Abstract
Vertebrates’ gut microbial communities can be altered by the hosts’ parasites. Helminths inhabiting the gut lumen can interact directly with their host’s microbiota via physical contact, chemical products, or competition for nutrients. Indirect interactions can also occur, for instance when helminths induce or suppress host immunity in ways that have collateral effects on the microbiota. If there is genetic variation in host immune responses to parasites, we would expect such indirect effects to be conditional on host genotype. To test for such genotype by infection interactions, we experimentally exposed Gasterosteus aculeatus to their naturally co-evolved parasite, Schistocephalus solidus. The host microbiota differed in response to parasite exposure, and between infected and uninfected fish. The magnitude and direction of microbial responses to infection differed between host sexes, and also differed between variants at autosomal quantitative trait loci. These results indicate that host genotype and sex regulate the effect of helminth infection on a vertebrate gut microbiota. If this result holds in other taxa, especially humans, then helminth-based therapeutics for dysbiosis might need to be tailored to host genotype and sex.
Similar content being viewed by others
Introduction
Helminth infection is often associated with changes in hosts’ gut microbiota [1, 2]. For example, Hymenolepis diminuta infection reduces Bacilli abundance in mammalian guts, in the lab and wild [3, 4]. Such infection-induced changes interest both evolutionary ecologists and biomedical researchers. For ecologists, host-parasite–microbiota interactions represent a compelling example of multi-way symbiotic interactions. From an evolutionary view, these multi-way interactions are likely to affect fitness of all taxa concerned, driving multi-way coevolution. Biomedically, the modern elimination of helminths from many human populations has been potentially linked to an increase in dysbiosis and inflammatory bowel diseases [5, 6].
Despite widespread interest in host-parasite–microbiota interactions, results have been inconsistent. Some studies report no effects of infection on microbiota [7, 8]. Other studies find the gut microbiota protects its host against helminths [9, 10]. For example, Trichuris muris infection altered the gut microbiota in mice in ways that inhibited subsequent reinfection [11]. Other studies find microbiome facilitation of parasite infection [12, 13]. These inconsistent findings of host-helminth–microbiota studies suggest there exist additional variables that modulate the outcome of these interactions. Identifying these other variables may facilitate treatment of dysbiosis or helminthiasis [14,15,16].
The mechanisms underlying helminth–microbiota interactions are still being elucidated. Macroparasites can interact directly with the gut microbiota by secreting antibacterial peptides [17]. Parasites can also indirectly alter microbiota via changes in host physiology and immunity. H. polygyrus bakeri infection induces colonic regulatory T-cells in mice [18], which can protect mice from colitis [19]. H. polygyrus also negatively regulates host intestinal mucosal IL-22 and IL-17, decreasing host expression of epithelial antimicrobial peptides [20]. Nonimmune indirect effects also occur: helminth damage to the host gut epithelium can cause malnutrition that changes the microbiota [21].
Direct and indirect between helminths and microbiota can be difficult to disentangle. One strategy to focus on indirect effects is to study parasites living outside the gut, physically isolated from the gut microbiota (although secreted products might still diffuse into the gut lumen). Another strategy is to test whether host genotype modulates the helminth effect on microbiota. Direct interactions between parasites and microbes should primarily reflect parasite and bacterial genotypes, and be relatively insensitive to host genotype. In contrast, indirect interactions acting through host traits should be sensitive to host genetic variation in those intermediary traits. Hosts immune response to helminths varies due to immunogenetic polymorphism [22, 23], body condition [24, 25], and past infections [26]. Males and females within a species also respond differently to infections [27], because sex hormones modulate immune traits [28]. Until now, we lack evidence that genetic variation in host immune response to parasites also alters the microbiota. Therefore, we tested whether host genotype modulates the effect of parasite infection on the microbiota, which would imply an indirect effect via host traits.
We experimentally exposed laboratory-bred threespine stickleback (Gasterosteus aculeatus) fish to their native cestode parasite (Schistocephalus solidus). We assayed the gut microbial response to infection and host genotype, and nonadditive interactions between infection and genotype. This host-parasite system is especially valuable for such studies because the microbiota have little opportunity for direct interaction with the cestode. After the fish ingests an infected copepod (the first host), within hours the parasite penetrates the intestinal wall, establishing a long-term infection in the peritoneal cavity physically separated from the gut microbiota. Although we cannot rule out direct effects of helminth secretory/excretory products diffusing into the gut lumen, the most parsimonious explanation is indirect effects via host phenotypes. Stickleback populations exhibit genetically divergent immune responses to cestode infection, so host genotype should modify indirect effects of the cestode on the microbiota. In an initial exposure experiment (Experiment 1), we show that cestode exposure suffices to alter the gut microbiota, even when the infection subsequently fails, but this effect varied among host full-sibling families. In Experiment 2, we show that infection presence/absence also alters the gut microbiota (all the fish were exposed to cestodes, but some infections failed), but this effect varies between host sexes and autosomal genotypes. This result demonstrates that host genetic variation modifies an indirect interaction between helminth and gut microbiota.
Materials and methods
Stickleback breeding
Gosling Lake and Roberts Lake on Vancouver Island, British Columbia contain stickleback populations that differ in immune phenotypes, parasite growth rates, and cestode prevalence in the wild (>70% and 0%, respectively) [29, 30]. We generated pure Roberts (ROB), pure Gosling (GOS), and reciprocal F1 hybrid families (RG or GR), and raised these to maturity in the lab. We bred these families to generate second generation (F2) hybrids including intercrosses (F1 × F1 matings) and reciprocal backcrosses (ROB × F1, F1 × ROB, GOS × F1, and F1 × GOS). All fish were reared in freshwater conditions and housed with full-siblings in either 40-L or 10-L tanks at the University of Texas at Austin., with IACUC approval (protocol AUP-2015-00200).
Parasite collection and experimental exposures
As a pilot study (Experiment 1), we evaluated the effects of cestode exposure, and successful infection, on the host gut microbiota. We experimentally fed S. solidus to adult pure-bred GOS fish from six full-sib families. Within each family, five fish were controls fed uninfected copepods (sham exposure), while the rest received infected copepods. This design yielded three categories of fish per family: unexposed controls, exposed-but-uninfected controls, and infected fish (N = 30, Supplementary Fig. 1a and Supplementary Table 1). The infection exposures followed a standard procedure described by Weber et al. [29]. Briefly, we dissected mature S. solidus out of infected wild-caught fish from Gosling, and bred the tapeworms in culture media in dark waterbath (mimicking the interior of bird guts). We hatched the tapeworm eggs and fed them to copepods (Macrocyclops albidus). After 24 h fasting to induce hunger, each fish was put in an individual tank with copepods that we visually confirmed carried the cestode, and then the fish and strained the aquarium water was removed 12 h postfeeding to confirm there was zero remaining copepods uneaten (the parasite passed into all fish guts). Forty-three days after parasite exposure, we euthanized fish with MS-222 and dissected the fish to visually inspect for the presence of successful tapeworms. These are large macroparasites that can grow to be greater than 50% the body mass of their host, so infection is easily confirmed. We froze the entire intestine in ethanol for subsequent analysis.
In Experiment 2, we focus instead on host genotype control of the gut microbiota, and its response to infection. We experimentally exposed adult F2 hybrid fish (intercross and backcross, N = 711, Supplementary Fig. 1b and Supplementary Table 2) to S. solidus, then assayed infection outcomes and sequenced the gut microbiota. All fish from this experiment were exposed to S. solidus, but not all were infected successfully, providing a contrast between fish with versus without a successfully established parasite, controlling for initial exposure history. We seek to specifically examine the effect of cestode presence, physically isolated in the body cavity, on the microbiota in the gut lumen, compared with fish where the cestode was absent, controlling for any direct effect arising from the tapeworm’s brief passage through the gut. If we compared infected fish to unexposed controls, we would be confounding the (likely indirect) effects of an established cestode in the body cavity, with the (potentially direct) effects of a transient cestode passing through the gut (assayed in Experiment 1). By focusing our statistical power on the comparison of infected versus exposed-but-uninfected fish, we control for that initial transient phase. Thus, the inferences we draw can be more confidently attributed to the effect of an established tapeworm that is physically separate from the microbiota. Moreover, the success rate of experimental infections is low for lake fish (often ~20%), so we needed to expose all available fish in the lab crosses, to maximize the number of successfully infected fish.
Amplification, sequencing, and analysis of 16S rRNA amplicons
We extracted DNA from the entire intestine (both content and mucosa) of stickleback (N = 30 for Experiment 1; N = 693 for Experiment 2) using the MoBio Powersoil DNA Isolation Kits, as previously described [31, 32]. For Experiment 1, we used the V4–V5 primers [33]. For Experiment 2, 16S rRNA amplicons were generated for the V4 hypervariable region [31]. Sample-specific barcodes, PCR amplification, and test for contamination were used as previously described [31]. Amplicon pools were paired-end sequenced on an IlluminaMiseq platform at GSAF (Genomic Sequencing and Analysis Facility, University of Texas at Austin).
Paired-end reads were demultiplexed, then processed using the mothur software package (v.1.39.1), following standard procedures [34, 35] (https://www.mothur.org/wiki/MiSeq_SOP). Briefly, sequences were trimmed for 16S rRNA gene primer sequences, then assembled into contigs and aligned with 16S rRNA gene sequences from the ARB Silva v128 reference database [36]. Chimeric sequences were removed using VSEARCH within mothur. The remaining sequences were classified using a Bayesian classifier with a training set (version 16) from the Ribosomal Database Project (http://rdp.cme.msu.edu) [37]. Operational Taxonomic Units (OTUs) were identified using the UCLUST algorithm based on 97% similarity. We rarefied the data to 2000/4000 sequences per sample to calculate unweighted/weighted UniFrac distance and PCoA scores by phyloseq library in R [38].
Analyses of pure GOS fish experimentally exposed to S. solidus or a negative control
We used mixed linear models to analyze effects of family (random effect) and infection status (fixed effect) on unweighted/weighted PCoA axis 1, with a random family × infection interaction. First, we contrasted unexposed (sham infections) versus exposed fish (regardless of the outcome of exposure). Second, we compared uninfected fish (sham or failed infections) versus infected fish. For a retrospective power analysis, we drew 10,000 randomly generated datasets with increasing effect size (difference in PCoA 1) given our sample size, family structure, and observed data variance, and calculated the proportion of simulations in which we were able to accurately reject the null hypothesis of no effect of infection versus failed exposure.
Analyses of F2 hybrid fish experimentally exposed to S. solidus
General linear model analyses
We used quasibinomial general linear models (GLMs), implemented in R, to examine the effects of infection and host genotype on microbial composition in Experiment 2. We focus on the relative abundance of each commonly observed Order or Family, arbitrarily set to be those taxa found in at least 20 fish [N = 84 Orders, 181 Families]. Previous analysis of stickleback gut microbiota (from the laboratory and wild populations) found that most microbe taxa exhibited significantly overdispersed read counts, relative to a binomial distribution, that was better fit with a quasibinomial [31, 32]. For each taxon, we estimated a quasibinomial GLM in which the fraction of reads attributed to that taxon (out of all reads) depended on effects of host cross (backcross or intercross), sex, mas, and S. solidus presence/absence, as well as key interactions: infection × cross, infection × sex, and infection × mass. Fish were reared in two rooms, which we used as a covariate. We used sequential Bonferroni correction when evaluating P values. For a random subset of taxa, we confirmed that the quasibinomial fit better than the binomial, as described by Bolker [39].
Discriminant function analyses
To test the whole-microbiota effects of infection, sex, and cross direction, we applied linear discriminant analysis (LDA) to the top 50 weighted/unweighted microbial PCoA axes (which cumulatively account for 99.99% of variance in microbial alpha diversity). We first used LDA to distinguish four groups (combinations of sex and infection status). Then, we used LDA to distinguish six groups (factorial combinations of three host cross types and infection status). We used ANOVAs to test for effects of sex, cross, and infection status on each LDA axis. We also confirmed these statistical effects using a MANOVA applied directly to the top 50 PCoA axes to test for effects of sex, cross, infection, host mass, and interactions among these variables.
QTL mapping: ddRAD genotyping
We used a Promega Wizard SV 96-well plate kit to extract DNA from alcohol-preserved fin clips from all intercross and backcross hybrids, and their grandparents. We genotyped them to obtain SNPs for quantitative trait locus (QTL) mapping, using the ddRADseq protocol described in Peterson et al. [40], using bioinformatics steps to identify SNPs described in Stuart et al. [41]. We retained SNPs exhibiting fixed differences between the Roberts and Gosling Lake grandparents (e.g., fully informative for QTL mapping).
QTL mapping: analyses
We mapped QTL for several microbiota measures: alpha diversity (using 2000/4000 read depth normalization), the top 10 weighted/unweighted PCoA axes, and the relative abundance of the common microbial Orders. We built linkage maps for each cross separately in R/qtl [42], and used the scanone function with “hk” interval mapping (using rank-based nonparametric tests for microbial Order relative abundance). To account for the complex cross design (with backcrosses and F2 intercrosses), we built separate QTL maps for each cross types, then summed their LOD scores. This yields one summary statistic measuring a locus’ association with the focal trait, accounting for between-cross differences in QTL effect size or marker linkage. We compared this summed LOD against null expectations obtained by 1000 permutations of the focal phenotype across fish within each cross, followed by reanalysis to generate null LOD scores. Conservatively, we consider an observed QTL significant when its summed LOD exceeded the 99.99% quantile from that marker’s null values at that same marker. We double-checked each significant QTL with a GLM testing for fish genotype effect on the focal microbiota variable.
Results
Effects of cestode exposure or infection
Experiment 1 showed that cestode-exposed fish (infected/uninfected) had different microbial communities than the sham-exposed control fish, though this trend varied among host full-sib families (Fig. 1a and Supplementary Table 3; fish family P < 0.0001, exposure P = 0.2589, exposure × family P = 0.0363). The significant exposure by family interaction occurs because one full-sib family showed an atypical microbial response to exposure. Omitting this one family, the remaining families all showed a consistent microbiota response to parasite exposure (exposure P = 0.0378, family P = 0.0579, exposure × family P = 0.1675). In contrast, we observed no difference between infected versus exposed-but-uninfected fish (Fig. 1b and Supplementary Table 3). However, this pilot experiment had modest statistical power to detect effects of successful infection (80% power to detect a simulated 1.2 standard deviation shift in PCoA1) whereas we observed only a nonsignificant 0.45 standard deviation shift.
For the purpose of plotting, microbiota community is measured here by scoring fish along the first linear discriminant axis trained on three groups of fish (control, exposed, and infected). Each point is an individual fish, color coded by the full-sibling family to which they belong. Lines connect family level means for control, exposed, or infected individuals. One family has an atypical response to exposure that generated a significant family by exposure interaction; without this family there is a significant exposure main effect. a microbiota community structure differs between stickleback that were experimentally exposed to S. solidus (“exposed”) versus control fish that were sham infected. b No significant effect of cestode infection success on experimentally exposed sticklebacks’ gut microbiota.
Gut microbial community composition
In Experiment 2, we observed 256 successful infections out of 711 cestode-exposed lab-raised adult F2 sticklebacks. We obtained 16S sequences for 693 of these fish (a few individuals’ intestines were not retained), yielding 11,586 OTUs. We retained an average of 11,066 sequence reads per fish (s.d. = 13,394; median = 7575; range from 48 to 126,943). We excluded all individual stickleback with fewer than 500 sequence reads (N = 43 removed, 650 retained, Supplementary Table 4). By using quasibinomial GLMs for analysis, we explicitly account for variation in read depth among individuals. A summary of the gut microbial community composition is provided in Supplementary Dataset 1. The five most abundant Phyla in the sticklebacks’ intestines were Firmicutes (61.07% of reads on average), Proteobacteria (28.68%), Actinobacteria (5.13%), Bacteroidetes (0.93%), and Planctomycetes (0.67%). As previously noted [31, 32], the stickleback gut microbiota differed dramatically among individuals (Supplementary Fig. 2), despite their similar ages, diet, and rearing facility.
Host sex, cross, and infection jointly affect the gut microbiota
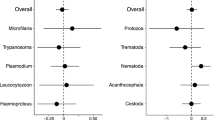
GLM analyses of the 84 most common microbial Orders (present in at least 20 fish) indicate that cestode infection and host genotype jointly affect the stickleback gut microbiota. All models included room effect as a covariate to account for the large effect of rearing room (Supplementary Fig. 3). Host sex, cross, and mass each had significant main effects on nearly a third of microbial Orders (32.1%, 35.7%, and 28.6%, respectively; Fig. 2; effect sizes and P values in Supplementary Dataset 2). However, main effects of cestode infection were relatively uncommon (only 7.1% of microbial Orders).
Here, we identify which microbial Orders exhibit various main and interaction effects. For each Order, a cell is colored if the relevant effect is significant at P < 0.05. Note that there is a strong excess of significant results at this threshold above the null expectation of a 5% false positive rate (χ2 tests P < 0.0001 for all but the mass × infection interaction). Generalized linear model results for all common Orders (found in at least 20 fish) are summarized in Supplementary Dataset 1. Color denotes an effect direction. For cross, gray denotes higher abundance with more Roberts Lake alleles (ROB backcross), black denotes higher abundance in Gosling Lake alleles (GOS backcross). For mass, gray denotes higher abundance in larger fish, black in smaller fish. For sex, gray denotes higher abundance in males, black higher in females. For infection, gray denotes higher abundance in infected than in uninfected fish, black is the reverse. For the cross × infection interaction, gray indicates instances where infection increases microbe abundance in ROB backcrosses while black indicates infection increases abundance in GOS. For mass × infection, gray indicates that infection increases microbe abundance more strongly in larger fish. For sex × infection, gray denotes cases where infection increases microbe abundance most strongly in males, whereas black implies infection increases microbe abundance mostly in females.
Although few microbial Orders exhibited a main effect of cestode infection (consistent with Experiment 1), substantially more Orders (33.3%) exhibit interactions between infection and other host traits. Specifically, 22.6% of microbial Orders exhibited significant infection × cross interactions (contrasting ROB backcross, F2 intercross, or GOS backcross), and 21.4% of Orders depend on infection × sex interactions. Fish mass had little effect on the cestode–microbe interaction: only 11.9% of microbial Orders exhibited a mass × infection interaction, significantly more than the 5% false positive rate (χ2 = 7.0, P = 0.008). Thus, the statistical main effect of infection (our focus in Experiment 1) underestimates the impacts of cestode presence, the majority of which were contingent on host genotype (sex, cross). Qualitatively similar trends were observed if we instead used other taxonomic ranks. Results for 181 common microbial Families are reported in Supplementary Fig. 4 and Dataset 2, also revealing numerous infection × cross and infection × sex interactions.
Examples of interactions between helminth infection and host genotype
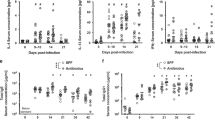
Figure 3 illustrates a few illustrative examples of microbial variation among hosts (see also Supplementary Fig. 5). We observed a strong main effect of infection for some microbe Orders, such as Fusobacteriales (Fig. 3a and Supplementary Fig. 5a, infection effect t = −2.65, P = 0.0083), which were less abundant in infected sticklebacks’ gut regardless of sex or genotype. As an example of a main effect of host sex, Lactobacillales were more abundant in male than in female stickleback regardless of infection status (Fig. 3b and Supplementary Fig. 5b, sex effect t = 3.37, P < 0.001).
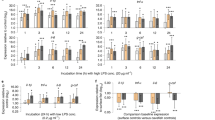
To visualize these effects we first adjusted for variation due to rearing location (room), by calculating residuals from a quasibinomial general linear model of the focal Order’s read counts (out of the total reads per fish) regressed on rearing room. Here we plot the mean (and ±1 s.e. confidence intervals) for this residual abundance, for various groups of fish to illustrate infection and host effects. a Fusobacteriales exhibit a significant decrease in cestode-infected fish, both males (black) and females (gray). b Lactobacillales are more abundant in males (black) than in females (gray) regardless of infection status. c Clostridiales abundance is reduced in infected males, but not infected females (black = males, gray = females). d Rhodobacterales are more common in females for uninfected fish, but more common in males for infected fish (black = males, gray = females). e Caulobacterales are more abundant in F2 hybrids than in either backcross, regardless of infection status (black = uninfected, gray = infected). f Rhodobacterales are more abundant in fish with a greater fraction of Roberts lake ancestry, but only among uninfected fish; infection leads to a higher Rhodobacterales abundance that is similar across fish genotypes (black = uninfected, gray = infected). Supplementary Fig. 5 shows the same plots, but with all data points included to show the small effect size relative to high among-individual variation.
Clostridiales illustrate the contingent effect of parasite infection (Fig. 3c and Supplementary Fig. 5c), with a sex main effect (t = 5.56, P < 0.0001) and a sex × infection interaction (t = −7.46, P < 0.0001). The relative abundance of Clostridiales was insensitive to infection in female hosts, but strongly reduced in S. solidus-infected males. Consequently, the relative abundance of Clostridiales differed strongly between uninfected males and females, but did not differ between infected males and females. Rhodobacterales also show a sex × infection interaction Fig. 3d and Supplementary Fig. 5d; their relative abundance increased with infection in males but decreased with infection in females. As a result, among uninfected fish Rhodobacterales were most abundant in females, whereas among infected fish this taxon was most abundant in males.
Host autosomal genotype also influenced the gut microbiota, as indicated by widespread differences between backcross versus F2 hybrids. Caulobacterales were significantly more common in F2 hybrids (Fig. 3e and Supplementary Fig. 5e, cross main effect F2,629 = 71.6, P < 0.0001) than in either backcross. Other Orders were more common in one particular cross, or exhibited an additive trend (F2s intercrosses being intermediate between backcrosses). This autosomal effect altered the impact of cestode infection. Rhodobacterales were more abundant in hosts with greater ancestry from Roberts Lake, but only in the absence of the cestode (Fig. 3f and Supplementary Fig. 5f). In contrast, in cestode-infected fish Rhodobacterales were uniformly common in all genotypes.
Discriminant function analyses of host cross and sex effects on microbiota
We next used multivariate analyses to evaluate the response of the overall gut microbiota community to infection and host genotype (Supplementary Table 5). LDA separating the four combinations of sex and infection confirmed that helminth infections alter the gut microbiota, but sex modifies the effects of helminthiasis (Fig. 4a). The two leading LDA axes, respectively, exhibited a significant effect of sex (F1,466 = 90.8, P < 0.0001), and an effect of infection (F1,466 = 4.64, P = 0.0317). For LDA axis 1 (LDA1), these variables also interacted (F1,466 = 9.6, P = 0.0021). Overall, cestode infection changed female gut microbiota composition more strongly than it changed male microbiota.
a Results of LDA to separate all combinations of sex and infection status. LDA axis 1 (LDA1) separates infected (filled points) from uninfected (open points; P < 0.0001). LDA2 separates males and females (blue and red points; P < 0.0001), and exhibits a significant interaction effect (P = 0.002). b Results of LDA to separate combinations of cross (blue = Gosling backcross, purple = F2, red = Roberts backcross) and infection status (open circles = uninfected, filled circles = infected). LDA1 (68% of variation) separates F2 hybrids from both backcrosses. LDA2 (16% of variation; not shown here) separates Roberts from Gosling backcross fish, with F2 hybrids being intermediate. LDA3 explains 7% of the microbial variation and is most strongly associated with infection status, but in a manner that depends on host cross: the three host crosses are on average almost identical along LDA3 when uninfected (open circles), but diverge when infected with F2s intermediate as expected from additive genetic control. Raw points are shown in faded colors, overlain by larger darker circles representing bivariate averages with ± 1 s.e. error bars.
A separate discriminant function analysis of fish cross type and infection status (six combinations) revealed several insights. The first LDA axis separated F2 hybrid from both reciprocal backcross populations (Fig. 4b), suggesting that there may be transgressive genetic effects on the microbiota (e.g., Fig. 3e). F2 hybrids were intermediate between the backcrosses along LDA2, consistent with additive genetic control of other aspects of the microbiota (Supplementary Fig. 6). The third axis separated infected and uninfected fish, but in a highly genotype-dependent manner (Fig. 4b and Supplementary Fig. 6). In F2 intercross fish, LDA3 scores were insensitive to infection, whereas in both backcross populations the LDA3 scores changed strongly in response to infection. All three crosses had similar LDA3 scores when uninfected, but Gosling and Roberts backcross fishes’ microbiota diverged in response to infection.
QTL mapping of gut microbiota composition
Based on the effects of cross type, we expected to be able to locate host loci that explain variation in gut microbiota composition. We also expected these QTL to differ in effect between infected versus uninfected fish. ddRADseq identified 234 SNPs with fixed differences between the Roberts and Gosling populations and sufficiently deep coverage within and among the F2 hybrid individuals, yielding approximately ten markers per linkage group. Another paper will describe QTL mapping of infection outcomes and host immune traits (Weber et al. 2020, manuscript); here we focus only on mapping the gut microbiota. We located no significant QTL for any of the top 10 weighted PCoA axes, and no significant QTL for microbial diversity. But, we did detect weak-effect QTL for unweighted PCoA axis 5, with three QTLs that narrowly exceeded our stringent threshold for significance (Supplementary Fig. 7a–c). We had a stronger signal when we combined the many PCoA axes into a single metric using LDA (trained to distinguish the two backcrosses, then applied to all samples). We detected a single well-supported QTL on Chr9 (Supplementary Fig. 7d, e) that affects microbial community as measured by the LDA axis scores. Because QTL mapping was done within each cross, then summed, using LDA to define an axis that distinguishes between crosses is not tautological. Chromosome 9 does not have any significant QTL effect on cestode infection success (Weber et al. 2020).
The lack of strong QTL for microbial ordination metrics led us to hypothesize that host control of the whole microbial community is highly polygenic. If host genetic variation acts on particular microbe taxa, it might act only weakly on PCoA scores and be correspondingly hard to map. So, we next mapped microbial Orders separately, revealing numerous taxon-specific QTL. Qualitatively similar results were obtained mapping microbial Families. To illustrate, Fusobacterales exhibit two strong autosomal QTL, plus an association with the X chromosome (ChrX) indicating a sex effect (Fig. 5). Other examples are plotted in Supplementary figures (Supplementary Fig. 8), and their overlapping distributions in the genome are presented in Fig. 6. This summary reveals genomic “hotspots” for QTL affecting the microbiota, on Chr1, Chr2, and Chr3. Many of these microbial Orders also mapped to the sex chromosome (Chr19), consistent with the common main effect of sex.
Analyses were run separately for each cross, with rearing room as an additive covariate, then the LOD scores from the three maps were summed. The observed summed LOD scores are plotted in black for each linkage group, measuring statistical association between the focal trait and the chromosomal region. Marker locations are indicated as tick marks along the horizontal axis. Thin gray lines represent null summed LOD scores from within-cross permutations of traits. The horizontal dashed line indicates the upper 99.99% quantile for the null LOD scores. Three QTL exceed this threshold, on chromsomes 4, 9, and X. Two of these are plotted in the lower panels (left, locus X109 on Chr 4, genotype P = 0.0013; right, locus X156 on Chr9, genotype P = 0. 0060). The y-axis in these effect plots are the residuals from a regression of unweighted PCoA5 on rearing room. We plot the means microbial abundance with ± 1 standard error bars, for each of the three genotypes at each locus.
Each chromosome is plotted as a vertical gray bar, and to its right we plot microbial QTL located on that chromosome. A dot symbol indicates the location of maximum LOD score for each QTL, and thin vertical lines indicate the inferred width of the QTL. For sex-linked microbes, their QTL span the entire X chromosome, so we omit that linkage group. The key to the right lists the locations of the significant QTL for each focal taxon.
We next tested whether these QTL are contingent on cestode presence/absence. We mapped QTL separately for infected fish and for uninfected fish, and then looked for loci with QTL in only one of these groups. We found numerous host loci that are only associated with microbial variation in uninfected fish, whereas the same locus is unrelated to the microbiota among infected fish (e.g., Supplementary Fig. 9a–f). In fewer instances, we identified host QTL for microbiota in infected fish only (Supplementary Fig. 9g, h), though because fewer fish were infected our power in this group is lower. In several cases, stratifying by cestode infection revealed QTL that would not have been detected otherwise. For example, Solirubrobacterales (Supplementary Fig. 9e, f) have a QTL on Chr3 that is observed in all fish, but which in fact only acts on the uninfected fish (a larger sample size). When we ignored infection status, the same microbial taxon had no significant QTL, because the lack of Solirubrobacterales in infected fish was masking the effect of host genetic variation at that site.
Discussion
The interaction between hosts, parasites, and gut microbes represents a rich opportunity for experimental studies of multi-way ecological interactions. A growing literature suggests that helminth infection can alter vertebrates’ gut microbiota [1, 43]. Our experiment adds an important twist: the magnitude and direction of helminth-induced microbial changes depend on both the sex of the host, and the host’s autosomal genotype. By examining F2 hybrids between two recently diverged host populations, we identified autosomal loci that contribute to variation in microbiota composition, as well as variation in microbiota response to helminth infection. The implication is that genetic variation within one species (the host) alters the ecological effect of another species (the cestode parasite) on a third party (the microbial community). It remains to be seen whether this holds true in other hosts, particularly in humans. Previous findings of sex × diet interactions on stickleback gut microbiota have been widely confirmed in other hosts including humans [31].
If our results hold in more taxa, they would have an inconvenient implication: biologists may not readily generalize helminth-microbe effects beyond the genotypes (and sex) being studied. But, this effect also presents an opportunity: host genetic variation might help us identify the mechanisms by which helminths alter the gut microbiota (or, vice versa). Doing so may allow us explain why individual hosts differ in their microbial response to infection.
To date, many studies have evaluated effects of helminth infection on animals’ gut microbiota [1, 15]. Yet, we remain largely ignorant as to whether these helminth effects depend on other biological variables (environmental effects or host genotype). Here, we provide evidence of such interactions: host genotype (sex and autosomal) and helminth infection synergistically alter the microbiota. One third of the common microbial Orders exhibited significant interactions between infection and host genotype (Fig. 2), whereas infection by itself affected only 7% of the Orders. For example, the relative abundances of some microbial Orders in female hosts were significantly different from males when uninfected, but the sexes were not different when infected. Discriminant analysis suggested that females’ microbiota were generally more sensitive to infection than males’ were. This sex-dependent effect of infection is consistent with other recent papers on stickleback suggesting that sex modifies the gut microbiota’s response to diet [31], and MHC genotype [44]. Similar sex-dependent environmental effects occur in other vertebrates as well [45].
Host autosomal genotype affected gut microbial composition, and modified microbes’ responses to helminth infection. Many other studies of vertebrates have found host genetic control of microbiota composition [46, 47]. Our results add a new dimension to this conclusion: host genotype also alters the microbiota’s response to a third (parasitic) species. We coarsely localize these autosomal effects to a modest number of chromosomal loci. These QTLs do not contain MHCIIb, which has previously been associated with natural variation in stickleback gut microbiota [44]. At present we lack the resolution to fine-map down to specific candidate genes, but some intriguing possibilities exist. The QTL on Chr2, which affect multiple microbial Orders, contains transcription factor c-MAF, which activates the expression of IL-4 in Th2 cells and attenuates Th1 differentiation [48]. Interestingly, recently Xu et al. reported c-MAF-dependent regulatory T cells mediate immunological tolerance to intestinal microbiota [49]. The QTL on this chromosome also contains reticulocalbin and synaptotagmin IX. These genes are associated with calcium ion binding, which can affect intestinal microbiota composition [50,51,52]. The QTL on Chr3 contains cadherins, which play a key role in intestinal homeostasis and barrier function [53]. These candidate genes are just speculative, at present: substantial future work is needed to fine-map candidate genes and experimentally validate their suspected effects on the cestode–microbiota interaction.
Interactions between host genotype and infection likely arise from host genetic variation in immune response to cestodes. The presence of helminths can interfere with TLR or downstream signaling pathways [54], which may have ancillary effects on the microbiota. Adaptive immune responses may be involved as well. Helminths can trigger a Th2 response that increases mucus production and epithelial cell turnover, altering the mucosal microbiota [55]. Any polymorphism in immune genes involved in these pathways may lead to an exaggerated (or, reduced) immune response to helminth infection, and thereby modify the effect of helminths on gut microbes. This logic applies equally to immune differences between males and females [56, 57].
In the specific case examined here, helminth effects on the microbiota are especially likely to involve indirect effects via host immunity, rather than a direct microbe–helminth interaction. This is because S. solidus is not an intestinal parasite. After being ingested, it transits within hours into the body cavity where it persists for months but cannot directly contact the microbiota. Prior studies confirm that stickleback vary in their ability to detect S. solidus, mount an effective immune response, and susceptibility to cestode immune suppression [58,59,60]. Polymorphism in these immune responses may is likely to impose collateral effects on the microbiota. It remains possible, in principle, that despite their physical separation the helminth is directly affecting the microbiota by secreting compounds that diffuse from the peritoneum into the gut lumen. However, these direct effects are unlikely to be modulated by host genotype, and should therefore show up as statistical main effects of infection (which were rare) rather than infection by genotype interactions (which were common). This point, combined with extensive prior evidence that S. solidus does modify host immune traits [59], makes indirect effects the most parsimonious explanation.
Our Experiment 1 suggests that stickleback gut microbiota depended most strongly on parasite exposure (regardless of outcome), rather than infection itself. Apparently, there is a lasting effect of the transient presence of a tapeworm in the stickleback gut lumen, which outweighs a relatively weak effect of tapeworm presence or absence in the peritoneum (which we lacked power to detect in Experiment 1, but had more power to find in Experiment 2). If indeed brief parasite exposure sends lasting ripples through the microbial community, then the study of the wild microbiome will be still more difficult. We rarely if ever know wild animals’ history of unsuccessful infections, so the ‘ghosts’ of past failed infections might generate substantial and untraceable variation among wild individuals. Yet, infection success also alters the gut microbiota, as we then revealed in the follow-up Experiment 2.
At present we do not know the fitness consequences of the altered microbiota that we document here. Some studies have reported detrimental helminth effects on the microbiota. Experimental T. muris infection in mice alters the microbiota composition leading to reduced availability of microbial metabolomics products needed by the host [61]. Conversely, helminth infection can be beneficial, such as reducing the risk of H. pylori-induced gastric lesions [62]. Our results suggest that these positive or negative effects of helminth–microbiota interactions will be contingent on the specific sex and genotype of the hosts considered.
Data availability
Sequence data have been deposited in the Sequence Read Database (SRA) under project IDs SRP115642 (BioProject PRJNA398629) and SRP115678 (BioProject PRJNA398630) for experimental 1 and 2, respectively. All other relevant data are available in this article and its Supplementary Information files, or from the corresponding authors upon request.
References
Reynolds LA, Finlay BB, Maizels RM. Cohabitation in the intestine: Interactions among helminth parasites, bacterial microbiota, and host immunity. J Immunol. 2015;195:4059–66.
Peachey LE, Jenkins TP, Cantacessi C. This gut ain’t big enough for both of us. Or is it? Helminth-microbiota interactions in veterinary species. Trends Parasitol. 2017;33:619–32.
McKenney EA, Williamson L, Yoder AD, Rawls JF, Bilbo SD, Parker W. Alteration of the rat cecal microbiome during colonization with the helminth Hymenolepis diminuta. Gut Microbes. 2015;6:182–93.
Aivelo T, Norberg A. Parasite-microbiota interactions potentially affect intestinal communities in wild mammals. J Anim Ecol. 2018;87:438–47.
Leung JM, Loke P. A role for IL-22 in the relationship between intestinal helminths, gut microbiota and mucosal immunity. Int J Parasitol. 2013;43:253–7.
Loke P, Lim YAL. Can helminth infection reverse microbial dysbiosis? Trends Parasitol. 2015;31:534–5.
Cooper P, Walker AW, Reyes J, Chico M, Salter SJ, Vaca M, et al. Patent human infections with the whipworm, Trichuris trichiura, are not associated with alterations in the faecal microbiota. PLoS ONE. 2013;8:e76573.
Cantacessi C, Giacomin P, Croese J, Zakrzewski M, Sotillo J, McCann L, et al. Impact of experimental hookworm infection on the human gut microbiota. J Infect Dis. 2014;210:1431–4.
Jaenike J, Unckless R, Cockburn SN, Boelio LM, Perlman SJ. Adaptation via symbiosis: recent spread of a Drosophila defensive symbiont. Science. 2010;329:212–5.
Oliveira-Sequeira TC, David ÉB, Ribeiro C, Guimarães S, Masseno AP, Katagiri S, et al. Effect of Bifidobacterium animalis on mice infected with Strongyloides venezuelensis. Rev Inst Med Trop Sao Paulo. 2014;56:105–9.
White EC, Houlden A, Bancroft AJ, Hayes KS, Goldrick M, Grencis RK, et al. Manipulation of host and parasite microbiotas: survival strategies during chronic nematode infection. Sci Adv. 2018;4:eaap7399.
Dea-Ayuela MA, Rama-Iniguez S, Bolas-Fernandez F. Enhanced susceptibility to Trichuris muris infection of B10Br mice treated with the probiotic Lactobacillus casei. Int Immunopharmacol. 2008;8:28–35.
Reynolds LA, Smith KA, Filbey KJ, Harcus Y, Hewitson JP, Redpath SA, et al. Commensal-pathogen interactions in the intestinal tract: Lactobacilli promote infection with, and are promoted by, helminth parasites. Gut Microbes. 2014;5:522–32.
Loke P, Lim YA. Helminths and the microbiota: parts of the hygiene hypothesis. Parasite Immunol. 2015;37:314–23.
Zaiss MM, Harris NL. Interactions between the intestinal microbiome and helminth parasites. Parasite Immunol. 2016;38:5–11.
Rapin A, Harris NL. Helminth-bacterial interactions: cause and consequence. Trends Immunol. 2018;39:724–33.
Hewitson JP, Harcus Y, Murray J, van Agtmaal M, Filbey KJ, Grainger JR, et al. Proteomic analysis of secretory products from the model gastrointestinal nematode Heligmosomoides polygyrus reveals dominance of venom allergen-like (VAL) proteins. J Proteom. 2011;74:1573–94.
Geuking MB, Cahenzli J, Lawson MA, Ng DC, Slack E, Hapfelmeier S, et al. Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity. 2011;34:794–806.
Hang L, Blum AM, Setiawan T, Urban JP Jr, Stoyanoff KM, Weinstock JV. Heligmosomoides polygyrus bakeri infection activates colonic Foxp3+ T cells enhancing their capacity to prevent colitis. J Immunol. 2013;191:1927–34.
Su L, Su CW, Qi Y, Yang G, Zhang M, Cherayil BJ, et al. Coinfection with an intestinal helminth impairs host innate immunity against Salmonella enterica serovar Typhimurium and exacerbates intestinal inflammation in mice. Infect Immun. 2014;82:3855–66.
Glendinning L, Nausch N, Free A, Taylor DW, Mutapi F. The microbiota and helminths: sharing the same niche in the human host. Parasitology. 2014;141:1255–71.
Grant AV, Araujo MI, Ponte EV, Oliveira RR, Cruz AA, Barnes KC, et al. Polymorphisms in IL10 are associated with total Immunoglobulin E levels and Schistosoma mansoni infection intensity in a Brazilian population. Genes Immun. 2011;12:46–50.
Costa RD, Figueiredo CA, Barreto ML, Alcantara-Neves NM, Rodrigues LC, Cruz AA, et al. Effect of polymorphisms on TGFB1 on allergic asthma and helminth infection in an African admixed population. Ann Allergy Asthma Immunol. 2017;118:483.e1–8.
Janet K, Tommy LFL. Flying with diverse passengers: greater richness of parasitic nematodes in migratory birds. Oikos. 2015;124:399–405.
Guivier E, Bellenger J, Sorci G, Faivre B. Helminth interaction with the host immune system: short-term benefits and costs in relation to the infectious environment. Am Nat. 2016;188:253–63.
Bourke CD, Maizels RM, Mutapi F. Acquired immune heterogeneity and its sources in human helminth infection. Parasitology. 2011;138:139–59.
Fischer J, Jung N, Robinson N, Lehmann C. Sex differences in immune responses to infectious diseases. Infection. 2015;43:399–403.
Klein SL. The effects of hormones on sex differences in infection: from genes to behavior. Neurosci Biobehav Rev. 2000;24:627–38.
Weber JN, Steinel NC, Shim KC, Bolnick DI. Recent evolution of extreme cestode growth suppression by a vertebrate host. Proc Natl Acad Sci USA. 2017;114:6575–80.
Lohman BK, Steinel NC, Weber JN, Bolnick DI. Gene expression contributes to the recent evolution of host resistance in a model host parasite system. Front Immunol. 2017;8:1071.
Bolnick DI, Snowberg LK, Hirsch PE, Lauber CL, Org E, Parks B, et al. Individual diet has sex-dependent effects on vertebrate gut microbiota. Nat Commun. 2014;5:4500.
Bolnick DI, Snowberg LK, Hirsch PE, Lauber CL, Knight R, Caporaso JG, et al. Individuals’ diet diversity influences gut microbial diversity in two freshwater fish (threespine stickleback and Eurasian perch). Ecol Lett. 2014;17:979–87.
Smith CC, Snowberg LK, Gregory CJ, Knight R, Bolnick DI. Dietary input of microbes and host genetic variation shape among-population differences in stickleback gut microbiota. ISME J. 2015;9:2515–26.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol. 2013;79:5112–20.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–6.
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 2014;42:D633–42.
McMurdie PJ, Holmes S. Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE. 2013;8:e61217.
Bolker BR. Tools for general maximum likelihood estimation. 2017. https://cran.r-project.org/web/packages/bbmle/.
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE. Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE. 2012;7:e37135.
Stuart YE, Veen T, Weber JN, Hanson D, Ravinet M, Lohman BK, et al. Contrasting effects of environment and genetics generate a continuum of parallel evolution. Nat Ecol Evol. 2017;1:158.
Broman KW, Wu H, Sen S, Churchill GA. R/qtl: QTL mapping in experimental crosses. Bioinformatics. 2003;19:889–90.
Gaulke CA, Martins ML, Watral VG, Humphreys IR, Spagnoli ST, Kent ML, et al. A longitudinal assessment of host-microbe-parasite interactions resolves the zebrafish gut microbiome’s link to Pseudocapillaria tomentosa infection and pathology. Microbiome. 2019;7:10.
Bolnick DI, Snowberg LK, Caporaso JG, Lauber C, Knight R, Stutz WE. Major Histocompatibility Complex class IIb polymorphism influences gut microbiota composition and diversity. Mol Ecol. 2014;23:4831–45.
Min Y, Ma XG, Sankaran K, Ru Y, Chen LJ, Baiocchi M, et al. Sex-specific association between gut microbiome and fat distribution. Nat Commun. 2019;10:2408.
Benson AK, Kelly SA, Legge R, Ma F, Low SJ, Kim J, et al. Individuality in gut microbiota composition is a complex polygenic trait shaped by multiple environmental and host genetic factors. Proc Natl Acad Sci USA. 2010;107:18933–8.
Goodrich JK, Davenport ER, Waters JL, Clark AG, Ley RE. Cross-species comparisons of host genetic associations with the microbiome. Science. 2016;352:532–5.
Ho IC, Lo D, Glimcher LH. c-maf promotes T helper cell type 2 (Th2) and attenuates Th1 differentiation by both interleukin 4-dependent and -independent mechanisms. J Exp Med. 1998;188:1859–66.
Xu M, Pokrovskii M, Ding Y, Yi R, Au C, Harrison OJ, et al. c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature. 2018;554:373–7.
Mai V, McCrary QM, Sinha R, Glei M. Associations between dietary habits and body mass index with gut microbiota composition and fecal water genotoxicity: an observational study in African American and Caucasian American volunteers. Nutr J. 2009;8:49.
Mann E, Schmitz-Esser S, Zebeli Q, Wagner M, Ritzmann M, Metzler-Zebeli BU. Mucosa-associated bacterial microbiome of the gastrointestinal tract of weaned pigs and dynamics linked to dietary calcium-phosphorus. PLoS ONE. 2014;9:e86950.
Gomes JM, Costa JA, Alfenas RC. Could the beneficial effects of dietary calcium on obesity and diabetes control be mediated by changes in intestinal microbiota and integrity? Br J Nutr. 2015;114:1756–65.
Zhu Y, Michelle LT, Jobin C, Young HA. Gut microbiota and probiotics in colon tumorigenesis. Cancer Lett. 2011;309:119–27.
Venugopal PG, Nutman TB, Semnani RT. Activation and regulation of toll-like receptors (TLRs) by helminth parasites. Immunol Res. 2009;43:252–63.
Broadhurst MJ, Ardeshir A, Kanwar B, Mirpuri J, Gundra UM, Leung JM. et al. Therapeutic helminth infection of macaques with idiopathic chronic diarrhea alters the inflammatory signature and mucosal microbiota of the colon. PLoS Pathog. 2012;8:e1003000.
Scotland RS, Stables MJ, Madalli S, Watson P, Gilroy DW. Sex differences in resident immune cell phenotype underlie more efficient acute inflammatory responses in female mice. Blood. 2011;118:5918–27.
Klein SL, Flanagan KL. Sex differences in immune responses. Nat Rev Immunol. 2016;16:626–38.
Scharsack JP, Koch K, Hammerschmidt K. Who is in control of the stickleback immune system: interactions between Schistocephalus solidus and its specific vertebrate host. Proc Biol Sci. 2007;274:3151–8.
Barber I, Scharsack JP. The three-spined stickleback-Schistocephalus solidus system: an experimental model for investigating host-parasite interactions in fish. Parasitology. 2010;137:411–24.
Hendry AP, Peichel CL, Matthews B, Boughman JW, Nosil P. Stickleback research: the now and the next. Evol Ecol Res. 2013;15:111–41.
Houlden A, Hayes KS, Bancroft AJ, Worthington JJ, Wang P, Grencis RK, et al. Chronic Trichuris muris Infection in C57BL/6 mice causes significant changes in host microbiota and metabolome: effects reversed by pathogen clearance. PLoS ONE. 2015;10:e0125945.
Whary MT, Muthupalani S, Ge Z, Feng Y, Lofgren J, Shi HN, et al. Helminth co-infection in Helicobacter pylori infected INS-GAS mice attenuates gastric premalignant lesions of epithelial dysplasia and glandular atrophy and preserves colonization resistance of the stomach to lower bowel microbiota. Microbes Infect. 2014;16:345–55.
Acknowledgements
We thank the Genome Sequencing and Analysis Facility (GSAF) at the University of Texas at Austin for sequencing support, and the Texas Advanced Computing Center (TACC) and High Performance Computing platform of Northwest A&F University for computational resources. This research was supported by funding from National Natural Science Foundation of China to FL, NSFC(Grant 31672680), and the Howard Hughes Medical Institute to DB, NIH (Grant 1R01AI123659-01A1).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ling, F., Steinel, N., Weber, J. et al. The gut microbiota response to helminth infection depends on host sex and genotype. ISME J 14, 1141–1153 (2020). https://doi.org/10.1038/s41396-020-0589-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-0589-3
This article is cited by
-
Sex—the most underappreciated variable in research: insights from helminth-infected hosts
Veterinary Research (2022)
-
Parasite reliance on its host gut microbiota for nutrition and survival
The ISME Journal (2022)
-
Dysbiosis of fish gut microbiota is associated with helminths parasitism rather than exposure to PAHs at environmentally relevant concentrations
Scientific Reports (2022)