Abstract
Vibrio cholerae is a well-studied human pathogen that is also a common inhabitant of marine habitats. In both environments, the bacterium is subject to interbacterial competition. A molecular nanomachine that is often involved in such competitive behavior is the type VI secretion system (T6SS). Interestingly and in contrast to non-pandemic or environmental isolates, the T6SS of the O1 El Tor clade of V. cholerae, which is responsible for the ongoing 7th cholera pandemic, is largely silent under standard laboratory culture conditions. Instead, these strains induce their full T6SS capacity only under specific conditions such as growth on chitinous surfaces (signaled through TfoX and QstR) or when the cells encounter low intracellular c-di-GMP levels (TfoY-driven). In this study, we identified a single nucleotide polymorphism (SNP) within an intergenic region of the major T6SS gene cluster of V. cholerae that determines the T6SS status of the cell. We show that SNP conversion is sufficient to induce T6SS production in numerous pandemic strains, while the converse approach renders non-pandemic/environmental V. cholerae strains T6SS-silent. We further demonstrate that SNP-dependent T6SS production occurs independently of the known T6SS regulators TfoX, QstR, and TfoY. Finally, we identify a putative promoter region adjacent to the identified SNP that is required for all forms of T6SS regulation in V. cholerae.
Similar content being viewed by others
Competition between microbes occurs frequently in nature. A widespread molecular nanomachine in Gram-negative bacteria that is involved in interbacterial competition is the type VI secretion system (T6SS). T6SSs resemble inverted contractile phage tails, which, upon contraction, propel a molecular spear out of the bacterium and into neighboring cells. They thereby deliver cocktails of effector proteins for prey intoxication. The producing bacterium and its siblings are protected from self-harm through cognate antitoxins/immunity proteins [1].
The dual lifestyle of Vibrio cholerae makes it an ideal model organism to study T6SS-mediated interbacterial competition. Apart from being an important human pathogen, it is also a natural inhabitant of aquatic environments, where it frequently associates with zooplankton and their molted chitinous exoskeletons [2]. Importantly, chitin is a potent inducer of several phenotypes in V. cholerae such as natural competence for transformation and type VI secretion [3, 4]. Furthermore, the coupling of competence and T6S fosters the horizontal transfer of prey-released DNA [4, 5].
Chitin-dependent induction of competence and T6SS is triggered through the TfoX signaling pathway, which includes three principal regulatory proteins: TfoX, whose production is tightly linked to growth on chitin [3]; HapR, the master regulator of quorum sensing (QS), which signals high cell density [6]; and the downstream-acting transcription factor QstR [7, 8] (reviewed in [9]). In addition to chitin-dependent regulation, the T6SS is also inducible in V. cholerae and other Vibrios by the regulatory protein TfoY [10,11,12]. However, although TfoY translation occurs at low intracellular c-di-GMP levels [10, 11], the natural trigger for its production remains unknown.
TfoX-, QstR-, and TfoY-dependent T6SS induction was primarily demonstrated for V. cholerae strains that are responsible for the ongoing 7th cholera pandemic (O1 El Tor strains; referred to as 7PET clade), hereinafter referred to as “inducible strains”. Indeed, since its discovery, it is well known that the T6SS of 7PET strains is largely silent under standard laboratory growth conditions [13]. For this reason, the structural and mechanistic investigation of the T6SS machinery over the past 15 years has primarily relied on strains that are constitutively T6SS active under laboratory conditions, using either non-pandemic toxigenic strains (e.g., O37 serogroup strains V52 and ATCC25872) or various environmental isolates [14,15,16] (hereinafter referred to as “constitutive strains”). Notably, while we categorize them as constitutively active, we cannot exclude that these strains retain means to still regulate their T6SS and turn it off under yet to be discovered conditions. In this study, we asked the question “What caused the switch from seemingly constitutive to inducible T6SS activity in 7PET strains compared to their non-pandemic or environmental relatives?”.
To address this question, we generated a library of 800 hybrid strains in which each clone carried a mosaic genome between the 7PET pandemic strain A1552 (referred to as “T6SS OFF”, as the strain’s T6SS is mostly silent under the tested conditions) and the non-pandemic strain ATCC25872 (referred to as “T6SS ON” given the strain’s constitutive T6SS activity under standard laboratory growth conditions) (Fig. 1A). The library design was based on our previous work on transformation-mediated horizontal gene transfer (HGT) in V. cholerae whereby exchanges above 100 kb occurred frequently [5]. Specifically, we first created derivatives of strain A1552 bearing an antibiotic resistance marker (aph) at 40 different positions throughout its genome, spaced ~100 kb apart (Fig. 1A). Isolated genomic DNA of these 40 strains was then used to transform strain ATCC25872, and 20 transformants were isolated per reaction (Fig. 1A). The resulting 800 hybrid transformants were then tested for T6SS activity using a fluorescence imaging-based E. coli killing experiment (adapted from [12]). Strikingly, 19/20 hybrid clones that had transferred the aph marker at position #32 (aph#32) had lost their T6SS activity (T6SS ON > OFF; Fig. S1A). The reverse experiment (i.e., transfer of aph#32 from ATCC25872 to A1552) resulted in 7/20 transformants that had gained T6SS activity (T6SS OFF > ON; Fig. S1A). Since the aph#32 cassette was located ~15 kb upstream of the major/large T6SS cluster (Fig. 1B), we hypothesized that the genomic region that drives constitutive T6SS production under the tested conditions might be close to or inside this cluster. Indeed, when we repeated the transfer experiments using strains carrying an insertion (aph#42) immediately upstream of paar1 (first gene in this cluster), this resulted in 20/20 T6SS phenotypic conversion events in the ON > OFF direction and 19/20 events in the OFF > ON direction (Fig. S1A).
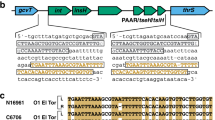
A Scheme of the hybrid strain library construction. 40 input strains in the A1552 strain background (pandemic isolate) were genetically engineered to each carry an aph resistance marker at a different genomic location (~every 100 kb; aph#1 to aph#40). Genomic DNA (gDNA) of these strains was used to transform the non-pandemic strain ATCC25872. 20 transformants of each reaction were kept resulting in a final hybrid strain library consisting of 800 strains. B Scheme of the large T6SS cluster of V. cholerae and the location of aph#32 and aph#42. The zoomed 431-bp intergenic region between the second (VCA0106) and third (vipA) gene of the cluster is shown below together with an alignment covering the start of this region and comparing the sequence of the pandemic strain A1552, the non-pandemic strain ATCC25872, and four aph#42 transformants of strain A1552. The T6SS-ON version of SNP45 (“T”) is highlighted in yellow and the strains’ T6SS activity status is shown on the right. C Sequence alignment of the same region as in (B) comparing the pandemic/non-pandemic control strains with 15 environmental V. cholerae isolates. D–F The SNP45-converted pandemic/non-pandemic clones and their parental strains were scored for (D) T6SS assembly (by imaging structures made of the T6SS sheath protein VipA-sfGFP; Scale bars: 2 µm); (E) T6SS activity (through Western Blot-based detection of the secreted T6SS tube protein Hcp); and (F) interbacterial killing of E. coli prey. Numbers of surviving prey are depicted on the Y-axis (CFU/ml) and each bar represents the average of four independent biological replicates, as indicated by individual dots (±SD). d.l. detection limit. G SNP45 conversion to T, C, or A in the pandemic strain A1552. Bacterial killing assay as in (F) with three biologically independent replicates. Statistical significance using a one-way ANOVA followed by a Šídák’s multiple comparisons test is indicated comparing each WT with its SNP-converted derivative (F) or each of the SNP45 convertants with the WT (G). ****p < 0.0001; n.s. not significant.
We compared the genome sequences surrounding aph#42 in strains A1552 [17] and ATCC25872 (see Supplementary Material and Methods). In addition, we Sanger sequenced the respective region of three T6SS OFF > ON-converted transformants plus the single non-converted T6SS OFF hybrid clone as a negative control. As shown in the alignment (Fig. 1B), this comparison revealed a perfect correlation between the T6SS status and a single nucleotide polymorphism (SNP) at position 45 of the intergenic region (hereon referred to as SNP45), downstream of the second gene of the T6SS cluster (VCA0106), whereby “G” resulted in a silenced T6SS, and “T” rendered the transformant T6SS active (Fig. 1B). This finding is strongly supported by the status of SNP45 in 15 environmental V. cholerae strains (Fig. 1C) as well as all examined 7PET strains, as was also recently confirmed in a preprint by Ng et al. [18].
To prove causality between SNP45 and T6SS activity, we investigated nucleotide 45 using site-directed mutagenesis. SNP45 conversion (G→T) in the pandemic strain A1552 (T6SS OFF) led to expression of the T6SS genes (Fig. S1B and Table S1), assembly of the T6SS machinery, secretion of the T6SS inner tube protein Hcp, and, ultimately, to killing of the E. coli prey (Fig. 1D–F). Conversely, SNP45 conversion (T→G) in the non-pandemic strain ATCC25872 (T6SS ON) silenced T6SS activity (Figs. 1D–F, S1C, and Table S2). We confirmed these SNP45-conversion data in five additional 7PET strains (Fig. S2A, B) that were isolated over the past ~40 years from three different continents (Table S3) as well as in a selection of environmental isolates (Fig. S2C, D). Finally, SNP45 conversion G→C in strain A1552 resulted in an intermediate activation of T6SS killing, whereas the G→A conversion remained T6SS silent (Fig. 1G).
In order to get a first insight into the nature of the SNP45-based regulation, we tested whether T6SS activation in the SNP45-converted pandemic strain was dependent on the known regulators TfoX, QstR, and TfoY, which turned out not to be the case (Fig. 2A). We therefore reasoned that an additional regulatory element(s) might be present in the SNP45-harboring intergenic region and therefore stepwise shortened this region (Fig. 2B). Interestingly, while deletion of 276 bp upstream of vipA did not impair killing, T6SS activity was completely abolished in a strain lacking 336 bp upstream of vipA, despite the presence of SNP45 (Fig. 2C). Consistent with this finding, deletion of the region that differed between these two constructs (60 bp in size) was sufficient to abolish T6SS activity in the SNP45-converted (G→T) pandemic strain as was the deletion of the almost complete intergenic region (∆full; Fig. 2C). Visual inspection of the intergenic region surrounding the SNP and encompassing the 60 bp region revealed a putative promoter with appropriately positioned −35 and −10 elements [19] (Fig. 2D). This putative promoter is located >300 bp upstream of the next annotated gene (vipA), which results in an unusually long 5′ untranslated region (5’ UTR). Indeed, previous analysis of 2,454 genes indicated that the length of the majority of 5′ UTRs ranged between 20 to 40 nucleotides in V. cholerae [20]. We nonetheless hypothesized that this putative promoter might be involved in T6SS regulation. Site-directed mutations designed to disable the −10 element (TAGAAT to TAGGCT) eliminated bacterial killing of the SNP45-converted (G→T) strain, confirming its importance in T6SS regulation. The exact mechanistic interplay between these two regulatory elements remains, however, unknown and might include unidentified transcriptional regulators, sRNA(s), the putative long 5’UTR of vipA, or other players. Importantly, this promoter region was also necessary for TfoX-, QstR-, or TfoY-driven T6SS production in the wild-type pandemic strain A1552 (Fig. 2E), which is consistent with increased vipA transcript levels under such conditions [4, 8, 10].
A Strains lacking known T6SS regulators maintain T6SS activity of the SNP45-converted pandemic strain. Pandemic strain A1552 or its SNP45-converted derivate were genetically engineered to delete tfoX, qstR, tfoY, hapR, or the two T6SS structural genes vipB and vasK as controls. All strains were tested in an E. coli killing assay. B Scheme of truncations introduced within the intergenic region. The intergenic region was shortened by 276 bp, 336 bp or entirely deleted (396 bp deleted; ∆full), leaving solely 10 bp downstream of VCA0106 and 25 bp upstream of vipA intact (gray boxes). C T6SS activity of strains with a truncated intergenic region as depicted in (B) was assessed in an E. coli killing assay. D A promoter signature is located close to SNP45. Comparison of SNP45-converted (yellow label) intergenic region in the Δ276 and Δ336 mutants with the putative −35 and −10 promoter elements highlighted in purple. The 60 bp region deleted in the ∆60 mutant is underlined in the left scheme. The “AA” nucleotides within the −10 element that were changed to “GC” in the respective mutants in (E) are shown in boldface. E No T6SS activation occurs in strains with a mutated −10 element, as assessed in an E. coli killing assay. Neither SNP45-conversion nor arabinose (0.2%)-induced production of TfoX, QstR, and TfoY (from the respective Tn constructs, as described [8, 10]) led to T6SS-mediated prey killing in the mutant strain carrying a defective −10 element (mut-10) in contrast to the WT background. Details for the E. coli killing assays in panels (A, C, E) are as described in Fig. 1. Bar plots represent the average of at least three independent biological replicates as shown by the individual dots (±SD). Only statistically significant comparisons are indicated in the plots using one- or two-way ANOVA followed by a Šídák’s multiple comparisons test. The comparisons were: (A) each mutant with its parental strain for WT or SNP45-converted A1552; (C) all strains against the parental WT strain of A1552 shown in boldface; and (E) without or with arabinose conditions for each strain. *p < 0.05; ***p < 0.001; ****p < 0.0001.
Collectively, we have identified a SNP in V. cholerae that is deterministic of T6SS production. Notably, the SNP nucleotide of the 7th pandemic V. cholerae clade renders the bacterium’s T6SS silent under standard laboratory conditions. This loss-of-function finding is in contrast to a gain-of-function single nucleotide mutation that was recently described for an epidemic African clade of Salmonella, which was proposed to promote bacterial survival and dissemination during human infection [21]. Interestingly, recent work has shown that strains responsible for earlier cholera pandemics, and specifically those belonging to the classical clade of 6th pandemic strains, contain multiple frameshift mutations/deletions in their structural T6SS genes that render them T6SS silent [22]. It is therefore tempting to speculate that the displacement of the classical clade by 7PET clade strains was in part driven by their superior T6SS regulation. Indeed, by keeping their T6SS mostly silent under non-inducing conditions, 7PET strains might keep intestinal inflammation to a minimum [23], while maintaining the ability to produce their T6SS machine “on demand” (e.g., during competition and HGT on chitinous surfaces). Our work will therefore prompt future studies on T6SS regulation in V. cholerae and its involvement in a disease context.
References
Galan JE, Waksman G. Protein-injection machines in bacteria. Cell. 2018;172:1306–18.
Lipp EK, Huq A, Colwell RR. Effects of global climate on infectious disease: the cholera model. Clin Microbiol Rev. 2002;15:757–70.
Meibom KL, Blokesch M, Dolganov NA, Wu C-Y, Schoolnik GK. Chitin induces natural competence in Vibrio cholerae. Science. 2005;310:1824–7.
Borgeaud S, Metzger LC, Scrignari T, Blokesch M. The type VI secretion system of Vibrio cholerae fosters horizontal gene transfer. Science. 2015;347:63–7.
Matthey N, Stutzmann S, Stoudmann C, Guex N, Iseli C, Blokesch M. Neighbor predation linked to natural competence fosters the transfer of large genomic regions in Vibrio cholerae. eLife. 2019;8:e48212.
Papenfort K, Bassler BL. Quorum sensing signal-response systems in Gram-negative bacteria. Nat Rev Microbiol. 2016;14:576–88.
Lo Scrudato M, Blokesch M. A transcriptional regulator linking quorum sensing and chitin induction to render Vibrio cholerae naturally transformable. Nucleic Acids Res. 2013;41:3644–58.
Jaskólska M, Stutzmann S, Stoudmann C, Blokesch M. QstR-dependent regulation of natural competence and type VI secretion in Vibrio cholerae. Nucleic Acids Res. 2018;46:10619–34.
Veening JW, Blokesch M. Interbacterial predation as a strategy for DNA acquisition in naturally competent bacteria. Nat Rev Microbiol. 2017;15:621–9.
Metzger LC, Stutzmann S, Scrignari T, Van der Henst C, Matthey N, Blokesch M. Independent Regulation of Type VI Secretion in Vibrio cholerae by TfoX and TfoY. Cell Rep. 2016;15:951–8.
Metzger LC, Matthey N, Stoudmann C, Collas EJ, Blokesch M. Ecological implications of gene regulation by TfoX and TfoY among diverse Vibrio species. Environ Microbiol. 2019;21:2231–47.
Ben-Yaakov R, Salomon D. The regulatory network of Vibrio parahaemolyticus type VI secretion system 1. Environ Microbiol. 2019;21:2248–60.
Pukatzki S, Ma AT, Sturtevant D, Krastins B, Sarracino D, Nelson WC, et al. Identification of a conserved bacterial protein secretion system in Vibrio cholerae using the Dictyostelium host model system. Proc Natl Acad Sci USA. 2006;103:1528–33.
Basler M, Pilhofer M, Henderson GP, Jensen GJ, Mekalanos JJ. Type VI secretion requires a dynamic contractile phage tail-like structure. Nature. 2012;483:182–6.
Unterweger D, Miyata ST, Bachmann V, Brooks TM, Mullins T, Kostiuk B, et al. The Vibrio cholerae type VI secretion system employs diverse effector modules for intraspecific competition. Nat Commun. 2014;5:3549.
Drebes Dörr NC, Blokesch M. Interbacterial competition and anti-predatory behavior of environmental Vibrio cholerae strains. Environ Microbiol. 2020;22:4485–504.
Matthey N, Drebes Dörr NC, Blokesch M. Long-Read-Based Genome Sequences of Pandemic and Environmental Vibrio cholerae Strains. Microbiol Resour Announc. 2018;7:e01574–18.
Ng SL, Kammann S, Steinbach G, Hoffmann T, Yunker PJ, Hammer BK. Evolution of a cis-acting SNP that controls Type VI Secretion in Vibrio cholerae. bioRxiv. 2022. https://doi.org/10.1101/2022.01.11.475911.
Hawley DK, McClure WR. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983;11:2237–55.
Papenfort K, Forstner KU, Cong JP, Sharma CM, Bassler BL. Differential RNA-seq of Vibrio cholerae identifies the VqmR small RNA as a regulator of biofilm formation. Proc Natl Acad Sci USA. 2015;112:E766–75.
Hammarlöf DL, Kröger C, Owen SV, Canals R, Lacharme-Lora L, Wenner N, et al. Role of a single noncoding nucleotide in the evolution of an epidemic African clade of Salmonella. Proc Natl Acad Sci USA. 2018;115:E2614–23.
Kostiuk B, Santoriello FJ, Diaz-Satizabal L, Bisaro F, Lee KJ, Dhody AN, et al. Type VI secretion system mutations reduced competitive fitness of classical Vibrio cholerae biotype. Nat Commun. 2021;12:6457.
Zhao W, Caro F, Robins W, Mekalanos JJ. Antagonism toward the intestinal microbiota and its effect on Vibrio cholerae virulence. Science. 2018;359:210–3.
Acknowledgements
The authors thank C. Stoudmann for technical assistance, Eve Rahbé for a transposon mutagenesis screen in strain ATCC25872, A. Vanhove for assisting with the stocking of the hybrid strain library, and members of the Blokesch group for fruitful discussions. The authors also acknowledge the staff of the Lausanne Genomic Technologies Facility at the University of Lausanne for sample processing, PacBio sequencing, and genome assembly and S. Strempel (Microsynth) for the RNA-seq analysis.
Funding
This work was supported by the Swiss National Science Foundation (310030_185022), the Novartis Foundation for medical-biological Research (#18C178), and a Consolidator grant by the European Research Council (724630). MB is a Howard Hughes Medical Institute (HHMI) International Research Scholar (grant 55008726). Open access funding was provided by EPFL Lausanne.
Author information
Authors and Affiliations
Contributions
NCDD and MB conceived the study and designed the experiments; NCDD, AP, MJ, SS, LB, and MB performed experiments; MJ identified the promoter region; NCDD, and MB analyzed the data; MB secured funding and supervised the study; NCDD and MB drafted the paper; AP and MJ, commented on the paper, and all authors approved its final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Drebes Dörr, N.C., Proutière, A., Jaskólska, M. et al. Single nucleotide polymorphism determines constitutive versus inducible type VI secretion in Vibrio cholerae. ISME J 16, 1868–1872 (2022). https://doi.org/10.1038/s41396-022-01234-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-022-01234-7
This article is cited by
-
Community composition and the environment modulate the population dynamics of type VI secretion in human gut bacteria
Nature Ecology & Evolution (2023)