Abstract
Protozoan predators form an essential component of activated sludge communities that is tightly linked to wastewater treatment efficiency. Nonetheless, very little is known how protozoan predation is channelled via bacterial communities to affect ecosystem functioning. Therefore, we experimentally manipulated protozoan predation pressure in activated-sludge communities to determine its impacts on microbial diversity, composition and putative functionality. Different components of bacterial diversity such as taxa richness, evenness, genetic diversity and beta diversity all responded strongly and positively to high protozoan predation pressure. These responses were non-linear and levelled off at higher levels of predation pressure, supporting predictions of hump-shaped relationships between predation pressure and prey diversity. In contrast to predation intensity, the impact of predator diversity had both positive (taxa richness) and negative (evenness and phylogenetic distinctiveness) effects on bacterial diversity. Furthermore, predation shaped the structure of bacterial communities. Reduction in top-down control negatively affected the majority of taxa that are generally associated with increased treatment efficiency, compromising particularly the potential for nitrogen removal. Consequently, our findings highlight responses of bacterial diversity and community composition as two distinct mechanisms linking protozoan predation with ecosystem functioning in activated sludge communities.
Similar content being viewed by others
Introduction
The treatment of wastewater using activated sludge communities represents arguably the largest single biotechnological process world-wide [1]. This crucial ecosystem service is provided by diverse communities of bacteria, protozoans and metazoan grazers [2,3,4,5]. Past research has highlighted that the effective biological treatment of wastewater critically depends on the composition and diversity of bacterial assemblages [6, 7]. However, also protozoan predators play a key role in maintaining treatment efficiency in activated sludge [8,9,10,11]. Characteristic predators, such as ciliates and heterotrophic nanoflagellates (HNFs) express dynamic changes in their densities and complex successional patterns [12, 13]. Their total density is, nonetheless, often positively associated with essential bacterial functions, such as denitrification and the reduction of biological oxygen demand (BOD) in treatment plant effluent [9].
The positive impacts of protozoan predation on ecosystem functioning have been traditionally explained by stimulating effects on bacterial physiology [8, 10]. For example, protozoa may excrete growth-stimulating substances that boost bacterial activity [4]. Predation plays also an important role maintaining high bacterial growth rates enhancing nutrient re-mineralisation and carbon respiration [10, 14, 15]. In contrast, direct impacts of predation on prey community composition are much less studied in activated sludge communities [16, 17]. However, the strength of direct predator–prey interactions [18] and their importance for ecosystem functioning is well demonstrated in other systems [16, 19, 20], highlighting a potential route for further optimisations of biological wastewater treatments.
One link with potentially considerable consequences for ecosystem functioning is the relationship between protozoan predation and bacterial diversity. Diversity is well-known to increase the rate of ecosystem functioning [21,22,23] and promote multiple aspects of ecosystem stability [24, 25], including a greater toxin resistance of more diverse activated sludge communities [26]. However, the relationship between predation pressure and prey diversity is not always positive [27, 28], and both positive and negative effects of predation on prey diversity have been documented [29,30,31]. This has led to the postulation of a hump-shaped relationship between prey diversity and the strength of predation pressure [27, 32].
This hump-shaped relationship is thought to emerge because intermediate predation pressure facilitates the co-existence of multiple prey strategies [28, 33]. More predation resistant K- and opportunistic r-strategists may equally persist at intermediate levels of top-down control (Fig. 1A). Predator-mediated prey co-existence is particularly favoured in systems where predator densities fluctuate over time [34], as frequently observed in activated sludge communities [4, 35]. The strength of predation pressure that maintains such peak prey diversity is believed to be mediated by nutrient concentrations and resulting ecosystem productivity [27]. Higher productivity is reflected in higher prey population growth rates, which requires a stronger top-down control of opportunistic r-strategists to facilitate prey coexistence (Fig. 1A). Activated sludge reactors are engineered ecosystems characterised by high nutrient concentrations and microbial carrying capacities [e.g. [8]]. The predation pressure required to maintain peak prey diversity is therefore expected to be much higher than in many natural ecosystems, potentially resulting in almost linear relationships between predation pressure and prey diversity (Fig. 1A). This conceptual framework may thus explain the frequently observed positive knock-on effects of predator density on treatment efficiency in activated sludge communities [9, 10].
A At low pressure (I), predators are unable to control rapidly growing r-strategists resulting in the exclusion of slower growing prey taxa [27]. At intermediate predation pressure (II) more K-strategists resistant to predation start to emerge and the co-existence of different strategies leads to a peak in prey diversity. A further increase in predation pressure (III) benefits K-strategists as it promotes the exclusion of less defended, opportunistic prey. This relationship is proposed to be mediated by ecosystem productivity [i.e. nutrient level, [28, 33]]. In extremely nutrient rich water treatment reactors (dotted line), this is expected to lead to a largely positive impact of protozoan predation on bacterial diversity. We performed eight experiments (B) in which we manipulated predation pressure by diluting activated sludge (AS) communities with growth media (M). The dilutions resulted in a reduction in predator-prey encounter rates and hence predation pressure, while growth conditions remained relatively constant, effectively shifting conditions to the left on the x-axis in panel A. Four out of 8 experiments where pre-conditioned in chemostats and three of them were pre-filtered to remove the largest fraction of predators from experimental communities and to diversify the types of communities tested.
In addition to impacting prey diversity, protozoans can alter the identity of dominant bacterial taxa [17, 36] and selective predation may change the relative densities of functionally important bacteria in water treatment reactors. Indeed, different protozoans such as bacterivorous Chilodonella and Colpidium are associated with higher treatment efficiency [37], whereas others (e.g. the HNFs Bodo and Polytoma) appear to have predominantly negative impacts [9]. Currently, the mechanisms that underlie such shifts in functional identity and the direct impacts of protozoan predation on bacterial community composition remain unexplored. Moreover, the relationship between prey and predator diversity is conceptually poorly understood [38, 39], limiting our potential to further optimise sewage treatment by activated sludge communities.
Our aim was to determine the effect of protozoan predation intensity on bacterial diversity and community composition in activated sludge. We used a series of dilution experiments, developed to quantify the impacts of predation pressure on plankton communities [40, 41], in order to experimentally control the strength of protozoan predation. Metabarcoding and flow cytometric analyses of prey and predators allowed us to characterise microbial communities and responses to reductions in top-down control. Specifically, we quantified changes in bacterial alpha and beta diversity in response to reduced levels of predation pressure. Furthermore, we investigated relationships between bacterial and protozoan diversity to evaluate inter-trophic linkages in richness, evenness and genetic diversity. Finally, we examined whether reduced top-down control resulted in systematic shifts in community composition, gauging potential consequences for the efficiency of wastewater treatment plants.
Materials and methods
Sample collection and preparation
Activated sludge samples were collected from the Severn Trent wastewater treatment plant in Derby (UK) between 9:30 and 11:30 am on 14th February 2019. Aeration tanks contained four fully separated lanes (no water exchange). We collected 800 mL of suspended activated sludge from each of the four lanes as inocula for laboratory experiments. We also collected 40 L of influent to the biological treatment tank, i.e. wastewater that had already undergone primary treatment. These 40 L were filtered on site (75 uL mesh sieves to remove debris), autoclaved and used for the preparation of experimental growth media. All samples were stored in insulated coolers, kept in the dark and transported to the laboratory within 3 h.
Priming of communities prior to experiments
In total, we conducted eight dilution experiments (Fig. 1B). Four of these experiments (labelled as experiments 1–4) were directly inoculated with microbial communities from one of the four treatment plant lanes (all four lanes at Derby Treatment plant were sampled). The other four experiments (experiments 5–8) were established from the outflow of four different continuous flow-through chemostats, which were inoculated with activated sludge (the same sample from lane 1; see Fig. S1 for details about chemostat design and operation). Chemostat were run for two weeks before the start of dilution experiments and they were implemented for two reasons as conditioning pre-treatments for microbial communities. First, activated sludge community composition can be substantially influenced by bacteria entering over the inflow [42]. The experiments with cultures from chemostats that used filtered and sterilised media, marginalised the impact of inflow bacteria and allowed to control for potentially confounding effects on community composition. Second, the use of chemostats allowed to diversify experimental communities, which allowed us to double the number of experiments and increase the generality of our findings. Dilution rates in chemostats impose unselective background mortality rates on predator and prey taxa and filtration of inocula selectively excludes certain community members (e.g. rotifers and larger, tentatively carnivorous ciliates). We therefore initiated chemostats with either unfiltered or prefiltered (50 µm mesh size) activated sludge samples, and operated chemostats at different dilution rates in order to prime different predator assemblages (chemostat for experiment 5: unfiltered and a dilution rate of 0.35 d−1; chemostats for experiments 6-8: pre-filtered with dilutions rates of 0.35, 0.5, 0.2 d-1, respectively). The use of autoclaved treatment plant influent, which is rich in organic substrates [43], as growth media helped to maintain a high microbial diversity over the course of the conditioning phase (Fig. S2).
Experimental set-up and sampling
Dilution experiments are based on the principle of diluting microbial communities with organism free ambient water [40]. The impact of predation on prey community composition and diversity can be assessed by this method because predation pressure is reduced (lowered encounter rates), whereas growth conditions for prey species are relatively unaffected [40]. For each of our eight experiments, we established six duplicated dilution treatments in 50 mL falcon tubes (in total 96 microcosms with 5 mL volume). Microcosms were established by combining an inoculum with autoclaved and filtered (0.2 µm nylon filters) influent. The six dilution treatments per experiment included 100%, 60%, 30%, 10%, 5% and 1% of inoculum. Experiment 4 was inadvertently set up with a slightly altered dilution series including 100%, 38%, 24%, 10%, 6.6%, 2.4% of inoculum. To obtain enough DNA for next-generation sequencing, additional microcosms for the 100% and 1% inoculum treatments were set up containing larger volumes (20 mL and 200 mL total volume, respectively; two replicates each). Microcosms were continuously homogenised on a shaking table (120 rotations min−1) and kept in the dark at 20 ± 0.5 ˚C. After 24 h, all microcosms were sampled for flow cytometry and the lowest and highest dilution were sampled for next-generation sequencing. Prior to the experiment, all inocula were also sampled in triplicates to determine starting conditions.
For flow cytometry, 0.9 mL from each microcosm were sampled to measure ratios of high nucleic acid (HNA) to low nucleic acid (LNA) bacterial cells, and 2.7 mL were taken to enumerate HNF densities. Samples were fixed with paraformaldehyde and glutaraldehyde, shock frozen in liquid nitrogen and stored at −80 ˚C following protocols by Gasol and Morán [Fig. S3; [44]]. Samples for DNA extraction were collected by pressure filtration and material was collated until filters clogged (20 mL from undiluted communities, 100 mL from diluted communities; 0.2 µm polycarbonate filters, Cyclopore Whatman, UK). All filters were shock frozen and stored at −80 ˚C.
Flow cytometry and high-throughput sequencing
In all experiments, we assessed prey and predator community composition applying a meta-barcoding approach. Additionally, we used flow cytometry to evaluate HNA–LNA ratios of bacteria, which are interpreted as a potential indicator of bacterial cell activity [45]. Enumeration of bacterial density with flow cytometry was not reliable as many taxa were particle-associated confounding accurate quantification. Moreover, we quantified HNF densities in undiluted samples (deemed technically not feasible in undiluted samples), representing one important fraction of grazer communities.
HNF densities and HNA–LNA bacteria ratios were analysed on a BD Accuri C6 automatic flow cytometer (BD Biosciences, USA) following largely the protocol by Gasol and Morán [[44]; for further details see SI, section S1]. DNA for meta-barcoding analyses was extracted with the QIAGEN DNeasy Blood and Tissue Kit, following the manufacturer’s protocols. The 16S rRNA gene (V3-V4 region) from the DNA samples were amplified using the universal bacterial primers [46], 515F (5′-GTGYCAGCMGCCGCGGTAA-3′) and 806R (5′-GGACTACNVGGGTWTCTAAT-3′). Additionally, we targeted eukaryotic sequences amplifying the 18S rRNA gene using the primers 574*f (5′-CGGTAAYTCCAGCTCYV-3′) + 1132r - (5′-CCGTCAATTHCTTYAART-3′) based on Hugerth et al. [47]. Barcodes were added via PCR and the amplicons were then cleaned up using a bead-based kit (AMPure XP, Beckman Coulter, US), pooled and sequenced (2 × 250 bp) on the MiSeq (Illumina, US) platform [48].
Sequence and statistical analysis
Raw sequence reads were first quality controlled for chimera and sequence fragments (72% and 64% of raw sequences remained for prokaryotes and protozoa respectively) in QIIME2 [49]. DNA-polymerase sequencing errors were accounted for using the dada2 algorithm [50] to attain relative frequencies of amplicon sequence variants (ASVs). The mean number of reads per sample was 69,018 ± 12,345 (SD) for prokaryotes and 29,561 ± 18,502 for protozoa. Total number of reads in some protozoan samples were relatively low due to primer or PCR inhibition. We eliminated samples with low total copy number (<15,000) from further analysis before rarefication, resulting in a replication of 10, 12 and 15 samples from the reduced grazing, the ambient grazing and start samples, respectively. The taxonomic identity of prokaryote ASVs was determined using the SILVA RNA database at 99% similarity [release 138; [51]] and a multinomial Naive Bayes classifier trained for the selected V4 sequence in QIIME 2. However, we maintained the recently challenged family of the Comamonadaceae to aid comparability with earlier studies. All non-assigned ASVs at the Kingdom level, and all chloroplast ASVs, were removed from the analyses. As bacteria dominated our samples (only 0.12% of ASVs were Archea), we henceforth refer to prokaryote as ‘bacterial’ ASVs. Taxonomic identity of numerically important ASVs was confirmed by blast-searching and checking manually the 100 most abundant ASVs across all samples on the NCBI database. Protozoan sequences were analogously classified using the SILVA database at 99% similarity [51]. To assure that we only considered bacterial predators and avoided contamination (e.g. mammalian DNA), we considered only taxa that were affiliated to the classes Alveolata, Rhizaria, Discoba, Discosea or Holozoa. Within Holozoa, we also included the potentially bacterivorous taxa Chromadorea, Bdelloidea and Phyllopoda. However, as Holozoa comprised only a small subfraction of all taxa and reads, we refer hereafter to ‘predator ASVs’ as protozoans. Phylogenetic trees were constructed using the FastTree software [52]. All samples were uploaded to NCBI database (PRJNA726629).
The effect of dilution on alpha diversity was assessed by comparing ASV richness, ASV evenness (Pielou’s evenness) and genetic diversity measured by the Faith index [53], after rarefaction to standardise sampling effort to the lowest sequencing depth. We also assessed mean phylogenetic distinctiveness of ASVs following Tsirogiannis and Sandel [54]. Phylogenetic distinctiveness is a measure based on the Faith index, which removes the effect of species richness on genetic diversity using a bootstrapping approach (1000 iterations). We applied a linear mixed effects model to determine differences in diversity metrics among communities at the start of incubations as well as in diluted and undiluted communities (also referred to as reduced-predation and predation treatment, respectively) at the end of incubations. Experiment identity (experiment 1–8) was accounted for as random effect. We also compared relative abundances of ASVs between predation and reduced predation treatments at the end of the experiments using a non-parametric factorial analysis after Wobbrock et al. [55], again including experiment number as random effect.
A community similarity matrix was established based on Bray–Curtis similarity and visualised using non-metric multidimensional scaling (NMDS; stress value of 0.08). We then applied ANOVA with subsequent Tukey post-hoc tests to evaluate whether (i) communities in the predation or reduced-predation treatments at the end of the experiments were more similar in composition to the starting (inocula) communities and (ii) beta diversity (i.e. dissimilarity among communities) was different among the communities in the start inocula, predation or reduced predation treatments. Non-parametric tests were used when variance-homogeneity could not be achieved through transformation. Finally, we used ordinary least squares regressions to test the effect of HNF densities on prey alpha diversity within treatments (i.e. a separate analysis for communities with reduced and normal predation pressure) to assess whether this relationship is consistent at low and high predation pressure. Because we were able to measure HNF densities in undiluted samples only, we used the starting HNF densities for these within treatment assessments. We examined whether regression model residuals met the assumptions of normality, equal variances, and were not autocorrelated. All implemented regression models met these requirements. Nonlinearity between dependent and explanatory variables was assessed visually and by comparing models with log-transformed, exponentially-transformed and untransformed independent variables based on the smallest Akaike’s Information Criterion [AIC, corrected for small sample size; [56]].
Finally, we applied two complementary approaches to examine how shifts in bacterial community composition affected their putative functionality. First, we used an automated, taxonomy inferred approach to predict potential functional differences between treatments [METAGENassist [57]; results only presented in SI]. Second, we related our results to a global meta-analysis of activated sludge communities [7], which provides the functional association of commonly occurring taxa (>20% occurrence across samples in meta-analysis). We compared all ASVs related to those taxa and evaluated significant responses in relative abundance to microcosm dilution. All analyses were performed in R, version 3.6 [58], and all R-scripts are provided in Annex 1.
Results
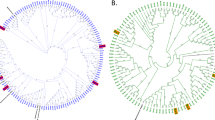
Experimental predator communities had a mean ASV-richness of 72 ± 28 (SD) and were dominated in richness and relative abundance by ciliates (mainly Peritrichia and Suctoria) and amoeba (primarily Rhizaria; Fig. 2, Fig. S4). Both treatment implementation (i.e. dilution to reduce prey encounter rates and thus predation pressure) and filtration, during the experimental conditioning phase, had significant impacts on predator diversity (Tables 1 and TS1). However, they affected different components of predator diversity. Whereas filtration significantly reduced taxa richness, dilution lowered phylogenetic diversity of predators (Fig. 2C, Table S1). Filtration during the conditioning phase also had a marked impact on predator community composition, significantly reducing relative densities of Haptoria, Phyllopharyngea and other rare protozoan families (paired Wilcox-Test, W > 326, p < 0.001). Yet, overall protozoan taxonomy was not well resolved as 31.1% of ASVs could only be assigned to class level.
Protozoan community differed between unfiltered (A) and pre-filtered (B) communities. In each panel, box plots for each taxonomic class in microcosms with ambient predation pressure (P), reduced predation pressure (RP) and at starting conditions (S) are illustrated. In C, responses of protozoan diversity (i.e. taxa richness, evenness and phylogenetic distinctiveness) to treatment implementation and filtration in the priming phase of the experiment (50 μm) are displayed. Points represent sample means, bars represent ±1 standard error of the mean.
The diversity of bacterial prey communities was strongly influenced by the experimental dilution and filtration during the conditioning phase (Fig. 3). Both manipulations additively reduced different bacterial diversity components, including richness (R2 = 0.82, p < 0.001), evenness (R2 = 0.56, p < 0.001) and phylogenetic distinctiveness (R2 = 0.55, p < 0.001). Notably, communities with high richness were less sensitive to negative effects of dilution highlighted by their lower loss rates in phylogenetic distinctiveness in diluted microcosms (p = 0.003, R2 = 0.75, y = 0.004x -4.7; Fig. 2D). Prey diversity was also linked to the diversity of protozoan predators (Table S2), although predator diversity impacts were additive to and not underlying filtration and dilution effects. Further, the impact of predator diversity was variable in effect direction and neither consistently negative nor positive. E.g., bacterial phylogenetic distinctiveness was affected positively by protozoan richness, but negatively by protozoan evenness and phylogenetic distinctiveness of predators. Protozoan phylogenetic distinctiveness also had a weak but significant negative effect on bacterial evenness.
Prokaryotic ASV richness (A), evenness (B) and phylogenetic distinctiveness (C) decreased in the diluted treatment (red) compared starting conditions (blue and the undiluted trated (yellow). Results for each of the 8 experiments are plotted separately to account for systematic differences in starting conditions across experiments. D The decrease in phylogenetic distinctiveness in the treatments with reduced predation was positively related to the starting ASV richness of experiments (linear regression; R2 = 0.75, p = 0.03, y = 0.004x − 4.7). Grey line denotes the predicted relationship and the shaded grey area represents the 95% confidence interval of the slope.
We further tested whether predator densities were related to prey diversities within individual dilution treatments (Fig. 4A, B). The densities of HNFs, i.e. the predator group that was quantifiable by flow cytometry, were positively associated with prey diversity components in the reduced predation treatment (regression for prey richness: R2 = 0.30, p = 0.02; evenness: R2 = 0.32, p = 0.01; phylogenetic distinctiveness: R2 = 0.23, p = 0.03; Fig. S5). Further, during the course of the experiments, prey richness decreased less in diluted microcosms that had higher HNF densities (linear regression: R2 = 0.23, p = 0.03, Fig. 4A). By contrast, there was no relationship between HNF densities and richness or genetic diversity in undiluted microcosms (Fig. 4B, p > 0.10), and only prey evenness was positively associated with HNF densities (R2 = 0.45, p = 0.003).
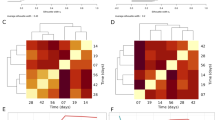
Bacterial beta diversity was strongly influenced by dilution and associated reduction in predation pressure. Bacterial community composition was predominantly driven by differences in inocula, but the composition of bacterial communities also changed over time (Fig. 5A). These temporal changes were more pronounced in the diluted microcosms (Fig. 5B, C; ANOVA; F(1,56) = 103, p < 0.001), leading to a homogenisation of communities illustrated as drop in beta diversity (Bray-Curtis dissimilarity) from 0.80 to 0.68 (ANOVA, F(2,327) = 15.83, p < 0.001). Protozoan beta diversity, however, significantly increased from 0.76 to 0.86 in diluted microcosms (Kruskal-Wallis Test, W = 3140, p < 0.01).
A Non-metric multidimensional scaling (NMDS) representation of Bray-Curtis community similarity. B Similarity between communities at start and in undiluted (i.e. high predation pressure) samples from the same experiment was significantly higher (p < 0.001) than the similarity between communities at start and in diluted (i.e. reduced grazing) samples. C Community similarity within treatments was significantly higher for the reduced predation treatment (p < 0.001), indicating reduced beta diversity and community homogenisation. Grey points in B and C represent pairwise community comparisons, black points represent means of community comparisons and the black horizontal lines are ±1 standard deviation.
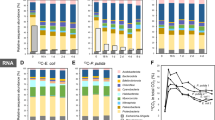
Bacterial communities in all treatments were dominated by Proteobacteria, but experimental dilution shifted dominance from Betaproteobacteriales to Pseudo- and Alteromonadales (Fig. 6A–C). Experimental dilution resulted also in an increase in HNA-LNA ratios (i.e., an increase in the relative abundance of more active cells; paired t test, t value = 3.8, p = 0.002; Figs. S6–8). Shifts in bacterial community composition had a substantial effect on the putative functionality of activated sludge communities. The comparison of our results with a global meta-analysis (Table 1) revealed that relative densities of many bacterial taxa associated with increased treatment efficiency, significantly declined in the low predation treatment. This included numerous taxa belonging to the Rhodocyclaceae (e.g. Canditatus Accumulibacter), Comamonadaceae and Nitrospiraceae families (Table 1). An exception from this observation were the families of Moraxcellaceae and Xanthomonadaceae. Whereas Xanthomonadaceae did not show much of a net change, Moraxcellaceae, a group often associated with improved aggregate formation and phosphorus removal, benefited from the experimental dilution. These findings were also corroborated by a METAGENassist analysis, showing a strong reduction in N-removal potential and a tentative reduction in C remineralisation in the reduced predation treatment (Fig. S9).
A A phylogenetic three showing all taxa with a mean relative abundance of >0.35% across all microcosms (n = 37). Circles present presence (red: reduced predation; yellow: ambient predation), size of the circle reflects relative densities. Taxonomic affiliation is expressed at the order level (bold) and at the lowest taxonomic level that could be associated to ASVs. B The relative contribution of different orders to the total number of reads in reduced predation and ambient predation treatments. C Differences in relative abundance of all taxa (summed at class level) that significantly differed between predation and reduced-predation treatments. For each order, ASVs that expressed positive and negative change were summed separately. Numbers denote the counts of ASVs with a significant difference between treatments. Bars represent standard deviation of class sums per treatment. Cytophagales did not include any ASVs that significantly differed between treatments are not displayed in C.
Discussion
Despite the importance of protozoan predation for maintaining treatment efficiency in activated sludge communities [3, 4], the mechanisms governing this process are poorly understood. We demonstrated that the manipulation of protozoan predators has profound impacts on bacterial diversity and community composition with potentially far-reaching implications for ecosystem functioning. Both the decrease of prey encounter rates through dilution and the removal of top predators via filtration substantially altered bacterial prey diversity, whereas predator diversity per se had only lesser and ambiguous impacts. Moreover, reductions in predator-prey encounter rates via dilution altered bacterial community composition and triggered the decline of multiple taxa that support wastewater treatment efficiency. This suggests that protozoan predation may enhance functioning of activated sludge communities through diversity and compositional effects, which are at least partly mediated by the identity of dominant predators.
The impact of predation pressure on prey diversity
Dilution experiments to regulate predator-prey encounter rates are common tools in plankton ecology [40, 59], but comparable, manipulative predation experiments are almost non-existent in activated sludge research. In our study, reduced encounter rates, which are well known to weaken top-down control [40], caused marked declines in richness, evenness and phylogenetic diversity of bacterial prey communities. This positive effect of predation on prey diversity is likely governed by preventing the competitive exclusion of slower growing bacteria that invest more resources in antipredator defences [Fig. 1; [32]].
Predators themselves have adapted to antipredator defences of their prey [60] causing a diversification of defence strategies such as increases in prey body size, movement speed or toxin production [61,62,63]. The emerging positive impact on prey diversity is often maintained by predator and prey population fluctuations, density-dependent predation and diversity-enhancing ‘kill the winner’ dynamics [i.e. reducing the dominance of successful competitors; [39]]. Specialist predators can support such ‘kill the winner‘ dynamics because of their high susceptibility to food limitation. Therefore, changes in prey population can cause even at the very high food densities found in activated sludge reactors that predators enter the non-linear part of their functional response curves, enforcing density-dependent prey control [64,65,66]. Generalist predators, on the other hand, often preferentially feed on the most common prey types, again triggering ‘kill the winner‘ dynamics [67, 68]. Hence, a positive response of prey diversity to predation is not only based on the resulting co-existence of K- and r-strategists, but also emerges from density-dependent predation and from the co-existence of multiple K-strategists with alternative predator-defence mechanisms.
However, an increase in predation pressure does not necessarily result in a linear, positive impact on prey diversity [27, 28]. We found the effect of predation on prey diversity to vary along a gradient of predation intensity. Whereas HNF densities were positively associated with bacterial diversity in the reduced predation treatment, there was no clear association in undiluted microcosms with high predation pressure. Even though HNFs represent only one group of predators in activated sludge communities, these findings support previous hypotheses of a hump-shaped relationship between prey diversity and predation pressure [28, 32]. The predation intensity that results in maximal prey diversity (i.e. the peak of the hump) has been suggested to increase with ecosystem productivity [Fig. 1; [27]]. In highly productive activated sludge communities, this may result in an overall positive impact of protozoan biomass on prey diversity. However, protozoans can account for very high proportion of community biomass, reaching up to 20% of total activated sludge mass [69]. Such elevated predator biomass may eventually exceed limits of beneficial top-down control and trigger negative responses in prey diversity.
Diversity effects on ecosystem functioning
Positive impacts of diversity on functioning are well supported across ecosystem types and taxonomic groups [21, 70] and hence high bacterial diversity can be expected to also increase wastewater treatment efficiency [e.g. enhanced nutrient-uptake, reduced biological oxygen demand in outflow; [7, 71]]. Research about diversity and ecosystem functioning traditionally relied on species richness as biodiversity indicator [72]. However, it has been argued that phylogenetic diversity is a better predictor of functionality as it better reflects niche complementarity, a key mechanism linking biodiversity to ecosystem functioning [73]. Here, we used phylogenetic distinctiveness as a measure of phylogenetic diversity because of its mathematical independence from taxa richness [54]. Nevertheless, we showed that losses of phylogenetic diversity resulting from reduced predation pressure were mitigated by high taxa richness (Fig. 3D). These findings agree with the insurance hypothesis, postulating that high taxa richness mitigates the erosion of functionality in stressed ecosystems [74]. Therefore, the insurance hypothesis may be an important mechanism enhancing treatment efficiency in activated sludge reactors with high bacterial diversity.
Beta diversity represents another biodiversity component that can improve ecosystem functioning, particularly at larger spatial and temporal scales [75, 76]. We showed that beta diversity was positively related to high predation pressure (Fig. 5). By contrast, conceptual frameworks [32] and experiments with fish communities [29, 77] suggested a negative impact of predation on beta diversity. In this context, predation is suggested to reduce stochasticity and increase the relative importance of deterministic community assembly processes [29]. The contrasting results in our study may result from our focus on complex and highly variable predator assemblages compared to the previous work that investigated the impacts of a single top predator [29, 77]. Protozoan predators show a high functional diversity in their feeding modes [63, 78] and therefore impose different selection pressures on their prey [e.g. ambush vs. filter feeding predators; [60]]. Hence, predation in our study may still have enhanced the importance of deterministic assembly processes [29]. However, diverging selection pressures across our experiments would “push” prey communities in different directions, explaining the observed increase in beta diversity in our study.
The effects of community composition on ecosystem functioning
Dilution of microcosms resulted in strong changes in the identity of dominant bacterial ASVs in our experiments. These changes can in principle emerge from reductions in predator-prey encounter rates and predation pressure or from an increased resource supply in diluted communities. Dilution experiments are designed to maintain an equal initial resource availability across treatments [40], which together with the high resource concentration in the growth media counteracts resource limitation. Moreover, if nutrient limitation was an important driver of community changes, it should have had a stronger impact in undiluted microcosms. Yet, these differences were small compared to temporal changes in community composition in diluted microcosms and therefore differences in resource availability likely played a subordinate role in driving community shits.
At higher taxonomic levels, ASVs belonging to the same taxon exhibited partly contrasting responses to reduction in predation pressure (Fig. 6). Diverse responses can generally be expected because of the high functional diversity within higher taxonomic groups (e.g. Betaproteobacteriales) and predation-mediated changes in the outcome of competition among closely related prey species. Despite these sometime bi-directional changes, our assessment of putative functionality in sludge communities, a topic that currently gains rapidly in attention [79], indicated decreases of treatment efficiency at lower levels of predation pressure. Relative densities of many taxa that are associated with high wastewater treatment efficiency, such as Comamonadaceae, Nitrospira and Candidatus Accumulibacter [6, 7] increased in treatment with high predation pressure (Table 1). Compositional changes resulted in a tendency of a decreasing potential for carbon degradation and phosphorus uptake and a strong reduction in nitrogen removal at low predator-prey encounter rates (Table 1, Fig. S9). Although these findings are restricted to putative functionality, they highlight the large potential impacts that changes in predation may have on wastewater processing in activated sludge communities.
Outlook
The overarching goal of many recent studies and research applications is to maximise the positive impacts of bacterial communities on wastewater treatment efficiency [2, 5, 7]. Our findings demonstrate the critical role of protozoan predation in governing diversity and composition of activated sludge communities and suggest their indirect consequences for treatment efficiency. We call for more community-level experiments that directly manipulate mechanisms linking predator and prey density, identity, and multiple aspects of diversity with specific functions of activated sludge ecosystems. Such mechanistic research represents a crucial step forward in advancing general ecological theory as well as improving the capacity of biological treatments in activated sludge reactors.
References
Seviour RJ, Kragelund C, Kong Y, Eales K, Nielsen JL, Nielsen PH. Ecophysiology of the Actinobacteria in activated sludge systems. Antonie Van Leeuw J Microb. 2008;94:21–33.
Jiang X-T, Ye L, Ju F, Wang Y-L, Zhang T. Toward an intensive longitudinal understanding of activated sludge bacterial assembly and dynamics. Environ Sci Technol. 2018;52:8224–32.
Fiałkowska E, Pajdak-Stós A. The role of Lecane rotifers in activated sludge bulking control. Water Res. 2008;42:2483–90.
Madoni P. Protozoa in wastewater treatment processes: a minireview. Ital J Zool. 2011;78:3–11.
Ye L, Mei R, Liu W-T, Ren H, Zhang X-X. Machine learning-aided analyses of thousands of draft genomes reveal specific features of activated sludge processes. Microbiome. 2020;8:16.
Peces M, Astals S, Jensen P, Clarke W. Deterministic mechanisms define the long-term anaerobic digestion microbiome and its functionality regardless of the initial microbial community. Water Res. 2018;141:366–76.
Wu L, Ning D, Zhang B, Li Y, Zhang P, Shan X, et al. Global diversity and biogeography of bacterial communities in wastewater treatment plants. Nat Microbiol. 2019;4:1183–95.
Cox HH, Deshusses MA. Biomass control in waste air biotrickling filters by protozoan predation. Biotechnol Bioeng. 1999;62:216–24.
Madoni P. A sludge biotic index (SBI) for the evaluation of the biological performance of activated sludge plants based on the microfauna analysis. Water Res. 1994;28:67–75.
Ratsak C, Maarsen K, Kooijman S. Effects of protozoa on carbon mineralization in activated sludge. Water Res. 1996;30:1–12.
Pogue AJ, Gilbride KA. Impact of protozoan grazing on nitrification and the ammonia- and nitrite-oxidizing bacterial communities in activated sludge. Can J Microbiol. 2007;53:559–71.
Esteban G, Tellez C, Bautista LM. Dynamics of ciliated protozoa communities in activated-sludge process. Water Res. 1991;25:967–72.
Madoni P, Davoli D, Chierici E. Comparative analysis of the activated sludge microfauna in several sewage treatment works. Water Res. 1993;27:1485–91.
Otto S, Harms H, Wick LY. Effects of predation and dispersal on bacterial abundance and contaminant biodegradation. FEMS Microbiol Ecol. 2017;93:fiw241.
Peralta-Maraver I, Reiss J, Robertson AL. Interplay of hydrology, community ecology and pollutant attenuation in the hyporheic zone. Sci Total Environ. 2018;610:267–75.
Yang JW, Wu W, Chung C-C, Chiang K-P, Gong G-C, Hsieh C-H. Predator and prey biodiversity relationship and its consequences on marine ecosystem functioning—interplay between nanoflagellates and bacterioplankton. ISME J. 2018;12:1532–42.
Seiler C, van Velzen E, Neu TR, Gaedke U, Berendonk TU, Weitere M. Grazing resistance of bacterial biofilms: a matter of predators’ feeding trait. FEMS Microbiol Ecol. 2017;93:fix112.
Burian A, Nielsen JM, Winder M. Food quantity-quality interactions and their impact on consumer behavior and trophic transfer. Ecol Monogr. 2020;90:e01395.
Schmitz OJ. Effects of predator functional diversity on grassland ecosystem function. Ecology. 2009;90:2339–45.
Estes JA, Terborgh J, Brashares JS, Power ME, Berger J, Bond WJ, et al. Trophic downgrading of planet Earth. Science. 2011;333:301–6.
Cardinale BJ, Duffy JE, Gonzalez A, Hooper DU, Perrings C, Venail P, et al. Biodiversity loss and its impact on humanity. Nature. 2012;486:59–67.
Isbell F, Calcagno V, Hector A, Connolly J, Harpole WS, Reich PB, et al. High plant diversity is needed to maintain ecosystem services. Nature. 2011;477:199–202.
Delgado-Baquerizo M, Maestre FT, Reich PB, Jeffries TC, Gaitan JJ, Encinar D, et al. Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat Commun. 2016;7:10541.
McCann KS. The diversity–stability debate. Nature. 2000;405:228.
Pennekamp F, Pontarp M, Tabi A, Altermatt F, Alther R, Choffat Y, et al. Biodiversity increases and decreases ecosystem stability. Nature. 2018;563:109–12.
Saikaly PE, Oerther DB. Diversity of dominant bacterial taxa in activated sludge promotes functional resistance following toxic shock loading. Microb Ecol. 2011;61:557–67.
Worm B, Lotze HK, Hillebrand H, Sommer U. Consumer versus resource control of species diversity and ecosystem functioning. Nature. 2002;417:848–51.
Gauzens B, Legendre S, Lazzaro X, Lacroix G. Intermediate predation pressure leads to maximal complexity in food webs. Oikos. 2016;125:595–603.
Chase JM, Biro EG, Ryberg WA, Smith KG. Predators temper the relative importance of stochastic processes in the assembly of prey metacommunities. Ecol Lett. 2009;12:1210–8.
Paine RT. Food web complexity and species diversity. Am Nat. 1966;100:65–75.
Gliwicz ZM, Wursbaugh WA, Szymanska E. Absence of predation eliminates coexistence: experience from the fish–zooplankton interface. Fifty years after the “Homage to Santa Rosalia”: old and new paradigms on biodiversity in aquatic ecosystems. Springer; 2010. p. 103–17.
Terborgh JW. Toward a trophic theory of species diversity. Proc Natl Acad Sci USA. 2015;112:11415–22.
Kondoh M. Unifying the relationships of species richness to productivity and disturbance. Proc R Soc B-Biol Sci. 2001;268:269–71.
Hutchinson GE. The paradox of the plankton. Am Nat. 1961;95:137–45.
Al-Shahwani S, Horan N. The use of protozoa to indicate changes in the performance of activated sludge plants. Water Res. 1991;25:633–8.
Torsvik V, Øvreås L, Thingstad TF. Prokaryotic diversity-magnitude, dynamics, and controlling factors. Science. 2002;296:1064–6.
Papadimitriou C, Papatheodoulou A, Takavakoglou V, Zdragas A, Samaras P, Sakellaropoulos G, et al. Investigation of protozoa as indicators of wastewater treatment efficiency in constructed wetlands. Desalination. 2010;250:378–82.
Rossberg AG. Food webs and biodiversity: foundations, models, data. John Wiley & Sons; 2013.
Vage S, Bratbak G, Egge J, Heldal M, Larsen A, Norland S, et al. Simple models combining competition, defence and resource availability have broad implications in pelagic microbial food webs. Ecol Lett. 2018;21:1440–52.
Landry M, Hassett R. Estimating the grazing impact of marine micro-zooplankton. Mar Biol. 1982;67:283–8.
Dolan J, Gallegos C, Moigis A. Dilution effects on microzooplankton in dilution grazing experiments. Mar Ecol Prog Ser. 2000;200:127–39.
Dottorini G, Michaelsen TY, Kucheryavskiy S, Andersen KS, Kristensen JM, Peces M, et al. Mass-immigration determines the assembly of activated sludge microbial communities. Proc Natl Acad Sci USA; 2021;118:e2021589118.
Stevens-Garmon J, Drewes JE, Khan SJ, McDonald JA, Dickenson ERV. Sorption of emerging trace organic compounds onto wastewater sludge solids. Water Res. 2011;45:3417–26.
Gasol JM, Morán XAG. Flow cytometric determination of microbial abundances and its use to obtain indices of community structure and relative activity. Hydrocarbon and lipid microbiology protocols. Springer; 2015. p. 159–87.
Ram AP, Chaibi-Slouma S, Keshri J, Colombet J, Sime-Ngando T. Functional responses of bacterioplankton diversity and metabolism to experimental bottom-up and top-down forcings. Microb Ecol. 2016;72:347–58.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA. 2011;108:4516–22.
Hugerth LW, Muller EE, Hu YO, Lebrun LA, Roume H, Lundin D, et al. Systematic design of 18S rRNA gene primers for determining eukaryotic diversity in microbial consortia. Plos One. 2014;9:e95567.
D’Amore R, Ijaz UZ, Schirmer M, Kenny JG, Gregory R, Darby AC, et al. A comprehensive benchmarking study of protocols and sequencing platforms for 16S rRNA community profiling. BMC Genom. 2016;17:55.
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol. 2019;37:852–7.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2012;41:D590–D596.
Price MN, Dehal PS, Arkin AP. FastTree 2-approximately maximum-likelihood trees for large alignments. Plos One. 2010;5:10.
Faith DP. Conservation evaluation and phylogenetic diversity. Biol Conserv. 1992;61:1–10.
Tsirogiannis C, Sandel B. PhyloMeasures: a package for computing phylogenetic biodiversity measures and their statistical moments. Ecography. 2016;39:709–14.
Wobbrock JO, Findlater L, Gergle D, Higgins JJ, Acm. The aligned rank transform for nonparametric factorial analyses using only ANOVA procedures. Association Computing Machinery: New York; 2011.
Burnham KP, Anderson DR. Model selection and multimodel interference: a practical information—theoretic approach. Springer: New York, USA; 2002.
Arndt D, Xia J, Liu Y, Zhou Y, Guo AC, Cruz JA, et al. METAGENassist: a comprehensive web server for comparative metagenomics. Nucleic Acids Res. 2012;40:W88–W95.
R Development Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria; 2015. ISBN 3-900051-07-0, http://wwwR-projectorg.
Calbet A, Landry MR. Phytoplankton growth, microzooplankton grazing, and carbon cycling in marine systems. Limnol Oceanogr. 2004;49:51–57.
Kiorboe T. How zooplankton feed: mechanisms, traits and trade-offs. Biol Rev. 2011;86:311–39.
Juergens K, Matz C. Predation as a shaping force for the phenotypic and genotypic composition of planktonic bacteria. Antonie Van Leeuw J Microb. 2002;81:413–34.
Hammill E, Kratina P, Beckerman A, Anholt BR. Precise time interactions between behavioural and morphological defences. Oikos. 2010;119:494–9.
Pernthaler J. Predation on prokaryotes in the water column and its ecological implications. Nat Rev Microbiol. 2005;3:537–46.
Visser MD, Muller‐Landau HC, Wright SJ, Rutten G, Jansen PA. Tri‐trophic interactions affect density dependence of seed fate in a tropical forest palm. Ecol Lett. 2011;14:1093–1100.
Bagchi R, Gallery RE, Gripenberg S, Gurr SJ, Narayan L, Addis CE, et al. Pathogens and insect herbivores drive rainforest plant diversity and composition. Nature. 2014;506:85–88.
Kratina P, Vos M, Anholt BR. Species diversity modulates predation. Ecology. 2007;88:1917–23.
Jaworski CC, Bompard A, Genies L, Amiens-Desneux E, Desneux N. Preference and prey switching in a generalist predator attacking local and invasive alien pests. Plos One. 2013;8:e82231.
Coblentz KE, DeLong JP. Predator‐dependent functional responses alter the coexistence and indirect effects among prey that share a predator. Oikos. 2020;129:1404–14.
Madoni P. Estimates of ciliated protozoa biomass in activated sludge and biofilm. Bioresour Technol. 1994;48:245–9.
Tilman D, Knops J, Wedin D, Reich P, Ritchie M, Siemann E. The influence of functional diversity and composition on ecosystem processes. Science. 1997;277:1300–2.
Sato Y, Hori T, Navarro RR, Habe H, Ogata A. Functional maintenance and structural flexibility of microbial communities perturbed by simulated intense rainfall in a pilot-scale membrane bioreactor. Appl Microbiol Biot. 2016;100:6447–56.
Cardinale BJ, Wright JP, Cadotte MW, Carroll IT, Hector A, Srivastava DS, et al. Impacts of plant diversity on biomass production increase through time because of species complementarity. Proc Natl Acad Sci USA. 2007;104:18123–8.
Srivastava DS, Cadotte MW, MacDonald AAM, Marushia RG, Mirotchnick N. Phylogenetic diversity and the functioning of ecosystems. Ecol Lett. 2012;15:637–48.
Yachi S, Loreau M. Biodiversity and ecosystem productivity in a fluctuating environment: the insurance hypothesis. Proc Natl Acad Sci USA. 1999;96:1463–8.
Mori AS, Isbell F, Seidl R. β-diversity, community assembly, and ecosystem functioning. Trends Ecol Evol. 2018;33:549–64.
Hammill E, Hawkins CP, Greig HS, Kratina P, Shurin JB, Atwood TB. Landscape heterogeneity strengthens the relationship between β‐diversity and ecosystem function. Ecology. 2018;99:2467–75.
Ellingsen KE, Yoccoz NG, Tveraa T, Frank KT, Johannesen E, Anderson MJ, et al. The rise of a marine generalist predator and the fall of beta diversity. Glob Change Biol. 2020;26:2897–907.
Weisse T. The significance of inter-and intraspecific variation in bacterivorous and herbivorous protists. Antonie Van Leeuw J Microb. 2002;81:327–41.
Nierychlo M, Andersen KS, Xu Y, Green N, Jiang C, Albertsen M, et al. MiDAS 3: an ecosystem-specific reference database, taxonomy and knowledge platform for activated sludge and anaerobic digesters reveals species-level microbiome composition of activated sludge. Water Res. 2020;182:115955.
Acknowledgements
We thank Severn Trent Waters for providing an access to the Derby waste-water treatment plant, and Peter Vale for his help in facilitating waste-water sampling at the site. Further, many thanks go to Richard Duff for his essential support during media preparation and laboratory assistance. We also thank two anonymous reviewers for their constructive feedback. This project was funded by internal support from Derby University to AB and from Queen Mary University of London to PK.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
AB, DP and KP conceptually developed the study, DP and AB implemented the experiments, IPM conducted to flowcytometric measurements and QM the metabarcoding analyses. AB, MB and TR performed statistical analyses and all co-authors contributed to the manuscript establishment.
Corresponding authors
Ethics declarations
Competing interests
DP is employed by Thames Waters, none of the other co-authors have to declare a competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Burian, A., Pinn, D., Peralta-Maraver, I. et al. Predation increases multiple components of microbial diversity in activated sludge communities. ISME J 16, 1086–1094 (2022). https://doi.org/10.1038/s41396-021-01145-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01145-z
This article is cited by
-
Genetic variation in Zea mays influences microbial nitrification and denitrification in conventional agroecosystems
Plant and Soil (2024)
-
Environmental changes associated with drying climate are expected to affect functional groups of pro- and microeukaryotes differently in temporary saline waters
Scientific Reports (2023)
-
Active predation, phylogenetic diversity, and global prevalence of myxobacteria in wastewater treatment plants
The ISME Journal (2023)