Abstract
Coexistence of microaerophilic Fe(II)-oxidizers and anaerobic Fe(III)-reducers in environments with fluctuating redox conditions is a prime example of mutualism, in which both partners benefit from the sustained Fe-pool. Consequently, the Fe-cycling machineries (i.e., metal-reducing or –oxidizing pathways) should be most affected during co-cultivation. However, contrasting growth requirements impeded systematic elucidation of their interactions. To disentangle underlying interaction mechanisms, we established a suboxic co-culture system of Sideroxydans sp. CL21 and Shewanella oneidensis. We showed that addition of the partner’s cell-free supernatant enhanced both growth and Fe(II)-oxidizing or Fe(III)-reducing activity of each partner. Metabolites of the exometabolome of Sideroxydans sp. CL21 are generally upregulated if stimulated with the partner´s spent medium, while S. oneidensis exhibits a mixed metabolic response in accordance with a balanced response to the partner. Surprisingly, RNA-seq analysis revealed genes involved in Fe-cycling were not differentially expressed during co-cultivation. Instead, the most differentially upregulated genes included those encoding for biopolymer production, lipoprotein transport, putrescine biosynthesis, and amino acid degradation suggesting a regulated inter-species biofilm formation. Furthermore, the upregulation of hydrogenases in Sideroxydans sp. CL21 points to competition for H2 as electron donor. Our findings reveal that a complex metabolic and transcriptomic response, but not accelerated formation of Fe-end products, drive interactions of Fe-cycling microorganisms.
Similar content being viewed by others
Introduction
Chemical communication is often the driving force of mutualistic interactions within microbial communities [1, 2]. The excreted metabolites can positively affect the growth of co-existing organisms by providing key metabolites produced by one partner and needed by the other [3,4,5,6,7,8]. The reciprocal benefits resulting from the excretion of metabolites indicate that while each partner invests metabolic resources, they both gain something from the other mutualistic partner [9,10,11]. Such mechanisms can not only control growth but also spatial distribution within the community in cases of heterogeneous environments. Synchronization of lifestyles is particularly important for mutualism interaction based on the partner’s end products for energy production [7, 12, 13].
Fe-cycling microorganisms are often found coexisting in nature. This is explained by the needs of both the ferrous iron [Fe(II)] oxidizer and ferric iron [Fe(III)] reducer to have sufficient exposure to electron donors and acceptors. For lithoautotrophic Fe(II)-oxidizers, including Gallionella sp. and Sideroxydans lithotrophicus ES-1, Fe(II) serves as an electron donor under micro-oxic conditions, thereby providing ample reducing equivalents for the assimilation of carbon [14,15,16,17]. Microbial Fe(II) oxidation facilitates the rapid precipitation of Fe(III) oxy-hydroxide nanoparticles [18, 19], which are preferred by anaerobic heterotrophic Fe(III)-reducers, such as Shewanella spp. or Geobacter spp., due to their small mineral size [20,21,22,23]. For Fe-cycling, at least temporary access to oxygen, or to oxidized compounds, is necessary for the renewal of the Fe(III) pool [14, 16]. In wetlands or peatlands, this renewal is mediated by water table fluctuations, bioturbation, or the release of oxygen via plant roots [24] which allows interaction between Fe-cycling microorganisms regardless of their different demands for O2. Production of Fe(III)-chelating ligands like riboflavins by Shewanella spp. or the presence of natural organic matter with Fe-complexing or electron shuttling properties can overcome spatial restrictions and speed up microbial activities [25, 26].
Interestingly, a shared homology is observed between the genes encoding the Fe(III) reduction machinery located at the cell surface in S. oneidensis and the genes encoding the Fe(II) oxidation machinery of Sideroxydans lithotrophicus ES-1 [17, 27]. In S. oneidensis, the Mtr machinery, comprised of the mtrABC gene cluster, is a porin–cytochrome complex in the outer membrane enabling the electron transport through the outer membrane via various hemes to Fe(III) [28, 29]. The MtoAB complex of Sideroxydans is homologous to MtrA and MtrB and functions in the same manner, forming a porin–cytochrome complex. Thus, we hypothesized that in parallel to the exchange of genetic information, additional mutualistic interactions based on the exchange of each partner’s exometabolites may have been established between these Fe-cycling partner organisms.
To elucidate these interaction mechanisms, co-culture systems are needed to facilitate downstream analytics on both the transcriptome and metabolome levels. But, co-cultivation of Fe-cycling microorganisms is very challenging due to their different lifestyles for O2. We selected two model microorganisms, the microaerophilic, autotrophic Fe(II)-oxidizer Sideroxydans sp. CL21 isolated from Schlöppnerbrunnen fen [15] and closely related to Sideroxydans lithotrophicus ES-1 [17], and the facultative anaerobic Fe(III)-reducer S. oneidensis isolated from Lake Oneida, NY [30]. First, we optimized incubation conditions that allow both partners to thrive simultaneously. The co-cultivation design allowed us to explore the influence of exposure to the partner’s end product for energy production by analyzing changes in 16S rRNA gene copies, RNA-seq analysis, and metabolomics. We hypothesized that co-cultivation will lead to an accelerated iron wheel (Fe(II) ↔ Fe(III)), because the Fe-cycling machinery would be the most affected during co-cultivation. In addition, we tested the effect of the exometabolome of one partner organism on the activity of the other.
Materials and methods
Bacterial strains and cultivation conditions
Sideroxydans sp. CL21 and S. oneidensis were cultivated in liquid supernatant exchange, monoculture, and co-culture incubations [15, 30,31,32] using ATCC medium 2672 (modified Wolfe’s minimal media (MWMM)) amended with 10 mM MES buffer (pH 6.3) unless otherwise noted. Sideroxydans sp. CL21 stock cultures grown in semi-solid gradient tubes containing 0.15% agarose-stabilized MWMM (Biozyme LE Agarose; Biozyme Scientific GmbH, Hessisch Oldendorf, DE) and zero-valent Fe (Fe(0)) were used as inoculum (1 mL inoculum 100 mL−1 media) in these incubations. Zero-valent iron powder is often used as the source of Fe(II) to cultivate Fe(II)-oxidizing bacteria [33], and we observed enhanced biomass during cultivation of Sideroxydans sp. CL21 in semi-solid gradient tubes, in comparison to FeS or FeCl2. Fe(0) also provided the best Fe source for the liquid co-culture incubations. S. oneidensis overnight cultures were grown aerobically in Luria-Bertani medium. 1 mL was transferred into 125 mL serum bottles containing 60 mL MWMM amended with 18 mM lactate and grown aerobically at room temperature. 2 mL of MWMM-grown S. oneidensis was harvested after ~14 h (late exponential growth), centrifuged (10,000 g, 5 min), washed twice, resuspended in 2 mL MWMM, and used as inoculum (final concentration: ~2 × 105 cells mL-1) for the above mentioned incubations.
Monoculture and co-culture incubations
Monocultures and co-cultures were set up in triplicates in 125 mL serum bottles with an Fe(0)-containing bottom-layer plug (10 mL MWMM, 3% agarose (Biozyme LE Agarose), 1 g L−1 Fe(0)). Following solidification, 60 mL MWMM amended with 1 mM lactate was dispensed on top and continuously flushed with an N2:CO2:O2, 78:20:2, (flow rate: 300 mL min−1) gas mixture to maintain a micro-oxic environment (Fig. 1a). Cells were harvested at four time-points (6 h, 3 d, 6 d, 9 d), centrifuged) and stored at −80 °C. 6 d samples were selected for RNA-Seq.
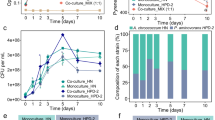
a Pre-cultures were grown under standard optimized conditions either in gradient tubes or in liquid media prior to inoculation of co-culture incubations. b Both organisms can also grow in monoculture in this incubation set-up. 16S rRNA gene copy abundances of Sideroxydans CL21 or S. oneidensis showing growth in monoculture and in co-culture over 9 days of incubation. Data points represent mean abundances (n = 3) from either Sideroxydans CL21 monocultures (open squares) or Sideroxydans CL21 in co-culture (closed squares) and mean abundances (n = 3) from S. oneidensis monocultures (open circles) or S. oneidensis in co-culture (closed circles). In b, a box with dashed lines is used to indicate that samples from the 6 d time point were used for RNA-seq. Error bars represent standard deviation in triplicate incubations. In some cases, error bars are smaller than the symbols.
Cell-free supernatant exchange experiment
Supernatant exchange experiments were set up in triplicates according to Mori et al. [34], with the following modifications: Sideroxydans sp. CL21 and S. oneidensis cultures were grown in 200 mL MWMM amended with 1 mM lactate and 1.2 mM FeSO4 or 10 mM hydrous Fe(III) oxyhydroxide, respectively, for 4 days with shaking at room temperature. Cultures were centrifuged, filtered (0.22 µm), cell-free supernatant was collected, and supernatant-exchange experiments were set up (1 mL inoculum 100 mL−1 media) (Fig. 2a). Fe(II) concentrations were measured using the phenanthroline method [35]. Cells were harvested at four time-points (0d, 1 d, 1.5 d, 2 d), centrifuged, and stored at −80 °C.
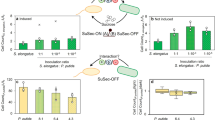
a Briefly, monocultures of Sideroxydans CL21 (Sid) and S. oneidensis (Shew) were amended with cell-free supernatants obtained from either the same or partner strain. b Fe(II) concentrations were measured to follow Fe(II) oxidation in Sideroxydans CL21 incubations amended with either Sideroxydans CL21 [Sid (Sid)] or S. oneidensis [Sid (Shew)] cell-free supernatant or to monitor Fe(III) reduction in incubations with S. oneidensis amended with either Sideroxydans CL21 [Shew (Sid)] or S. oneidensis [Shew (Shew)] cell-free supernatant. c qPCR assays were used to determine mean abundances of 16S rRNA gene copies in triplicate samples from cell-free supernatant exchange incubations. Error bars represent standard deviation in triplicate incubations (n = 3). In some cases, error bars are smaller than the symbols.
Genomic DNA extraction
Cells were harvested via centrifugation (5000 g, 10 min, 4 °C), extracted using the DNeasy PowerSoil kit (Qiagen, Hilden, DE) according to manufacturer’s instructions, and frozen at −20 °C. Sideroxydans sp. CL21 genomic DNA was used for whole genome sequencing (PacBio sequencing) [36], and the assembled genome used as the reference genome for RNA-Seq analysis.
Quantitative PCR
qPCR analysis using primer pairs Sid-120F/Sid-167R [18] or She12F/She220R was performed to determine total 16S rRNA gene copies of Sideroxydans sp. CL21 and S. oneidensis from supernatant exchange, monoculture, and co-culture incubations. 2–20 ng of genomic DNA was used as template for analyzes on a Mx3000P qPCR system (Agilent, Waldbronn, DE) with Maxima SYBR Green Master Mix (Thermo Scientific, Schwerte, DE). Standard curves produced using serial dilutions of representative plasmid mixtures (5 × 108 to 5 × 102 copies; R2 value = 0.999–1.000) were linear for both primer sets. The qPCR performed with efficiencies ranging from 80 to 90%.
Sample prep for RNA-Seq analysis
Total RNA was extracted from three biological replicates from Sideroxydans sp. CL21 and S. oneidensis monoculture and co-culture incubations using the phenol-chloroform extraction method adapted from Wegner et al. [37]. Equimolarly pooled cDNA libraries were sequenced in paired-end mode (2 × 150 bp) on an Illumina NextSeq 500 platform by LGC Genomics (Berlin, DE).
Transcriptome data processing and analysis
The quality of raw, demultiplexed RNA-Seq datasets was inspected using FastQC (v0.11.7) [38]. Quality trimming (settings: minlen=75, qtrim=rl, ktrim=rl, k = 25, mink=11, trimq=20, qtrim=rl) and adapter sequence removal was done with bbduk (v38.26) [39] using the included set of common sequence contaminants and adapters. rRNA-derived and non-coding RNA sequences were filtered out with sortmerna (v2.1) [40] and its pre-compiled SILVA databases [41] and Rfam [42]. The remaining, putatively mRNA-derived sequences were mapped onto the available S. oneidensis reference genome [43] and the newly sequenced and annotated Sideroxydans. sp. CL21 genome [36] using bbmap (v28.26) [39] (settings: slow, k = 11). Resulting bam files were sorted and indexed with samtools (v1.3.1) [44]. Genome annotations were used for generating simplified annotation format files for subsequent read counting (http://bioinf.wehi.edu.au/featureCounts/). Read counts (number of mapped reads per coding gene) were deduced from the generated bam files using the program featureCounts implemented in subread (v1.6.3) [45, 46]. The number of reads mapped to either Sideroxydans sp. CL21 and S. oneidensis in both monoculture and co-culture incubations can be found in Table S1. Differential gene expression analysis was carried out in the R framework for statistical analysis (v3.5.1) [47], using the package edgeR (v3.20.9) [48], including all dependencies.
Metabolomics methods
Sample preparation for metabolomic profiling
Sample preparation was conducted in triplicate as described previously [34], unless otherwise noted. Briefly, 50 mL of liquid cultures were centrifuged (4000 g) and extracted using Strata-X® polymeric reverse phase cartridges (200 mg adsorbent; Phenomenex, Torrance, CA, USA). Sample preparation was performed in an anaerobic chamber to minimize Fe(II) oxidation and subsequent Fe(III) precipitation. (NOTE: in previous experiments, precipitation of Fe(III) during sample preparation resulted in completely dissimilar metabolite profiles—data not shown). Samples were measured in a randomized sample list immediately after preprocessing.
GC/MS measurement
GC/MS metabolomics analysis was conducted on 3 and 6 d supernatant exchange samples using a high resolution Q-Exactive-GC electron impact (EI) orbitrap mass spectrometer (Thermo Scientific), with gas-chromatographic separations implemented on a Trace 1310 equipped with TriPlus RSH autosampler. A TG-5SILMS column (length = 30 m; inner diameter = 0.25 mm, 0.25 μm film; Thermo Scientific) was used. Column operation parameters and GC-Orbitrap settings used are described in the supplementary methods.
XCMS metabolomic analysis
The XCMS data processing was carried out using –cdf files, which were converted from the Thermo RAW-files using the Xcalibur 3.0.63 (Thermo Scientific) onboard file converter. Processing was carried out using XCMS Server version 3.01.01. Pre-defined settings were used for GC-measurement “Single Quad (matched filter),” except, retention time correction was removed [49]. Parameters used are listed in Table S2.
UHPLC/HRMS measurement and spectra acquisition
Ultra high performance liquid chromatography (UHPLC) coupled with high resolution MS (HRMS) of both for untargeted and targeted (γ-aminobutanoic acid (GABA) and zinc) metabolites was performed using an UltiMate HPG-3400 RS binary pump (Thermo Scientific, Bremen, Germany) and WPS-3000 autosampler (Thermo Scientific) equipped with a 25 µL injection syringe and 100 µL sample loop. Detailed parameters used are described in the supplementary methods.
GC/MS data processing and annotation
Using an in-house R-script [50], detected masses were deconvoluted via peak-shape comparison to potential metabolites and reduced in number to only include potential metabolites containing ≥20 fragments. Potential metabolites present in blanks and media controls (<fivefold increase or decrease relative to bacterial samples) were removed (Table S3). The remaining potential metabolites were log-transformed and auto-scaled via Metaboanalyst 4.0 [51] to yield heatmaps with the 15 most dysregulated metabolites. Unknown metabolites were identified by comparing fragmentation patterns to known compounds in the NIST/EPA/NIH EI Mass Spectral Library NIST 17 (copyright United States Department of Commerce) searched with the NIST Mass Spectral Search Program v.2.3a nd Golm Metabolome Database library [52]. Metabolites with match scores >700 were tentatively identified and compounds with multiple match score hits >600 were labeled by compound class. During the analysis of supernatant exchange samples, an S. oneidensis control (Shew(Shew)) sample (6 d) was also removed from the analysis due to errors during spectral profiling.
UHPLC/HRMS data processing and quantification
Quan Browser (XCalibur 3.0.63) (Thermo Scientific) was used for peak detection and integration for GABA and a detected zinc compound with the following settings: mass tolerance = 10 ppm, peak detection algorithm=ICIS; smoothing points=1; baseline window = 40; area noise factor=15; peak noise factor = 10; peak high = 5%, tailing factor = 2.5 and peak detection method = nearest RT. Each biological sample was measured in triplicate using a randomized sample list. The presence of GABA in the samples was confirmed by comparison to GABA standards. In order to identify metalloenzyme complexes, for example zinc complexes, MICP was applied to the UHPLC/HRMS data of the supernatant extracts and the zinc compounds were identified based on the natural isotopic pattern. The single isotopes of the metal-complexes were resolved due to a resolution of 280.000 at m/z = 200, e.g., for 64Zn-, 66Zn-, 67Zn-, and 68Zn-complexes for one substance. The identification is based on the mass differences between the single isotopic peaks of the substance (with an error <1 ppm) and the intensity of the mass peaks according to the abundance of the natural zinc isotopes.
Results
Merging lifestyles of S. oneidensis and Sideroxydans sp. CL21
S. oneidensis can be grown easily anaerobically in liquid media with Fe(III) as a terminal electron acceptor. However, the microaerophilic Sideroxydans sp. CL21 prefers growing in so-called “gradient” tubes filled with semi-solid media, an Fe(II) source in the bottom, and O2 supplied in the headspace. Opposing Fe and O2 gradients allow growth at a depth with optimal O2 concentrations of 20–40 µM [53]. Microbial Fe(II)-oxidation leads to the formation of a distinct orange ring of precipitated Fe(III) (oxy)hydroxides. However, the semi-solid media poses complications for the envisioned metabolomics profiling. To identify metabolites produced and secreted by each individual partner, we had to establish a suboxic liquid media setup that enables the growth of both Sideroxydans sp. CL21 and S. oneidensis in monocultures and co-cultures. In total we tested 48 combinations of media, carbon sources, buffer type, and Fe sources, bottle sizes, and headspace conditions to land on the best setup for our incubations (Fig. 1a): MWMM supplemented with MES buffer, lactate, and Fe(0) provided in a semi-solid media plug at the bottom, and a headspace of N2:CO2:O2 (78:20:2) that is constantly flushed through the bottles. Under these conditions, Fe(0) reacts slowly with water forming Fe(II) and H2 (anaerobic corrosion) and the released Fe(II) is slowly abiotically oxidized to Fe(III). Consequently, both Fe(II) and Fe(III) are continuously provided, and microbial activity should speed up the production of each Fe species. Unlike other Fe minerals, Fe(0) is not ubiquitous in environmental systems and Fe(0) is most often applied to groundwater and soil environments for in situ remediation of metalloid-contaminated sites. However, amendment with Fe(0) results in enrichment of Fe(III)-reducing and Fe(II)-oxidizing microorganisms [54,55,56,57,58].
Exometabolome stimulates growth and activity of each partner
First, we performed supernatant exchange experiments (Fig. 2a). Amendments with the partner’s cell-free supernatant led to enhanced rates of the production and consumption of Fe(II). Rates of Fe(II) oxidation by Sideroxydans sp. CL21 over 2 days increased from 4.3 to 5.5 μM h−1 (t-test, p = 0.002) when amended with S. oneidensis cell-free supernatant and Fe(III) reduction rates of S. oneidensis increased from 4.9 to 7.3 μM h-1 (t-test, p = 0.007) following amendment with Sideroxydans sp. CL21 cell-free supernatant (Fig. 2b, Supplementary Table S4). 16S rRNA gene copy numbers increased one order of magnitude in Sideroxydans sp. CL21 [Sid(Shew)] and two orders of magnitude in S. oneidensis [Shew(Sid)] after 3 days of batch incubation compared with its control Sid(Sid) or Shew(Shew), respectively (Fig. 2c).
These stimulatory effects show that diffusive metabolites are beneficial for this mutualistic relationship, such that metabolites from one species promotes growth and other metabolic processes, such as iron oxidation or reduction, of the other species. To identify potential chemical mediators, comparative (untargeted) metabolomic profiling was performed after 3 and 6 days. After three days of incubation with S. oneidensis supernatant, an increase in Sideroxydans sp. CL21 excretion was detected (Fig. 3a, Supplementary Table S3). A phenylketone was putatively identified by analyzing the accurate mass (309.0972 m/z) as a dehydrated fragment of a hydrated phenylketone with accurate mass (327.1077 m/z) via comparison with the NIST database (match scores of 631 and 673 respectively). The accurate mass of the dehydrated fragment fits within 1.4 ppm error according to exact mass. After 6 days of incubation with S. oneidensis cell-free supernatant, 88% of the top 25 dysregulated metabolites, meaning metabolites with concentrations affected by treatment, (out of 473 annotated peaks), were more abundant in the exometabolome of Sideroxydans sp. CL21 exposed to S. oneidensis supernatant (Fig. 3b, Supplementary Table S3). S. oneidensis exposed to the supernatant of Sideroxydans sp. CL21 also exhibited increased excretion with a total of 321 annotated compounds. A mixed metabolic response was observed, with only 52% of the top 25 most dysregulated compounds more abundant after 3 days. After 6 days, all 25 of the top dysregulated compounds were less abundant in S. oneidensis exposed to Sideroxydans sp. CL21 supernatant suggesting that these metabolites were not being produced at similar rates as in monocultures. For example, a dihydroxyindole was putatively identified (accurate mass 365.1655 m/z, which fits −0.8 ppm according to exact mass; NIST database R. match 674 and match score 658). An overall decrease in excretion at day 6 relative to day 3 was also exhibited, as evidenced by only 151 annotated compounds detected at day 6. Efforts to identify additional specific metabolites whose concentrations differed most were unsuccessful, which is in accordance with a high degree of secondary metabolite production that are not annotated in standard libraries (Fig. 3, Supplementary Table S3).
Comparative analyses of the exometabolites produced by Sideroxydans sp. CL21 incubated with its’ own cell-free supernatant (50% v/v) (Sid(Sid)) versus Sideroxydans sp. CL21 incubated with the cell-free supernatant of S. oneidensis (50% v/v) (Sid(Shew)) after a 3 d (a) and 6 d (b) incubation period were conducted. Similarly, comparative analyses of the exometabolites produced by S. oneidensis incubated with its’ own cell-free supernatant (50% v/v) (Shew(Shew)) versus S. oneidensis incubated with the cell-free supernatant of Sideroxydans sp. CL21 (50% v/v) (Shew(Sid)) after a 3 d (c) and 6 d (d) incubation period were conducted. “_A, _B, or _C” corresponds to one replicate (n = 3).
RNA-Seq profiling of co-cultures show unexpected patterns of differentially expressed genes
Sideroxydans sp. CL21 and S. oneidensis were grown together over 9 days to cover potential mutually induced responses and effects of physical contact with the partner (Fig. 1a). Co-culturing was beneficial to growth, as the increase in 16S rRNA gene copies of both organisms was higher in co-cultures compared with monocultures (Fig. 1b) with the highest increase between 3 and 6 days. Thus, maximal 16S rRNA gene copies in co-cultures coupled with increased Fe-cycling rates in supernatant exchange experiments (Table S4) led to the selection of day 6 for RNA-Seq analyses.
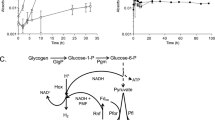
The overall transcriptome profiles of the monocultures demonstrated that genes involved in Fe(II) oxidation (i.e., the Sideroxydans sp. CL21 mto gene cluster) or Fe(III) reduction (i.e., the S. oneidensis mtr gene cluster) were highly expressed (ranging from 6.2–9.5 log2CPM), showing that not only Sideroxydans sp. CL21, but also S. oneidensis were utilizing Fe under suboxic conditions. Both monoculture gene expression profiles were dominated by genes encoding proteins linked to core metabolic functions, ribosomal proteins, RNA polymerase subunits, and ATP synthase subunits (Fig. 4a, Supplementary Tables S5, S6). Genes encoding a lactate permease and lactate dehydrogenase were highly expressed, which indicated S. oneidensis is using lactate as the electron donor (Supplementary Table S5). Genes involved in flagellar motility were also highly expressed in both organisms (Supplementary Tables S5, S6). In the case of S. oneidensis, we detected notably high gene expression values (7.5–9.0 log2CPM) for genes of the putrescine degradation pathway (Table 1, Supplementary Table S5). In the Sideroxydans sp. CL21 gene expression profiles, we identified multiple genes for cation efflux system proteins that featured particular high gene expression values (log2CPM > 10) (Supplementary Table S5). A cluster of NiFe-hydrogenases-related genes, including genes for hydrogenase subunits and proteins involved in hydrogenase assembly and maturation was uniformly and highly expressed with log2CPM values ranging between 7 and 10 (Table 2, Supplementary Table S6). Genes involved in the Calvin–Benson–Bessham pathway, which reduces 3-phosphoglycerate via RubisCO (ribulose-1,5-bisphosphate carboxylase/oxygenase), were highly expressed in Sideroxydans sp. CL21, such that genes encoding the proteins CbbO and CbbQ were upregulated (log2FC 1.02–1.31). The presence of both the small and large forms of RubisCO suggests a high tolerance to fluctuating O2 concentrations [17]. Also, genes encoding a lactate dehydrogenase and permease were highly expressed, thus implicating possible simultaneous use of inorganic and organic carbon during co-cultivation (Supplementary Table S6).
Graphical representation of overall gene expression in Sideroxydans sp. CL21 and S. oneidensis monocultures (a) compared wiith the differential gene expression during co-cultivation b are shown. Differentially expressed genes were determined by analysis of RNA-seq data from triplicate co-culture (Sideroxydans CL21 + S. oneidensis) and monoculture batch incubations at the 6 d time point (see Fig. 1). log2CPM (CPM = counts per million) values represent a normalized measure for gene expression based on the mRNA-derived sequences that were mapped onto genes.
For co-culture incubations, 15.7 and 21.3% of the total genes were differentially expressed in Sideroxydans sp. CL21 and S. oneidensis, respectively. 927 genes were differentially expressed of which 428 were up- and 499 down-regulated in S. oneidensis, compared with 588 genes (249 up-, 338 down-regulated), which were differentially expressed in Sideroxydans sp. CL21 when grown in co-culture (Supplementary Fig. S1). Surprisingly, genes involved in Fe(II) oxidation or Fe(III) reduction in both partners were not upregulated in co-culture, but were still highly expressed, indicating growth in steady-state (Tables 1, 2; Supplementary Tables S5, S6). The most upregulated genes in S. oneidensis include genes encoding putrescine breakdown and an EPS biosynthetic gene cluster, as well as substrate-binding and permease components of zinc and tungstate ABC-type uptake systems. The most downregulated genes encode proteins linked to pilus retraction, type IV pili, and chemotaxis signal transduction (Table 1, Supplementary Table S5). In Sideroxydans sp. CL21, the most upregulated genes code for biopolymer and lipoprotein transport, NiFe hydrogenase incorporation and metallo-center assembly proteins, and hydrogenase-1 and -2 components. The first couples H2 oxidation in the periplasm to reduction of quinones in the inner membrane, whereas the second is involved in O2 reduction. Motility-related genes and genes associated with NAD-dependent formate dehydrogenase alpha, beta, gamma, and delta subunits and the sulfur carrier protein FdhD were among the most downregulated in Sideroxydans sp. CL21 (Table 2, Supplementary Table S6).
Targeted candidate analyses
Based on gene expression changes in S. oneidensis, which include gene clusters involved in putrescine and amino acid breakdown, an ABC-type tungsten- and an ABC-type zinc uptake system, we used targeted analyses to examine metabolites linked to these differentially expressed pathways. We detected GABA and evidence of ornithine and arginine [59, 60], all of which are metabolites comprising the putrescine biosynthesis and breakdown pathways. The upregulated pathways are thus responsible for the production of excreted intermediates, but putrescine itself and agmatine, another biosynthetic precursor, were not detected. We also detected GABA, ornithine, and arginine in the supernatant exchange experiment both on day 3 and day 6. We quantified GABA in supernatant exchange experiments on day 6 (Fig. 5). The amendment of the Sideroxydans sp. CL21 supernatant to S. oneidensis did not influence GABA amounts. In contrast, addition of the S. oneidensis supernatant to Sideroxydans sp. CL21 led to a significant decrease of GABA (t-test, p < 0.001, n = 3) (Fig. 5). No tungsten complexes were identified based on the evaluation of the isotope patterns of the molecular ions. Isotope pattern analysis indicated a metabolite containing complexed zinc. The identification of the metabolite containing complexed Zn was based on the mass differences between the single isotopic peaks of the substance (with an error <1 ppm) and the intensity of the mass peaks was determined according to the abundance of the natural Zn isotopes. Addition of S. oneidensis supernatant to Sideroxydans CL21 as well as the amendment of Sideroxydans CL21 supernatant to S. oneidensis led to reduced levels of the Zn compound (Fig. S2).
Targeted metabolomics used to quantify the presence of γ-butanoic acid (GABA), a degradation product in the putrescine biosynthesis pathway, in S. oneidensis MR-1 and Sideroxydans CL21 supernatant exchange experiment. Sid and Shew correspond to Sideroxydans sp. CL21 and S. oneidensis monoculture control incubations, respectively. The labeling of Sid(Sid), Sid(Shew), Shew(Shew) and Shew(Sid) are described in the legend of Fig. 3. Area under the curve of GABA in the supernatant exchange experiment was measured on day 6. Error bars represent the standard deviation of three biological replicates (n = 3). Stars indicate significance: *p < 0.05, **p < 0.01 and ***p < 0.001.
Discussion
Systematic investigations of Fe-cycling microorganisms were hampered in the past due to their different requirements for oxygen. After methodical optimization, we could merge the growth requirements of a microaerophilic Fe(II)-oxidizer and an anaerobic Fe(III)-reducer to study interactions of organisms driving the complete iron wheel. The overall expression levels of genes involved in Fe(III) reduction in S. oneidensis and Fe(II) oxidation in Sideroxydans sp. CL21 were high under suboxic conditions. Although cell densities of standard monoculture incubations were not reached using this co-culture approach [61,62,63,64], we argue that such “candy shop” laboratory conditions do not mimic conditions in nature. Only co-culturing allows investigating the dynamics of interaction mechanisms of naturally co-occurring bacteria.
By combining transcriptomics and metabolomics, we disproved the hypothesis that the expression of genes involved in Fe(III) reduction and Fe(II) oxidation is the most affected during co-cultivation. Our data further suggest no acceleration of the iron wheel at all under these conditions. Excreted metabolites promoting growth and Fe-cycling were the greatest benefit of co-existence. Exometabolites from both S. oneidensis and Sideroxydans sp. CL21 elicit a positive effect on growth of their partners, suggesting a mutualistic interaction (Fig. 2). The physical presence of the partner seemed to be less important than the presence of the partner’s exometabolome.
Untargeted metabolomics revealed an overall upregulated metabolism of Sideroxydans sp. CL21 from day 3 to 6 when exposed to S. oneidensis exudates compared with Sideroxydans sp. CL21 alone (Fig. 3a, b). In contrast, S. oneidensis exposed to Sideroxydans sp. CL21 exudates showed a mixed response on day 3 and a predominately downregulated metabolism on day 6 (Fig. 3c, d). Here, the increased performance did not correlate with increased metabolite excretion. Alternatively, S. oneidensis might take up metabolites from the spent medium more efficiently if triggered with exudates from the partner. We found evidence for regulated phenylketone metabolism in Sideroxydans sp. CL21 exposed to S. oneidensis supernatant at day 3 and indole metabolism in S. oneidensis exposed to Sideroxydans sp. CL21 at day 6. Sugars, amino acids and carboxylic acids were differentially regulated at day 3, indicating a balanced response of primary metabolism by both partners.
The most differentially expressed genes of both partners grown in co-culture were involved in polyamine biosynthesis and degradation, biofilm formation, motility (i.e., flagella assembly and chemotaxis), utilization of inorganic and organic carbon substrates, and the formation of hydrogenases and dehydrogenases (Fig. 4b). These genes are known to be involved in physiological responses to environmental cues and nutrient availability, however, they are not uniquely related to Fe(III) reduction or Fe(II) oxidation.
The genomes of both S. oneidensis and Sideroxydans sp. CL21 encode NiFe hydrogenases, implicating a potential competition for H2. Periplasmic NiFe hydrogenases in S. oneidensis play a role in both H2 production and oxidation [65, 66], while the NiFe hydrogenases in Sideroxydans sp. CL21 enable only H2 oxidation [67]. In iron-rich peatlands, like the Schlöppnerbrunnen fen, H2 concentrations range from 4.5 to 6.0% in depths up to 40 cm [68, 69]. Here H2 appears to be produced by fermenters, with the rhizospheres providing a prime location, and is scavenged by anaerobic secondary fermenters, Fe(III) reducers, methanogens, and acetogens [68, 70,71,72]. As Sideroxydans sp. CL21 might be also able to utilize H2 in this fen as an alternative electron donor, variable oxygen concentrations released by plant roots in the rhizosphere and redox fluctuations due to variations in the water table might favor this microaerophile in the interspecies H2 competition. The differential gene expression patterns observed during co-cultivation confirmed interspecific H2 competition. Upregulation of genes encoding various NiFe hydrogenases in Sideroxydans sp. CL21 suggests the preferred utilization of H2, which serves here as an alternative electron donor to Fe(II). Simultaneous downregulation of genes encoding hydrogenases in S. oneidensis implicates synchronization between these Fe-cycling microorganisms. To compensate for the lack of H2 as the electron donor, S. oneidensis invests in accelerated lactate permease and dehydrogenase expression (Fig. 4b). These orchestrated responses in an environment offering multiple energy sources can even lead to a slowdown of the oxidative part of the iron wheel.
Differential gene expression patterns linked to biofilm formation suggest that Sideroxydans sp. CL21 and S. oneidensis communicate in co-culture using excreted metabolites to coordinate physiology. Biofilm formation was likely triggered by the upregulation of genes involved in the biosynthesis of putrescine and EPS, as well as the downregulation of genes involved in chemotaxis, motility, and type IV pili biosynthesis and activation in S. oneidensis. Detection of GABA, a metabolic product of putrescine, provides additional evidence that polyamines do play a role in microbial interactions [34]. In Sideroxydans sp. CL21, upregulation of genes involved in production and transport of biopolymers and lipoproteins and downregulation of genes involved in flagella machinery biosynthesis and motility are also relevant for biofilm formation during co-cultivation. Polyamine signaling in co-culture [73,74,75] might regulate biofilm formation activity of Sideroxydans sp. CL21. In conclusion, both partners invest in biofilm formation, thus initiating and supporting mutualistic interactions.
Polyamines, including norspermidine, spermidine, and putrescine are known to be required for cell growth and are important signaling molecules. Norspermidine enhances biofilm formation as an extracellular signaling molecule in Vibrio cholerae by interacting with a periplasmic sensor protein and a transmembrane protein [74, 76, 77]. Spermidine and putrescine transporters play a role in surface-associated growth of Agrobacterium tumefaciens and Pseudomonas putida [78, 79]. In Yersinia pestis, endogenous putrescine is required for the formation and development of biofilms. Amendment with putrescine rescues biofilm-deficient phenotypes in Y. pestis, implicating a role for both endogenous and exogenous putrescine in the activation of biofilm formation [80]. The latter finding is especially important for the current study, such that putrescine produced by S. oneidensis might play a dual role as an endogenous signal for biofilm development in S. oneidensis and an exogenous signal supporting the same process in Sideroxydans sp. CL21.
Other highly upregulated genes in S. oneidensis include those associated with ABC-type Zn transporters and ABC-type tungstate uptake systems (Table 1). Zn, an essential micronutrient, is also known to be toxic in excess as being a highly competitive divalent metal capable of displacing weakly bound transition metals in the active sites of metalloenzymes [81]. The observed upregulation of ABC-type Zn transporters suggests dynamic regulation of Zn homeostasis to preserve metabolic functioning. The upregulation of genes involved in ABC-type tungstate uptake systems hints that tungsten may function as a cofactor in place of molybdenum for formate dehydrogenases (FDHs) [82, 83]. Interestingly, FDHs can also use FeS clusters, flavins, or cytochromes as cofactors, all of which play a role in Fe(III) reduction and electron transfer [84]. While the links between tungsten-dependent enzymes and electron transfer in co-cultures are poorly understood, tungsten-containing FDHs are beneficial in certain syntrophic interactions under electroactive conditions [83].
Sideroxydans sp. CL21 and S. oneidensis share homologous Fe-cycling machinery: mto genes in Sideroxydans sp. CL21 and mtr genes in S. oneidensis [17, 27]. In both microorganisms, these gene clusters function as a porin–cytochrome complex forming an electron conduit through the outer membrane via various hemes. This cluster ultimately results in either reduction of Fe (III) outside the cell, coupled to the oxidation of organic carbon or H2 in case of S. oneidensis [28, 29], or the transfer of electrons from Fe(II) via MtoAB to a molybdoenzyme and subsequently to the respiratory chain (i.e., cbb3 or bd oxidase) via CymA [85] in Sideroxydans sp. CL21. The genes encoding MtoAB are located in a gene cluster also containing mtoD, a soluble periplasmic cytochrome encoding the protein MtoD, and cymA (homologous to cymA found in S. oneidensis), a quinol oxidoreductase encoding the protein CymA [17, 27]. The mtoAB cluster is also homologous to the pioABC gene cluster in Rhodopseudomonas palustris, which similarly forms a porin-cytochrome conduit [85].
Homologs to MtrAB and PioAB are widespread amongst Fe(III)-reducers and Fe(II)-oxidizers widely distributed across Alpha-, Beta-, Gamma-, and Deltaproteobacteria [85] pointing toward possible horizontal gene transfer events. It is believed that Fe(III) respiration is one of the oldest respiration processes on Earth [86] and aerobic microbial Fe(II) oxidation became possible billions of years ago when oxygen was released as a byproduct by photosynthetic organisms [85]. Various mechanisms to optimize mutual dependencies including communication might have developed over time when colonizing the same habitat despite seemingly different preferred ecological niches (i.e., microoxic vs. anoxic) [14, 87, 88]. Thus, the homologous Fe-cycling machinery, the ability of both bacteria to positively respond to the partner’s diffusive exometabolites concerning activity and growth, and the observation that gene expression is in flux (differentially expressed genes) when these microorganisms are active in co-culture, hints at potential co-evolution.
Our co-cultivation setup provides evidence that S. oneidensis is respiring Fe(III) under suboxic conditions without switching to its preferred terminal electron acceptor, O2. This flexibility allows adjustment to redox gradients and helps to explain the spatial co-existence of Fe-cycling bacteria in a variety of habitats [89,90,91,92]. Under fluctuating redox conditions in nature in which these microorganisms must continuously adapt, regulation of exometabolites can become key for Fe-cycling. Acceleration of the iron wheel via optimization of electron donor and electron acceptor availability may not be the most important factor driving their interaction, since Fe is the fourth most abundant element on Earth [93, 94]. Instead, both partners invest in shaping their environment via biosynthesis and detection of diffusive metabolites. Joint biofilm formation might be more important to ensure access to common goods, such as riboflavin, that function as electron shuttles [95,96,97,98,99]. High abundances of FeOB and FeRB have been reported from microbial mats, one of the most well-studied types of biofilms [100]. Simultaneous activity, which we mimicked in our setup, may not be as crucial in nature, where fluctuating redox conditions favor alternately one half of the iron wheel. Temporally decoupled redox processes are the rule and not the exception in nature.
Conclusion
The results of this study have important implications for the understanding of microbial metal-cycling. Using an integrative transcriptomic and metabolomic approach, we revealed that this Sideroxydans sp. CL21 and S. oneidensis co-culture system can be used as a model system for future investigations focused on the interactions of Fe(II) oxidizers and Fe(III) reducers that likely occur in nature and the diffusive metabolites that play a role in these interactions. We provide quantitative evidence that Sideroxydans sp. CL21 and S. oneidensis benefit from growth in co-culture, and that diffusive metabolites play a more important role in biogeochemical processes than previously thought. Shaping the environment by a regulated inter-species biofilm formation appears to be the key mechanism underlying interactions of Fe-cycling microorganisms. Characterization of biofilms formed by Sideroxydans sp. CL21 and S. oneidensis during co-cultivation will provide further evidence about the regulation of these interactions, specifically in regards to producing, sharing, and utilizing common goods, such as diffusive chemical mediators and electron shuttles. Our data further demonstrated that competition for H2 between Fe-cycling partners provides another level of metabolic complexity when both partners are simultaneously active. Future incubation studies are needed to prove that Sideroxydans sp. CL21 does, in fact, prefer H2.
Data availability
The RNA-seq datasets used in this study have been deposited in ArrayExpress under the accession number E-MATB-9015 and the Metabolomics datasets have been deposited in MetaboLights under the accession number MTBLS1733.
References
Ponomarova O, Patil KR. Metabolic interactions in microbial communities: untangling the Gordian knot. Curr Opin Microbiol. 2015;27:37–44.
D’Souza G, Shitut S, Preussger D, Yousif G, Waschina S, Kost C. Ecology and evolution of metabolic cross-feeding interactions in bacteria. Nat Prod Rep. 2018;35:455–88.
Schink B. Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev. 1997;61:262–80.
Pernthaler A, Dekas AE, Brown CT, Goffredi SK, Embaye T, Orphan VJ. Diverse syntrophic partnerships from deep-sea methane vents revealed by direct cell capture and metagenomics. Proc Natl Acad Sci USA. 2008;105:7052–7.
Men Y, Feil H, Verberkmoes NC, Shah MB, Johnson DR, Lee PKH, et al. Sustainable syntrophic growth of Dehalococcoides ethenogenes strain 195 with Desulfovibrio vulgaris Hildenborough and Methanobacterium congolense: global transcriptomic and proteomic analyses. ISME J. 2012;6:410–21.
Zelezniak A, Andrejev S, Ponomarova O, Mende DR, Bork P, Patil KR. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc Natl Acad Sci USA. 2015;112:6449–54.
Marchal M, Goldschmidt F, Derksen-Müller SN, Panke S, Ackermann M, Johnson DR. A passive mutualistic interaction promotes the evolution of spatial structure within microbial populations. BMC Evol Biol. 2017;17:106.
Thompson AW, Foster RA, Krupke A, Carter BJ, Musat N, Vaulot D, et al. Unicellular cyanobacterium symbiotic with a single-celled eukaryotic alga. Science. 2012;337:1546–50.
Zengler K, Palsson BO. A road map for the development of community systems (CoSy) biology. Nat Rev Microbiol. 2012;10:366–72.
Sachs JL, Hollowell AC. The origins of cooperative bacterial communities. MBio. 2012;3:1–3.
Johnson DR, Goldschmidt F, Lilja EE, Ackermann M. Metabolic specialization and the assembly of microbial communities. ISME J. 2012;6:1985–91.
Bull JJ, Rice WR. Distinguishing mechanisms for the evolution of co-operation. J Theor Biol. 1991;149:63–74.
Foster KR, Wenseleers T. A general model for the evolution of mutualisms. J Evol Biol. 2006;19:1283–93.
Weber KA, Achenbach LA, Coates JD. Microorganisms pumping iron: anaerobic microbial iron oxidation and reduction. Nat Rev Microbiol. 2006;4:752–64.
Lüdecke C, Reiche M, Eusterhues K, Nietzsche S, Küsel K. Acid-tolerant microaerophilic Fe(II)-oxidizing bacteria promote Fe(III)-accumulation in a fen. Environ Microbiol. 2010;12:2814–25.
Emerson D, Fleming EJ, McBeth JM. Iron-oxidizing bacteria: an environmental and genomic perspective. Annu Rev Microbiol. 2010;64:561–83.
Emerson D, Field EK, Chertkov O, Davenport KW, Goodwin L, Munk C, et al. Comparative genomics of freshwater Fe-oxidizing bacteria: implications for physiology, ecology, and systematics. Front Microbiol. 2013;4:254.
Fabisch M, Beulig F, Akob DM, Küsel K. Surprising abundance of Gallionella-related iron oxidizers in creek sediments at pH 4.4 or at high heavy metal concentrations. Front Microbiol. 2013;4:390.
Fleming EJ, Cetinić I, Chan CS, Whitney King D, Emerson D. Ecological succession among iron-oxidizing bacteria. ISME J. 2014;8:804–15.
Byrne JM, van der Laan G, Figueroa AI, Qafoku O, Wang C, Pearce CI, et al. Size dependent microbial oxidation and reduction of magnetite nano- and micro-particles. Sci Rep. 2016;6:1–13.
Byrne JM, Klueglein N, Pearce C, Rosso KM, Appel E, Kappler A. Redox cycling of Fe(II) and Fe(III) in magnetite by Fe-metabolizing bacteria. Science. 2015;347:1473–6.
Braunschweig J, Bosch J, Meckenstock RU. Iron oxide nanoparticles in geomicrobiology: from biogeochemistry to bioremediation. N. Biotechnol. 2013;30:793–802.
Bosch J, Heister K, Hofmann T, Meckenstock RU. Nanosized iron oxide colloids strongly enhance microbial iron reduction. Appl Environ Microbiol. 2010;76:184–9.
Küsel K, Blöthe M, Schulz D, Reiche M, Drake HL. Microbial reduction of iron and porewater biogeochemistry in acidic peatlands. Biogeosci Discuss. 2008;5:2165–96.
Marsili E, Baron DB, Shikhare ID, Coursolle D, Gralnick JA, Bond DR. Shewanella secretes flavins that mediate extracellular electron transfer. Proc Natl Acad Sci USA. 2008;105:3968–73.
Royer RA, Burgos WD, Fisher AS, Unz RF, Dempsey BA. Enhancement of biological reduction of hematite by electron shuttling and Fe(II) complexation. Environ Sci Technol. 2002;36:1939–46.
Beckwith CR, Edwards MJ, Lawes M, Shi L, Butt JN, Richardson DJ, et al. Characterization of MtoD from Sideroxydans lithotrophicus: a cytochrome c electron shuttle used in lithoautotrophic growth. Front Microbiol. 2015;6:332.
Hartshorne RS, Reardon CL, Ross D, Nuester J, Clarke TA, Gates AJ, et al. Characterization of an electron conduit between bacteria and the extracellular environment. Proc Natl Acad Sci USA. 2009;106:22169–74.
White GF, Shi Z, Shi L, Wang Z, Dohnalkova AC, Marshall MJ, et al. Rapid electron exchange between surface-exposed bacterial cytochromes and Fe(III) minerals. Proc Natl Acad Sci USA. 2013;110:6346–51.
Venkateswaran K, Moser DP, Dollhopf ME, Lies DP, Saffarini DA, MacGregor BJ, et al. Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp. nov. Int J Syst Bacteriol. 1999;49:705–24.
Myers CR, Nealson KH. Bacterial manganese reduction and growth with manganese oxide as the sole electron acceptor. Science. 1988;240:1319–21.
Myers CR, Nealson KH. Respiration-linked proton translocation coupled to anaerobic reduction of manganese(IV) and iron(III) in Shewanella putrefaciensMR-1. J Bacteriol. 1990;172:6232–8.
McBeth JM, Little BJ, Ray RI, Farrar KM, Emerson D. Neutrophilic iron-oxidizing ‘Zetaproteobacteria’ and mild steel corrosion in nearshore marine environments. Appl Environ Microbiol. 2011;77:1405–12.
Mori JF, Ueberschaar N, Lu S, Cooper RE, Pohnert G, Küsel K. Sticking together: inter-species aggregation of bacteria isolated from iron snow is controlled by chemical signaling. ISME J. 2017;11:1075–86.
Tamura H, Goto K, Yotsuyanagi T, Nagayama M. Spectrophotometric determination of iron(II) with 1,10-phenanthroline in the presence of large amounts of iron(III). Talanta. 1974;21:314–8.
Cooper RE, Wegner C-E, McAllister SM, Shevchenko O, Chan CS, Küsel K. Draft genome sequence of Sideroxydanssp. Strain CL21, an Fe(II)-oxidizing bacterium. Microbiol Resour Announc. 2020;9:1–2.
Wegner C-E, Gaspar M, Geesink P, Herrmann M, Marz M, Küsel K. Biogeochemical regimes in shallow aquifers reflect the metabolic coupling of the elements nitrogen, sulfur, and carbon. Appl Environ Microbiol. 2019;8:1–19.
Andrews S. FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc..40.
Bushnell B. BBMap short read aligner. https://www.sourceforge.net/projects/bbmap/..41.
Kopylova E, Noé L, Touzet H, Noe L, Touzet H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics. 2012;28:3211–7.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–6.
Burge SW, Daub J, Eberhardt R, Tate J, Barquist L, Nawrocki EP, et al. Rfam 11.0: 10 years of RNA families. Nucleic Acids Res. 2013;41:D226–32.
Heidelberg JF, Paulsen IT, Nelson KE, Gaidos EJ, Nelson WC, Read TD, et al. Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis. Nat Biotechnol. 2002;20:1118–23.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAM tools. Bioinformatics. 2009;25:2078–9.
Liao Y, Smyth GK, Shi W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 2013;41:e108.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30.
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2018.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Tautenhahn R, Patti GJ, Rinehart D, Siuzdak G. XCMS Online: a web-based platform to process untargeted metabolomic data. Anal Chem. 2012;84:5035–9.
Stettin D, Poulin RX, Pohnert G. Metabolomics benefits fom orbitrap GC-MS—Comparison of low- and high-resolution GC-MS. Metabolites. 2020;10:1–16.
Chong J, Yamamoto M, Xia J. MetaboAnalystR 2.0: from raw spectra to biological insights. Metabolites. 2019;9:1–10.
Hummel J, Strehmel N, Bölling C, Schmidt S, Walther D, Kopka J. Mass Spectral search and analysis using the golm metabolome database. In: Weckwerth W, Kahl G (eds). The handbook of plant metabolomics. 2013. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, Germany, p. 321–43.
Lueder U, Druschel G, Emerson D, Kappler A, Schmidt C. Quantitative analysis of O2 and Fe2+ profiles in gradient tubes for cultivation of microaerophilic Iron(II)-oxidizing bacteria. FEMS Microbiol Ecol. 2018;94:1–15.
Lefevre E, Bossa N, Wiesner MR, Gunsch CK. A review of the environmental implications of in situ remediation by nanoscale zero valent iron (nZVI): behavior, transport and impacts on microbial communities. Sci Total Environ. 2016;565:889–901.
Kirschling TL, Gregory KB, Minkley EG Jr, Lowry GV, Tilton RD. Impact of nanoscale zero valent iron on geochemistry and microbial populations in trichloroethylene contaminated aquifer materials. Environ Sci Technol. 2010;44:3474–80.
Wu S, Cajthaml T, Semerád J, Filipová A, Klementová M, Skála R, et al. Nano zero-valent iron aging interacts with the soil microbial community: a microcosm study. Environ Sci: Nano. 2019;6:1189–206.
Auffan M, Rose J, Wiesner MR, Bottero J-Y. Chemical stability of metallic nanoparticles: a parameter controlling their potential cellular toxicity in vitro. Environ Pollut. 2009;157:1127–33.
Anza M, Salazar O, Epelde L, Alkorta I, Garbisu C. The application of nanoscale zero-valent iron promotes soil remediation while negatively affecting soil microbial biomass and activity. Front Environ Sci. 2019;7:1–6.
Friedrich B, Magasanik B. Enzymes of agmatine degradation and the control of their synthesis in Klebsiella aerogenes. J Bacteriol. 1979;137:1127–33.
Kurihara S, Oda S, Kato K, Kim HG, Koyanagi T, Kumagai H, et al. A novel putrescine utilization pathway involves gamma-glutamylated intermediates of Escherichia coli K-12. J Biol Chem. 2005;280:4602–8.
Hädrich A, Taillefert M, Akob DM, Cooper RE, Litzba U, Wagner FE, et al. Microbial Fe(II) oxidation by Sideroxydans lithotrophicus ES-1 in the presence of Schlöppnerbrunnen fen-derived humic acids. FEMS Microbiol Ecol. 2019;95:1–19.
Cooper RE, Eusterhues K, Wegner C-E, Totsche KU, Küsel K. Ferrihydrite-associated organic matter (OM) stimulates reduction by Shewanella oneidensis MR-1 and a complex microbial consortia. Biogeosciences. 2017;14:5171–88.
Liu J, Wang Z, Belchik SM, Edwards MJ, Liu C, Kennedy DW, et al. Identification and characterization of MtoA: a decaheme c-type cytochrome of the neutrophilic Fe(II)-oxidizing bacterium Sideroxydans lithotrophicus ES-1. Front Microbiol. 2012;3:37.
DiChristina TJ, Moore CM, Haller CA. Dissimilatory Fe(III) and Mn(IV) reduction by Shewanella putrefaciens requires ferE, a homolog of the pulE (gspE) type II protein secretion gene. J Bacteriol. 2002;184:142–51.
Meshulam-Simon G, Behrens S, Choo AD, Spormann AM. Hydrogen metabolism in Shewanella oneidensis MR-1. Appl Environ Microbiol. 2007;73:1153–65.
Shi L, Belchik SM, Plymale AE, Heald S, Dohnalkova AC, Sybirna K, et al. Purification and characterization of the [NiFe]-hydrogenase of Shewanella oneidensis MR-1. Appl Environ Microbiol. 2011;77:5584–90.
Vignais PM, Billoud B, Meyer J. Classification and phylogeny of hydrogenases. FEMS Microbiol Rev. 2001;25:455–501.
Reiche M, Torburg G, Küsel K. Competition of Fe(III) reduction and methanogenesis in an acidic fen. FEMS Microbiol Ecol. 2008;65:88–101.
Reiche M, Haedrich A, Lischeid G, Kuesel K, Hädrich A, Lischeid G, et al. Impact of manipulated drought and heavy rainfall events on peat mineralization processes and source-sink functions of an acidic fen. J Geophys Res-Biogeosci. 2009;114:1–13.
Hädrich A, Heuer VB, Herrmann M, Hinrichs K-U, Küsel K. Origin and fate of acetate in an acidic fen. FEMS Microbiol Ecol. 2012;81:339–54.
Hamberger A, Horn MA, Dumont MG, Murrell JC, Drake HL. Anaerobic consumers of monosaccharides in a moderately acidic fen. Appl Environ Microbiol. 2008;74:3112–20.
Wüst PK, Horn MA, Drake HL. Trophic links between fermenters and methanogens in a moderately acidic fen soil. Environ Microbiol. 2009;11:1395–409.
Tabor CW, Tabor H. Polyamines. Annu Rev Biochem. 1984;53:749–90.
Karatan E, Watnick P. Signals, regulatory networks, and materials that build and break bacterial biofilms. Microbiol Mol Biol Rev. 2009;73:310–47.
Bachrach U, Heimer YM. The physiology of polyamines. 1989. CRC Press Taylor and Francis Group, Boca Raton, FL, USA.
Karatan E, Duncan TR, Watnick PI. NspS, a predicted polyamine sensor, mediates activation of Vibrio cholerae biofilm formation by norspermidine. J Bacteriol. 2005;187:7434–43.
Cockerell SR, Rutkovsky AC, Zayner JP, Cooper RE, Porter LR, Pendergraft SS, et al. Vibrio cholerae NspS, a homologue of ABC-type periplasmic solute binding proteins, facilitates transduction of polyamine signals independent of their transport. Microbiology. 2014;160:832–43.
Matthysse AG, Yarnall HA, Young N. Requirement for genes with homology to ABC transport systems for attachment and virulence of Agrobacterium tumefaciens. J Bacteriol. 1996;178:5302–8.
Sauer K, Camper AK. Characterization of phenotypic changes in Pseudomonas putida in response to surface-associated growth. J Bacteriol. 2001;183:6579–89.
Patel CN, Wortham BW, Lines JL, Fetherston JD, Perry RD, Oliveira MA. Polyamines are essential for the formation of plague biofilm. J Bacteriol. 2006;188:2355–63.
Capdevila DA, Wang J, Giedroc DP. Bacterial strategies to maintain zinc metallostasis at the host–pathogen interface. J Biol Chem. 2016;291:20858–68.
Schoepp-Cothenet B, van Lis R, Philippot P, Magalon A, Russell MJ, Nitschke W. The ineluctable requirement for the trans-iron elements molybdenum and/or tungsten in the origin of life. Sci Rep. 2012;2:263.
Reda T, Plugge CM, Abram NJ, Hirst J. Reversible interconversion of carbon dioxide and formate by an electroactive enzyme. Proc Natl Acad Sci USA. 2008;105:10654–8.
Hartmann T, Schwanhold N, Leimkühler S. Assembly and catalysis of molybdenum or tungsten-containing formate dehydrogenases from bacteria. Biochim Biophys Acta. 2015;1854:1090–1100.
Ilbert M, Bonnefoy V. Insight into the evolution of the iron oxidation pathways. Biochim Biophys Acta. 2013;1827:161–75.
Lovley DR, Holmes DE, Nevin KP. Dissimilatory Fe(III) and Mn(IV) reduction. Adv Micro Physiol. 2004;49:219–86.
Melton ED, Swanner ED, Behrens S, Schmidt C, Kappler A. The interplay of microbially mediated and abiotic reactions in the biogeochemical Fe cycle. Nat Rev Microbiol. 2014;12:797–808.
Maisch M, Lueder U, Laufer K, Scholze C, Kappler A, Schmidt C. Contribution of microaerophilic Iron(II)-oxidizers to Iron(III) mineral formation. Environ Sci Technol. 2019;53:8197–204.
Hong Y, Wu J, Wilson S, Song B. Vertical stratification of sediment microbial communities along geochemical gradients of a subterranean estuary located at the Gloucester Beach of Virginia, United States. Front Microbiol. 2018;9:3343.
Liptzin D, Silver WL. Spatial patterns in oxygen and redox sensitive biogeochemistry in tropical forest soils. Ecosphere. 2015;6:art211.
Borer B, Tecon R, Or D. Spatial organization of bacterial populations in response to oxygen and carbon counter-gradients in pore networks. Nat Commun. 2018;9:769.
Widder S, Allen RJ, Pfeiffer T, Curtis TP, Wiuf C, Sloan WT, et al. Challenges in microbial ecology: building predictive understanding of community function and dynamics. ISME J. 2016;10:2557–68.
Frey PA, Reed GH. The ubiquity of iron. ACS Chem Biol. 2012;7:1477–81.
Edwards KJ, Bach W, McCollom TM, Rogers DR. Neutrophilic iron-oxidizing bacteria in the ocean: their habitats, diversity, and roles in mineral deposition, rock alteration, and biomass production in the deep-sea. Geomicrobiol J. 2004;21:393–404.
Bondici VF, Khan NH, Swerhone GDW, Dynes JJ, Lawrence JR, Yergeau E, et al. Biogeochemical activity of microbial biofilms in the water column overlying uranium mine tailings. J Appl Microbiol. 2014;117:1079–94.
Lee AK, Newman DK. Microbial iron respiration: impacts on corrosion processes. Appl Microbiol Biotechnol. 2003;62:134–9.
Glasser NR, Saunders SH, Newman DK. The colorful world of extracellular electron shuttles. Annu Rev Microbiol. 2017;71:731–51.
Philips J, Verbeeck K, Rabaey K, Arends JBA. Electron transfer mechanisms in biofilms. In: Scott K, Yu EH (eds). Microbial electrochemical and fuel cells. 2016. Woodhead Publishing, Sawston, Cambridge, United Kingdom, p. 67–113.
Gao L, Lu X, Liu H, Li J, Li W, Song R, et al. Mediation of extracellular polymeric substances in microbial reduction of hematite by Shewanella oneidensis MR-1. Front Microbiol. 2019;10:575.
Roden EE, McBeth JM, Blöthe M, Percak-Dennett EM, Fleming EJ, Holyoke RR, et al. The microbial ferrous wheel in a neutral pH groundwater seep. Front Microbiol. 2012;3:172.
Acknowledgements
This study was part of the Collaborative Research Centre Chemical Mediators in Complex Biosystems (CRC 1127 ChemBioSys) of the Friedrich Schiller University Jena, funded by the Deutsche Forschungsgemeinschaft. We gratefully acknowledge the Fonds der Chemischen Industrie. Additional financial support was provided by the German Centre for Integrative Biodiversity Research (iDiv) Halle–Jena–Leipzig funded by the Deutsche Forschungsgemeinschaft and by the state of Thuringia (2015 FGI 0021) with means of the EU in the framework of the EFRE program. RXP acknowledges funding by the Alexander von Humboldt Foundation. Open access funding provided by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
KK conceived the study. REC, KK, and GP designed the experiments. REC, SK, NU, and JDW performed the experiments. REC, CEW, KK, SK, GP, RXP, DS, NU, and TW performed the data analyses and/or helped interpret the results. REC wrote the manuscript, with contributions from CEW, KK, SK, GP, NU, and RXP. All co-authors critically reviewed the manuscript and approved the final version.
Corresponding author
Ethics declarations
Conflict of interest
The research presented in this manuscript was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cooper, R.E., Wegner, CE., Kügler, S. et al. Iron is not everything: unexpected complex metabolic responses between iron-cycling microorganisms. ISME J 14, 2675–2690 (2020). https://doi.org/10.1038/s41396-020-0718-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-0718-z
This article is cited by
-
Characterization and environmental applications of soil biofilms: a review
Environmental Chemistry Letters (2024)
-
Pulsed electric field treatment of seeds altered the endophytic bacterial community and promotes early growth of roots in buckwheat
BMC Microbiology (2023)
-
Redox signaling-driven modulation of microbial biosynthesis and biocatalysis
Nature Communications (2023)
-
Integrating microbial DNA community analyses into time-integrated suspended sediment sampling methods
Journal of Soils and Sediments (2023)