Abstract
Corals house a variety of microorganisms which they depend on for their survival, including endosymbiotic dinoflagellates (Symbiodiniaceae) and bacteria. While cnidarian–microorganism interactions are widely studied, Symbiodiniaceae–bacteria interactions are only just beginning to receive attention. Here, we describe the localization and composition of the bacterial communities associated with cultures of 11 Symbiodiniaceae strains from nine species and six genera. Three-dimensional confocal laser scanning and electron microscopy revealed bacteria are present inside the Symbiodiniaceae cells as well as closely associated with their external cell surface. Bacterial pure cultures and 16S rRNA gene metabarcoding from Symbiodiniaceae cultures highlighted distinct and highly diverse bacterial communities occur intracellularly, closely associated with the Symbiodiniaceae outer cell surface and loosely associated (i.e., in the surrounding culture media). The intracellular bacteria are highly conserved across Symbiodiniaceae species, suggesting they may be involved in Symbiodiniaceae physiology. Our findings provide unique new insights into the biology of Symbiodiniaceae.
Similar content being viewed by others
Introduction
Virtually all eukaryotes associate with symbiotic bacteria [1,2,3], and unicellular algae are no exception. Algae–bacteria associations are widespread in the marine environment, including the pelagic zone [4, 5] and coral reefs [6]. Extensive interactions in the phycosphere, an extracellular boundary layer composed of algal exudates, such as fixed organic carbon, may attract and nurture bacteria [7, 8]. Moreover, bacteria have been observed inside many types of algae [9,10,11,12,13], indicating the existence of intracellular interactions. Bacteria are known to provide their algal hosts with vitamin B12 [14,15,16,17], increased iron bioavailability [18], carotenoids [19], and growth-promoting hormones [20], though the array of functions is likely much greater.
One major group of marine microalgae, the Symbiodiniaceae (Suessiales, Dinophyta) is well known for its obligate endosymbiosis with cnidarians, including reef-building corals and sea anemones [21, 22]. The Symbiodiniaceae are highly diverse, with nine described genera and three additional evolutionary lineages that currently lack formal nomenclature [23, 24], encompassing free-living species and species associated with a wide range of marine organisms, including cnidarians, mollusks, and protists [21]. By translocating photosynthate to the cnidarian tissues, Symbiodiniaceae provide cnidarian hosts with most of their carbon requirements [25,26,27]. This photosymbiosis allows corals to build the reef’s three-dimensional structure, thus providing habitat to many other reef-dwelling organisms. Coral reefs are rapidly declining globally because of increasingly frequent episodes of mass coral bleaching caused by heatwaves [28, 29]. Coral bleaching is the breakdown of the symbiosis between Symbiodiniaceae and coral hosts. It is thought to be the result of excess production of reactive oxygen species (ROS) by Symbiodiniaceae under excessive temperature and/or light exposure, and leads to coral starvation and mortality [30,31,32]. Thus, unlocking the ecology and physiology of Symbiodiniaceae is a priority in global efforts to mitigate coral bleaching and minimize the impacts of climate change on coral reefs.
Aside from diverse Symbiodiniaceae, cnidarians associate with abundant and diverse bacterial communities [22, 33, 34]. Specific bacteria have been shown to be involved in nutrient cycling within the coral holobiont [35,36,37], such as metabolic exchanges linked to the sulfur and nitrogen cycles between Symbiodiniaceae and bacteria [35, 38]. More recently, a bacterial strain closely related to Muricauda was shown to provide zeaxanthin to cultured Symbiodiniaceae (Durusdinium sp.), thereby enhancing its resistance to light and temperature stresses [19]. Hence, investigating the bacterial communities that naturally associate with Symbiodiniaceae is a key step toward a broader understanding of the impact of Symbiodiniaceae–bacteria interactions on cnidarian health.
To date, few studies have investigated the microbiome of Symbiodiniaceae [39,40,41]. Lawson et al. reported the composition of the bacterial communities associated with 18 Symbiodiniaceae cultures [40]. Communities were dominated by Alpha- and Gammaproteobacteria with Labrenzia, Marinobacter, and Chromatiaceae found in all cultures. Camp et al. studied the impact of a thermal stress event on the bacterial community composition of four Symbiodiniaceae cultures, finding again a high prevalence of Alphaproteobacteria, and particularly Labrenzia [41]. However, neither of these studies distinguished the planktonic bacteria from the bacteria closely associated with the algae. Furthermore, only extracellular bacteria have been unequivocally reported through scanning electron microscopy (SEM) [39, 42]. Actinobacteria and Ralstonia were reported to occur inside in hospite Symbiodiniaceae of the coral Acropora granulosa, using 2-dimensional confocal microscopy [43]. However, these observations remain unverified without 3-dimensional imaging to confirm whether such bacteria are truly inside the cell.
Here, we use a broad range of Symbiodiniaceae strains from six genera and unequivocally show the presence of intracellular and extracellular bacteria closely associated with both cultured and freshly isolated algae by 3-dimensional fluorescence in situ hybridization (FISH) confocal laser scanning microscopy (CLSM) and SEM. By combining culture-dependent and -independent techniques, we further characterize the composition of extracellular and intracellular bacteria in 11 Symbiodiniaceae cultures from nine different species. Our findings extend current knowledge of intracellular symbiosis, providing unique new insights into the biology of Symbiodiniaceae, and set the stage for the study of bacterial functions in Symbiodiniaceae, and more broadly in the cnidarian holobiont.
Material and methods
Additional methods (CLSM, SEM, flow cytometry, and bacteria pure culturing) are available in the supplementary information.
Symbiodiniaceae cultures
Symbiodiniaceae cultures (Table S1) were provided by the Australian Institute of Marine Science (AIMS, Townsville, QLD, Australia), except for Breviolum minutum which was isolated at the Marine Microbial Symbiont Facility (University of Melbourne, VIC, Australia) [44], and Symbiodinium tridacnidorum which was obtained from the Australian National Algae Culture Collection, CSIRO. Nine of the 11 strains were isolated from scleractinian corals, one from a clam (Tridacna maxima) and one from the anemone Exaiptasia diaphana. Symbiodiniaceae were grown in sterile 50 mL polypropylene (Falcon) culture flasks in 15 mL of sterile 1× IMK liquid media (Diago’s IMK Medium for marine microalgae, 1% w/v, NovaChem) prepared with filter sterilized Red Sea Salt water (fRSS). Growth of Symbiodiniaceae cultures was maintained by fortnightly transfers of cultures into fresh media and incubation at 26 °C on a 12:12 h light:dark cycle, with lighting at 50–70 μmol photons m−2 s−1 (Taiwan Hipoint Corporation, model 740FHC LED, light chambers).
Symbiodiniaceae sampling and fixation for fluorescence in situ hybridization (FISH)
Symbiodiniaceae cells were sampled and twice washed by centrifugation at 5000 × g for 5 min, supernatant removed, and resuspended in phosphate buffered saline (PBS, pH 7.4) before being fixed in ice-cold 66% ethanol for 2 h at 4 °C. To minimize Symbiodiniaceae autofluorescence during downstream observations, cells were then photobleached by placing the samples under bright white light (LED) of ~400 µmol photons m−2 s−1 for at least 2 h at ambient temperature. Cells were washed in PBS and stored in 70% ethanol at −20 °C.
Freshly isolated Symbiodiniaceae from Exaiptasia diaphana and Galaxea fascicularis
Five E. diaphana polyps [45] were placed in a glass tank and submerged in a solution of 0.39 M magnesium chloride in PBS for 30 min to anesthetize them. Tentacles were then removed with scissors and placed in 1.5 mL microcentrifuge tubes in 200 μL fRSS. The tissue was homogenized with a sterile micro-pestle.
G. fascicularis colonies were purchased from Cairns Marine in February 2020. These colonies were collected mid-February on Sudbury Reef in the Great Barrier Reef (GBR). They were kept in the Cairns Marine aquaria for 2 weeks and subsequently shipped to the University of Melbourne. In the University of Melbourne laboratory, corals were kept in RSS (34 ppt salinity) at 25.5 °C, on a 10:14 h light:dark cycle with lighting at 160–180 μmol photons m−2 s−1. In July 2020, four G. fascicularis polyps were detached from a single colony. To remove tissues from the skeleton, polyps were sprayed with pressured seawater (sterilized through a 0.22-µm filter) using a water flosser (Waterpik) with a Pik Pocket tip. Tissues were centrifuged at 825 × g for 5 min and homogenized with a sterile micro-pestle.
Freshly isolated Symbiodiniaceae cells were thrice washed by centrifugation at 825 × g for 5 min and resuspension in fRSS. After the last wash, pelleted cells were fixed in ice-cold 66% ethanol. Cells were then photobleached by placing the samples under bright white light (LED) of ~400 µmol photons m−2 s−1 for at least 2 h at ambient temperature. Cells were washed in PBS and stored in 70% ethanol at −20 °C.
The E. diaphana genotype used in this study [45] was previously shown to harbor B. minutum [44], while corals of the Galaxea genus from the GBR usually harbor Symbiodiniaceae of the Cladocopium and Durusdinium genera [46, 47].
FISH
Teflon-printed microscope slides (ProSciTech) were coated with poly-L-lysine solution (0.01%) in PBS by aliquoting the solution onto printed wells, incubated at 37 °C for 3 h, then thrice washed in sterile Milli-Q water and air dried. Aliquots of 5–10 μL were pipetted into a well on treated ten-well slides. FISH was performed as previously described [48]. Samples on glass slides were air dried and permeabilized for 10 min in Triton X-100 solution (0.1% v/v) in PBS, and dehydrated in an ethanol series (3 min each in 50%, 80%, and 98% ethanol). Hybridization was in 18 μL buffer (0.9 M NaCl, 20 mM Tris-HCL pH 7.2, formamide [varied to accommodate optimal stringency for different probes—Table S2], 0.01% SDS) and 2 μL 16S rRNA targeting oligonucleotide probes at a final concentration of 5 ng/μL for 2 h at 46 °C. All probe sequences are listed in Table S2 [49,50,51,52]. When two different probes were used on the same sample, both probes had a final concentration of 5 ng/μL. Samples were then washed in a prewarmed buffer (NaCl [varied to accommodate optimal stringency for different probes—Table S2], 20 mM Tris-HCL pH 7.2, 0.01% SDS) for 15 min at 48 °C, briefly washed in ice-cold water and carefully dried. Slides were mounted with CitiFluor™ CFM3 mounting medium (Hatfield, PA, USA) and stored in the dark at −20 °C until observation in confocal microscopy.
Symbiodiniaceae sampling for 16S rRNA gene metabarcoding
The experimental design for this part of the study is outlined in Fig. 1. Sampling was performed between 2 and 4 p.m. (8–10 h into the light cycle) on the same day for every strain, during the exponential growth phase. For each culture used (Table S1), a total of six samples of 105 cells were collected and filtered through individual 5 µm mesh strainers (pluriSelect, Germany). Strainers were sealed and centrifuged for 5 min at 12,000 × g to separate planktonic bacteria (<5 µm) from Symbiodiniaceae cells (>5 µm). For three of those samples, loosely attached bacteria were separated from Symbiodiniaceae cells by pipetting 500 µL fRSS onto the Symbiodiniaceae cells collected in the strainer, followed by centrifugation for 5 min at 12,000 × g. Filtrates (initial culture medium containing planktonic bacteria + fRSS wash) were snap frozen and represent ‘loosely associated bacteria’ (Fig. 1, Sample 1). Filters, having retained algal cells, intracellular bacteria and bacteria tightly attached to the algal cell’s exterior, were extracted from the strainer with sterile forceps, put into a sterile 1.5 mL microcentrifuge tube, and snap frozen in liquid nitrogen, and represent ‘closely associated bacteria’ (Fig. 1, Sample 2). For each culture used, the other three samples of 105 cells were collected, filtered through a 5 µm mesh strainer (pluriSelect, Germany), sealed, and centrifuged for 5 min at 12,000 × g. To kill and remove all extracellular bacteria, retained Symbiodiniaceae cells were washed by pipetting 500 µL 6% sodium hypochlorite (bleach) onto the strainers, which were then sealed and centrifuged for 5 min at 12,000 × g. Filtrates containing bleach were discarded. Filters, having retained Symbiodiniaceae cells and only intracellular bacteria, were removed from the strainer with sterile forceps, put into a sterile 1.5 mL microcentrifuge tube, and snap frozen (Fig. 1, Sample 3). Hence, each culture yielded nine samples: three ‘loosely associated bacteria’ true replicates, three ‘closely associated bacteria’ true replicates, and three ‘intracellular bacteria’ true replicates. Three clean filters (onto which no algae cells were added) were extracted and snap frozen, as negative controls.
Aliquots of 105 Symbiodiniaceae cells were transferred to 5 µm mesh size strainers and centrifuged to remove planktonic bacteria. To remove loosely associated bacteria (left), Symbiodiniaceae cells stuck on the strainer were washed with fRSS. Both successive filtrates, including planktonic bacteria, are referred to as ‘loosely associated bacteria’. Filters were recovered with sterile forceps, having kept Symbiodiniaceae cells and bacteria attached to the Symbiodiniaceae cell exterior and intracellular (closely associated). Another aliquot (right) was washed with sodium hypochlorite 6% to kill and remove extracellular bacteria, so that only intracellular bacteria remained in the sample. Three filters onto which no Symbiodiniaceae cells were deposited were sampled as negative controls.
DNA extraction for 16S rRNA gene metabarcoding
DNA extractions were performed using a salting-out method [53] with modifications as previously described [54]. Two blank DNA extractions were conducted as negative controls. A mock community, ZymoBIOMICS Microbial Community DNA Standard (Zymo Research), was included to check sequencing and processing quality.
16S rRNA gene PCR amplification, library preparation, and sequencing
Hypervariable regions V5–V6 of the 16S rRNA genes were amplified using the primer set 784F (5ʹ GTGACCTATGAACTCAGGAGTCAGGATTAGATACCCTGGTA 3ʹ) and 1061R (5ʹ CTGAGACTTGCACATCGCAGCCRRCACGAGCTGACGAC 3ʹ) [55]. Illumina adapters were attached to the primers and are shown as underlined. Bacterial 16S rRNA genes were PCR-amplified on a SimpliAmp Thermal Cycler (Applied Biosystems, ThermoFisher Scientific). Each reaction contained 1 μL of DNA template, 1.5 μL of forward primer (10 μM stock), 1.5 μL of reverse primer (10 μM stock), 7.5 μL of MyTaq HSRed MasterMix (BioLine), and 3.5 μL of nuclease-free water (Thermofisher), with a total volume of 15 μL per reaction. Three triplicate PCRs were conducted for each sample and three no template PCRs were conducted as negative controls. PCR conditions for the 16S rRNA genes were as follows: initial denaturation at 95 °C for 3 min, then 18 cycles of: denaturation at 95 °C for 15 s, annealing at 55 °C for 30 s, and extension at 72 °C for 30 s; with a final extension at 72 °C for 7 min. Samples were then held at 4 °C. Following PCR, triplicates were pooled, resulting in 45 μL per sample.
Metabarcoding library preparation and sequencing was performed at the Walter and Eliza Hall Institute (WEHI) in Melbourne, Victoria on one MiSeq V3 system (Illumina) with 2 × 300 bp paired-end reads. Library preparation involved addition of 20 μL of next-generation sequencing magnetic beads to 20 μL of PCR product (1:1), for clean-up to ensure high quality downstream sequencing. Beads were washed twice with 70% ethanol, and DNA was resuspended with 40 μL of nuclease-free water. Ten microliters of cleaned-up PCR products was combined with 10 μL 2× Taq MasterMix (M0270L, New England BioLabs) and 0.25 μM of forward and reverse indexing primers. The second PCR conditions were as follows: initial denaturation at 95 °C for 3 min, 24 cycles of: 9 °C for 15 s, 60 °C for 30 s, and 72 °C for 30 s; followed by a final extension at 72 °C for 7 min. Product size and specificity of two replicates of representative 16S rRNA gene amplifications were assessed using a TapeStation (2200 TapeStation, Agilent Technologies). A final bead clean-up (0.8 bead/PCR product ratio) was performed on a pool of 5 μL from each well per plate. Pooled libraries were checked for quality control, size determination, quantity, and purity of each sample, to inform pool normalization by using the TapeStation (2200 TapeStation, Agilent Technologies).
Bacterial 16S rRNA gene analysis
QIIME2 v 2019.4.0 [56] was used for processing 16S rRNA gene sequences. The plugin demux [56] was used to create an interactive plot to visualize the data and assess the quality, for demultiplexing and quality filtering of raw sequences. The plugin cutadapt [57] was used to remove the primers and MiSeq adapters. Plugin DADA2 [58] was used for denoising and chimera checking, trimming, dereplication, generation of a feature table, joining of paired-end reads, and correcting sequencing errors and removing low quality reads (Q-score < 30). Summary statistics were obtained using the feature table to ensure processing was successful. Taxonomy was assigned by training a naive Bayes classifier with the feature-classifier plugin [56], based on a 99% similarity to the V5–V6 region of the 16S rRNA gene in the SILVA 132 database to match the 784F/1061R primer pair used [59]. Alignment [60] and phylogeny [61] packages enabled the production of a phylogenetic tree for later analyses in R Studio. Mitochondria and chloroplast reads were filtered out. Metadata file, phylogenetic tree, and tables with Amplicon Sequence Variant (ASV) taxonomic classifications and counts were imported into R for statistical analyses.
Statistical analyses in R studio
Statistical analyses and graphs were performed using R version 3.5.0 [62], and the packages phyloseq [63], vegan [64], RVAideMemoire [65], ggplot2 [66], tidyverse [67], indicspecies [68]. Statistical tests were considered significant at α = 0.05, unless otherwise stated. Metadata file, taxonomy table, phylogenetic tree, and ASV table were imported into R and mitochondria and chloroplast sequences were removed. Contaminant ASVs, arising from kit reagents and sample manipulation, were identified manually based on their abundance in negative controls: any ASV that was five times more abundant in the mean abundance of either filter blanks, extraction blanks or no template PCRs compared to the mean of all Symbiodiniaceae samples, and that represented at least 1000 reads in all Symbiodiniaceae samples, was considered a contaminant and removed from the dataset. Known contaminants (e.g., Cutibacterium) were also removed manually. Thirty putative contaminant ASVs were identified and removed, constituting 3.7% relative abundance of the bacterial communities in Symbiodiniaceae samples (Table S3). The mock community sample results validated the sequencing run.
Alpha-diversity metrics (observed ASVs, Simpson index, Shannon index) were calculated after rarefying the samples to 4800 reads per sample (the lowest read number for a given sample), a depth that was sufficient to capture diversity across samples, as the number of ASVs had reached an asymptote at this read number (Fig. S1). Alpha-diversity data were then analyzed for overall differences using the non-parametric Kruskal–Wallis test. Differences in community composition (β-diversity) were computed using Bray–Curtis dissimilarity matrices and tested via permutational multivariate analysis of variance (PERMANOVA). Variation in community composition among samples was visualized with PCoA. A test for multivariate homogeneity of group dispersions (PERMDISP) was used to check for homogeneity of variances and pairwise comparisons were performed between groups. Core genera were identified as genera that were present in all Symbiodiniaceae strains within one location. The indicator value analysis [68] was applied to detect genera that were significantly associated with a specific treatment when both specificity and fidelity had probabilities >75%.
Results
Intra- and extracellular bacteria of diverse taxonomic affiliations associate with cultured Symbiodiniaceae
To characterize the bacterial communities associated with Symbiodiniaceae, we first used FISH and CSLM to localize bacteria in seven different Symbiodiniaceae cultures (Table S1). The use of the universal bacterial probes (EUB338-mix) revealed the presence of bacteria closely associated with cells of all Symbiodiniaceae species examined (Fig. 2A and S2). Orthogonal projections of Z-stacks showed that bacteria were localized both intra- and extracellularly (Fig. 2B). Extracellular bacteria occurred both close to the cell surface, and free in the culture (Figure S2). SEM confirmed the occurrence of extracellular bacteria attached to the algal surface in six tested cultures (Fig. 2C and S3). Dividing extracellular bacteria were observed (Fig. S3A), suggesting they are active and viable, and some bacteria occurred within extracellular polymeric substances (Fig. S3D), potentially indicating very close physiological association, despite still being extracellular.
A Summary table of bacterial occurrence in seven Symbiodiniaceae species based on FISH observations. Intra: intracellular; Extra: extracellular; ± signifies present/absent. B Orthogonal projections of a Z-stack of two Gerakladium sp. (G3) cells stained with the EUBmix probe observed in CLSM, highlighting the presence of both intra- and extracellular bacterial associates. C SEM photo of two Gerkladium sp. (G3) cells, highlighting the presence of bacteria attached to the cell exterior. Localization of FISH-stained Alphaproteobacteria (D), Gammaproteobacteria (E), and Flavobacteriia (F) in Gerkladium sp. (G3), observed in CLSM. Scale bar in B, D, E, F is 10 µm. Arrows point at extracellular bacteria and arrowheads point at intracellular bacteria. Green: Symbiodiniaceae; red: EUB338-mix probe (all bacteria); blue: Alf1B probe (Alphaproteobacteria); white: Gam42a probe (Gammaproteobacteria); magenta: CF319 probe (Flavobacteriia). G FISH negative control using the nonEUB probe. All scale bars are 10 µm (color figure online).
Furthermore, FISH on the Symbiodiniaceae cultures using class-specific bacterial probes (Table S2) revealed that all Symbiodiniaceae species examined were associated with extracellular bacteria of Alphaproteobacteria, Gammaproteobacteria, and Flavobacteriia, but the taxonomic affiliation of the intracellular bacteria differed among Symbiodiniaceae species (Figs. 2A, D–F, and S2). Flavobacteriia were the most commonly occurring intracellular bacteria, with only Durusdinium glynnii not harboring this class. B. minutum, Fugacium sp. (F5.1) and Gerakladium sp. (G3) were the only Symbiodiniaceae cultures where intracellular bacteria hybridized with all probe types, indicating that divergent bacterial species co-occur intracellularly. Such co-occurrence was confirmed by combining class-specific probes: Gammaproteobacteria were found to co-occur intracellularly with both Alphaproteobacteria and Flavobacteriia in B. minutum (Fig. S4A, B), and with Flavobacteriia in Gerakladium sp. (G3) (Fig. S4C).
Bacteria also associate with Symbiodiniaceae in hospite
To assess whether the observed bacteria–Symbiodiniaceae associations were not artefacts of extended laboratory culture, we freshly isolated Symbiodiniaceae from two cnidarians: the sea anemone E. diaphana and the scleractinian coral G. fascicularis. FISH probing and CSLM of freshly isolated Symbiodiniaceae from E. diaphana showed that Alphaproteobacteria, Gammaproteobacteria, and Flavobacteriia were all found both intra- and extracellularly (Fig. 3A). In G. fascicularis’ Symbiodiniaceae, only Flavobacteriia were found intracellularly, while all three bacterial classes were observed extracellularly (Fig. 3B). These findings indicate that in hospite Symbiodiniaceae also closely associate and interact with bacteria.
Localization of FISH-stained bacteria in Symbiodiniaceae freshly isolated from the anemone Exaiptasia diaphana (A) and the coral Galaxea fascicularis (B), observed with CLSM. Scale bar is 10 µm in all photos. Arrows point at extracellular bacteria and arrowheads point at intracellular bacteria. Green: Symbiodiniaceae; red: EUB338-mix probe (all bacteria); blue: Alf1B probe (Alphaproteobacteria); white: Gam42a probe (Gammaproteobacteria); magenta: CF319 probe (Flavobacteriia) (color figure online).
Cell cycling and sampling time influences bacterial prevalence in Cladocopium goreaui
Bacterial abundances differed between cells within samples and between Symbiodiniaceae species. Symbiodiniaceae follow a day–night cell cycle with cell division happening at dawn [69]. We hypothesized that the different physiological, metabolic, and morphological cellular states might impact the abundance, location, and composition of associated bacteria. Thus, we sampled C. goreaui at regular intervals for 24 h, combined FISH with propidium iodide (PI) staining to compare bacterial prevalence in the G1 and G2 phases of the algal cell cycle, and processed samples by flow cytometry to quantify the proportion of Symbiodiniaceae associated with bacteria (Fig. S5). PI staining revealed a higher proportion of Symbiodiniaceae cells in G2 or M phase during the night (Fig. 4A), consistently with cell division occurring at dawn. Within a sample, not all cells were stained by FISH (Fig. 4B, dotted black lines), suggesting that not all cells are associated with bacteria at any given time. The proportion of stained cells varied between 33% at 2:30 p.m. and 82% at 6:30 p.m. immediately after the lights were turned off and was consistently higher at night. Time of sampling was a statistically significant factor in these variations (ANOVA; F(6,14) = 16.1; p < 0.0001). When separating samples by their cell cycle phase as determined by the PI staining, the same trend was observed, with the proportion of stained cells being higher at night for both groups (Fig. 4B, red and blue lines). In addition, cells in G1 or G0 phase showed a higher proportion of stained cells than cells in G2 or M phase. Both sampling time and cell cycle phase were statistically significant factors, but their interaction was not (ANOVA; Ftime(6,28) = 34.8, ptime < 0.0001; Fcell cycle(1,28) = 5.8, pcell cycle = 0.02; Finteraction(6,28) = 1.1, pinteraction = 0.41). Thus, while it is evident that Symbiodiniaceae cells tend to more readily associate with bacteria during the night, the cell cycle phase does not explain this variability, as cells in G1 or G0 phase show a higher proportion of stained cells regardless of sampling time.
A Proportion of cells either in G1 or G0 phase (red) and G2 or M phase (blue) across a day, as assessed by PI staining analyzed by flow cytometry. B Proportion of cells stained by FISH in either all cells (black), cells in G1 and G0 phase (red), or cells in G2 and M phase (blue), as analyzed by flow cytometry. For each time point, three independent replicates from three cultures are represented, with dotted lines linking the median of each condition. Gray-shaded areas represent dark time, while white areas represent light time (color figure online).
Culture-dependent and -independent methods reveal diverse intracellular and extracellular bacterial communities
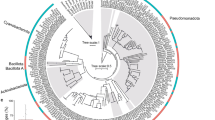
Pure culturing of bacteria associated with Symbiodiniaceae cultures revealed the existence of diverse bacterial communities. Pure bacterial isolates (n = 141) from six Symbiodiniaceae species were identified through phylogenetic analysis of their 16S rRNA gene sequences (Fig. S6, Dataset S1) and found to span 20 genera within four classes: Alpha- and Gammaproteobacteria, Flavobacteriia, and Bacilli. Only Muricauda (Flavobacteriia) was cultured from all Symbiodiniaceae species. Because the bacterial diversity revealed from pure-culturing methods likely under-represents the actual diversity, we also implemented a culture-independent method—16S rRNA gene metabarcoding. Each algal culture was fractionated through washes of different stringency in three sub-components, called loosely associated, closely associated (intracellular and tightly attached to the outer surface of Symbiodiniaceae cells), and intracellular bacteria (see Fig. 1 and material and methods for detailed experimental design). These three groups are hereafter referred to as the ‘location’ factor. SEM of B. minutum and Fugacium sp. (F5.1) cells confirmed that washing the cells with sodium hypochlorite removed any extracellular bacteria while keeping the Symbiodiniaceae cells intact, while washing them with fRSS removed some, but not all, cell surface-attached bacteria, suggesting that only closely attached bacteria remain (Fig. S7). Sequencing produced 7,747,433 reads across 11 Symbiodiniaceae cultures (n = 3 per culture per treatment; 3 treatments; 99 samples in total), filter controls (n = 3), DNA extraction blanks (n = 2), and no template PCRs (n = 4). Four samples were removed because they had low-read counts (<1000 reads). After merging, denoising and chimera filtering 4,831,092 reads remained. After removal of contaminants, 884 ASVs were observed across the remaining samples.
One out of three alpha-diversity metrics (observed ASVs) showed significant differences among Symbiodiniaceae locations (Kruskal–Wallis test; χ2ObsASBs(2) = 8.7, p = 0.01) and Symbiodiniaceae strains (Kruskal–Wallis test; χ2ObsASBs(10) = 37.3, p < 0.001), while Shannon’s index only showed differences among Symbiodiniaceae locations (Kruskal–Wallis test; χ2Shan(2) = 8.8, p = 0.01) (Fig. S8A, B). The Simpson index did not show any difference among locations (Kruskal–Wallis test; χ2Simp(2) = 2.5, p = 0.28) or Symbiodiniaceae strains (Kruskal–Wallis test; χ2Simp(10) = 14.9, p = 0.13). Principal coordinate analysis (PCoA) visualization of β-diversity using the Bray–Curtis dissimilarity index revealed a clear overlap between closely and loosely associated communities, while intracellular communities formed a separate cluster (Fig. 5A). Statistical analyses showed that community structure varied with Symbiodiniaceae location and strain, and their interaction (PERMANOVA, Bray–Curtis method, 999 permutations; Fstrain(10,62) = 11.5, p < 0.001; Flocation(2,62) = 27.4, p < 0.001; Finteraction(20,62) = 5.3, p < 0.001). Pairwise PERMANOVA comparisons between locations confirmed all three were statistically different from each other (p = 0.001 for each pairwise comparison). However, the PCoA representation highlighted that, for a given strain, closely and loosely associated communities were often similar. PCoAs confirmed that intracellular communities from different Symbiodiniaceae strains overlap (Fig. S8C), while closely and loosely associated communities are clearly separated based on strain (Fig. S8D, E). Community composition at the family level highlighted those dissimilarities (Fig. 5B). Intracellular communities were dominated by Hyphomicrobiaceae, Beijerinckiaceae, and Sphingomonadaceae (Fig. 5B, upper panel); closely associated communities were dominated by Rhodobacteraceae, Marinobacteraceae, Phycisphaeraceae, Rhizobiaceae, and Stappiaceae (Fig. 5B, middle panel); loosely associated communities were dominated by Rhodobacteraceae, Marinobacteraceae, Algiphilaceae, and Balneolaceae (Fig. 5B, lower panel).
A PCoA visualization of beta-diversity of the bacterial communities in 11 Symbiodiniaceae strains and three locations, based on Bray–Curtis dissimilarity matrices. Each point is an individual sample. Both location and Symbiodiniaceae strain, as well as their interaction, had significant effects based on PERMANOVA testing. B Relative abundance of bacterial families in Symbiodiniaceae’s intracellular (upper panel), closely associated (middle panel), and loosely associated (lower panel) communities. For each strain × location combination, three independent replicates were merged. UC unclassified.
Genera that yielded culturable isolates from at least four of the six Symbiodiniaceae species (Fig. S6) were specifically investigated in the metabarcoding data. Marinobacter, Labrenzia, and Roseitalea were consistently abundant in most samples (Figure S9). Muricauda was present in most samples, but with a relative abundance below 1% in all samples (Figure S9). The prevalence and abundance of Bacillus and Roseovarius were very low, with Bacillus being present in only three Symbiodinaceae strains, contrasting with their high prevalence in cultured isolates. Intriguingly, typically intracellular taxa, such as Simkaniaceae and Rickettsiales, were found in high abundances both in closely and loosely associated communities, particularly in C. goreaui and Durusdinium glynii, but in low abundance in intracellular communities (Fig. 5B, Dataset S2). Simkaniaceae alternate between infectious extracellular elementary bodies, and intracellular replicative reticulate bodies [70, 71]. Hence, high numbers of Simkaniaceae elementary bodies might have been released, through vesicles or host cell lysis, at the time of sampling, explaining their high abundance in extracellular communities.
Investigation of core bacterial communities in Symbiodiniaceae
A large number of genera (325) were present among the 884 ASVs observed across all samples. Seven genera were found in intracellular communities of all 11 Symbiodiniaceae strains, seven in closely associated communities of all strains, and five in loosely associated communities of all strains (Fig. 6A). These genera are hereafter referred to as ‘core genera’ for their respective location. No genus was found to occur in all Symbiodiniaceae strains and all locations. Intracellular core genera accounted for an average of 57% of all reads in intracellular communities across all strains (Fig. 6B, left panel, Table S4, and Dataset S3A), with Hyphomicrobium, Methylobacterium, and Sphingomonas being the most abundant. Intracellular core genera had very low abundance in closely and loosely associated communities (<1% of all reads for both locations in all strains) (Fig. 6B, left panel, Table S4, and Dataset S3A), suggesting they are specifically present in intracellular communities. Conversely, closely associated core genera accounted for 44%, 26%, and 6% in closely associated, loosely associated and intracellular communities, respectively (Fig. 6B, middle panel, Table S4, and Dataset S3B); likewise, loosely associated core genera accounted for 33%, 20%, and 5% in closely associated, loosely associated and intracellular communities, respectively (Fig. 6B, right panel, Table S4, and Dataset S3C). This suggests that closely and loosely associated communities show a degree of similarity that they do not share with intracellular communities, which is consistent with the PCoA results. Finally, an indicator value analysis was performed to identify genera that represent indicators for each location. Eight genera were identified: Pseudomonas, Hyphomicrobium, Methylobacterium, and Sphingomonas for intracellular communities; Labrenzia, uncultured Phycisphaeraceae, and uncultured Kiloniellaceae for closely associated communities; Pseudohongiella for loosely associated communities. All but uncultured Kiloniellaceae were already present in the core genera analysis and are underlined in Fig. 6A. Uncultured Kiloniellaceae had an abundance lower than 1% in all but two samples (C. goreaui SCF049.01—closely associated; Fugacium kawagutii—Closely associated).
A Relative abundance of genera found in every Symbiodiniaceae strains in either intracellular, closely associated, or loosely associated samples—‘core genera’. For each value, three independent replicates were merged. Asterisk in front of an underlined genus signifies that the genus was also detected through an indicator genera analysis. B Relative abundance of the sum of all core genera from intracellular (left panel; seven genera), closely associated (middle panel; seven genera), or loosely associated (right panel; five genera) communities.
Discussion
Despite coral-Symbiodiniaceae interactions having been extensively studied, and coral–bacteria interactions recently gaining traction, Symbiodiniaceae–bacteria interactions remain understudied. Deciphering these interactions in ex hospite Symbiodiniaceae is the first step toward understanding Symbiodiniaceae–bacteria cooperation within the coral holobiont. Here, we describe the bacterial microbiome of a wide diversity of cultured Symbiodiniaceae. Notably, through three-dimensional CLSM and SEM on long-term cultures of six genera, we provide unequivocal evidence that bacteria are present inside Symbiodiniaceae cells, as well as on the cell’s exterior. We show the occurrence of three different classes within, and outside, Symbiodiniaceae cells: Alpha- and Gammaproteobacteria, and Flavobacteriia. In addition, different classes were found to co-occur within the same Symbiodiniaceae cell, suggesting the possibility of inter-class, intracellular cooperation in this symbiosis. We also show the presence of intra- and extracellular bacteria in Symbiodiniaceae freshly isolated from two cnidarians, suggesting that Symbiodiniaceae–bacteria interactions might be similar within cnidarian hosts. Additional work will be needed on in hospite Symbiodiniaceae to confirm that our results obtained in long-term cultures are generalizable to cnidarian holobionts.
Metabarcoding data indicated that bacterial communities associated with Symbiodiniaceae cultures were largely comprised of Hyphomicrobium, Methylobacterium, Sphingomonas, Marinobacter, Labrenzia, Balneola, Algiphilus, Rhodobacteraceae, Phycisphaeraeae, and Roseitalea. Marinobacter, Labrenzia, and Muricauda were previously found to be abundant in a wide range of Symbiodiniaceae cultures [39,40,41], indicating the conservation of specific bacterial associates across Symbiodiniaceae species, coral hosts, and coral host location. Marinobacter enhances the growth of Scrippsiella trochoidea, presumably by producing vibrioferrin, a siderophore that promotes iron uptake by the dinoflagellate [18]. Labrenzia has the ability to produce and degrade DMSP [72, 73], a compound that has ROS-scavenging abilities [74], and thus could have functions in overall ROS regulation, or excess ROS scavenging during coral bleaching events. Muricauda produces zeaxanthin, a carotenoid with ROS-scavenging abilities, and is known to increase heat and light resistance in cultured Symbiodiniaceae [19]. Through these functions, it is thus likely that Marinobacter, Labrenzia, and Muricauda are beneficial symbionts of Symbiodiniaceae.
Unlike previous studies, our metabarcoding experiment distinguished between intracellular, closely associated, and loosely associated bacterial communities. Closely and loosely associated communities were similar to each other in any given Symbiodiniaceae strain, while intracellular communities were different from those in both extracellular locations. Furthermore, intracellular communities showed high similarity among strains, as evidenced by the fact that the seven intracellular core genera accounted on average for more than 50% of all intracellular reads. Interestingly, core genera from closely and loosely associated communities largely overlapped within a given Symbiodiniaceae strain. This indicates that the closely associated communities were dominated by extracellular bacteria that were also present in the loosely associated communities. It is possible that loosely associated bacteria are derived from closely associated bacteria and can live freely in the algal culture medium, reinforcing the importance of the phycosphere in shaping extracellular bacterial assemblages. Whether the phycosphere promotes the establishment of intracellular bacteria remains unclear, and the factors attracting bacteria and allowing them to enter Symbiodiniaceae cells, in culture but also in hospite, need to be addressed. While extracellular associates could come from extracellular bacteria naturally associating with Symbiodiniaceae in cnidarian hosts or in seawater, they also might have been accidentally introduced, and subsequently maintained, during the culturing process. Comparing bacterial communities between cultured and in hospite Symbiodiniaceae will shed light on this question.
The core genera analysis allowed us to identify conserved bacterial genera across Symbiodiniaceae strains, which might support crucial functions for the Symbiodiniaceae. Among intracellular core genera, Hyphomicrobium, Methylobacterium, and Sphingomonas were the most abundant. Hyphomicrobium can metabolize dimethylsulfate [75], and could thus be involved in sulfur cycling. Incidentally, this process is improved when co-cultured with Pseudomonas [75], another intracellular core genus. Hyphomicrobium species can denitrify using methanol as a carbon source [76]. Interestingly, Methylobacterium is methylotrophic, and can also use methanol, and other short carbon compounds, as a carbon source. Whether there is sufficient methanol production within Symbiodiniaceae cells to support such activity is unknown. Sphingomonas has previously been found intracellularly in the toxic dinoflagellate Alexandrium minutum [77]. Some Sphingomonas species are able to fix nitrogen in rice plants [78], and hence could be involved in nitrogen cycling within Symbiodiniaceae cells. Interestingly, Ralstonia, a core intracellular genus here, was found to closely associate with in hospite Symbiodiniaceae in the coral A. granulosa [43], although its role remains elusive.
Unexpectedly, intracellular communities showed the highest richness. Nonetheless, community composition analysis indicates that some cultures are dominated by very few families: only two families show relative abundance above 3% in the intracellular communities of C. goreaui SCF055.01-10 and D. trenchii SCF082, and these two families account for more than 60% of the reads. Low-abundance bacteria detected in intracellular communities might therefore not be present in all the cells in the sample and could be the result of heterotrophy. Indeed, Symbiodiniaceae have been shown to engulf and feed on bacteria [79]. While it is possible that low-abundance families are the result of unspecific heterotrophic feeding, several lines of evidence point at a significant proportion of bacteria being true intracellular symbionts: (1) intracellular bacteria were detected in Symbiodiniaceae freshly isolated from cnidarian hosts, where a bacterium encounter and engulfment is less likely than in culture; (2) flow cytometry and FISH showed a high proportion of cells associated with intracellular bacteria, and sometimes had high abundances of intracellular bacteria, during the day when heterotrophy is minimal (Jeong et al. [79] show a maximum of 6 Synechococcus cells being ingested per Symbiodiniaceae cell per hour); (3) we show the existence of a specific and conserved intracellular community that is low in abundance in extracellular communities.
Finally, we provide evidence that the Symbiodiniaceae microbiome is dynamic. We demonstrate that daily cycling and sampling time, especially light, influence the proportion of Symbiodiniaceae cells that are closely associated with bacteria. This is concordant with previous studies that showed Symbiodiniaceae bacterial communities change with temperature [41], and that bacterial density increases with culture age in the dinoflagellates Heterocapsa circularisquama, Alexandrium catenella, and Protoceratium reticulatum [11, 80]. It was also shown in H. circularisquama that the density of intracellular bacteria increases when the dinoflagellate culture is switched to continuous darkness [80], an increase we also report here during the night period. Several factors could explain such an increase when these dinoflagellates do not have access to light and thus cannot photosynthesize: (1) dark mitochondrial respiration modifies oxygen levels, pH and overall metabolite profiles within the cell as well as within the phycosphere, factors that could impact bacterial density and diversity; and (2) heterotrophy [79], which is higher in the dark [81].
In conclusion, the data presented in this study provide important new insights into Symbiodiniaceae–bacteria interactions in long-term cultures and to some extent in hospite. Future studies should focus on identifying the in hospite Symbiodiniaceae-associated bacterial communities in greater detail and assess whether Symbiodiniaceae-bacteria communities are similar in and ex hospite. Importantly, we show the existence of a complex intracellular consortium of bacteria, which is highly conserved across six cultured Symbiodiniaceae genera. Determining the function of these endosymbionts, and whether they might contribute to coral health, and potentially to bleaching resilience, will be key in fully understanding Symbiodiniaceae–bacteria interactions. Bacterial probiotics are increasingly brought forward as a potential solution to mitigate coral bleaching [33, 82,83,84]. ROS-scavenging bacteria able to intimately associate with Symbiodiniaceae may be strong candidates for probiotics as excessive levels of ROS, the putative cause of coral bleaching, originate within Symbiodiniaceae cells. Further, intracellular bacteria could be considered as targets for genetic modification, because their intracellular lifestyle might prevent them from spreading to the environment and other non-targeted organisms.
Data availability
Genbank accession numbers for 16S rRNA sequences of cultured bacteria are MT840521-MT840661 (see Dataset S1). Raw MiSeq data are available under NCBI BioProject ID PRJNA650221.
Change history
14 April 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41396-021-00970-6
References
McFall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Domazet-Lošo T, Douglas AE, et al. Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci USA. 2013;110:3229–36.
Moran NA. Symbiosis as an adaptive process and source of phenotypic complexity. Proc Natl Acad Sci USA. 2007;104:8627–33.
Gilbert SF, Sapp J, Tauber AI. A symbiotic view of life: we have never been individuals. Q Rev Biol. 2012;87:325–41.
Amin SA, Parker MS, Armbrust EV. Interactions between diatoms and bacteria. Microbiol Mol Biol Rev. 2012;76:667–84.
Ramanan R, Kim B-H, Cho D-H, Oh H-M, Kim H-S. Algae–bacteria interactions: evolution, ecology and emerging applications. Biotechnol Adv. 2016;34:14–29.
Matthews JL, Raina J, Kahlke T, Seymour JR, van Oppen MJH, Suggett DJ. Symbiodiniaceae‐bacteria interactions: rethinking metabolite exchange in reef-building corals as multi-partner metabolic networks. Environ Microbiol. 2020;22:1675.
Bell W, Mitchell R. Chemotactic and growth responses of marine bacteria to algal extracellular products. Biol Bull. 1972;143:265–77.
Seymour JR, Amin SA, Raina JB, Stocker R. Zooming in on the phycosphere: the ecological interface for phytoplankton-bacteria relationships. Nat Microbiol. 2017;2:17065.
Bird C, Darling KF, Russell AD, Davis CV, Fehrenbacher J, Free A, et al. Cyanobacterial endobionts within a major marine planktonic calcifier (Globigerina bulloides, Foraminifera) revealed by 16S rRNA metabarcoding. Biogeosciences. 2017;14:901–20.
Schulz F, Horn M. Intranuclear bacteria: inside the cellular control center of eukaryotes. Trends Cell Biol. 2015;25:339–46.
Córdova JL, Escudero C, Bustamante J. Bloom inside the bloom: intracellular bacteria multiplication within toxic dinoflagellates. Rev Biol Mar Oceanogr. 2003;38:57–67.
Kirchner M, Sahling G, Schütt C, Döpke H, Uhlig G. Intracellular bacteria in the red tide-forming heterotrophic dinoflagellate Noctiluca scintillans. Arch fur Hydrobiol Spec Issue Adv Limnol. 1999;54:297.
Biegala IC, Kennaway G, Alverca E, Lennon J, Vaulot D, Simon N. Identification of bacteria associated with dinoflagellates (Dinophyceae) Alexandrium spp. using tyramide signal amplification-fluorescent in situ hybridization and confocal microscopy. J Phycol. 2002;38:404–11.
Croft MT, Lawrence AD, Raux-Deery E, Warren MJ, Smith AG. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature. 2005;438:90–3.
Kazamia E, Czesnick H, Nguyen TTVan, Croft MT, Sherwood E, Sasso S, et al. Mutualistic interactions between vitamin B12-dependent algae and heterotrophic bacteria exhibit regulation. Environ Microbiol. 2012;14:1466–76.
Xie B, Bishop S, Stessman D, Wright D, Spalding MH, Halverson LJ. Chlamydomonas reinhardtii thermal tolerance enhancement mediated by a mutualistic interaction with vitamin B12-producing bacteria. ISME J. 2013;7:1544–55.
Grant MAA, Kazamia E, Cicuta P, Smith AG. Direct exchange of vitamin B12 is demonstrated by modelling the growth dynamics of algal-bacterial cocultures. ISME J. 2014;8:1418–27.
Amin SA, Green DH, Hart MC, Küpper FC, Sunda WG, Carrano CJ. Photolysis of iron-siderophore chelates promotes bacterial-algal mutualism. Proc Natl Acad Sci USA. 2009;106:17071–6.
Motone K, Takagi T, Aburaya S, Miura N, Aoki W, Ueda M. A Zeaxanthin-producing bacterium isolated from the algal phycosphere protects coral endosymbionts from environmental stress. MBio. 2020;11:e01019–19.
Amin SA, Hmelo LR, Van Tol HM, Durham BP, Carlson LT, Heal KR, et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature. 2015;522:98–101.
Trench RK. Microalgal-invertebrate symbioses: a review. Endocytobiosis Cell Res. 1993;9:135.
Blackall LL, Wilson B, van Oppen MJH. Coral-the world’s most diverse symbiotic ecosystem. Mol Ecol. 2015;24:5330–47.
LaJeunesse TC, Parkinson JE, Gabrielson PW, Jeong HJ, Reimer JD, Voolstra CR, et al. Systematic revision of Symbiodiniaceae highlights the antiquity and diversity of coral endosymbionts. Curr Biol. 2018;28:2570–2580.e6.
Nitschke MR, Craveiro SC, Brandão C, Fidalgo C, Serôdio J, Calado AJ, et al. Description of Freudenthalidium gen. nov. and Halluxium gen. nov. to formally recognize Clades Fr3 and H as genera in the family Symbiodiniaceae (Dinophyceae). J Phycol. 2020;56:923–40.
Muscatine L, Porter JW. Reef corals: mutualistic symbioses adapted to nutrient-poor environments. Bioscience 1977;27:454–60.
Muscatine L, McCloskey LR, Marian RE. Estimating the daily contribution of carbon from zooxanthellae to coral animal respiration. Limnol Oceanogr. 1981;26:601–11.
Muscatine L, Falkowski PG, Porter JW, Dubinsky Z. Fate of photosynthetic fixed carbon in light- and shade-adapted colonies of the symbiotic coral Stylophora pistillata. Proc R Soc Lond Ser B Biol Sci. 1984;222:181–202.
Hughes TP, Barnes ML, Bellwood DR, Cinner JE, Cumming GS, Jackson JBC, et al. Coral reefs in the Anthropocene. Nature. 2017;546:82–90.
Hughes TP, Anderson KD, Connolly SR, Heron SF, Kerry JT, Lough JM, et al. Spatial and temporal patterns of mass bleaching of corals in the Anthropocene. Science. 2018;359:80–3.
Weis VM. Cellular mechanisms of Cnidarian bleaching: stress causes the collapse of symbiosis. J Exp Biol. 2008;211:3059–66.
Suggett DJ, Smith DJ. Coral bleaching patterns are the outcome of complex biological and environmental networking. Glob Change Biol. 2020;26:68–79.
Davy SK, Allemand D, Weis VM. Cell biology of cnidarian-dinoflagellate symbiosis. Microbiol Mol Biol Rev. 2012;76:229–61.
van Oppen MJH, Blackall LL. Coral microbiome dynamics, functions and design in a changing world. Nat Rev Microbiol. 2019;17:557.
Bourne DG, Morrow KM, Webster NS. Insights into the coral microbiome: underpinning the health and resilience of reef ecosystems. Annu Rev Microbiol. 2016;70:317–40.
Ceh J, Kilburn MR, Cliff JB, Raina J-B, van Keulen M, Bourne DG. Nutrient cycling in early coral life stages: Pocillopora damicornis larvae provide their algal symbiont (Symbiodinium) with nitrogen acquired from bacterial associates. Ecol Evol. 2013;3:2393–400.
Lema KA, Clode PL, Kilburn MR, Thornton R, Willis BL, Bourne DG. Imaging the uptake of nitrogen-fixing bacteria into larvae of the coral Acropora millepora. ISME J. 2016;10:1804–8.
Rädecker N, Pogoreutz C, Voolstra CR, Wiedenmann J, Wild C. Nitrogen cycling in corals: the key to understanding holobiont functioning? Trends Microbiol. 2015;23:490–7.
Raina JB, Clode PL, Cheong S, Bougoure J, Kilburn MR, Reeder A, et al. Subcellular tracking reveals the location of dimethylsulfoniopropionate in microalgae and visualises its uptake by marine bacteria. Elife. 2017;6:e23008.
Frommlet JC, Sousa ML, Alves A, Vieira SI, Suggett DJ, Serôdio J. Coral symbiotic algae calcify ex hospite in partnership with bacteria. Proc Natl Acad Sci USA. 2015;112:6158–63.
Lawson CA, Raina J-B, Kahlke T, Seymour JR, Suggett DJ. Defining the core microbiome of the symbiotic dinoflagellate, Symbiodinium. Environ Microbiol Rep. 2018;10:7–11.
Camp EF, Kahlke T, Nitschke MR, Varkey D, Fisher NL, Fujise L, et al. Revealing changes in the microbiome of Symbiodiniaceae under thermal stress. Environ Microbiol. 2020:1462-2920.14935.
Shoguchi E, Shinzato C, Kawashima T, Gyoja F, Mungpakdee S, Koyanagi R, et al. Draft assembly of the Symbiodinium minutum nuclear genome reveals dinoflagellate gene structure. Curr Biol. 2013;23:1399–408.
Ainsworth TD, Krause L, Bridge T, Torda G, Raina J-B, Zakrzewski M, et al. The coral core microbiome identifies rare bacterial taxa as ubiquitous endosymbionts. ISME J. 2015;9:2261–74.
Tortorelli G, Belderok R, Davy SK, Mcfadden GI, Van Oppen MJH. Host genotypic effect on algal symbiosis establishment in the coral model, the anemone Exaiptasia diaphana, from the Great Barrier Reef. Front Mar Sci. 2020;6:833.
Dungan AM, Hartman LM, Tortorelli G, Belderok R, Lamb AM, Pisan L, et al. Exaiptasia diaphana from the great barrier reef: a valuable resource for coral symbiosis research. Symbiosis. 2020;80:195–206.
Bongaerts P, Sampayo EM, Bridge TCL, Ridgway T, Vermeulen F, Englebert N, et al. Symbiodinium diversity in mesophotic coral communities on the Great Barrier Reef: a first assessment. Mar Ecol Prog Ser. 2011;439:117–26.
LaJeunesse TC, Bhagooli R, Hidaka M, DeVantier L, Done T, Schmidt GW, et al. Closely related Symbiodinium spp. differ in relative dominance in coral reef host communities across environmental, latitudinal and biogeographic gradients. Mar Ecol Prog Ser. 2004;284:147–61.
Hugenholtz P, Tyson GW, Blackall LL. Design and evaluation of 16S rRNA-targeted oligonucleotide probes for fluorescence in situ hybridization. Methods Mol Biol. 2002;179:29–42.
Manz W, Amann R, Ludwig W, Wagner M, Schleifer KH. Phylogenetic oligodeoxynucleotide probes for the major subclasses of Proteobacteria: problems and solutions. Syst Appl Microbiol. 1992;15:593–600.
Manz W, Amann R, Ludwig W, Vancanneyt M, Schleifer KH. Application of a suite of 16S rRNA-specific oligonucleotide probes designed to investigate bacteria of the phylum cytophaga-flavobacter-bacteroides in the natural environment. Microbiology. 1996;142:1097–106.
Daims H, Brühl A, Amann R, Schleifer KH, Wagner M. The domain-specific probe EUB338 is insufficient for the detection of all bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol. 1999;22:434–44.
Wallner G, Amann R, Beisker W. Optimizing fluorescent in situ hybridization with rRNA-targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry. 1993;14:136–43.
Wilson K, Li Y, Whan V, Lehnert S, Byrne K, Moore S, et al. Genetic mapping of the black tiger shrimp Penaeus monodon with amplified fragment length polymorphism. Aquaculture. 2002;204:297–309.
Hartman LM, van Oppen MJH, Blackall LL. Microbiota characterization of Exaiptasia diaphana from the Great Barrier Reef. Anim Microb. 2020;2:10.
Aubrey BJ, Kelly GL, Kueh AJ, Brennan MS, O’Connor L, Milla L, et al. An inducible lentiviral guide RNA platform enables the identification of tumor-essential genes and tumor-promoting nutations in vivo. Cell Rep. 2015;10:1422–32.
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol. 2019;37:852–7.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581–3.
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome. 2018;6:90.
Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80.
Price MN, Dehal PS, Arkin AP. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE. 2010;5:e9490.
R Core Team. R: A language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2018. http://www.R-project.org/.
McMurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE. 2013;8:e61217.
Oksanen J, Guillaume Blanchet F, Kindt R, Legendre P, Minchin PR, O’Hara RB, et al. Package “vegan” community ecology package. 2018. https://cran.r-project.org.
Hervé M. RVAideMemoire: testing and plotting procedures for biostatistics. R package version 0.9–69. 2018. https://cran.r-project.org.
Wickham H. ggplot2: elegant graphics for data analysis. New York: Springer-Verlag; 2009.
Wickham H. tidyverse: easily install and load the ‘Tidyverse’. R package version 1.2.1. 2017. https://tidyverse.tidyverse.org.
De Cáceres M, Legendre P. Associations between species and groups of sites: Indices and statistical inference. Ecology. 2009;90:3566–74.
Fitt WK, Trench RK. The relation of diel patterns ofr cell division to diel patterns of motility in the symbiotic dinoflagellate Symbiodinium microadriaticum Freudenthal in culture. N Phytol. 1983;94:421–32.
Horn M. Chlamydiae as symbionts in Eukaryotes. Annu Rev Microbiol. 2008;62:113–31.
Collingro A, Köstlbacher S, Horn M. Chlamydiae in the environment. Trends Microbiol. 2020;28:877.
Curson ARJ, Liu J, Bermejo Martínez A, Green RT, Chan Y, Carrión O, et al. Dimethylsulfoniopropionate biosynthesis in marine bacteria and identification of the key gene in this process. Nat Microbiol. 2017;2:17009.
Hatton AD, Shenoy DM, Hart MC, Mogg A, Green DH. Metabolism of DMSP, DMS and DMSO by the cultivable bacterial community associated with the DMSP-producing dinoflagellate Scrippsiella trochoidea. Biogeochemistry. 2012;110:131–46.
Sunda W, Kieber DJ, Kiene RP, Huntsman S. An antioxidant function for DMSP and DMS in marine algae. Nature. 2002;418:317–20.
Zhang L, Hirai M, Shoda M. Removal characteristics of dimethyl sulfide by a mixture of Hyphomicrobium sp. I55 and Pseudomonas acidovorans DMR-11. J Ferment Bioeng. 1992;74:174–8.
Martineau C, Mauffrey F, Villemur R. Comparative analysis of denitrifying activities of Hyphomicrobium nitrativorans, Hyphomicrobium denitrificans, and Hyphomicrobium zavarzinii. Appl Environ Microbiol. 2015;81:5003–14.
Lu YH, Chai TJ, Hwang D. Isolation of bacteria from toxic dinoflagellate Alexandrium minutum and their effects on algae toxicity. J Nat Toxins. 2000;9:409–17.
Videira SS, de Araujo JLS, da Silva Rodrigues L, Baldani VLD, Baldani JI. Occurrence and diversity of nitrogen-fixing Sphingomonas bacteria associated with rice plants grown in Brazil. FEMS Microbiol Lett. 2009;293:11–9.
Jeong HJ, Yoo YDU, Kang NS, Lim AS, Seong KA, Lee SY, et al. Heterotrophic feeding as a newly identified survival strategy of the dinoflagellate Symbiodinium. Proc Natl Acad Sci USA. 2012;109:12604–9.
Maki T, Imai I. Relationships between intracellular bacteria and the bivalve killer dinoflagellate Heterocapsa circularisquama (Dinophyceae). Fish Sci. 2001;67:794–803.
Xiang T, Hambleton EA, DeNofrio JC, Pringle JR, Grossman AR. Isolation of clonal axenic strains of the symbiotic dinoflagellate Symbiodinium and their growth and host specificity. J Phycol. 2013;49:447–58.
van Oppen MJH, Oliver JK, Putnam HM, Gates RD. Building coral reef resilience through assisted evolution. Proc Natl Acad Sci USA. 2015;112:2307–13.
Peixoto RS, Rosado PM, Leite DC, de A, Rosado AS, Bourne DG. Beneficial microorganisms for corals (BMC): proposed mechanisms for coral health and resilience. Front Microbiol. 2017;8:341.
Blackall LL, Dungan AM, Hartman LM, van Oppen MJ. Probiotics for corals. Microbiol Aust. 2020;41:100–4.
Acknowledgements
This research was supported by the Australian Research Council Laureate Fellowship FL180100036 (to MJHvO) and the Australian Research Council (grant ID: DP160101468) to MJHvO and LLB. We thank the Biosciences Microscopy Unit (University of Melbourne) and the Biological Optical Microscopy Platform for the use of their confocal and scanning electron microscopes, and particularly Dr Allison van de Meene and Dr Gabriela Segal for their valuable assistance. We are grateful to: Dr. Katarina Damjanovic and Dr. Leon M. Hartman for their valuable help in developing codes for metabarcoding data analysis; Giada Tortorelli for assistance in Symbiodiniaceae maintenance; Dr. Wing Chan and Talisa Döring for providing Galaxea fascicularis polyps; Dr. Cindy Castelle for her help in creating phylogenetic trees.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to some errors in Figures 5 and 6 and in Supplementary Material
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maire, J., Girvan, S.K., Barkla, S.E. et al. Intracellular bacteria are common and taxonomically diverse in cultured and in hospite algal endosymbionts of coral reefs. ISME J 15, 2028–2042 (2021). https://doi.org/10.1038/s41396-021-00902-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-00902-4
This article is cited by
-
Shining light on the coral photosymbiont family Symbiodiniaceae
Nature Microbiology (2024)
-
A genome-centric view of the role of the Acropora kenti microbiome in coral health and resilience
Nature Communications (2024)
-
The coral microbiome in sickness, in health and in a changing world
Nature Reviews Microbiology (2024)
-
Metabolic relationships between marine red algae and algae-associated bacteria
Marine Life Science & Technology (2024)
-
Genomic exploration of coral-associated bacteria: identifying probiotic candidates to increase coral bleaching resilience in Galaxea fascicularis
Microbiome (2023)