates analysis (PCoA). To identify nematode and microbial taxa with significantly different relative abundances between mutants and parental lines, we performed differential OTU abundance analysis using DESeq2 v1.30 [59]. To determine whether the relative abundances of nematode taxa at species rank were significantly different between genotypes, we used ANOVA followed by pairwise comparisons between the individual mutants and their respective parental line using the t-test in the STAMP software v2.1.3 [60].
We used an R script described previously for Arabidopsis root microbial- and nematode co-occurrence analysis [61]. In brief, all nematode, fungal, and bacterial OTUs were pooled, subjected to trimmed means of M transformation, and normalized as relative abundance counts per million using the “edgeR” package [62, 63]. We used OTUs that were present in at least 10 samples with Spearman’s rank correlations > 0.4 for positive correlations and < −0.4 for negative correlations, and p-values < 0.01. Correlations were visualized in a heatmap.
Results
Sequence data characteristics
In Experiment 1, we obtained 223 041, 660 481, and 529 372 sequence reads that could be assigned to Nematoda, bacteria, and fungi, respectively, from the roots of the thirteen Arabidopsis mutants and their respective parental lines. Nematode, bacterial, and fungal reads were assigned to 248, 3 378, and 756 OTUs, respectively. PCoA plots of beta-diversities of the communities are shown in Supplementary Fig. 5 and Supplementary Table 1. In Experiment 2, we retrieved 1 902 879 bacterial and 1 148 226 fungal sequence reads belonging to 6 919 bacterial and 548 fungal OTUs from the roots of eight Arabidopsis mutants and their respective parental lines.
Experiment 1: Genetic disruption of glucosinolate, camalexin, and flavonoid biosynthesis pathways affect nematode communities
PCoA ordination plots clearly separated nematode communities from the glucosinolate mutants, except myb51, and the camalexin mutants, from Col-0 (Fig. 1a, b). Communities from tt3-1 and tt3-1tt5-1 disrupted in flavonoid synthesis were distinct from Ler-0, while communities from the flavonoid overexpressing pap1-D did not separate notably from Col-0 (Fig. 1c). Corroborating the PCoA plots, PERMANOVA showed significant differences in nematode communities between glucosinolate (Adonis, R2 = 0.60, p < 0.001), camalexin (Adonis, R2 = 0.76, p < 0.001), and flavonoid (Adonis, R2 = 0.56, p < 0.001) mutants, and their parental lines (Table 2). The glucosinolate mutants explained 33% to 76% of the observed variation in the nematode community composition (Table 2), and the highest variation (81%) was explained in the camalexin defective mutant line pad3-1. Contrarily, only 29% of the variation was explained in the over-expression flavonoid mutant line pap1-D (Table 2).
Secondary metabolites specifically affect PPN taxa
Overall, patterns of enrichment and depletion of individual nematode taxa visually separated glucosinolate and flavonoid disruption mutants, as some taxa were depleted in the flavonoid deficient lines, while others were enriched only in the glucosinolate mutants compared to their parental lines (Fig. 2). No nematode taxa were significantly altered in the flavonoid overexpressing pap1-D mutant compared to Col-0. Meloidogyne was enriched in all secondary metabolite mutants except pap1-D (Fig. 2). The root-lesion nematode Pratylenchus and the root ectoparasite Paratylenchus were both depleted in most of the mutants, while the ectoparasite Bitylenchus was enriched in most of the glucosinolate mutants but not in the flavonoid mutants (Fig. 2). We observed both enrichment and depletion of free-living nematode taxa in the mutants. For instance, the bacterial feeding nematode Pseudoacrobeles was enriched in the glucosinolate and camalexin mutants and depleted in the flavonoid disrupted mutants (Fig. 2).
Nematode taxa that were affected in the individual mutants were determined by DESeq2. Only nematode taxa with significantly different relative abundances in the mutants compared to their parental lines are depicted. Blue color indicates enrichment in the mutant, red color indicates depletion and white indicates 0-fold changes. Color codes prefix taxa name indicates feeding habits of soil nematodes. Star symbol in front of mutant lines indicates deficiency in camalexin (Cmlx). Ler-0 is the parental line of tt3-1 and tt3-1tt5-1; Col-0 is the parental line of the other mutants.
Corroborating the differential abundance analysis, the relative abundance of Meloidogyne was higher in most mutant lines compared to parental lines, but not in pap1-D (Fig. 3a). To verify whether the variation in relative abundances reflected variation in absolute abundances, we performed a qPCR assay targeting M. hapla. This confirmed that M. hapla abundance varied between genotypes (ANOVA, p < 0.001) and was significantly higher in five of the glucosinolate mutants compared to the parental line (Fig. 3b).
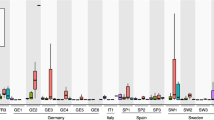
A Relative abundances of the ten most abundant nematode taxa. Plant parasitic nematode taxa are labelled with asterisks. For each taxon, relative read abundances were tested using ANOVA, followed by a t-test for pairwise comparisons between mutants and their respective parental lines in STAMP software. The significance of test indicated as ***p < 0.001, **p < 0.01, and *p < 0.05. B DNA quantities (ng/mg dry root) of Meloidogyne hapla were determined by qPCR. The effect of the different Arabidopsis lines on M. hapla DNA abundance (qPCR data) was tested using ANOVA (p-value presented in top right corner), followed by post-hoc pairwise comparisons between the mutants and their respective parental line with a Tukey test in R. The significance of test is indicated as ***p < 0.001 and **p < 0.01. Star symbol in front of mutant lines indicates deficiency in camalexin (Cmlx). Ler-0 is the parental line of tt3-1 and tt3-1tt5-1; Col-0 is the parental line of the other mutants.
Secondary metabolites affect microbial taxa and co-occurrence patterns between soil nematode and microbial taxa
PCoA ordination plots clearly separated microbial communities in the glucosinolate, camalexin, and flavonoids mutants from Col-0/Ler-0, except in the pap1-D line (Supplementary Fig. 5). We detected both negative and positive correlations between root-associated nematode taxa and fungal and bacterial taxa (Fig. 4). Allorhizobium-Neorhizobium-Pararhizobium-Rhizobium (hereafter Rhizobium) correlated negatively with several nematode operational taxonomic units (OTUs) belonging to different genera. Notably, Meloidogyne correlated negatively with Rhizobium, Adhaeribacter, Deinococcus, Gemmatimonas, Iamia, Nocardioides, and Oceanobacillus, and positively with Bacillus, Castellaniella, Gemmata, Haliangium, Leptolyngbya, Pirellula, Rhodopirellula, and Taibaiella (Fig. 4a). Several fungal genera such as Acremonium, Dichotomopilus, Exophiala, Lachnum, Meliniomyces, Monocillium, Metapochonia, Pseudogymnoascus, and Trichoderma correlated negatively with several nematode taxa, including Meloidogyne. Clonostachys correlated negatively with Filenchus and Pratylenchus, while Mortierella correlated negatively with Bitylenchus and Acrobeles. Auxarthron, Paraphoma, Oidiodendron, and Metarhizium correlated positively with Meloidogyne among other nematode taxa (Fig. 4b).
Plant parasitic nematode taxa are labelled with asterisks. A Blue color indicates positive correlation and red color indicates negative correlation between nematode and bacterial taxa. Rhizobium indicates Allorhizobium-Neorhizobium-Pararhizobium-Rhizobium. B Blue color indicates positive correlation and red color indicates negative correlation between nematode and fungal taxa.
Experiment 2: Microbiomes from roots altered in flavonoid synthesis significantly affect M. incognita invasion of tomato roots
When exposed to microbiomes from the flavonoid overexpressing line pap1-D we found significant increases in the number of J2s invading tomato roots, and at the same time, microbiomes harvested from the flavonoid deficient ttg1ttg2 and tt3-1tt5-1 lines significantly reduced J2 invasion (Fig. 5), suggesting a remarkable flavonoid-induced microbiota effect on nematode invasion capability. We note that the biomass of the microbiota used was similar for all Arabidopsis lines (Supplementary Fig. 6). The bacterial communities from pap1-D did not notably separate from the parental line in a PCoA ordination plot, whereas ttg1ttg2 and tt3-1tt5-1 separated more clearly from their respective parental lines (Supplementary Fig. 7b), as also indicated in a PERMANOVA (Supplementary Table 2). For the fungal communities, PCoA plots did not clearly separate the flavonoid mutants from their parental lines (Supplementary Fig. 7d), as also confirmed in the PERMANOVA (Supplementary Table 2).
Arabidopsis mutant lines cyp79B2, cyp79B2cyp79B3, gsm1-1, tgg1tgg2, pad3-1, ttg1ttg2, tt3-1tt5-1, pap1-D, and their parental lines were used. Control treatment was sterile 0.085% NaCl plus 1/300 TSB. An ANOVA test was performed (p-value presented in top right corner), followed by Tukey’s test in R and the significance of tests was indicated as ***p < 0.001 and **p < 0.01. Ler-0 is the parental line of tt3-1tt5-1; Col-0 is the parental line of the other mutants, Cmlx Camalexin.
The DEseq2 analysis performed on class level for bacteria showed that most bacterial classes were not significantly affected in pap1-D (Supplementary Fig. 8). However, a t-test at the OTU level showed that Chitinimonas was depleted in pap1-D, while Halomonas, Chitinophagaceae, Terracidiphilus, Pseudogulbenkiania, Herminiimonas, and Salinimicrobium were enriched in pap1-D (Supplementary Table 3). Likewise, DEseq2 revealed several taxa enriched in tt3-1tt5-1 and some were also enriched in the other flavonoid-deficient mutant ttg1ttg2 (Supplementary Fig. 8a).
For the fungal communities, DEseq2 showed that Cladosporium was strongly enriched in pap1-D and strongly depleted in tt3-1tt5-1, while Vishniacozyma was highly depleted in pap1-D (Supplementary Fig. 8b). Using a pairwise t-test at OTU level, we detected significant depletions of Gibellulopsis nigrescens, Keithomyces, Solicoccozyma terricola, Tetracladium, Clonostachys rosea, Mortierella, Acremonium, and Acrostalagmus in pap1-D (p < 0.05), while no significant enrichment of fungal taxa was observed (Supplementary Table 4). Most of the depleted taxa in pap1-D were also depleted in ttg1ttg2 (Supplementary Table 4), while Fusicolla, Trichoderma, Chloridium, Pyrenochaetopsis, and Monocillium griseo-ochraceum were depleted in tt3-1tt5-1 (Supplementary Table 4). However, only a few sequence reads were retrieved belonging to the taxa that were differentially abundant in the mutants (Supplementary Tables 3 and 4).
Discussion
With the intention to improve our understanding of root-nematode community interactions in plant-soil systems, we grew a range of Arabidopsis mutants genetically altered in glucosinolate, camalexin or flavonoid synthesis pathways in arable soil and characterized their root-associated nematode communities. In addition, we tested the potential modulating effects of fungal and bacterial communities on these root-nematode interactions. Our study showed that disruption of the genes radically affected root-associated nematode communities, with the most significant effects observed for M. hapla. Furthermore, we found that microbiota from roots with genetically altered flavonoid synthesis had significant effects on the ability of M. incognita to invade tomato roots.
Secondary metabolites affected nematode communities, particularly Meloidogyne hapla
Genetic disruption of the biosynthetic pathways leading to a range of secondary metabolites had distinct effects on the root-associated nematode communities. The separation in PCoA plots of nematode communities from almost all mutant lines demonstrated that these groups of metabolites play significant roles in the intricate plant-nematode interaction. We detected, however, only minor effects of the myb51 and pap1-D mutants on the communities. This could be caused by the moderate effects of myb51 on glucosinolate contents [64], and that, in pap1-D, despite the enhanced accumulation of flavonoids, anthocyanidins, hydroxycinnamic acids, syringyl, and guaiacyl lignin [65, 66], these metabolites were already present in sufficient quantities in Col-0 to prevent M. hapla infection.
Relative abundances of M. hapla increased significantly in all mutants except pap1-D. This suggests stronger effects of secondary metabolites on sedentary endoparasitic nematodes such as M. hapla living in intimate contact with roots. Arabidopsis is not considered a good host for M. hapla (Tina Kyndt, unpublished results), and our results indicate that this may result from the production of secondary metabolites. Previous in vitro assays similarly showed higher numbers of M. incognita galls, higher H. schachtii female and cysts numbers, and larger syncytia, associated with glucosinolate and camalexin disruption mutants [15,16,17]. Distinct nematode taxa were affected in the flavonoid and glucosinolate defective mutants. M. hapla and the stubby root nematode Paratrichodorus were enriched in the tt3-1tt5-1 double mutant, while several other nematode taxa were depleted, suggesting taxon-specific effects of flavonoids. Flavonoids have been shown to act as defense metabolites against the endoparasitic M. incognita, H. glycines, and Radopholus similis [28, 32, 67]. However, confirming the PCoA plots, no nematode taxa were affected in the pap1-D mutant despite a massive activation of the phenylpropanoid pathway and thus enhanced accumulation of flavonoids, anthocyanidins, hydroxycinnamic acids, syringyl, and guaiacyl lignin in this mutant [65, 66]. The root-lesion nematode P. neglectus was depleted in several mutants, which could have been caused by displacement by the massively enriched M. hapla in those mutants (Figs. 2, 3a). The ectoparasitic Paratylenchus was depleted, and Bitylenchus was enriched in most of the mutants, while Merlinius and Mesocriconema were depleted in the flavonoid defective mutants. However, our results suggest that the metabolites investigated in this study only had minor effects on ectoparasites. Similarly, free-living taxa were only slightly affected, although a few alterations of relative abundances were observed. These observations probably reflect that ectoparasites and free-living taxa are in less intimate contact with host tissues and thus plant-synthesized metabolites than endoparasitic nematodes.
Several microbial taxa are potential antagonists of PPNs
Several microbial taxa are closely associated with nematodes and have been shown to affect nematodes, for instance, as antagonists of PPN [39, 68], but as recently suggested, specific microbial taxa may also alleviate the effects of nemato-antagononists [69]. In our study, the composition of both nematode, bacterial, and fungal communities varied between Arabidopsis parental lines and mutants. It is challenging to disentangle whether nematode community changes are caused directly by altered plant secondary metabolite synthesis or indirectly by plant metabolite modulation of the microbial communities. Vice versa, changes in nematode community composition may also influence the assembly of microbial communities [70]. Moreover, direct metabolite effects on specific taxa within communities may disrupt e.g. competitive or predator-prey interactions, eventually changing the abundance of other taxa. Evidently, we cannot identify causal links between secondary plant metabolites and abundances of individual microbial or nematode taxa or between individual microbial and nematode taxa in this complex system. However, the analysis of co-occurrence patterns between microbial and nematode taxa elucidates putative interactions between taxa, which can be further tested under more controlled conditions.
Bacterial genera such as Gemmatimonas, Deinococcus, Nocardiodes, and Rhizobium were negatively co-occurring with several nematode genera. In a previous study, Gemmatimonas was associated with the surface of J2s of M. hapla in nematode suppressive soils [71], suggesting an antagonistic potential of this taxon. Other bacterial taxa include Leptolyngbya, Castellania, Taibaiella, Rhodopiruellula, Pirullula, Pseudoflavitelea, Halangium, and Gemmata were positively co-occurring with some of the nematode genera. In contrast others such as Bacillus, Iamia, and Adhaeribacter showed both positive and negative co-occurrences with individual nematode genera. Several Bacillus species are known as antagonists against PPNs [72, 73], although the potential of Bacillus to control PPNs may vary between individual species or strains.
The fungal taxa Trichoderma, Metapochonia, Monocillium, Exophiala, and Acremonium were negatively co-occurring with Meloidogyne, while Clonostachys and Mortierella were negatively correlating with other PPNs. All fungal genera that were negatively co-occurring with PPNs have previously been reported to parasitize PPNs [74,75,76,77,78]. For instance, Acremonium is an endophytic fungus that can reduce M. incognita populations in tomato roots [79], suggesting that some of the identified microbial taxa may have potential for use in biocontrol strategies.
Secondary metabolites modulate nematode invasion through the associated microbiota
J2 invasion into the tomato roots increased several-fold when the nematodes were exposed to the pap1-D-derived microbiome, although this mutant did not affect nematode relative abundances (or microbiome composition) in Experiment 1. We speculate that the enhanced production of anthocyanins, hydroxycinnamic acids, syringyl, or guaiacyl lignin in pap1-D could affect specific microorganisms interacting with Meloidogyne and thus affect invasion. In line with the enhanced invasion of nematodes exposed to pap1-D-microbiota, the microbiota isolated from the flavonoid deficient ttg1ttg2 and tt3-1tt5-1 suppressed J2 invasion, indicating a strong flavonoid effect on nematode invasion via the associated microbiota.
The sequencing of microbial communities from the mutants used in Experiment 2 revealed a significantly lower abundance of several bacterial genera, for instance, Chitinimonas, in the pap1-D microbiome. Chitinimonas metabolizes chitin, one of the major components of the nematode cuticle and eggshells [80]. Although sequence read numbers for this taxon were low, we speculate that the depletion of this nematode-antagonistic microorganism in the pap1-D microbiome could partly explain the increase in M. incognita J2 invasion of tomato. Likewise, the fungus Gibellulopsis nigrescens was depleted in the pap1-D microbiome. G. nigrescens is associated with nematode-suppressive soils and is reported to function as a nematode antagonist [38, 81]. Acremonium, Paecilomyces, and Mortierella were likewise depleted in the microbiota from pap1-D and ttg1ttg2. The fungal genus Acremonium correlated negatively with Meloidogyne in Experiment 1, suggesting an antagonistic role of this fungus. Likewise, several bacterial taxa were enriched in the flavonoid defective tt3-1tt5-1 and or ttg1ttg2-derived microbiota. The enrichment/depletion of specific microorganisms in the flavonoid altered mutants could thus potentially explain the effects observed on M. incognita J2 invasion of tomato roots. However, further studies should aim to identify the responsible taxa conclusively and elucidate their antagonistic potentials against PPNs.
The microbiomes collected from the Arabidopsis lines were transferred to the tomato roots 11 days before the addition of M. incognita J2s to ensure microbiome the establishment of the tomato roots before M. incognita invasion. The composition of the inoculated microbiomes might have changed during establishment and growth on the tomato roots. As the metabolic profile of tomato roots differ from Arabidopsis roots, most likely, the tomato roots selected microbiomes with a different composition than Arabidopsis. Likewise, the pre-incubation of M. incognita J2s with Arabidopsis microbiomes in dilute TSB will also have selectively enriched specific microbial taxa. The microbiome composition thus changed during Experiment 2; however, the difference in M. incognita invasion success between microbiomes selected in flavonoid deficient (ttg1ttg2 and tt3-1tt5-1) and flavonoid, hydroxycinnamic acids, syringyl, and guaiacyl lignin overproducing (pap1-D) mutants compared to their parental lines indicate that flavonoids affect components of the microbiome that determine the invasion ability of M. incognita.
In Experiment 1, pap1-D did not affect M. hapla abundance (or nematode communities), whereas in Experiment 2, the pap1-D-microbiota significantly enhanced M. incognita invasion in tomato. One explanation for these contradicting results could be that the flavonoid, hydroxycinnamic acids, syringyl, and guaiacyl lignin concentrations in Col-0 were sufficient to prevent M. hapla invasion, and thus the higher concentrations in pap1-D did not have any additional effects on M. hapla. Similarly, it was too early for any microbiome-mediated effects of pap1-D on the nematodes to be observed. The results of Experiment 2, on the other hand, suggest that the higher flavonoid, hydroxycinnamic acids, syringyl, or guaiacyl lignin contents in pap1-D reduced Meloidogyne-suppressive microorganisms, consequently allowing more M. incognita J2s to enter the tomato roots. In line with this, the microbiomes from the flavonoid and anthocyanin defective mutants ttg1ttg2 and tt3-1tt5-1 reduced the invasion capacity of M. incognita.
This study thus shows that plant secondary metabolites exert complex effects on the tripartite interaction among plants, nematodes, and microbial taxa. This finding should be the foundation for further research to disentangle causal relationships between plant secondary metabolites, microbial taxa, and nematode responses.
Conclusions
We have demonstrated that genetic disruption of pathways leading to glucosinolates, camalexin, and flavonoids in Arabidopsis increases the susceptibility of Arabidopsis to PPNs, especially the sedentary endoparasitic nematode M. hapla that is in intimate contact with host roots. The microbial communities assembled by the flavonoid, hydroxycinnamic acids, syringyl, and guaiacyl lignin overexpressing pap1-D significantly enhanced M. incognita invasion of tomato roots, while microbiomes from the flavonoid deficient ttg1ttg2 and tt3-1tt5-1 mutants suppressed M. incognita invasion. These results strongly suggest that flavonoids indirectly affect PPN invasion by modulating the root-associated microbiome. In addition, direct flavonoid effects on nematode communities may also contribute to altered microbiome assembly. We identified specific microbial taxa that are known antagonists of PPNs that could potentially be responsible for this effect. However, further studies are needed to identify the causal microorganisms conclusively. For nematode management, the governing role of plant secondary metabolites for the recruitment of PPN antagonistic microbiomes is an interesting perspective. Future research coupling specific metabolites and PPN antagonistic microbiomes could lead to breeding crop cultivars with metabolic profiles facilitating microbial PPN suppression.
Data availability
The paired end sequence reads for the nematode 18 S rRNA gene, bacterial 16 S rRNA gene, and fungal ITS2 region have been deposited in the NCBI SRA database (https://www.ncbi.nlm.nih.gov/sra) under accession code PRJNA698344 (198 samples), and PRJNA698287 (roots of Arabidopsis parental lines only, 12 samples). Raw data are openly available in the public depository Zenodo at https://doi.org/10.5281/zenodo.5115230.
References
van den Hoogen J, Geisen S, Routh D. Soil nematode abundance and functional group composition at a global scale. Nature 2019;572:194–98.
Yeates GW, Bongers T, Degoede RGM, Freckman DW, Georgieva SS. Feeding habits in soil nematode families and genera - an outline for soil ecologists. J Nematol. 1993;25:315–31.
Nicol JM, Turner SJ, Coyne DL, Nijs Ld, Hockland S, Maafi ZT. Current nematode threats to world agriculture. In: Jones J, Gheysen G, Fenoll C, editors. Genomics and Molecular Genetics of Plant-Nematode Interactions. Dordrecht: Springer; 2011. p. 21–43.
Decraemer W, Hunt D. Structure and Classification. In: R. N. Perry, M. Moens, Eds. Plant Nematology. CABI, Wallingford, Oxfordshire, UK and Boston, USA, 2005, pp. 26–27.
Fleming TR, Maule AG, Fleming CC. Chemosensory responses of plant parasitic nematodes to selected phytochemicals reveal long-term habituation traits. J Nematol. 2017;49:462–71.
Murungi LK, Kirwa H, Coyne D, Teal PEA, Beck JJ, Torto B. Identification of key root volatiles signaling preference of tomato over spinach by the root knot nematode Meloidogyne incognita. J AgricFood Chem. 2018;66:7328–36.
Wang CL, Masler EP, Rogers ST. Responses of Heterodera glycines and Meloidogyne incognita infective juveniles to root tissues, root exudates, and root extracts from three plant species. Plant Dis. 2018;102:1733–40.
Sikder MM, Vestergård M. Impacts of root metabolites on soil nematodes. Front Plant Sci. 2020;10:1792.
van Dam NM, Tytgat TOG, Kirkegaard JA. Root and shoot glucosinolates: A comparison of their diversity, function and interactions in natural and managed ecosystems. Phytochem Rev. 2009;8:171–86.
Bressan M, Roncato MA, Bellvert F, et al. Exogenous glucosinolate produced by Arabidopsis thaliana has an impact on microbes in the rhizosphere and plant roots. ISME J. 2009;3:1243–57.
Mucha S, Heinzlmeir S, Kriechbaumer V, Strickland B, Kirchhelle C, Choudhary M, et al. The formation of a camalexin biosynthetic metabolon. Plant Cell. 2019;31:2697–710.
Kettles GJ, Drurey C, Schoonbeek HJ, Maule AJ, Hogenhout SA. Resistance of Arabidopsis thaliana to the green peach aphid, Myzus persicae, involves camalexin and is regulated by microRNAs. N. Phytol. 2013;198:1178–90.
Tsuji J, Jackson EP, Gage DA, Hammerschmidt R, Somerville SC. Phytoalexin accumulation in Arabidopsis thaliana during the hypersensitive reaction to Pseudomonas syringae pv. syringae. Plant Physiol. 1992;98:1304–9.
Thomma BPHJ, Nelissen I, Eggermont K, Broekaert WF. Deficiency in phytoalexin production causes enhanced susceptibility of Arabidopsis thaliana to the fungus Alternaria brassicicola. Plant J 1999;19:163–71.
Teixeira MA, Wei LH, Kaloshian I. Root-knot nematodes induce pattern-triggered immunity in Arabidopsis thaliana roots. N Phytol. 2016;211:276–87.
Shah SJ, Anjam MS, Mendy B, Anwer MA, Habash SS, Lozano-Torres JL, et al. Damage-associated responses of the host contribute to defence against cyst nematodes but not root-knot nematodes. J Exp Bot. 2017;68:5949–60.
Ali MA, Wieczorek K, Kreil DP, Bohlmann H. The beet cyst nematode Heterodera schachtii modulates the expression of WRKY transcription factors in syncytia to favour its development in Arabidopsis roots. PLoS One. 2014;9:e102360.
Lazzeri L, Curto G, Leoni O, Dallavalle E. Effects of glucosinolates and their enzymatic hydrolysis products via myrosinase on the root-knot nematode Meloidogyne incognita (Kofoid et White) Chitw. J Agric Food Chem. 2004;52:6703–07.
Avato P, D’Addabbo T, Leonetti P, Argentieri MP. Nematicidal potential of Brassicaceae. Phytochem Rev. 2013;12:791–802.
Mathesius U. Flavonoid functions in plants and their interactions with other organisms. Plants (Basel) 2018;7:30.
Weston LA, Mathesius U. Flavonoids: Their structure, biosynthesis and role in the rhizosphere, including allelopathy. J Chem Ecol. 2013;39:283–97.
Badri DV, Loyola-Vargas VM, Broeckling CD, De-la-Pena C, Jasinski M, Santelia D, et al. Altered profile of secondary metabolites in the root exudates of Arabidopsis ATP-binding cassette transporter mutants. Plant Physiol. 2008;146:762–71.
Cesco S, Neumann G, Tomasi N, Pinton R, Weisskopf L. Release of plant-borne flavonoids into the rhizosphere and their role in plant nutrition. Plant Soil. 2010;329:1–25.
Drewnowski A, Gomez-Carneros C. Bitter taste, phytonutrients, and the consumer: A review. Am J Clin Nutr. 2000;72:1424–35.
Chin S, Behm CA, Mathesius U. Functions of flavonoids in plant-nematode interactions. Plants (Basel) 2018;7:1–17.
Kaplan DT, Keen NT, Thomason IJ. Association of glyceollin with the incompatible response of soybean roots to Meloidogyne incognita. Physiol Plant Pathol. 1980;16:309–18.
Aoudia H, Ntalli N, Aissani N, Yahiaoui-Zaidi R, Caboni P. Nematotoxic phenolic compounds from Melia azedarach against Meloidogyne incognita. J AgricFood Chem. 2012;60:11675–80.
Kennedy MJ, Niblack TL, Krishnan HB. Infection by Heterodera glycines elevates isoflavonoid production and influences soybean nodulation. J Nematol. 1999;31:341–47.
Collingborn FMB, Gowen SR, Mueller-Harvey I. Investigations into the biochemical basis for nematode resistance in roots of three Musa cultivars in response to Radopholus similis infection. J Agric Food Chem. 2000;48:5297–301.
Cook R, Tiller SA, Mizen KA, Edwards R. Isoflavonoid metabolism in resistant and susceptible cultivars of white clover infected with the stem nematode Ditylenchus dipsaci. J Plant Physiol. 1995;146:348–54.
Kirwa HK, Murungi LK, Beck JJ, Torto B. Elicitation of differential responses in the root-knot nematode Meloidogyne incognita to tomato root exudate cytokinin, flavonoids, and alkaloids. J AgricFood Chem. 2018;66:11291–300.
Wuyts N, Swennen R, De, Waele D. Effects of plant phenylpropanoid pathway products and selected terpenoids and alkaloids on the behaviour of the plant-parasitic nematodes Radopholus similis. Pratylenchus penetrans Meloidogyne Incogn Nematol. 2006;8:89–101.
Hartwig UA, Joseph CM, Phillips DA. Flavonoids released naturally from alfalfa seeds enhance growth rate of Rhizobium meliloti. Plant Physiol. 1991;95:797–803.
Hassan S, Mathesius U. The role of flavonoids in root-rhizosphere signalling: Opportunities and challenges for improving plant-microbe interactions. J Exp Bot. 2012;63:3429–44.
Kudjordjie EN, Sapkota R, Nicolaisen M. Arabidopsis assemble distinct root-associated microbiomes through the synthesis of an array of defense metabolites. PLoS One. 2021;10:e0259171.
Rønn R, Vestergård M, Ekelund F. Interactions between bacteria, protozoa and nematodes in soil. Acta Protozool. 2012;51:223–35.
Thakur MP, Geisen S. Trophic regulations of the soil microbiome. Trends Microbiol. 2019;27:771–80.
Elhady A, Gine A, Topalovic O, Jacquiod S, Sorensen SJ, Sorribas FJ, et al. Microbiomes associated with infective stages of root-knot and lesion nematodes in soil. PLoS One. 2017;12:e0177145.
Toju H, Tanaka Y. Consortia of anti-nematode fungi and bacteria in the rhizosphere of soybean plants attacked by root-knot nematodes. R Soc Open Sci. 2019;6:181693.
Topalović O, Bredenbruch S, Schleker ASS, Heuer H. Microbes attaching to endoparasitic phytonematodes in soil trigger plant defense upon root penetration by the nematode. Front Plant Sci 2020;11:138.
Schaad NW, Walker JT. The use of density-gradient centrifugation for the purification of eggs of Meloidogyne spp. J Nematol. 1975;7:203–04.
Hooper DJ, Hallmann J, Subbotin SA. Methods for extraction, processing and detection of plant and soil nematodes. In: Luc M, Sikora RA, Bridge J, editors. Plant parasitic nematodes in subtropical and tropical agriculture. Second ed. Wallingford, UK: CABI Publishing; 2005. p. 53.
Topalovic O, Elhady A, Hallmann J, Richert-Poggeler KR, Heuer H. Bacteria isolated from the cuticle of plant-parasitic nematodes attached to and antagonized the root-knot nematode Meloidogyne hapla. Sci Rep. 2019;9:11477.
R Core Team. R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2019.
Porazinska DL, Giblin-Davis RM, Faller L, Farmerie W, Kanzaki N, Morris K, et al. Evaluating high-throughput sequencing as a method for metagenomic analysis of nematode diversity. Mol Ecol Resour. 2009;9:1439–50.
Sapkota R, Nicolaisen M. High-throughput sequencing of nematode communities from total soil DNA extractions. BMC Ecol. 2015;15:3.
Sikder MM, Vestergård M, Sapkota R, Kyndt T, Nicolaisen M. Evaluation of metabarcoding primers for analysis of soil nematode communities. Diversity (Basel) 2020;12:388.
Ihrmark K, Bodeker ITM, Cruz-Martinez K, Friberg H, Kubartova A, Schenck J, et al. New primers to amplify the fungal ITS2 region - evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol Ecol. 2012;82:666–77.
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013;41:e1.
Sapkota R, Skantar AM, Nicolaisen M. A TaqMan real-time PCR assay for detection of Meloidogyne hapla in root galls and in soil. Nematol. 2016;18:147–54.
Rognes T, Flouri T, Nichols B, Quince C, Mahe F. VSEARCH: A versatile open source tool for metagenomics. Peer J. 2016;4:e2584.
Bengtsson-Palme J, Ryberg M, Hartmann M, Branco S, Wang Z, Godhe A, et al. Improved software detection and extraction of ITS1 and ITS2 from ribosomal ITS sequences of fungi and other eukaryotes for analysis of environmental sequencing data. Methods Ecol Evol. 2013;4:914–19.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–D6.
Yilmaz P, Parfrey LW, Yarza P, Gerken J, Pruesse E, Quast C, et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res. 2014;42:D643–D8.
UNITE. UNITE QIIME release for Fungi [Internet]. UNITE Community. 2020.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–36.
McMurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One. 2013;8:e61217.
Oksanen J, Blanchet FG, Kindt R, Friendly M, Legendre P, McGlinn D, et al. Vegan: Community Ecology Package. Ordination methods, diversity analysis and other functions for community and vegetation ecologists. R Package Version 2.5-5 ed: The Comprehensive R Archive Network; 2019.
Love MI, Huber W, Anders S. Moderated estimation of fold change anddispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Parks DH, Tyson GW, Hugenholtz P, Beiko RG. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 2014;30:3123–24.
Kudjordjie EN, Sapkota R, Steffensen SK, Fomsgaard IS, Nicolaisen M. Maize synthesized benzoxazinoids affect the host associated microbiome. Microbiome 2019;7:59.
McCarthy DJ, Chen YS, Smyth GK. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 2012;40:4288–97.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010;26:139–40.
Frerigmann H, Gigolashvili T. MYB34, MYB51, and MYB122 distinctly regulate indolic glucosinolate biosynthesis in Arabidopsis thaliana. Mol Plant. 2014;7:814–28.
Schulz E, Tohge T, Zuther E, Fernie AR, Hincha DK. Flavonoids are determinants of freezing tolerance and cold acclimation in Arabidopsis thaliana. Sci Rep. 2016;6:34027.
Borevitz JO, Xia Y, Blount J, Dixon RA, Lamb C. Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell. 2000;12:2383–94.
Du SS, Zhang HM, Bai CQ, Wang CF, Liu QZ, Liu ZL, et al. Nematocidal flavone-C-glycosides against the root-knot nematode (Meloidogyne incognita) from Arisaema erubescens tubers. Molecules 2011;16:5079–86.
Zhou DM, Feng H, Schuelke T, De Santiago A, Zhang QM, Zhang JF, et al. Rhizosphere microbiomes from root knot nematode non-infested plants suppress nematode Infection. Micro Ecol. 2019;78:470–81.
Topalović O, Vestergård M. Can microorganisms assist the survival and parasitism of plant-parasitic nematodes? Trends Parasitol. 2021;37:947–58.
De Mesel I, Derycke S, Moens T, Van der Gucht K, Vincx M, Swings J. Top-down impact of bacterivorous nematodes on the bacterial community structure: a microcosm study. Environ Microbiol. 2004;6:733–44.
Adam M, Westphal A, Hallmann J, Heuer H. Specific microbial attachment to root knot nematodes in suppressive soil. Appl Environ Microbiol. 2014;80:2679–86.
Ramyabharathi S, Sankari Meena K, Rajendran L, Karthikeyan G, Jonathan EI, Raguchander T. Biocontrol of wilt-nematode complex infecting gerbera by Bacillus subtilis under protected cultivation. Egypt J Biol Pest Co. 2018;28:21.
Jamal Q, Cho JY, Moon JH, Munir S, Anees M, Kim KY. Identification for the first time of cyclo (D-Pro-L-Leu) produced by Bacillus amyloliquefaciens y1 as a nematocide for control of Meloidogyne incognita. Molecules 2017;22:1839.
Moosavi MR, Zare R. Fungi as biological control agents of plant-parasitic nematodes. In: Mérillon J-M, Ramawat KG, editors. Plant Defence: Biological Control. Progress in Biological Control 22. 2nd Edition ed. Switzerland: Springer; 2020. p. 333–84.
Ashrafi S, Stadler M, Dababat AA, Richert-Poggeler KR, Finckh MR, Maier W. Monocillium gamsii sp nov and Monocillium bulbillosum: two nematode-associated fungi parasitising the eggs of Heterodera filipjevi. Mycokeys. 2017;27:21–38.
Nuaima RH, Ashrafi S, Maier W, Heuer H. Fungi isolated from cysts of the beet cyst nematode parasitized its eggs and counterbalanced root damages. J Pest Sci. 2021;94:563–72.
Iqbal M, Dubey M, McEwan K, Menzel U, Franko MA, Viketoft M, et al. Evaluation of Clonostachys rosea for control of plant parasitic nematodes in soil and in roots of carrot and wheat. Phytopathology 2018;108:52–59.
DiLegge MJ, Manter DK, Vivanco JM. A novel approach to determine generalist nematophagous microbes reveals Mortierella globalpina as a new biocontrol agent against Meloidogyne spp. nematodes. Sci Rep. 2019;9:7521.
Goswami J, Pandey RK, Tewari JP, Goswami BK. Management of root knot nematode on tomato through application of fungal antagonists, Acremonium strictum and Trichoderma harzianum. J Environ Sci Health. 2008;43:237–40.
Chen Q, Peng D. Nematode chitin and application. In: Yang Q, Fukamizo T, editors. Targeting Chitin-containing Organisms. Advances in Experimental Medicine and Biology. 1142. Singapore: Springer; 2019. pp. 209–219.
Zhou WQ, Verma VC, Wheeler TA, Woodward JE, Starr JL, Sword GA. Tapping into the cotton fungal phytobiome for novel nematode biological control tools. Phytobiomes J 2020;4:19–26.
Alcazar R, von Reth M, Bautor J, Chae E, Weigel D, Koornneef M, et al. Analysis of a plant complex resistance gene locus underlying immune-related hybrid incompatibility and its occurrence in nature. PLoS Genet. 2014;10:e1004848.
Mikkelsen MD, Hansen CH, Wittstock U, Halkier BA. Cytochrome P450CYP79B2 from Arabidopsis catalyzes the conversion of tryptophan to indole-3-acetaldoxime, a precursor of indole glucosinolates and indole-3-acetic acid. J Biol Chem. 2000;275:33712–17.
Hull AK, Vij R, Celenza JL. Arabidopsis cytochrome P450s that catalyze the first step of tryptophan-dependent indole-3-acetic acid biosynthesis. Proc Natl Acad Sci USA. 2000;97:2379–84.
Zhao YD, Hull AK, Gupta NR, Goss KA, Alonso J, Ecker JR, et al. Trp-dependent auxin biosynthesis in Arabidopsis: involvement of cytochrome P450s CYP79B2 and CYP79B3. GenesDev. 2002;16:3100–12.
Schlaeppi K, Bodenhausen N, Buchala A, Mauch F, Reymond P. The glutathione-deficient mutant pad2-1 accumulates lower amounts of glucosinolates and is more susceptible to the insect herbivore Spodoptera littoralis. Plant J. 2008;55:774–86.
Schuhegger R, Nafisi M, Mansourova M, Petersen BL, et al. CYP71B15 (PAD3) catalyzes the final step in camalexin biosynthesis. Plant Physiol. 2006;141:1248–54.
Glawischnig E. The role of cytochrome P450 enzymes in the biosynthesis of camalexin. Biochem Soc Trans. 2006;34:1206–8.
Haughn GW, Davin L, Giblin M, Underhill EW. Biochemical genetics of plant secondary metabolites in Arabidopsis thaliana: The glucosinolates. Plant Physiol. 1991;97:217–26.
Kroymann J, Textor S, Tokuhisa JG, Falk KL, Bartram S, Gershenzon J, et al. A gene controlling variation in Arabidopsis glucosinolate composition is part of the methionine chain elongation pathway. Plant Physiol. 2001;127:1077–88.
Textor S, de Kraker JW, Hause B, Gershenzon J, Tokuhisa JG. MAM3 catalyzes the formation of all aliphatic glucosinolate chain lengths in Arabidopsis. Plant Physiol. 2007;144:60–71.
Barth C, Jander G. Arabidopsis myrosinases TGG1 and TGG2 have redundant function in glucosinolate breakdown and insect defense. Plant J. 2006;46:549–62.
Dong XY, Braun EL, Grotewold E. Functional conservation of plant secondary metabolic enzymes revealed by complementation of Arabidopsis flavonoid mutants with maize genes. Plant Physiol. 2001;127:46–57.
Peer WA, Brown DE, Tague BW, Muday GK, Taiz L, Murphy AS. Flavonoid accumulation patterns of transparent testa mutants of Arabidopsis. Plant Physiol. 2001;126:536–48.
Gonzalez A, Brown M, Hatlestad G, Akhavan N, Smith T, Hembd A, et al. TTG2 controls the developmental regulation of seed coat tannins in Arabidopsis by regulating vacuolar transport steps in the proanthocyanidin pathway. Dev Biol. 2016;419:54–63.
Walker AR, Davison PA, Bolognesi-Winfield AC, James CM, Srinivasan N, Blundell TL, et al. The TRANSPARENT TESTA GLABRA1 locus, which regulates trichome differentiation and anthocyanin biosynthesis in Arabidopsis, encodes a WD40 repeat protein. Plant Cell. 1999;11:1337–49.
Acknowledgements
We acknowledge Professor Judith Bender (Brown University, USA), Professor Wendy Ann Peer (University of Maryland, USA), and the Nottingham Arabidopsis Stock Centre (NASC, UK) for kindly providing the Arabidopsis lines. We are grateful to Mathilde Schiøtt Dige and Simone Ena Rasmussen for their laboratory technical assistance.
Funding
This research work was funded by Aarhus University (project number: 27747) and the Independent Research Fund Denmark DFF (grant numbers: 6111-00065B and 9041-00139B).
Author information
Authors and Affiliations
Contributions
MMS, MN, MV, and OT conceived and designed the experiments. MMS and OT executed the experiments. MMS and ENK analyzed the data. MN, TK, and MV supervised the experiments. MMS wrote the original draft, and all authors edited and reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sikder, M.M., Vestergård, M., Kyndt, T. et al. Genetic disruption of Arabidopsis secondary metabolite synthesis leads to microbiome-mediated modulation of nematode invasion. ISME J 16, 2230–2241 (2022). https://doi.org/10.1038/s41396-022-01276-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-022-01276-x
This article is cited by
-
The Arabidopsis holobiont: a (re)source of insights to understand the amazing world of plant–microbe interactions
Environmental Microbiome (2023)