Abstract
Termite mounds have recently been confirmed to mitigate approximately half of termite methane (CH4) emissions, but the aerobic CH4 oxidising bacteria (methanotrophs) responsible for this consumption have not been resolved. Here, we describe the abundance, composition and CH4 oxidation kinetics of the methanotroph communities in the mounds of three distinct termite species sampled from Northern Australia. Results from three independent methods employed show that methanotrophs are rare members of microbial communities in termite mounds, with a comparable abundance but distinct composition to those of adjoining soil samples. Across all mounds, the most abundant and prevalent methane monooxygenase sequences were affiliated with upland soil cluster α (USCα), with sequences homologous to Methylocystis and tropical upland soil cluster (TUSC) also detected. The reconstruction of a metagenome-assembled genome of a mound USCα representative highlighted the metabolic capabilities of this group of methanotrophs. The apparent Michaelis–Menten kinetics of CH4 oxidation in mounds were estimated from in situ reaction rates. Methane affinities of the communities were in the low micromolar range, which is one to two orders of magnitude higher than those of upland soils, but significantly lower than those measured in soils with a large CH4 source such as landfill cover soils. The rate constant of CH4 oxidation, as well as the porosity of the mound material, were significantly positively correlated with the abundance of methanotroph communities of termite mounds. We conclude that termite-derived CH4 emissions have selected for distinct methanotroph communities that are kinetically adapted to elevated CH4 concentrations. However, factors other than substrate concentration appear to limit methanotroph abundance and hence these bacteria only partially mitigate termite-derived CH4 emissions. Our results also highlight the predominant role of USCα in an environment with elevated CH4 concentrations and suggest a higher functional diversity within this group than previously recognised.
Similar content being viewed by others
Introduction
Termites are mound-building eusocial insects that live in colonies throughout the tropics and subtropics. These organisms completely degrade lignocellulose in a process primarily mediated by anaerobic symbiotic microorganisms in their hindgut [1]. During this process, hydrogenotrophic methanogens produce substantial amounts of methane (CH4) that is emitted from the termite into the atmosphere [2,3,4]. Production rates vary by three to four orders of magnitude depending on the termite species and their dietary preferences (i.e. wood-, grass-, soil- or fungus-feeding) [1, 4, 5]. Current models suggest that termites are responsible for between 1 and 3% of global CH4 emissions to the atmosphere [6].
Aerobic CH4 oxidizing bacteria (methanotrophs) significantly mitigate emissions of CH4 from termites [7]. Methanotrophs gain carbon and energy by oxidising CH4 to carbon dioxide, with the first step in this reaction being catalysed by particulate and soluble methane monooxygenases [8]. It is controversial whether termite hindguts harbour such organisms; while Methylocystis spp. were recently isolated from termites [9], other studies could not detect methanotroph functional gene markers or measurable amounts of 14CO2 during 14CH4 incubation experiments [10]. We also observed that the addition of inhibitors of CH4 oxidation did not increase direct termite CH4 emissions [7]. However, many termite colonies construct large mounds built from soil material or build their nest in soil, which is generally a sink for atmospheric CH4 [11]. While results from incubation experiments of mound material were conflicting [12,13,14], we recently presented clear evidence of widespread CH4 oxidation in North Australian termite mounds [7]: results from three different in situ methods to measure CH4 oxidation in mounds confirmed that methanotrophs mitigate between 20 and 80% of termite-derived CH4 before emission to the atmosphere. However, the community composition and CH4 oxidation kinetics of the methanotrophs responsible remain largely unknown.
Compared with soils, methanotrophs inhabiting termite mounds have received little attention. Ho et al. [14] investigated mound material of the African fungus-feeding termite Macrotermes falciger using a pmoA-based diagnostic microarray approach. Community composition differed between mound and soil, and at some locations within the large compartmentalised mounds. Slurry incubations confirmed potential CH4 oxidation at high and low CH4 concentrations. The mound community was reportedly dominated by gammaproteobacterial methanotrophs of the Jasper Ridge 3 (JR3) cluster, while the functional gene of the soluble methane monooxygenase could not be detected and nor could Verrucomicrobia and Methylomirabilota (NC10) methanotrophs. Beside this pioneering work, the mound methanotroph communities of no other termite species has been investigated. However, large differences might exist between mounds of different termite species given factors such as their different dietary preferences, as well as the impressive variety of mound sizes, shapes and internal structures [15, 16]. Reflecting this, in situ studies have shown that there is a large variation in methanotroph activity in mounds both within and between species; for example, Tumulitermes pastinator mounds appear to be largely inactive and still a high fraction of termite-derived CH4 can be oxidised in soil beneath mounds, due to facilitation of CH4 transport within the mound [7]. It remains unclear whether differences in methanotroph community abundance or composition account for these activity differences.
In this work, we aimed to resolve these discrepancies by conducting a comprehensive analysis of the composition and kinetics of the methanotroph communities within termite mounds of Australian termite species. Three mound-building termite species were selected, the wood-feeding Microcerotermes nervosus (Mn), soil-interface-feeding Macrognathotermes sunteri (Ms) and grass-feeding Tumulitermes pastinator (Tp), which represent the three main feeding groups present in Australia [17]. Mounds of these species were previously confirmed to oxidise a high fraction of termite-produced CH4 [7]. We comprehensively assessed the abundance, diversity and composition of methanotroph communities within 17 mounds and a subset of adjoining soils; to do so, we combined metagenomic sequencing with amplicon sequencing and quantitative PCR (qPCR) of both the 16S rRNA gene and the pmoA marker gene, encoding a subunit of the particulate methane monooxygenase present in most methanotrophs [18]. In parallel, we performed in situ studies using gas push–pull tests (GPPTs) to derive the apparent kinetic parameters of CH4 oxidation. We demonstrate that methanotrophic communities in termite mounds are derived from surrounding soils, but are compositionally and kinetically distinct, and primarily comprise upland soil cluster α (USCα) alphaproteobacteria with an apparent medium affinity for CH4.
Materials and methods
Field sites and sampling
Field tests and sampling were performed in April and May 2016 in a coastal savanna woodland on the campus of Charles Darwin University in Darwin, Northern Territory, Australia (12.370° S, 130.867° E). The site is described in detail in Nauer et al. [19], and is representative of large areas of tropical northern Australia in terms of vegetation and soil conditions. For this study, 29 termite mounds were first subject to in situ CH4 oxidation measurements using GPPTs (described below). For further investigations following field measurements, we selected 17 mounds of an appropriate size for processing in the laboratory (initially 18, but one was damaged during transport and had to be discarded). These mounds were first excavated but kept intact to measure internal structure, volume, densities and porosities as previously described [19]. They were then deconstructed to (1) sample termites for species identification, (2) collect mound material for physicochemical analyses and gravimetric water content measurements and (3) collect mound material for molecular analyses of methanotrophic community. For termite species identification, soldiers were individually picked and stored in pure ethanol as previously described [19]. For gravimetric water content measurements, ~200 g of mound material from both core and periphery locations were subsampled and oven-dried at 105 °C for >72 h; subsamples were measured before and after drying and the water content calculated based on mass loss. Subsamples for physicochemical parameters were oven-dried at 60 °C for 72 h, carefully homogenised into a composite sample for each termite species and location, and sent to an external laboratory for analyses according to standard protocols (CSBP Laboratory, Bibra Lake WA, Australia). For community analysis, mound and soil material was collected under sterile conditions using bleach- and heat-sterilised spatulas, and immediately stored in autoclaved 2 ml centrifuge tubes at −20 °C. For each of the 17 mounds, we collected pooled samples of mound core and periphery material derived from three different spots. Mound cores were sampled from within 20–30 cm from the approximate centroid of mound, whereas mound periphery was collected from the outer 5–10 cm of the mound. For a subset of the investigated mounds, soil was collected from beneath the mound immediately after mound excavation, and from the surrounding soil within a 1–2 m radius from the mound. This resulted in 34 mound and 14 soil samples processed for molecular analyses.
Genomic DNA extraction
Each individual sample of mound material and soil was homogenised. DNA was extracted from 0.25 to 0.5 g of each sample using the PowerLyzer PowerSoil DNA Isolation Kit (Qiagen, US), according to the manufacturer instructions. The purity and integrity of the DNA extracts was verified by spectrophotometry (NanoDrop ND-1000 spectrophotometer, Nanodrop Technologies Inc., USA) and PCR amplification of 16S rRNA genes. Good yields of high-quality, amplifiable genomic DNA were obtained from all 48 samples. PCR-grade water was extracted as a negative control.
Quantitative PCR
qPCR assays were used to estimate the abundance of the total bacterial community and methanotroph community. Methanotroph abundance was estimated by amplifying the pmoA gene using previously described degenerate primers (A189f 5′-GGNGACTGGGACTTCTGG-3′ and mb661 5′-CCGGMGCAACGTCYTTACC-3′) [20, 21] and cycling conditions [22]. These primers were chosen as a compromise to cover methanotrophs from environments with elevated CH4 concentrations and those of upland soil communities, while minimising amplification of closely related ammonium-oxidizing bacteria likely to be of high abundance in termite mounds [23]. Total bacterial abundance was estimated by amplifying the 16S rRNA gene using degenerate primers (515FB 5′-GTGYCAGCMGCCGCGGTAA-3′ and 806RB 5′-GGACTACNVGGGTWTCTAAT-3′) and cycling conditions as previously described [24,25,26]. Gene copy numbers were determined using a LightCycler 480 real-time PCR system (Roche, Basel, CH). Individual reactions contained 1 × PowerUp SYBR Green Master Mix (Thermo Fisher Scientific), 400 µM of each primer and 1 µl of diluted environmental DNA mixed to a final volume of 20 µl. Thermal profiles were adapted from those used for previous PCRs and included an acquisition step of 85 °C for 30 s at the end of each amplification cycle. Melting curve analysis was performed as follows: 95 °C for 15 s, 60 °C for 60 s, 95 °C for 30 s and 60 °C for 15 s. For each assay (96-well plate), duplicate serial dilutions of quantified DNA extract from Methylosinus trichosporium strain OB3b [27] were used for calibration curves to quantify 16S rRNA genes or pmoA genes. Each sample was analysed in triplicate, and a total of three assays were required for each gene to include all the samples. Amplification efficiencies calculated from the slopes of calibration curves were >70% and R2 values were >0.98. Using the copy numbers obtained, the total methanotroph community was estimated by assuming an average of two pmoA copies per cell. It is common for methanotrophs to possess two of the pmoA gene, and this average has been previously applied in studies investigating abundance of methanotrophs in upland soils [28, 29]. The total bacterial community was estimated by assuming an average of 4.2 16S rRNA gene copies per cell [30]. No significant amplification of the negative water extractions was observed in either the pmoA or 16S rRNA gene qPCR assays.
PmoA amplicon sequencing
The structure of the methanotroph community within each sample was primarily inferred from amplicon sequencing of community pmoA genes. Amplification reaction mixtures (25–50 µl final volume) were prepared using 1 µl of DNA extract as template, 1 × PCR buffer, 0.2 mM of each primer, 0.25 mM deoxynucleoside triphosphates and 0.025 U µl−1 of Taq polymerase (Takara Biotechnology Ltd., Japan). Different dilutions (from undiluted to 1:100 dilution in PCR-grade water) of DNA extracts were used as template during amplification, and the dilution resulting in the highest yield and quality of PCR product was used for further analyses. DNA extracts of all samples were sent to the Australian Genome Research Facility (Brisbane, QLD) for preparation of pmoA gene amplicon libraries using the above primer sets (A189f and mb661) and thermal conditions. Subsequent amplicon sequencing was performed on a MiSeq DNA-sequencing platform using a 600-cycle MiSeq Reagent Kit v3 (Illumina, San Diego, CA). Sequencing yielded 5,594,739 paired-end sequences, of which 3,078,335 passed quality checks and data processing, and were used for subsequent analyses. Sequence read counts spanned three orders of magnitude (105–102), with 54% of the samples exhibiting read counts above the average read count value (>65 K) and most mound periphery samples having read counts below 10 K. Six samples with read counts below 1000 were excluded from subsequent analyses. Sequencing data were processed according to our previously published pipeline [31], with minor modifications. Briefly, reads were 3′-trimmed to remove ambiguous or low-quality endings, then merged and primer-site trimmed. Quality filters included an amplicon-size selection (471 nt) and the removal of amplicons containing stop codons (i.e. TAA, TAG and TGA). Sequences were also checked for correct open reading frames using the FrameBot tool (http://fungene.cme.msu.edu/FunGenePipeline/framebot/form.spr). The centroid clustering method [32] identified 25 operational taxonomic units (OTUs) that shared 86% nucleotide sequence similarity [33] with sequences from a curated pmoA gene database derived from Dumont et al. [34]. Phylogenetic distances of the assigned OTUs in relation to reference pmoA sequences were assessed as previously described [31]. The phylogenetic tree of amino acid-deduced pmoA sequences was constructed in Seaview4 [35] using the maximum-likelihood method and the LG empirical amino acid substitution model, which showed the lowest Akaike information criterion (AIC) during substitution model testing, and was bootstrapped using 100 bootstrap replicates. All sequences affiliated with methanotrophs, with no clear amoA nor pxmA sequences detected.
16S rRNA gene amplicon sequencing
For each sample, the V4 hypervariable region of the 16S rRNA gene was amplified using the universal Earth Microbiome Project primer pairs F515 and R806 [24]. Samples were subject to Illumina MiSeq paired-end sequencing at the Australian Centre for Ecogenomics, University of Queensland. Using the QIIME 2 platform [36], the resultant raw sequences were subject to quality filtering, primer trimming, denoising and singleton removal. The final dataset contained 9998 high-quality 16S rRNA gene amplicon sequence variants (ASVs). For alpha diversity analysis, observed richness, Shannon index and Inverse Simpson index were analysed using the package phyloseq v1.30 [37]. For beta diversity analysis, Bray–Curtis community dissimilarity was visualised using non-parametric multidimensional scaling ordinations (nMDS) in phyloseq v1.30 [37]. To estimate the proportion of methanotrophs in the community via the 16S rRNA gene amplicon dataset, the obtained ASVs were aligned to a reference database containing 16S rRNA gene sequences from methanotrophs and related non-methanotrophic isolates and MAGs (Table S1). The V4 hypervariable region of the 16S rRNA gene was extracted from the reference sequences using QIIME 2 (extract-reads function in the feature-classifier plugin). Identical reference sequences were collapsed before alignment of V4-derived ASVs to this database with the global aligner VSEARCH v2.13.6 [38] (–id 0.90, –maxaccepts 5, –top_hits_only). The evolutionary relationships of the reference sequences and the 380 ASVs with a percentage sequence identity ≥90% were then visualised in phylogenetic trees to assess whether they most closely branch with known methanotrophs. It was found that 27 ASVs branched together with methanotrophic bacteria, all from the Methylocystaceae and Hyphomicrobiaceae families, whereas the remaining sequences were assumed to be from non-methanotrophic taxa (Table S2). A phylogenetic tree of these ASVs was constructed in Seaview4 [35] using a GTR-based model and bootstrapped with 100 replicates. The relative abundance of these ASVs was calculated compared with the rest of the 16S rRNA gene dataset. Note that this analysis cannot detect divergent uncultured methanotrophs with unknown 16S rRNA gene sequences, most notably TUSC.
Phylogenetic and diversity analyses
Alpha and beta diversity calculations, as well as read count normalisation of the pmoA sequences, were performed with the package phyloseq v1.12.2 [37] from the open source software Bioconductor. To account for differences in numbers of reads between samples, we rarefied OTU counts to an even sampling depth of 1611 read counts. Chao1, Shannon and Inverse Simpson indices were computed to assess the alpha diversity of MOB communities. Beta diversity of methanotroph communities was measured using the phylogenetic metric Unifrac weighted by the relative abundance of individual OTUs [39]. Distances between sample groups were visualised using nMDS ordinations. To determine whether the observed between-group distances were statistically significant, we performed permutational multivariate analysis of variance (PERMANOVA) with the software PRIMER-E v7 (PRIMER-E Ltd., Plymouth, United Kingdom). Negative binomial models were performed on the non-rarefied OTU dataset to assess the differential abundance of bacterial OTUs between sample groups, and the false discovery rate approach was used to account for multiple testing.
Shotgun metagenome sequencing and analysis
DNA extracts of mound cores and adjacent soils were pooled for each of the three termite species investigated (six samples in total). These samples were subject to metagenomic shotgun library preparation using the Nextera XT DNA Sample Preparation Kit (Illumina Inc., San Diego, CA, USA) at the Australian Centre for Ecogenomics. Sequencing was performed on an Illumina NextSeq500 platform with a 2 × 150 base pair High Output run. Raw metagenomic paired-end reads were inspected for quality with FastQC v0.11.7 [40] and MultiQC v1.0 [41]. BBDuk v38.51 from the BBTools suite [42] was used to trim adaptor sequences, remove reads aligning to the PhiX genome and trim low-quality bases (minimum quality score 20) and discard reads <50 bp in length. After preprocessing, an average of 91.2% of the raw read pairs per sample remained (76.6 M paired-end reads). To calculate the abundance of pmoA and mmoX (encoding for a subunit of the soluble methane monooxygenase) genes in the short read data, the pre-processed reads were aligned with the blastx function of DIAMOND v0.9.24 [43] against a manually curated database of deduced amino acid sequences for each gene (PmoA and MmoX). The employed PmoA database is the same one we used for the study of the pmoA gene amplicons and comprises sequences from methanotroph isolates, draft genomes and uncultivated sequence clusters. Forward reads at least 140 bp length were aligned with a query coverage threshold of 80% and a percentage identity threshold of 60% to PmoA and MmoX. Hits to the PxmA sequences were not included in subsequent estimates of methanotroph abundance. The abundance of each gene was converted to reads per kilobase million (RPKM). These adjusted gene abundance values were used to estimate the proportion of the community carrying each gene by normalising to the abundance for universal single-copy genes. The same reads were aligned to each of the 14 single-copy ribosomal marker genes available in SingleM (https://github.com/wwood/singlem) with DIAMOND (query coverage 80%). Alignments with a bitscore below 40 were removed, and the counts converted to RPKM and averaged across the 14 genes. The RPKM values for pmoA and mmoX were divided by the average RPKM value of the single-copy genes for each sample, and it was assumed that each methanotroph genome encoded two copies of pmoA per genome on average.
Genome assembly and analysis
Quality-filtered paired-end and singleton reads were assembled from each sample using metaSPAdes v3.13.0 [44] with k-mer sizes 43, 55, 77, 99 and 121. Contigs shorter than 1500 bp were removed from the assembly using seqmagick v0.6.1 (https://fhcrc.github.io/seqmagick/), then paired-end reads were mapped to the remaining contigs using BBMap to obtain coverage profiles. Differential coverage binning was performed for each sample using MetaBAT v2.12.1 [45], MaxBin v2.2.4 [46] and CONCOCT v1.1 [47]. All tools were run using default parameters, with the exception of MetaBAT2 where the minimum contig size was set to 1500 bp. Within each sample, bins were dereplicated using DAS_Tool v1.0 [48], resulting in a total of 136 bins. The reduced set of bins were then refined using a modified version of the VizBin protocol [49], performing a t-SNE projection which included coverage as a feature and weighted it to account for 50% of the distance calculation. Projections were performed for each sample and contigs with uncertain placement were manually removed from bins. The refined set of bins were combined into a single collection, assessed for completeness and contamination using CheckM v1.0.13 [50], then dereplicated using dRep v2.3.2 [51]. A total of 55 bins, with at least 50% completeness, were retained after this process. Taxonomy was assigned to all bins using the GTDB-TK v0.2.2 software [52]. Inspection of the classification summary revealed a single bin (MAG-47) classified as a member of the proteobacterial genus Methylocapsa (Alphaproteobacteria; Rhizobiales; Beijerinckiaceae), which was explored in detail. A robust phylogeny was inferred for this bin by extracting the GTDB-TK alignment for 135 bacterial genomes from the family Beijerinckiaceae. This selection was augmented with the MAG USCα AHI (VDMG00000000.1) [53] and confirmed to include the species M. acidiphila B2 (GCF_000427445.1), which was used in the original characterisation of the USCα pmo operon [54]. A selection of 20 bacterial species was randomly sampled from families within the order Rhizobiales, excluding Beijerinckiaceae, for use as an outgroup. Phylogenetic inference was performed using IQ-Tree v1.6.12 [55] with the Le and Gascuel model of amino acid evolution [56] with empirically determined amino acid frequencies and Gamma-distributed rate heterogeneity (LG + F + G). Bootstrap support was calculated using 100 resamplings, and the tree visualised using iToL v4 [57]. For metabolic pathway reconstruction of MAG-47, the genome was firstly annotated using MetaErg v1.2.2 [58]. The predicted amino sequences were then used as query for identification of key metabolism markers via METABOLIC v2.0 [59].
Gas push–pull tests
The GPPT was used to estimate in situ activity coefficients as described previously [7, 60]. Michaelis–Menten parameters estimated from in situ methods are integrated measures across a large mass of substrate and are thus better suited to characterise the kinetic potential of whole microbial communities in heterogeneous systems than laboratory microcosms, which suffer from inevitable sampling bias [61]. In brief, a gas mixture containing laboratory air, ~900 µl l−1 of CH4, and ~0.1 l l−1 argon (Ar) was injected at a rate of ~0.5 l min−1 into the lower centre of the termite mounds and then immediately extracted from the same location at the same flow rate. During extraction, the injected gas mixture was gradually diluted with termite mound air down to background levels; the tracer Ar accounted for this dilution due to its similar transport behaviour to CH4. A timeseries of CH4 and Ar concentrations was collected during the 24 min injection phase, and the 36 min extraction phase. Concentrations of CH4 were measured quasi-continuously (frequency of 1 Hz) using a field-portable spectrometer (Fast Greenhouse Gas Analyser Los Gatos Research, Mountain View, CA). For Ar, discrete samples were collected at fixed intervals during injection (n = 3) and extraction (n = 10–12), as well as prior to injection to determine background levels. Argon concentrations were analysed on a customised gas chromatography system (SRI 8610, SRI Instruments, Torrance, CA) with an external thermal conductivity detector (VICI Valco Instruments Co., Houston, TX). To improve separation of Ar, oxygen was removed from the sample gas stream prior to separation with a manually packed Pd-Al catalyst column [7, 62].
Kinetic analysis
First-order rate coefficients of CH4 oxidation (activity coefficient k) were estimated from GPPTs via the slope of the logarithm of relative CH4 vs Ar concentrations, plotted against a transformed reaction time, according to the plug-flow reactor model for simplified GPPT analysis [63]. These data also allowed the calculation of reaction rates at different CH4 concentrations from segments of the extraction timeseries, and thus the estimation of apparent Michaelis–Menten parameters [64]. The running average of CH4 concentrations (CCH4) over three consecutive extraction samples was multiplied with the corresponding activity k from the logarithmic plot to calculate an individual reaction rate (Rox) for each segment. The apparent Michaelis–Menten parameters (Km and Vmax) were then estimated for each individual GPPT, and for all combined pairs of concentrations and reaction rates, by fitting the Michaelis–Menten model Rox = Vmax × CCH4/(Km + CCH4) to the data using the non-linear regression routine nls() in R [65]. The AIC was calculated and compared with a linear regression model of Rox vs CCH4; if the AIC of the linear model was lower, no Michaelis–Menten parameters were reported. Cell-specific reaction rates were calculated from overall GPPT reaction rates based on total mound dry mass [7], divided by pmoA copy numbers and assuming two gene copies of pmoA per cell. Correlations between kinetic parameters (k, Km, Vmax), gene abundance (pmoA and 16S rRNA genes), and physical mound parameters (mound micro- and macro-porosity, total volume and water content) were tested for significance using linear regression (Table S3), after transformations (sqrt for kinetic parameters, log for gene abundances) and removal of outliers as indicated by diagnostic plots (qq- and Cook’s distance). Stepwise parameter elimination was performed using the step() function in R [65].
Results and discussion
Methanotrophic bacteria are in low abundance in termite mounds and associated soils
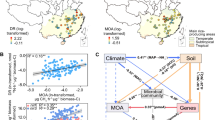
qPCR was used to estimate the abundance of the methanotroph community (via pmoA gene) and total bacterial community (via 16S rRNA gene) in mound and soil samples. Bacterial abundance was consistently high (av. 6.4 × 109 cells per gram of dry soil; range 6.0 × 107–8.2 × 1010) and did not significantly differ between sample locations (Figs. 1a, d and S1); an earlier study found higher microbial biomass in the mound compared with soil [66], but this may reflect different methodologies applied to each substrate. In contrast, methanotroph cell abundance was relatively low across the samples (av. 1.5 × 106 copies per gram of dry sample material; range: 1.0 × 104–9.0 × 106) and just 0.016% that of total bacterial cells (range: 0.0004–0.10%) (Figs. 1b, e and S1). Such values are comparable to those previously reported for the abundance of pmoA genes in upland soils that mediate atmospheric CH4 oxidation (~106 copy number, ~0.01% relative abundance [29]). However, while pmoA-based methods are a standard method for inferring methanotroph abundance and enable comparisons with previous studies [29, 31, 67], they can potentially underestimate the CH4-oxidizing community given pmoA primer mismatches can result in low amplification of certain methanotroph groups such as USCα [18].
Samples are grouped according to termite species (a–c) and sampling location (d–f). a, d show total bacterial cells estimated from 16S rRNA gene qPCR, assuming on average 4.2 16S rRNA gene copies per cell. b, e show methanotroph cells estimated from pmoA qPCR, assuming 2 pmoA copies per cell. c, f show methanotroph abundance estimated from 16S rRNA gene qPCR, using the ratio of identified methanotrophs to total 16S rRNA gene copies. Different letters denote significant differences in abundance between sample groups (p < 0.05, Kruskal–Wallis and Dunn test). Mn Microcerotermes nervosus mounds, Ms Macrognathotermes sunteri mounds, Tp Tumulitermes pastinator mounds, core termite mound core, periphery termite mound periphery, beneath soil beneath mound, surrounding soil surrounding mound, MOB methane-oxidising bacteria.
Two other methods were therefore used to predict the abundance of the methanotroph community. We used 16S rRNA gene amplicon sequencing, in conjunction with phylogenetic analysis, to identify CH4-oxidising taxa within the context of the total bacterial and archaeal community. Overall, the bacterial community of termite mound samples had lower richness and distinct composition to soil samples, including an increased relative abundance of Actinobacteria and decreased relative abundance of Gammaproteobacteria across the mounds of all three termite species sampled (Fig. S2; Table S4). On average, 0.24% of the community (range: 0.004–2.37%) affiliated with known methanotrophs, all within the class Alphaproteobacteria (Figs. 1c, f and S1; Table S4). In parallel, we performed shotgun metagenome sequencing and used homology-based searches of quality-filtered short reads to estimate the abundance of genes encoding particulate (pmoA) and soluble (mmoX) methane monooxygenases relative to single-copy ribosomal marker genes; on average, 0.21% of the community (range: 0.13–0.53%) are predicted to encode methane monooxygenases, with 82% of the hits to the particulate type (Table S2). While the results of 16S rRNA gene and metagenome analyses are concordant, the values obtained are an order of magnitude higher than estimates based on pmoA qPCR data. It is unclear which results are most reliable given all three methods have intrinsic detection biases that can lead to the underestimation and overestimation of particular methanotroph groups. However, regardless of the method used to infer their relative abundance, methanotrophs appear to be members of the rare biosphere in mounds and soils alike despite the elevated CH4 concentrations available in the mound environment.
Some differences in methanotroph abundance were observed between sample locations and termite species (Table S3). Based on pmoA qPCR, methanotroph abundance was 3.5-fold higher in mound core and 1.5-fold higher in soil beneath than in surrounding soil, though differences were below the threshold of significance (Fig. 1e). In contrast, methanotroph abundance was significantly lower in mound periphery samples of all species (p < 0.01; Kruskal–Wallis and Dunn test; Fig. 1e) and in mound samples of T. pastinator compared with the other two species tested (p < 0.01; Kruskal–Wallis and Dunn test; Fig. 1b); the latter observation is in line with the finding that CH4 oxidation occurs at low rates in T. pastinator mound material [7]. Similar but weaker trends were also observed based on estimating methanotroph abundance via 16S rRNA gene amplicon sequencing (Fig. 1c, f). We performed linear regression on how environmental parameters correlated with methanotroph abundance, including stepwise elimination of parameters (Table S3). Micro-porosity and sampling location were the only significant parameters remaining (p < 0.001 for location and micro-porosity, p = 0.08 for water content). With the exception of macro-porosity in periphery samples, other physical parameters did not correlate significantly with methanotroph or total bacterial abundance (Table S3). This suggests that denser mound material, as found in mound periphery and T. pastinator mounds, limits methanotroph abundance and activity [7, 19]. These differences may also reflect the relatively harsh conditions in the mound periphery, with its strong fluctuations of temperature and water content, compared with the core with termite-engineered homoeostasis [68, 69].
Termite mounds contain compositionally distinct methanotroph communities that are derived from surrounding soils
We analysed the diversity and composition of the methanotroph community. To do so, we focused on pmoA amplicon sequencing given this method enables higher-resolution methanotrophic community analysis, but verified findings using 16S rRNA gene amplicon and metagenomic sequencing. Across the samples, 25 OTUs were detected (Fig. 2). Observed and estimated richness of these OTUs was higher in soil samples compared with mound samples (p < 0.001; Wilcoxon rank-sum test), and mound core compared with periphery (p < 0.001; Wilcoxon rank-sum test) (Fig. 3a; Table S3); however, these differences were driven primarily by rare OTUs in soil samples, with Shannon and inverse Simpson indices were similar between sample locations (Fig. S3; Table S3). Beta diversity of the samples was analysed by weighted Unifrac and visualised on an nMDS ordination plot (Fig. 3b). PERMANOVA analysis confirmed communities significantly differed between sample locations (p = 0.001); communities in mound core and periphery samples were similar and compositionally distinct from soil communities, and methanotroph communities in soils beneath mounds were more similar to mounds than surrounding soils (Fig. 3b, c). In addition, methanotroph communities differed significantly between termite species (p = 0.022), while soil samples did not. Specifically, mound communities differed between Ms and Mn in both mound core (p = 0.005) and periphery (p = 0.032), and between Ms and Tp for mound core (p = 0.015).
The 25 OTUs are depicted in bold and numbered according to decreasing relative abundance among all samples. The tree was built using the LG empirical amino acid substitution model and bootstrapped using 100 bootstrap replicates. Node numbers indicate bootstrap branch support ≥60. Genbank accession numbers for the sequences at individual node tips are given in parentheses. The scale bar displays 0.2 changes per amino acid position.
a Estimated richness of the methanotroph community, based on Chao1 index of the pmoA gene as determined by amplicon sequencing. b Non-metric multidimensional scaling (nMDS) ordination of the methanotrophic community structure (beta diversity) measured by weighted UniFrac distance metric of the pmoA gene, and differentiation according to termite species and sample location. c Heatmap of the relative abundance of the pmoA OTUs in all samples. OTUs are ordered according to their position on the phylogenetic tree shown in Fig. 2. Differential abundance of pmoA OTUs between sample groups was assessed from negative binomial models of the OTU read counts; p values were corrected with the false discovery rate approach to account for multiple testing. Only significant tests are shown (p < 0.05).
The 25 OTUs detected were visualised on a phylogenetic tree (Fig. 2) against reference sequences from a curated pmoA gene database [34]. Across the mound and soil samples, over 80% of the sequences were affiliated with USCα, a recently cultivated lineage of alphaproteobacterial methanotrophs known to mediate atmospheric CH4 oxidation [70, 71] (Figs. 2 and 3c). The dominance of USCα was confirmed by less-resolved 16S rRNA gene and metagenome-based analyses, where they were estimated to comprise an average of 74% and 88% of the alphaproteobacterial methanotroph community, respectively (Fig. S4; Table S2); in both analyses, most sequences detected were closely related to those encoded by two USCα bacteria, Candidatus Methyloaffinis lahnbergensis [71] and Methylocapsa gorgona MG08 [70]. Based on pmoA amplicon sequencing and metagenome sequencing, the second most dominant taxonomic groups were affiliated with the alphaproteobacterial lineage Methylocystis in mound samples (<10% relative abundance) and the uncultivated tropical upland soil cluster (TUSC) [18] in soil samples (<10% relative abundance) (Fig. 2; Table S2). There was a large proportion of shared taxa across the samples, with the three most abundant OTUs (USCα-affiliated) present in all samples, regardless of type (mound vs soil), location and termite species (Fig. 3c). However, differential abundance analysis supported the differences between sample type and termite species observed by Unifrac analysis (Fig. 3b, c). USCα OTUs were more abundant in mound core, mound periphery and soils beneath, whereas TUSC OTUs were more abundant in surrounding soils. Significant differential abundance was also observed for certain OTUs between mounds of different termite species (Fig. 3b, c). Overall, these findings suggest that termite mounds contain methanotroph communities that are primarily derived from soil, but have a distinct composition overall.
Methanotroph communities are kinetically adapted to elevated CH4 concentrations
We determined the kinetics of CH4 oxidation in 29 mounds from all three species by performing in situ GPPTs. Methane oxidation rate was high across the investigated mounds (Fig. 4a). The relationship between CH4 concentration and reaction rate best fitted a Michaelis–Menten model for 18 mounds and a linear model for 11 mounds based on AIC values (Fig. 4a). For the former group of mounds, apparent Michaelis–Menten coefficients (Km, Vmax) were calculated. Estimated Km values for the 18 mounds ranged from 0.32 to 47 µmol (l air)−1, and Vmax from 8.4 to 280 µmol (l air)−1 h−1. These parameters did not significantly differ between mounds of different termite species. The overall mean values for Km and Vmax were 17.5 ± 3.5 µmol (l air)−1 and 78.3 ± 17 µmol (l air)−1 h−1, respectively (standard error of the mean); such values were close to the optimal parameters when fitting a Michaelis–Menten model to combined GPPT data (excluding mounds with linear behaviour): Km = 13.2 ± 3.5 µmol (l air)−1 and Vmax = 55.4 ± 8.5 µmol (l air)−1 h−1 (Fig. 4a, b). Thus, the methanotroph communities within termite mounds have an apparent medium (µM) affinity for CH4. The apparent Km is approximately one to two orders of magnitude higher than high-affinity (nM) uptake observed in upland soils [72,73,74], but one to two orders of magnitude lower than the low-affinity (mM) uptake measured in landfill cover soils [75]. Similar Michaelis–Menten values were estimated from GPPTs in the vadose zone above a contaminated aquifer, which featured CH4 concentrations in a similar range to termite mounds (~1–40 μl l−1) [64].
a Optimal Michaelis–Menten (M–M) curve estimated for a combined dataset of in situ concentrations and reaction rates from 18 out of 29 mounds showing M–M behaviour, excluding 11 mounds with an apparent linear increase of rates based on AIC. b Individual M–M parameters of the 18 mounds showing M–M behaviour, including their mean and standard error, and the optimal M–M parameters of the combined dataset. c Positive correlation of activity coefficients k with pmoA gene abundance in mound core samples of the 17 mounds selected for detailed analysis (R2 = 0.44, p = 0.0069). Two outliers have been removed, indicated with red X. d Cell-specific reaction rates of CH4 oxidation, estimated from in situ oxidation rates in termite mounds divided by methanotroph abundance at the different sampling locations. For each mound, the same oxidation rate was divided by four different abundances for comparison. The dashed horizontal lines indicate a range of literature values estimated in upland soils [29].
It is noteworthy that 11 mounds showed an apparent linear increase of reaction rates with substrate concentrations (Fig. 4a). This could indicate that Vmax has not been reached during GPPTs with a maximum injected CH4 concentration of ~900 µl l−1 (~40 µM); indeed, the injection concentration is in the range of our highest Km, thus the capacity of some mounds to oxidise CH4 can be substantially higher. A linear increase could also indicate a shift in kinetics during the course of the GPPT. It is in the nature of the GPPT that different areas of the mound are exposed to different concentration ranges, depending on their distance to the gas injection/extraction point [63]. Hence, gas extracted at different times may have had a “history” of exposure to methanotroph communities with different kinetics. It is even conceivable that the 1 h long exposure to high injection concentrations around the injection/extraction point triggered the upregulation of low-affinity methane monooxygenase isozymes, such as those reported in Methylocystis sp. SC2 [76].
For the 17 selected mounds investigated in more detail, we analysed how the kinetics and abundance of the methanotroph communities were correlated. While a range of positive correlations were observed (Table S3), the most robustly supported was the relationship between the first-order activity coefficient k and mound core methanotroph abundance (R2 = 0.40, p = 0.007) (Fig. 4c). This relationship also held true for total bacterial abundance (R2 = 0.36, p = 0.007). In contrast, correlations with methanotroph abundance in periphery and soils were not significant. These observations are in line with previous inferences that mound cores are primarily responsible for CH4 oxidation in M. nervosus and M. sunteri mounds [7]. On this basis, hypothetical cell-specific CH4 oxidation rates were calculated from in situ GPPT rates integrated across the whole mound, and divided by location-specific methanotroph abundances for comparing different mound and soil compartments. Mound core values ranged from 1.8 × 10−17 to 1.3 × 10−14 mol CH4 cell−1 h−1 (Fig. 4d), with the median of 1.3 × 10−15 mol CH4 cell−1 h−1 slightly higher than the observed range of 5–8 × 10−16 mol CH4 cell−1 h−1 in upland soils calculated by the same method [29]. This discrepancy was mainly driven by high cell-specific rates of T. pastinator, and the median dropped to 8.7 × 10−16 mol CH4 cell−1 h−1 when excluding this species. Also, cell-specific rates of periphery samples were unreasonably high for all species and particularly for T. pastinator (Fig. 4d), with some values close to landfill cover biofilters [77]. These high cell-specific rates in periphery (where there are significantly fewer methanotrophs) corroborate our findings of mound core methanotrophs being responsible of the bulk of the CH4 oxidation in mounds of M. nervosus and M. sunteri [7]. In addition, estimated rates for soils, particularly soil beneath mounds, were well aligned with upland soil literature values for all three species [29], and further emphasises that CH4 oxidation in T. pastinator mounds is driven by methanotrophs in the soil beneath.
Termite mound USCα bacteria are predicted to be mixotrophic or facultative methanotrophs
Finally, we performed metagenomic assembly and binning to gain a genome-resolved view of the taxonomic affiliation and metabolic capabilities of termite mound methanotrophs. We retrieved a medium-quality [78] metagenome-assembled genome (MAG-47; completeness 73%, contamination 1.7%) from pooled DNA extracted from M. sunteri mound cores. A genome tree confirmed this bacterium is a USCα alphaproteobacterium closely related to Candidatus Methyloaffinis lahnbergensis [71] and an Antarctic MAG [79] predicted to oxidise atmospheric CH4 (Fig. S5). This supports our conclusions from analysis of pmoA amplicons (Figs. 2 and 3), 16S rRNA gene amplicons (Fig. S4), and metagenomic short reads (Table S2) that USCα dominates the methanotrophic community of North Australian termite mounds. We inferred metabolic capabilities of the termite mound USCα based on the retrieved genome (Table S5). As depicted in Fig. 5, the bacterium encodes a particulate methane monooxygenase via a complete pmoCAB operon. The bacterium also has the capacity to assimilate carbon derived from CH4 through the H4MPT pathway and serine cycle, and conserve energy through aerobic respiration using two terminal oxidases. These core metabolic features are conserved with other recently sequenced USCα bacteria [70, 71].
H4MPT tetrahydromethanopterin pathway, pMMO particulate methane monooxygenase, MDH methanol dehydrogenase, FDH formate dehydrogenase, NiR copper-containing nitrite reductase, I NADH dehydrogenase (complex I), II succinate dehydrogenase (complex II), IV cytochrome aa3 oxidase and cytochrome bo3 oxidase (complex IV). Genes detected are shown in italics. Dashed black lines indicate diffusion. Dashed grey lines indicate reactions performed by three enzymes that were not detected, methanol dehydrogenase, hydroxylamine dehydrogenase and serine hydroxymethyltransferase. It is assumed that these enzymes are encoded by the bacterium, but were not detected given the moderate predicted completeness of the MAG (73%).
It is likely that termite mounds are inhabited by mixotrophic or facultative methanotrophs. Consistently, the reconstructed USCα genome encoded enzymes to assimilate acetate into biomass (acetate kinase, phosphotransacetylase); previous studies have experimentally shown that USCα can assimilate acetate together with atmospheric CH4 [71, 80]. However, in contrast to a recent study that suggested M. gorgona can consume atmospheric H2 and CO [70], we did not detect hydrogenase or carbon monoxide dehydrogenase genes in the MAG; this may reflect niche differentiation between USCα methanotrophs or alternatively the incompleteness of the MAG. Nevertheless, homology-based searches of metagenome short reads of mound cores showed 2.8% of HhyL reads (group 1 h [NiFe]-hydrogenase) [81] and 0.03% of CoxL reads (type I carbon monoxide dehydrogenase) [82] were most closely related to those in alphaproteobacterial methanotroph genomes. This suggests that at least some methanotrophs in termite mounds consume atmospheric H2 and CO, in line with recent studies showing mixotrophic trace gas metabolism enables niche expansion of methanotrophs [70, 83, 84]. Finally, we detected a copper-containing nitrite reductase (NirK) in the MAG. It has recently been shown that this enzyme enables rapid detoxification of metabolites formed by the promiscuous oxidation of ammonia by particulate methane monooxygenase [85]. This process may be particularly important in termite mounds, given ammonia accumulates at high levels due to excretion by termites [23]; ammonia levels were particularly high in M. sunteri mounds from which the MAG was assembled (220 mg kg−1; Table S6).
Conclusions and perspectives
Overall, our results imply that local environmental concentrations of CH4 shape the composition and kinetics of the methanotroph community in termite mounds. Elevated CH4 production from termites appears to have selected for a specialised methanotroph community within mounds with medium substrate affinity. The community analysis shows that the methanotroph communities within termite mounds are closely related to those of adjoining soils. However, termite activity has facilitated selection for USCα and Methylocystis OTUs, together with exclusion of TUSC OTUs, in mounds compared with adjoining soil. This selection is likely to be primarily driven by elevated CH4 availability, but other effects of mound formation and termite activity (e.g. on nutrient levels, moisture levels, temperature and vegetation cover) may also influence methanotroph community assembly. Concordant findings were made across three different species, with methanotroph communities showing high cell-specific reaction rates and medium affinities for CH4. They are thus ideally kinetically adapted to grow on termite-derived CH4 and in turn reduce atmospheric emissions of this greenhouse gas. These findings are also supported by the reconstruction of a metagenome-assembled genome of a USCα bacterium, confirming the capacity of mound methanotrophs to use CH4 as a carbon and energy source, together with acetate and potentially other substrates.
However, the apparent kinetic parameters of CH4 oxidation in termite mounds are atypical of the dominant groups of methanotrophs present. It is probable that USCα mediates most of the overall CH4 oxidation in the investigated mounds, given they were observed to be the most abundant and prevalent mound and soil methanotrophs across three separate analyses. This inference is also reflected by consistent cell-specific rates of CH4 oxidation in mound core and soil. However, USCα are typically high-affinity methanotrophs associated with the oxidation of CH4 at atmospheric concentrations in upland soils [70, 71]. There are several possible explanations for this apparent discrepancy. Selective pressures may have caused mound USCα to modulate their apparent CH4 affinity, for example through amino acid substitutions or gene expression changes of their particulate methane monooxygenases; the first explanation is unlikely given soil and mound USCα encode common PmoA subunits, though substitutions in PmoB or PmoC cannot be ruled out. An alternative explanation is community heterogeneity. Any apparent Michaelis–Menten parameter estimates from environmental samples are “bulk” values integrating across the whole microbial community and thus a vast array of potentially different enzymes. Indeed, the observed in situ kinetics are also compatible with the coexistence of slow-acting, high-affinity USCα methanotrophs alongside faster-acting, low-affinity other groups. Methylocystis-like OTUs, as the second most abundant group, are likely to be particularly competitive at higher CH4 concentrations during phases of high termite activity. Members of this group encode kinetically distinct methane monooxygenase isozymes [76] and have been identified in other soils with elevated CH4 concentrations [22]. They would therefore have a competitive advantage in termite mounds where CH4 availability is elevated and exhibits considerable temporal and spatial variation. Lastly, while CH4 concentrations in mounds are elevated compared with the atmosphere (2–100 ppmv) [7, 86], they are well within the accessible range for high-affinity methanotrophs such as USCα.
Despite elevated substrate availability, methanotrophs remain minor members of microbial communities in termite mounds at our sampling sites. We observed that methanotrophs comprised <0.5% of the bacterial community across three separate assays, although the exact proportion varied depending on the method used. While methanotrophs are moderately enriched in mound core compared to adjoining soil overall, they are actually diminished relative to soil in both the mound core of T. pastinator and the periphery of all three species. The observed strong correlation of methanotroph abundance with the porosity of the mound material for all species strengthens the case for habitat porosity as a crucial factor in regulating the methanotrophic community, likely by regulating local CH4 availability through diffusion. Another factor driving low methanotroph abundance could be the accumulation of ammonia, which is produced in high levels by termites [23]; it is known that ammonia competitively inhibits methane monooxygenase activity [87, 88] and ammonia levels are major environmental factors regulating methanotroph community in soils [89, 90]. However, the high ammonia concentrations in mounds of M. sunteri (Table S6) did not correlate with interspecies differences in methanotroph abundance, yet aligned well with differences in community composition (Fig. 3). High ammonia levels may potentially drive adaptation of the methanotroph community, rather than diminishing abundance, as supported by the potential mechanism to detoxify ammonia-derived intermediates in the USCα MAG found in M. sunteri mounds. Putative ammonia-oxidising bacteria and archaea were also detected in the 16S rRNA gene datasets, but like methanotrophs, were in relatively low abundance (<0.5%) despite substrate availability and hence are unlikely to mediate significant rates of nonspecific CH4 oxidation. Other possible factors limiting methanotrophs in termite mounds, such as the micronutrients (e.g. copper) required for methane monooxygenase activity, remain to be investigated.
It should be noted that it is unclear to what extent these observations are extendable to mounds from other regions or species. Broadly, our findings agree with a previous study that inferred mounds of the African fungus-growing termite Macrotermes falciger have a distinct methanotroph community from surrounding soil, likely in part due to selective pressures exerted by termite CH4 emissions [14]. However, given methanotroph communities are derived from surrounding soil, the specific communities mitigating CH4 uptake are likely to greatly vary between different soil types. Indeed, we observed a striking difference in community composition of the mounds from the three Australian termite species compared with those of the one African termite species [14]. The African mounds were dominated by the JR3 cluster, a gammaproteobacterial methanotroph lineage closely related to USCγ, which was detected in very low abundance in the Australian mounds (Fig. 2; Table S2). These differences likely reflect the distinct habitat specificity of USCγ and USCα methanotrophs. USCα often occurs in acidic to neutral upland soils ([67], reviewed in [18]), which match the pH values between 5 and 6 measured in the mounds of this study (Table S3). In contrast, USCγ and associated lineages are commonly found in upland soils of neutral to basic pH [91, 92], which corresponds well to pH values of 7–8 in Macrotermes falciger mounds [14]. Global sampling would be required to more broadly determine how soil and termite characteristics influence the methanotroph community and CH4 oxidation kinetics of termite mounds. Our findings are nevertheless likely to extend to many Australian mounds, given the sampling site is broadly representative of much of tropical northern Australia [7, 93] and the three termite species selected represent the three major feeding groups on the continent. Overall, we conclude that methanotrophs remain minor members of mounds at our sampling sites and hence are only able to mitigate a proportion of the large amounts of CH4 produced by mound-dwelling termites.
Data availability
The sequence dataset generated and/or analysed during the current study are publicly available under NCBI BioProject accession number PRJNA641804. All metadata is provided in the supplemental material.
References
Brune A. Methanogenesis in the digestive tracts of insects. In: Timmis KN (ed). Handbook of hydrocarbon and lipid microbiology. Berlin, Heidelberg: Springer-Verlag; 2010. pp. 707–728.
Zimmerman PR, Greenberg JP, Wandiga SO, Crutzen PJ. Termites: a potentially large source of atmospheric methane, carbon dioxide, and molecular hydrogen. Science. 1982;218:563–5.
Rasmussen RA, Khalil MAK. Global production of methane by termites. Nature. 1983;301:700.
Sugimoto A, Inoue T, Tayasu I, Miller L, Takeichi S, Abe T. Methane and hydrogen production in a termite-symbiont system. Ecol Res. 1998;13:241–57.
Brauman A, Kane MD, Labat M, Breznak JA. Genesis of acetate and methane by gut bacteria of nutritionally diverse termites. Science. 1992;257:1384–7.
Kirschke S, Bousquet P, Ciais P, Saunois M, Canadell JG, Dlugokencky EJ, et al. Three decades of global methane sources and sinks. Nat Geosci. 2013;6:813–23.
Nauer PA, Hutley LB, Arndt SK. Termite mounds mitigate half of termite methane emissions. Proc Natl Acad Sci USA. 2018;115:13306–11.
Hanson RS, Hanson TE. Methanotrophic bacteria. Microbiol Rev. 1996;60:439–71.
Reuß J, Rachel R, Kämpfer P, Rabenstein A, Küver J, Dröge S, et al. Isolation of methanotrophic bacteria from termite gut. Microbiol Res. 2015;179:29–37.
Pester M, Tholen A, Friedrich MW, Brune A. Methane oxidation in termite hindguts: absence of evidence and evidence of absence. Appl Environ Microbiol. 2007;73:2024–8.
Dunfield PF. The soil methane sink. In: Reay D, Hewitt K, Smith K, Grace J, editors. Greenhouse gas sinks. Wallingford: CABI; 2007. pp. 152–70.
Bignell DE, Eggleton P, Nunes L, Thomas KL. Termites as mediators of forest carbon fluxes in tropical forests: budgets for carbon dioxide and methane emissions. In: Watt AD, Stork NE, Hunter MD (eds). Forests and insects. London: Chapman & Hall; 1997. pp. 109–134.
Jamali H, Livesley SJ, Grover SP, Dawes TZ, Hutley LB, Cook GD, et al. The importance of termites to the CH4 balance of a tropical savanna woodland of northern Australia. Ecosystems. 2011;14:698–709.
Ho A, Erens H, Mujinya BB, Boeckx P, Baert G, Schneider B, et al. Termites facilitate methane oxidation and shape the methanotrophic community. Appl Environ Microbiol. 2013;79:7234–40.
Noirot C, Darlington JPEC. Termite nests: architecture, regulation and defence. In: Abe T, Bignell DE, Higashi M (eds). Termites: Evolution, Sociality Symbioses, Ecology. Dordrecht: Springer; 2000. pp. 121–139.
Korb J. Termite mound architecture, from function to construction. In: Bignell DE, Roisin Y, Lo N (eds). Biology of termites: a modern synthesis. Dordrecht: Springer Netherlands; 2010. pp. 349–373.
Jones DT, Eggleton P. Global biogeography of termites: a compilation of sources. In: Bignell DE, Roisin Y, Lo N, editors. Biology of termites: a modern synthesis. Dordrecht: Springer; 2011. pp. 1–576.
Knief C. Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front Microbiol. 2015;6:1346.
Nauer PA, Chiri E, Souza D, de, Hutley LB, Arndt SK. Rapid image-based field methods improve the quantification of termite mound structures and greenhouse-gas fluxes. Biogeosciences. 2018;15:3731–42.
Holmes AJ, Costello A, Lidstrom ME, Murrell JC. Evidence that participate methane monooxygenase and ammonia monooxygenase may be evolutionarily related. FEMS Microbiol Lett. 1995;132:203–8.
Costello AM, Lidstrom ME. Molecular characterization of functional and phylogenetic genes from natural populations of methanotrophs in lake sediments. Appl Environ Microbiol. 1999;65:5066–74.
Henneberger R, Chiri E, Bodelier PEL, Frenzel P, Lüke C, Schroth MH. Field‐scale tracking of active methane‐oxidizing communities in a landfill cover soil reveals spatial and seasonal variability. Environ Microbiol. 2015;17:1721–37.
Ji R, Brune A. Nitrogen mineralization, ammonia accumulation, and emission of gaseous NH3 by soil-feeding termites. Biogeochemistry. 2006;78:267–83.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA. 2011;108:4516–22.
Parada AE, Needham DM, Fuhrman JA. Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol. 2016;18:1403–14.
Apprill A, McNally S, Parsons R, Weber L. Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Micro Ecol. 2015;75:129–37.
Heil JR, Lynch MDJ, Cheng J, Matysiakiewicz O, D’Alessio M, Charles TC. The completed PacBio single-molecule real-time sequence of Methylosinus trichosporium strain OB3b reveals the presence of a third large plasmid. Genome Announc. 2017;5:e01349–17.
Kolb S, Knief C, Stubner S, Conrad R. Quantitative detection of methanotrophs in soil by novel pmoA-targeted real-time PCR assays. Appl Environ Microbiol. 2003;69:2423–9.
Kolb S, Knief C, Dunfield PF, Conrad R. Abundance and activity of uncultured methanotrophic bacteria involved in the consumption of atmospheric methane in two forest soils. Environ Microbiol. 2005;7:1150–61.
Větrovský T, Baldrian P. The variability of the 16S rRNA gene in bacterial genomes and its consequences for bacterial community analyses. PLoS ONE. 2013;8:e57923.
Chiri E, Nauer PA, Rainer E-M, Zeyer J, Schroth MH. High temporal and spatial variability of atmospheric-methane oxidation in Alpine glacier-forefield soils. Appl Environ Microbiol. 2017;83:e01139–17.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
Wen X, Yang S, Liebner S. Evaluation and update of cutoff values for methanotrophic pmoA gene sequences. Arch Microbiol. 2016;198:629–36.
Dumont MG, Lüke C, Deng Y, Frenzel P. Classification of pmoA amplicon pyrosequences using BLAST and the lowest common ancestor method in MEGAN. Front Microbiol. 2014;5:34.
Gouy M, Guindon S, Gascuel O. SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol. 2010;27:221–4.
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol. 2019;37:852–7.
McMurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE. 2013;8:e61217.
Rognes T, Flouri T, Nichols B, Quince C, Mahé F. VSEARCH: a versatile open source tool for metagenomics. PeerJ. 2016;4:2584.
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R. UniFrac: an effective distance metric for microbial community comparison. ISME J. 2011;5:169.
Andrews S. FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics, Cambridge, UK: The Babraham Institute; 2010.
Ewels P, Magnusson M, Lundin S, Käller M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics. 2016;32:3047–8.
Bushnell B. BBMap: A fast, accurate, splice-aware aligner. Berkeley, CA, US: Lawrence Berkeley National Lab (LBNL); 2015.
Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND. Nat Methods. 2014;12:59.
Nurk S, Meleshko D, Korobeynikov A, Pevzner PA. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 2017;27:824–34.
Kang D, Li F, Kirton ES, Thomas A, Egan RS, An H, et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ. 2019;7:e7359.
Wu Y-W, Tang Y-H, Tringe SG, Simmons BA, Singer SW. MaxBin: an automated binning method to recover individual genomes from metagenomes using an expectation-maximization algorithm. Microbiome. 2014;2:26.
Alneberg J, Bjarnason BS, De Bruijn I, Schirmer M, Quick J, Ijaz UZ, et al. Binning metagenomic contigs by coverage and composition. Nat Methods. 2014;11:1144.
Sieber CMK, Probst AJ, Sharrar A, Thomas BC, Hess M, Tringe SG, et al. Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. Nat Microbiol. 2018;3:836–43.
Laczny CC, Sternal T, Plugaru V, Gawron P, Atashpendar A, Margossian HH, et al. VizBin—an application for reference-independent visualization and human-augmented binning of metagenomic data. Microbiome. 2015;3:1–7.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043–55.
Olm MR, Brown CT, Brooks B, Banfield JF. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 2017;11:2864.
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil P-A, et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36:996–1004.
Rusley C, Onstott TC, Vishnivetskaya TA, Layton A, Chauhan A, Pfiffner SM, et al. Metagenome-assembled genome of USCα AHI, a potential high-affinity methanotroph from Axel Heiberg Island, Canadian High Arctic. Microbiol Resour Announc. 2019;8:1–4.
Ricke P, Kube M, Nakagawa S, Erkel C, Reinhardt R, Liesack W. First genome data from uncultured upland soil cluster alpha methanotrophs provide further evidence for a close phylogenetic relationship to Methylocapsa acidiphila. Appl Environ Microbiol. 2005;71:7472–82.
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 2014;32:268–74.
Le SQ, Gascuel O. An improved general amino acid replacement matrix. Mol Biol Evol. 2008;25:1307–20.
Letunic I, Bork P. Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 2019;47:W256–9.
Dong X, Strous M. An integrated pipeline for annotation and visualization of metagenomic contigs. Front Genet. 2019;10:1–10.
Zhou Z, Tran P, Liu Y, Kieft K, Anantharaman K. METABOLIC: a scalable high-throughput metabolic and biogeochemical functional trait profiler based on microbial genomes. bioRxiv. 2019:761643.
Urmann K, Gonzalez-Gil G, Schroth MH, Hofer M, Zeyer J. New field method: gas push–pull test for the in-situ quantification of microbial activities in the vadose zone. Environ Sci Technol. 2005;39:304–10.
Reim A, Lüke C, Krause S, Pratscher J, Frenzel P. One millimetre makes the difference: high-resolution analysis of methane-oxidizing bacteria and their specific activity at the oxic–anoxic interface in a flooded paddy soil. ISME J. 2012;6:2128.
Raj SS, Sumangala RK, Lal KB, Panicker PK. Gas chromatographic analysis of oxygen and argon at room temperature. J Chromatogr Sci. 1996;34:465–7.
Schroth MH, Istok JD. Models to determine first‐order rate coefficients from single‐well push‐pull tests. Groundwater. 2006;44:275–83.
Urmann K, Schroth MH, Noll M, Gonzalez‐Gil G, Zeyer J. Assessment of microbial methane oxidation above a petroleum‐contaminated aquifer using a combination of in situ techniques. J Geophys Res Biogeosciences. 2008;113:G02006.
R Development Core Team. R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2017.
Holt JA. Microbial activity in the mounds of some Australian termites. Appl Soil Ecol. 1998;9:183–7.
Knief C, Lipski A, Dunfield PF. Diversity and activity of methanotrophic bacteria in different upland soils. Appl Environ Microbiol. 2003;69:6703–14.
King H, Ocko S, Mahadevan L. Termite mounds harness diurnal temperature oscillations for ventilation. Proc Natl Acad Sci USA. 2015;112:11589–93.
Bristow KL, Holt JA. Can termites create local energy sinks to regulate mound temperature? J Therm Biol. 1987;12:19–21.
Tveit AT, Hestnes AG, Robinson SL, Schintlmeister A, Dedysh SN, Jehmlich N, et al. Widespread soil bacterium that oxidizes atmospheric methane. Proc Natl Acad Sci USA. 2019;116:8515–24.
Pratscher J, Vollmers J, Wiegand S, Dumont MG, Kaster A-K. Unravelling the identity, metabolic potential and global biogeography of the atmospheric methane-oxidizing upland soil cluster α. Environ Microbiol. 2018;20:1016–29.
Bender M, Conrad R. Kinetics of CH4 oxidation in oxic soils exposed to ambient air or high CH4 mixing ratios. FEMS Microbiol Lett. 1992;101:261–70.
Nauer PA, Schroth MH. In situ quantification of atmospheric methane oxidation in near-surface soils. Vadose Zo J. 2010;9:1052–62.
Judd CR, Koyama A, Simmons MP, Brewer P, von Fischer JC. Co-variation in methanotroph community composition and activity in three temperate grassland soils. Soil Biol Biochem. 2016;95:78–86.
Schroth MH, Eugster W, Gómez KE, Gonzalez-Gil G, Niklaus PA, Oester P. Above-and below-ground methane fluxes and methanotrophic activity in a landfill-cover soil. Waste Manag. 2012;32:879–89.
Baani M, Liesack W. Two isozymes of particulate methane monooxygenase with different methane oxidation kinetics are found in Methylocystis sp. strain SC2. Proc Natl Acad Sci USA. 2008;105:10203–8.
Gebert J, Stralis‐Pavese N, Alawi M, Bodrossy L. Analysis of methanotrophic communities in landfill biofilters using diagnostic microarray. Environ Microbiol. 2008;10:1175–88.
Bowers RM, Kyrpides NC, Stepanauskas R, Harmon-Smith M, Doud D, Reddy TBK, et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat Biotechnol. 2017;35:725–31.
Ji M, Greening C, Vanwonterghem I, Carere CR, Bay SK, Steen JA, et al. Atmospheric trace gases support primary production in Antarctic desert surface soil. Nature. 2017;552:400–3.
Pratscher J, Dumont MG, Conrad R. Assimilation of acetate by the putative atmospheric methane oxidizers belonging to the USCα clade. Environ Microbiol. 2011;13:2692–701.
Greening C, Biswas A, Carere CR, Jackson CJ, Taylor MC, Stott MB, et al. Genomic and metagenomic surveys of hydrogenase distribution indicate H2 is a widely utilised energy source for microbial growth and survival. ISME J. 2016;10:761–77.
Cordero PRF, Bayly K, Leung PM, Huang C, Islam ZF, Schittenhelm RB, et al. Atmospheric carbon monoxide oxidation is a widespread mechanism supporting microbial survival. ISME J. 2019;13:2868–81.
Carere CR, Hards K, Houghton KM, Power JF, McDonald B, Collet C, et al. Mixotrophy drives niche expansion of verrucomicrobial methanotrophs. ISME J. 2017;11:2599–610.
Schmitz RA, Pol A, Mohammadi SS, Hogendoorn C, van Gelder AH, Jetten MSM, et al. The thermoacidophilic methanotroph Methylacidiphilum fumariolicum SolV oxidizes subatmospheric H2 with a high-affinity, membrane-associated [NiFe] hydrogenase. ISME J. 2020;14:1223.
Mohammadi SS, Pol A, van Alen T, Jetten MSM, Op, den Camp HJM. Ammonia oxidation and nitrite reduction in the verrucomicrobial methanotroph Methylacidiphilum fumariolicum SolV. Front Microbiol. 2017;8:1901.
Jamali H, Livesley SJ, Hutley LB, Fest B, Arndt SK. The relationships between termite mound CH4/CO2 emissions and internal concentration ratios are species specific. Biogeosciences. 2013;10:2229–40.
Schnell S, King GM. Mechanistic analysis of ammonium inhibition of atmospheric methane consumption in forest soils. Appl Environ Microbiol. 1994;60:3514–21.
Carlsen HN, Joergensen L, Degn H. Inhibition by ammonia of methane utilization in Methylococcus capsulatus (Bath). Appl Microbiol Biotechnol. 1991;35:124–7.
Bodelier PLE, Laanbroek HJ. Nitrogen as a regulatory factor of methane oxidation in soils and sediments. Fems Microbiol Ecol. 2004;47:265–77.
Veraart AJ, Steenbergh AK, Ho A, Kim SY, Bodelier PLE. Beyond nitrogen: the importance of phosphorus for CH4 oxidation in soils and sediments. Geoderma. 2015:259–60.
Chiri E, Nauer PA, Henneberger R, Zeyer J, Schroth MH. Soil–methane sink increases with soil age in forefields of Alpine glaciers. Soil Biol Biochem. 2015;84:83–95.
Angel R, Conrad R. In situ measurement of methane fluxes and analysis of transcribed particulate methane monooxygenase in desert soils. Environ Microbiol. 2009;11:2598–610.
de Caritat P, Cooper M, Wilford J. The pH of Australian soils: field results from a national survey. Soil Res. 2011;49:173–82.
Acknowledgements
We thank Lindsay Hutley and Matthew Northwood for technical and logistical assistance, Andreas Brune for an inspiring correspondence, and the three anonymous reviewers for suggestions that greatly improved the manuscript. This work was supported by Australian Research Council grants (DP120101735 and LP100100073 awarded to SKA), Swiss National Science Foundation Early Postdoc Mobility Fellowships (P2EZP3_178421 awarded to EC; P2EZP3_155596 awarded to PAN), an ARC DECRA Fellowship (DE170100310 awarded to CG), an NHMRC EL2 Fellowship (APP1178715; salary for CG), the Terrestrial Ecosystem Research Network (TERN) OzFlux, and the TERN Australian SuperSite Network.
Author information
Authors and Affiliations
Contributions
PAN, EC, CG and SKA designed the study. EC and PAN performed field and laboratory work. Different authors were responsible for methanotroph community analysis (EC, CG, TJ, RL, PAN and SKA), kinetic analysis (PAN, EC and SKA), phylogenetic analysis (EC and CG), metagenome binning (DWW) and metagenome analysis (RL, XD, DWW, CG and EC). CG, EC and PAN wrote and edited the paper with inputs from all authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chiri, E., Greening, C., Lappan, R. et al. Termite mounds contain soil-derived methanotroph communities kinetically adapted to elevated methane concentrations. ISME J 14, 2715–2731 (2020). https://doi.org/10.1038/s41396-020-0722-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-0722-3
This article is cited by
-
Bacterial diversity from soil-feeding termite gut and their potential application
Annals of Microbiology (2023)
-
Bark-dwelling methanotrophic bacteria decrease methane emissions from trees
Nature Communications (2021)
-
Trace gas oxidizers are widespread and active members of soil microbial communities
Nature Microbiology (2021)