Abstract
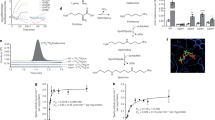
As a frequent inhabitant of sites polluted with toxic chemicals, the soil bacterium and plant-root colonizer Pseudomonas putida can tolerate high levels of endogenous and exogenous oxidative stress. Yet, the ultimate reason of such phenotypic property remains largely unknown. To shed light on this question, metabolic network-wide routes for NADPH generation—the metabolic currency that fuels redox-stress quenching mechanisms—were inspected when P. putida KT2440 was challenged with a sub-lethal H2O2 dose as a proxy of oxidative conditions. 13C-tracer experiments, metabolomics, and flux analysis, together with the assessment of physiological parameters and measurement of enzymatic activities, revealed a substantial flux reconfiguration in oxidative environments. In particular, periplasmic glucose processing was rerouted to cytoplasmic oxidation, and the cyclic operation of the pentose phosphate pathway led to significant NADPH-forming fluxes, exceeding biosynthetic demands by ~50%. The resulting NADPH surplus, in turn, fueled the glutathione system for H2O2 reduction. These properties not only account for the tolerance of P. putida to environmental insults—some of which end up in the formation of reactive oxygen species—but they also highlight the value of this bacterial host as a platform for environmental bioremediation and metabolic engineering.
Similar content being viewed by others
Introduction
Environmental bacteria are exposed to different types of stress in the niches they colonize. Pseudomonas putida is an archetypal instance of a strictly aerobic, Gram-negative soil bacterium characterized by a remarkable metabolic versatility [1,2,3,4], and equipped with the enzymatic machinery needed to catabolize both natural and recalcitrant aromatic compounds besides sugars and organic acids [5]. The metabolic pathways deployed for biodegradation of organic and xenobiotic chemicals [6] often involve harsh redox reactions prone to generate reactive oxygen species (ROS) [7]. Apart of this endogenous source of stress, many of the scenarios where bacteria meet environmental pollutants are characterized by the presence of both direct and indirect oxidative agents. Besides the presence of oxygen in the upper soil layers [8], cycles of inundation, elevation gradients, salinity fluctuations, sunlight, and other abiotic factors can generate ROS—thereby leading to an oxidative milieu [9,10,11]. Environmental bacteria have evolutionary acquired mechanisms to cope with such conditions and, besides Pseudomonas, species of the Burkholderia [12] and Sphingomonas [13] genera—typically used in bioaugmentation—are recognized to withstand harsh physicochemical conditions. Yet, how do these environmental microorganisms cope with redox insults? Peroxides and free radicals damage virtually all macromolecular components of the cell (e.g., proteins, lipids, and DNA [14, 15]), and mechanisms to prevent (or repair) the damage exerted by ROS are key components of virtually all biological systems [16]. Long-term transcriptional responses that lead to mechanisms of ROS scavenging, for instance, appear to be conserved across species [17]. In Escherichia coli, for instance, oxidative challenges cause upregulation of genes encoding ROS-quenching superoxide dismutases, catalase, and glutathione/glutaredoxin recycling systems [18,19,20,21]. However, until transcriptionally controlled defense mechanisms become fully operational, cell survival depends on the default activity of the above-mentioned enzymes and the availability of antioxidants (e.g. reduced glutathione [22, 23]) to scavenge ROS generated by endogenous reactions or external oxidative damage. Prior to full-fledged deployment of transcriptional responses (which can take from several minutes up to hours, depending on the species and the growth conditions [24, 25]), metabolic reactions are the first line of defense upon sudden exposure to oxidative stress [26,27,28,29], as long as they can provide reducing power to fuel ROS-detoxifying enzymes [30]. The specific metabolic mechanisms that underlay the tolerance of P. putida to oxidative stress, however, remain largely unknown.
The adaptability of P. putida KT2440 (type strain of this species [31]) to adverse conditions, including redox stress, is wired to a unique metabolic architecture. Glucose catabolism in P. putida relies on the Entner–Doudoroff (ED) pathway [32], which starts with 6-phosphogluconate (6PG) as the substrate, formed via separate and converging routes for hexose phosphorylation (in the cytoplasm) or oxidation (in the periplasm) [33, 34]. Additionally, the EDEMP cycle, a combination of enzymes from the ED, the pentose phosphate (PP) and the (incomplete) Embden–Meyerhof–Parnas (EMP) pathways [35], processes hexoses phosphate to feed the lower catabolism, i.e. downwards phosphoenolpyruvate (PEP), pyruvate (Pyr), and the tricarboxylic acid (TCA) cycle. In addition to catabolic processing of sugars, the operation of the EDEMP cycle entails redirecting part of the trioses phosphate back to hexoses phosphate via the gluconeogenic activity of the EMP branch of the cycle [36]. As such, cyclic sugar catabolism contributes to a slight catabolic overproduction of NADPH in glucose cultures [35]. Although mostly studied with model carbon sources, the EDEMP cycle is the authentic metabolic heart of P. putida and is key for the hierarchical consumption of sugars and aromatic compounds [37], also under fluctuating conditions of iron availability [38]. While the main sink of NADPH is the anabolic use of reducing power for biomass formation, this redox cofactor is the major metabolic currency to counterfeit oxidative stress, since it serves as the reductant for several ROS-detoxifying enzymes and mechanisms [39], e.g. glutathione regeneration. However, the connection between the operation of central carbon metabolism and oxidative stress responses have remained elusive thus far.
In this work, the reshaping of the central metabolism in P. putida KT2440 in response to a sub-lethal oxidative stress levels, elicited by exposure to hydrogen peroxide (H2O2), has been thoroughly inspected. The strategy encompassed 13C-tracer experiments, metabolomics, and metabolic flux analysis, combined with the determination of physiological parameters, in vitro measurement of key enzymatic activities and assessment of glutathione-mediated ROS-detoxifying mechanisms. The results revealed adaptions in the operation of the EDEMP cycle, concomitant with a decrease in the flux through sugar oxidation pathways for glucose processing, which helps replenishing the NADPH pool upon oxidative challenges. These metabolic adjustments are discussed at the light of the lifestyle of P. putida (and related environmental bacteria) and the value of this species as robust agent for both in situ remediation of environmental pollution and metabolic engineering.
Materials and methods
Bacterial strain and culture conditions
Wild-type P. putida strain KT2440 was used throughout this study [40], and it was routinely grown in LB medium [41]. Quantitative physiology experiments were carried out in M9 minimal medium containing 6 g L−1 Na2HPO4, 3 g L−1 KH2PO4, 1.4 g L−1 (NH4)2SO4, 0.5 g L−1 NaCl, 0.2 g L−1 MgSO4·7H2O and 2.5 mL L−1 of a trace elements solution [42]. Unless noted otherwise, either natural or 13C-labeled glucose was used as the sole carbon source at 20 mM. Growth was estimated by measuring the optical density at 600 nm (OD600) after diluting the culture with 9 g L−1 NaCl. Correlation factors between cell dry weight (CDW) and OD600 were determined in batch cultures [43]. All cultures were started with an isolated colony from an LB medium plate, suspended in 5 mL of M9 minimal medium in a test tube. After an 18-h incubation, this culture was used to inoculate fresh medium at an OD600 of 0.01. Working cultures were cultivated in 250-mL Erlenmeyer flasks containing medium up to one-fifth of their nominal volume. H2O2 was used as an oxidative stress agent at 1.5 mM, added at OD600 = 0.5 (mid-exponential phase). Cultures were incubated for an additional 1.5 h, and samples were taken for analyses as explained below.
Determination of physiological parameters
Regression analysis was applied during exponential growth to calculate [i] the maximum specific growth rate (μ), [ii] the biomass yield on substrate (YX/S), [iii] the specific rate of glucose consumption (qS), and [iv] the molar yield of organic acids on glucose (yP/S). CDW was quantified by harvesting cells by fast filtration in pre-weighed nitrocellulose filters (0.45 μm pore), washed twice with 9 g L−1 NaCl, and dried at 105 °C to a constant weight. Glucose was assayed using a commercial kit from R-Biopharm AG (Darmstadt, Germany). The concentration of gluconate and 2-ketogluconate in culture supernatants was measured enzymatically [44]. In order to assess the impact of different carbon sources in the growth phenotype of strain KT2440 upon oxidative challenges, batch cultures were developed in M9 minimal medium as indicated above, and either glucose (20 mM), α-ketoglutarate (24 mM), or glycerol (40 mM) were used as the sole carbon substrate. Note that these culture conditions provide a total carbon concentration of 120 mM. When the OD600 of the cultures reached 0.15, H2O2 was added at 3 mM, and the change in OD600 was monitored every 15 min until the stationary phase was reached. Normalized growth coefficients (i.e., the ratio between μ in stressed cultures and μ in control experiments) were calculated for each culture condition [45].
Determination of relative metabolite concentrations and 13C-labeling patterns by LC-MS/MS
Cultures were grown on either 100% [1-13C1]-glucose or 100% [6-13C1]-glucose as the sole carbon source. The biomass corresponding to 0.5–0.6 mg of CDW was collected in triplicates by fast centrifugation (13,000 × g, 30 s, −4 °C). Bacterial pellets were immediately frozen in liquid N2. Samples were then extracted three times with 0.5 mL of 60% (v/v) ethanol buffered with 10 mM ammonium acetate (pH = 7.2) at 78 °C for 1 min. After each extraction step, the biomass was separated by centrifugation at 13,000 × g for 1 min. The three liquid extracts were pooled and dried at 120 μbar, and stored at −80 °C thereafter. Samples were resuspended in 20 μL of MilliQ water, distributed in sealed 96-well microtiter plates, and injected into a Waters Acquity UPLC (Waters Corp., Milford, MA, USA) with a Waters Acquity T3 column (150 mm × 2.1 mm × 1.8 μm, Waters Corp.) coupled to a Thermo TSQ Quantum Ultra triple quadrupole instrument (Thermo Fisher Scientific Inc., Waltham, MA) with electrospray ionization. 13C-Labeling patterns of free intracellular metabolites were determined for dihydroxyacetone phosphate, fructose-6-phosphate (F6P), fructose-1,6-bisphosphate (FBP), glucose-6-phosphate (G6P), 6PG, PEP, ribose-5-phosphate (R5P), ribulose-5-phosphate (Ru5P), sedoheptulose-7-phosphate (S7P), xylulose-5-phosphate, and Pyr as described previously [46]. All other metabolites considered for the analysis are listed in Dataset S1. Relative abundances were calculated as the total ion count of all measured 13C isotopes for each metabolite across the different growth conditions, and presented as ratios whenever relevant. The raw data of labeling experiments is available in Dataset S1.
Determination of 13C-labeling patterns by GC-MS
Cultures were grown on either 100% [1-13C1]-glucose or a mixture of 20% (w/w) [U-13C6]-glucose and 80% (w/w) naturally labeled glucose, and 5 mL aliquots of cell broth were harvested by centrifugation at 1,200 × g and −4 °C for 10 min. Bacterial pellets were washed twice with 1 mL of 9 g L−1 NaCl, hydrolyzed in 1 mL of 6 M HCl for 24 h at 110 °C, and desiccated overnight at 85 °C under a constant air stream. The hydrolyzate was dissolved in 50 μL of 99.8% (w/v) dimethyl formamide and subsequently transferred into a new tube. For sample derivatization, 30 μL of N-methyl-N-(tert-butyldimethylsilyl)-trifluoroacetamide was added to the biomass hydrolyzate and incubated at 85 °C for 60 min. The 13C-labeling patterns of proteinogenic amino acids were determined on a 6890 N Network GC system with a 5975 inert XL mass selective detector (Agilent Technologies Inc., Santa Clara, CA, USA) [47, 48]. The raw GC-MS data from four independent experiments is presented in Dataset S2.
Metabolic flux ratio analysis
Mass distribution vectors of the proteinogenic amino acids were corrected for the natural abundance of all stable isotopes, and the relative metabolic flux ratios [i] oxaloacetate (OAA) from Pyr, [ii] glyoxylate shunt, [iii] PEP from OAA, and [iv] the lower and upper bound for Pyr from malate were calculated using the Fiat Flux software [49]. The mass distribution vectors of the free intracellular metabolites were corrected for the natural abundance of all stable isotopes using MatLab (The Mathworks Inc., Natick, MA, USA). Novel relative flux ratios were defined and calculated as described by Nikel et al. [35]. The fraction of G6P originating from glucose was estimated using data from the experiments using 100% [1-13C1]-glucose, whereas 6PG from G6P was calculated using data from either 100% [1-13C1]-glucose or 100% [6-13C1]-glucose experiments. Formation of F6P through the PP pathway was estimated using data from 100% [6-13C1]-glucose experiments, and Pyr through the ED pathway was assessed with data from 100% [1-13C1]-glucose experiments. The metabolic model used for net-flux analysis was based on a master reaction network with 45 reactions and 33 metabolites. Fluxes were calculated using [i] the stoichiometric reaction matrix, [ii] constraints accounting for the ratios from FiatFlux analysis and additionally for the ratios in the initial steps of glucose catabolism as described above, [iii] physiological data, and [iv] precursor requirements for biomass. The experimentally determined relative flux ratios were translated into constraints as follows (the reaction numbers, vx, are defined in Fig. S1). The fraction of G6P originating from glucose (a) was estimated as a = v3/(v3 + v16); the fraction of F6P originating through the PP pathway (b) was calculated as b = (v14 + v15)/(v14 + v15 + v17); and the fraction of 6PG originating from G6P (c) was determined as c = v7/(v4 + v6 + v7). For this ratio, the values from the experiments using data from either 100% [1-13C1]-glucose or [6-13C1]-glucose were averaged. The fraction of Pyr originating through the ED pathway (d) was determined as d = v9/(v9 + v22 + v33 + v34). The upper and lower bounds of Pyr originating from malate (e and f, respectively) were obtained according to e ≥ v34/(v9 + v22 + v33 + v34) and f ≤ v34/(v9 + v22 + v33 + v34). The fraction of OAA originating from Pyr (g) was obtained following g = v32/(v30 + v32). Finally, the fraction of Pyr originating from OAA (h) was derived from h = v33/(v9 + v22 + v33 + v34). The determined linear system of mass balances, flux ratios, quantitative physiology data, and biomass requirements was then solved with the fmincon function from MatLab using the Netto module from FiatFlux to obtain net metabolic fluxes [49].
Preparation of cell-free extracts and in vitro enzymatic assays
Cell-free extracts were prepared from cells harvested by centrifugation from an appropriate culture volume at 4,000 × g at 4 °C for 10 min. Pellets were suspended in 1 volume of 10 mM phosphate-buffered saline (PBS, pH = 7.5, and previously refrigerated) containing 10 mM 2-mercaptoethanol and centrifuged again. Cells were finally resuspended in 0.3–0.5 volume of the same buffer and sonicated intermittently for 6 min in an ice bath. Sonicated cells were centrifuged at 7,500 × g at 4 °C for 30 min, and the total protein concentration in cell extracts was measured by the Bradford method. The activities of Edd, Eda, Glk, Gcd, and Gad were assayed using standard protocols [34, 50]. The activity of Zwf and Gnd was assayed under both saturating and non-saturating, quasi in vivo conditions using concentrations reflecting intracellular experimentally determined abundance of cofactors and substrates. For reactions in which more than one enzyme catalyzes the corresponding transformation (e.g., Zwf, represented by three isozymes in P. putida KT2440), the total activity was calculated. In the latter case, the concentrations of the substrates (experimentally determined in cell-free extracts of glucose-grown KT2440 by means of LC-MS/MS), were 1.2 mM G6P and 2.4 mM 6PG.
Glutathione quantification
Glutathione (both oxidized and reduced forms) was quantified in cells harvested from 25 mL-culture samples using a modification of the procedure described by Michie et al. [51]. Cells were pelleted by centrifugation (4,000 × g at 4 °C for 10 min) and suspended in 2 mL of cold PBS. The cell suspension was mixed 1:1 (v/v) with a freshly prepared 10% (w/v) 5-sulfosalicylic acid solution, incubated on an ice bath for 10 min, and then sonicated. For the measurement of total glutathione, 40 μL of Tris(2-carboxyethyl)phosphine·HCl was added to an equal volume of the cell-free extract. For the measurement of reduced glutathione, 40 μL of H2O were used instead. Samples were centrifuged at 15,600 × g for 5 min after treatment, and 25 μL of the supernatant fluid was added to 100 μL of a buffer containing 200 mM N-ethylmorpholine and 20 mM NaOH. Following the addition of 50 μL of 0.5 N NaOH to this mixture, glutathione was derivatized by addition of 10 μL of 10 mM naphthalene-2,3-dicarboxaldehyde. The mixture was incubated at room temperature for 30 min, and the fluorescence intensity of the resulting naphthalene-2,3-dicarboxaldehyde–glutathione conjugate was measured at an excitation wavelength (λexcitation) = 472 nm and an emission wavelength (λexcitation) = 528 nm. A standard curve was constructed with known amounts of freshly prepared naphthalene-2,3-dicarboxaldehyde–glutathione conjugate.
Chemicals and enzymes
[1-13C1]-Glucose and [6-13C1]-glucose were purchased from Cambridge Isotope Laboratories Inc. (Tewksbury, MA, USA), and [U-13C6]-glucose was purchased from Sigma-Aldrich Co. (St. Louis, MO, USA). Other chemicals and enzymes used for in vitro assays were obtained from Sigma-Aldrich Co. and Merck KGaA (Darmstadt, Germany). H2O2 solutions were freshly prepared from a 30% (w/w) stock solution (Sigma-Aldrich Co.).
Data analysis and statistics
Unless otherwise indicated, all data are presented as mean values together with standard deviation, obtained from at least three independent experiments. Bacterial growth and physiological parameters were analyzed and plotted using GraphPad Prism 7 (GraphPad Software, San Diego, CA, USA) or Microsoft Excel 2019 (Microsoft Corp., Redmond, WA, USA), and statistical differences were calculated using the Student’s t test with Welch’s correction. The Student’s t test with Welch’s adjustment and the Games-Howell post-hoc test were used to calculate statistical differences for metabolomics and fluxome analyses, using R version 3.4.3, and a Bonferroni correction was applied, whenever relevant, to correct for multiple comparisons. Actual p values of these statistical comparisons are indicated in the figure legends.
Results
Quantitative physiology analysis of P. putida exposed to sub-lethal doses of oxidative stress
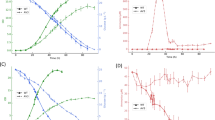
In order to capture the metabolic effects of oxidative stress in P. putida KT2440 without compromising cell viability, the experimental design (summarized in Fig. 1a) entailed exposure of the cells, grown in M9 minimal medium with glucose until mid-exponential phase [optical density at 600 nm (OD600) of ca. 0.5], to H2O2 at 1.5 mM. Cultures with different 13C-tracers ([1-13C1]-, [6-13C1]- and [U-13C6]-glucose, all of them added at 20 mM) were grown in parallel to resolve the relative contribution of cyclic EDEMP, PP, and ED pathway to sugar processing. At 1.5 h after the treatment, samples were collected to assess quantitative physiology parameters, flux ratio analysis, and to perform in vitro biochemical assays. This experimental setup resulted in similar growth patterns in the absence and presence of the oxidative stress agent (Fig. 1b), and neither the specific growth rate (μ) nor the final cell density (plateauing at ca. 10 h of cultivation) were significantly affected by the addition of H2O2 to the cultures. Besides the fact that growth was not significantly affected, this experimental setup was chosen to capture metabolic adaptions at the onset of transcriptional responses due to exposure to oxidative conditions [52, 53]. In particular, no significant changes in the level of expression of genes encoding enzymes of the central metabolism had been observed upon exposure of P. putida KT2440 to H2O2 in this time frame [52]. Quantitative analysis of physiology parameters confirmed that this was the case (Fig. 1c), with μ, the specific rate of glucose uptake (qS) and the yield of biomass on substrate (YX/S) being essentially the same in control and H2O2-treated experiments. The formation and secretion of organic acids to the extracellular milieu, a typical feature of glucose-grown P. putida, decreased significantly in the presence of the stressor. P. putida typically oxidizes glucose in the periplasm into gluconate and 2-ketogluconate [32], which are then taken up by the cell [54]. The about halved concentrations of the two products of sugar oxidation in H2O2-treated cultures indicates a decrease in periplasmic oxidation upon exposure to stressful conditions. The concentration of both acids peaked during exponential growth, and the corresponding molar yields of gluconate and 2-ketogluconate on glucose in control experiments were yG/S = 0.31 ± 0.09 and yK/S = 0.12 ± 0.02 C-mol C-mol−1, respectively. In the presence of oxidative stress, yG/S and yK/S were 0.19 ± 0.08 and 0.05 ± 0.01 C-mol C-mol−1, suggestive, again, of a redistribution of metabolic fluxes between oxidative (i.e. glucose → gluconate → 2-ketogluconate) or phosphorylative [i.e. glucose → G6P] processing routes of the carbon source. Since a shift from periplasmic to cytoplasmic glucose oxidation would entail higher NADPH formation, we next looked for direct evidence of increased intracellular fluxes through these pathways.
a Overview of the experimental design adopted in this study. All growth and quantitative physiology experiments were conducted at least in biological triplicates, and the oxidative challenge is indicated with a red symbol. OD600, optical density measured at 600 nm. b Representative growth curves for control (Ctrl.) experiments and H2O2-stressed cultures. Samples were taken 1.5 h after addition of H2O2, where the growth of the cells was still exponential. c Key physiological parameters of P. putida KT2440 in batch glucose cultures. All values represent the mean ± standard deviations from at least three biological replicates, and the differences between the growth parameters in H2O2-treated and control cultures was not significant (as assessed by means of the Student’s t test with Welch’s correction). The specific growth rate (μ) and the specific rate of carbon uptake (qS) were determined during exponential growth (including the treatment period) by linear regression of log-transformed OD600 data. The yield of biomass on substrate (YX/S) was calculated at 24 h, after glucose was completely exhausted. The extracellular concentration (concn.) of organic acids reported is the maximal reached during the whole culture period. CDW, cell dry weight.
PP pathway metabolite pools increase upon exposure of P. putida to oxidative stress
To assess the adaptions within the metabolic network of P. putida upon exposure to oxidative stress, the relative abundance of key metabolites was quantified by LC-MS/MS. Out of the metabolites measured, the species that showed the most significant changes in terms of abundance were intermediates of the PP pathway, the EMP route and the TCA cycle—albeit in opposite directions (Fig. 2a). In particular, the relative abundance of 6PG (key metabolic node where oxidative and phosphorylative pathways for glucose processing converge), R5P, Ru5P, Xu5P, and S7P augmented significantly in H2O2-stressed cells, ranging from a 2- (Xu5P) to 7.6-fold (Ru5P) increase. At the same time, the relative abundance of glycolytic intermediates of the EMP route decreased roughly by half (e.g., F6P) and up to 70% (e.g., Pyr) when cells were exposed to oxidative stress. Intermediates of the TCA cycle shared the same fate, and their relative abundance was reduced at least by 60% (e.g., isocitrate, ICT) and up to ca. 90% (e.g. α-ketoglutarate, succinate, and malate) in stressed cells. The changes in the relative abundance of other metabolites in the biochemical network of P. putida KT2440 upon exposure to the oxidative stress agent are listed in Dataset S1. Taken together, these results indicate increased fluxes through the PP pathway coupled to a decrease in the catabolic activities of the EMP route and the TCA cycle during oxidative stress. However suggestive, these metabolomic data cannot be used to infer relative or net fluxes, and flux ratio analysis was employed to piece together the contribution of fluxes within the upper metabolism of P. putida to the pool of glycolytic intermediates.
a Relative abundance of selected metabolites, grouped according to the biochemical block they belong to (i.e. PP pathway, EMP pathway, and TCA cycle). Relative metabolite abundance is expressed as the ratio between H2O2-induced oxidative stress and control (Ctrl.) conditions, derived from summed ion abundance of all isotopes (counts). Data from experiments in the presence of [1-13C1]- and [6-13C1]-glucose (Glc.) are averaged. Bars represent the mean value of metabolite abundance ± standard deviations obtained in triplicate measurements of samples from three independent experiments per labeled substrate. Statistical comparisons between the metabolite abundance ratios (with a ratio = 1 indicating no difference between stressed cultures and control experiments) were assessed by the Student’s t test with Welch’s correction. Single (*) and double asterisks (**) identify significant differences at the p < 0.05 and p < 0.01 levels, respectively. Actual p values for the metabolite ratios in the PP pathway (stressed versus control experiments) were p = 0.0052, 0.0031, 0.0092, 0.0043, and 0.0055, indicated in the same order as the bar graph. For the metabolites in the EMP pathway (stressed versus control experiments), the values were p = 0.0198, 0.0083, 0.0079, and 0.0412. For the metabolites in the TCA cycle (stressed versus control experiments), the values were p = 0.0322, 0.0049, 0.0931, and 0.0074. b Changes in selected metabolic flux ratios in upper metabolism upon exposure of the cells to H2O2. The 13C-labeled substrate used in each experiment is indicated. Bars represent averages from three independent experiments, and standard deviations were calculated using the covariance matrices of the respective mass distribution vectors by applying the Gaussian law of error propagation. c Relative pathway contribution to the G6P and F6P pools. The input of each of the metabolic pathways to the sugar phosphate pool under oxidative stress conditions is indicated with different colors. All abbreviations used in this figure are as indicated in the legend to Fig. S1. CDW, cell dry weight.
Exposure to oxidative stress increases cyclic operation of PP but not ED pathway
The relative contributions of reverse (i.e., gluconeogenic) flux from triose phosphates or through the PP pathway to the hexose phosphate pool was elucidated by using relative flux ratios derived from [1-13C1]- and [6-13C1]-glucose-labeling experiments. Calculation of relative ratios around the hexose node was possible given that the reaction catalyzed by 6-phosphofructo-1-kinase is absent in P. putida [55, 56], and by considering the reactions mediated by GntZ (6PG dehydrogenase) and Edd (6PG dehydratase) to be irreversible. In particular, using [6-13C1]-glucose as a tracer enables the resolution of fluxes from the cyclic PP and ED pathways back to hexose phosphates because the C6 position is maintained in both routes and can lead to double-labeled hexose phosphate molecules. Flux ratio analysis supported the notion that fluxes through the PP pathway significantly responded to H2O2 addition (Fig. 2b), in particular in the oxidative branch of this metabolic pathway.
The metabolic network of P. putida KT2440 and biochemical reactions considered in this study are summarized in Fig. S1 and Table S1, respectively. Experiments conducted in the presence of [1-13C1]-glucose suggested that the contribution of glucose kinase (Glk, v3 in Fig. S1) to the G6P pool was not significantly affected by oxidative stress, whereas the fraction of 6PG from G6P via G6P dehydrogenase (Zwf, v7 in Fig. S1) increased by at least 3.5-fold (as deduced from experiments conducted with either [6-13C1]- or [U-13C6]-glucose) in stressed P. putida (Fig. 2b). The increase was even more pronounced for the flux ratio reflecting the fraction of F6P from the PP pathway (4.5-fold higher in cultures treated with H2O2 than in control experiments). The full list of flux ratios, including those for reactions in lower catabolism, is presented in Table S2, and the GC-MS data are given in Dataset S2. Moreover, the contribution of the ED, EMP, and PP routes to the total G6P and F6P pool mirrored the data obtained by flux ratio analysis (Fig. 2c). The PP pathway took a predominant role in feeding the pool of both metabolites when cells were challenged with H2O2. Taken together, these results provided further evidence for a flux rerouting from periplasmic glucose oxidation to the NADPH-generating PP pathway as a direct metabolic response to oxidative stress. We next quantified net fluxes as the basis for accurate estimation of NADPH production in response to oxidative stress.
Redistribution of net fluxes in P. putida as a response to sub-lethal oxidative stress
Combining the quantitative physiology data (Fig. 1c) with the flux ratios as additional constraints (Fig. 2b and Table S2), we estimated net fluxes through the biochemical network (Fig. 3). The values for all the fluxes within the biochemical network are provided in Table S3. Since the relative contribution of the direct phosphorylation of gluconate to 6PG and that of the indirect (NADPH-dependent) route though 2-ketogluconate could not be resolved from 13C-labeling data, we determined the in vitro enzyme activities of gluconate kinase (GnuK, i.e. direct phosphorylation of the sugar) and 2-ketogluconate-6-phosphate reductase (KguD, i.e. 6PG from 2-ketogluconate) as a proxy. GnuK and KguD activities in cell-free extracts of P. putida KT2440 were similar under control and oxidative stress conditions, and we thus used a GnuK/KguD activity ratio of 0.87/0.13 to constrain the flux split at the gluconate branch point.
All fluxes, calculated under control conditions (a) and in the presence of H2O2-induced oxidative stress (b), were normalized to the specific glucose uptake rate (qS), and the width of each arrow is scaled to the relative flux. Flux values represent the mean ± standard deviations from at least three biological replicates, after integration of physiological and metabolomics data together with flux ratio analysis. Dashed lines indicate that no significant flux through the corresponding biochemical step was detected under the conditions tested. Abbreviations used for the metabolic intermediates and the main metabolic blocks within the biochemical network are given in the legend to Fig. S1. CDW, cell dry weight.
Under control conditions, glucose was processed mostly through the periplasmic pathway with about 90% of the sugar uptake rate channeled via Gcd (Fig. 3a). Most of the gluconate fed the 6PG node, with a very high flux through the ED route and a ca. 10% recycling of trioses phosphate through the EDEMP cycle. Upon exposure to H2O2, P. putida substantially rerouted its carbon fluxes (Fig. 3b). Periplasmic glucose oxidation decreased, and instead much more glucose was imported into the cytoplasm and converted to 6PG through the NADP+-dependent Zwf. Since the glucose uptake rate was essentially identical in the two experimental conditions, this large relative flux change reflects a likewise large change in absolute fluxes. In particular, a major change in flux distribution was detected in the non-oxidative branch of the PP pathway. Exposure of the cells to H2O2 increased the GntZ flux by more than sixfold, which fed the pools of other pentoses phosphate (as suggested by the measurement of their relative abundance increase, Fig. 2). The backward flux in this route fed the F6P and trioses phosphate node, resulting in carbon recycling via glucose-6-phosphate isomerase (Pgi, in the gluconeogenic F6P → G6P direction) and Zwf (G6P → 6PG). Accordingly, fluxes through Pgi and Zwf in H2O2-treated cells increased by 5.2- and 4.5-fold, respectively. The ED route was still the predominant catabolic pathway for 6PG under oxidative stress, albeit with a slight decrease in the flux values as compared to control conditions. Gluconeogenic fluxes via fructose-1,6-bisphosphate aldolase (Fda) and fructose-1,6-bisphosphatase (Fbp) had a small increase (1.5-fold) in cultures exposed to oxidative stress, and flux through the lower, catabolic branch of the EMP pathway (from glyceraldehyde-3-phosphate to PEP and Pyr) was reduced by ca. 10% under oxidative stress. The diminished input of the ED and the EMP catabolism under oxidative stress propagated into a relatively low flux towards acetyl-CoA (i.e., Pyr dehydrogenase) and fluxes within the TCA cycle. No significant activity of the glyoxylate shunt could be detected under any of the experimental conditions tested. Anaplerotic routes within the Pyr shunt, typical of glucose-grown P. putida, had a likewise lower activity in H2O2-stressed cells. The impact of oxidative stress-mediated re-arrangement of carbon fluxes on redox metabolism was investigated next.
The re-arrangement of net fluxes in P. putida under oxidative stress increases NADPH supply
Considering that the flux through Zwf and GntZ, two of the major NADP+-dependent dehydrogenases in the biochemical network, showed a significant increase under oxidative stress, the overall net NADPH production increased substantially. This reconfiguration presumably matches the increased NADPH demand to regenerate glutathione (and other antioxidants) for H2O2 reduction. To assess this impact quantitatively, the net rate of NADPH production (RNADPH) was calculated according to Σi (riF − riC), where riF and riC represent the rates of NADPH formation and consumption, respectively, of all major dehydrogenases potentially carrying significant flux in the biochemical network (Fig. S1). Zwf, GntZ, Gap, Icd, Mdh, and MaeB were considered as potential inputs to NADPH formation for this analysis, whereas biomass formation and the KguD activity were included as NADPH sinks. The values of ri were derived from the actual previously calculated net fluxes, and the cofactor specificity of all dehydrogenases under both saturating, in vitro conditions and non-saturating, quasi in vivo conditions (Table S4) was used to derive the ri values. By combining these parameters, RNADPH was estimated for P. putida in both control and H2O2-treated experiments (Fig. 4). When using cofactor specificities determined under saturating, in vitro conditions, NADPH formation and consumption in control experiments added up (i.e. RNADPH = 0, Fig. 4a), whereas adjusting these specificities according to non-saturating, quasi in vivo determinations revealed a slight catabolic overproduction of reducing power characteristic of strain KT2440 (RNADPH = 1.3 mmol gCDW−1 h−1, Fig. 4b). In H2O2-stressed cells, RNADPH was >0 irrespective of the cofactor specificity used to calculate the individual ri values (Fig. 4c, d). In particular, when applying the specificity coefficients derived from non-saturating, more realistic quasi in vivo determinations, RNADPH increased by 3.6-fold (Fig. 4d) as compared to the control condition (Fig. 4b). Taken together, these results expose a ca. 50% surplus in NADPH formation under oxidative stress conditions (i.e. the flux of NADPH generation increased to 14.4 ± 0.3 mmol gCDW−1 h−1 upon exposure of the cells to H2O2), that becomes available to counteract ROS accumulation.
Overall NADPH balances and sensitivity analysis for the wild-type strain under control (a, b, e, f) or H2O2-induced oxidative stress conditions (c, d, g, h) using cofactor specificities of major dehydrogenases under saturating conditions (left column) or quasi in vivo conditions (right column). NADPH formation was determined from carbon fluxes through redox cofactor-dependent reactions (Fig. 3 and Table S1) multiplied by experimentally determined relative cofactor specificities. NADPH consumption was calculated from the requirements for biomass production and the actual NADPH-dependent KguD activity. The overall rates of NADPH formation (green) and NADPH consumption (red) are individually indicated (a–d). Dependence of the overall rates of NADPH turnover on the relative GnuK/KguD flux ratio and the cofactor specificity of Zwf derived from saturating conditions or quasi in vivo conditions (e–h). Actual values for the net NADPH balance are given for the experimentally determined NADP+ specificity of Zwf (0.937) as a function of the relative GnuK/KguD flux ratio, with the calculated net rate for each condition indicated with a red dot. CDW, cell dry weight.
To ensure that our NADPH balance estimates are not biased by key assumptions, we assessed the sensitivity of net NADPH formation as a function of the GnuK/KguD activity ratio at the gluconate node and the NADP+-specificity of the dehydrogenase reaction catalyzed by Zwf. The Zwf activity of P. putida KT2440 is represented by three isozymes (i.e. Zwf, ZwfA, and ZwfB; Table S1). Even when we have determined NADP+ and NAD+ specificities for the overall Zwf reaction with in vitro assays, it is likely that the individual isozymes may have different cofactor preferences, which would have a direct effect on the overall redox balance if expressed differently. Irrespective of the cofactor preferences (0.672 and 0.937 either under saturating conditions or quasi in vivo conditions, respectively; see Table S4), RNADPH was relatively more responsive to changes in the GnuK/KguD flux ratio in control, non-stressed experiments (Fig. 4e, f). The opposite was true for H2O2-treated cells, with a strong dependence of RNADPH on the output of the Zwf flux (Fig. 4g, h) mainly because the relative flux through Zwf increased in comparison to the control experiment. These calculations demonstrate that the NADPH surplus observed in stressed cells largely stems from the NADP+-specificity of Zwf, with the GnuK/KguD flux ratio playing a minor role on RNADPH values.
Altered activities of enzymes involved in glucose catabolism under oxidative stress
To further validate the results of in vivo distribution of metabolic fluxes, in vitro measurements of key enzymatic activities were carried out in cell-free extracts of P. putida KT2440 incubated under the same conditions. In agreement with the relative flux distribution (Fig. 3), there was a significant decrease in the activities of enzymes within the peripheral reactions for glucose oxidation under oxidative stress conditions (Fig. 5a). The observed decrease was particularly noticeable in the case of the specific Gad activity (gluconate → 2-ketogluconate), which was roughly halved in cells exposed to H2O2. The hexose kinase activity, accordingly, increased by twofold. Changes in the activity of the two NADPH producing dehydrogenases of the PP pathway were more prominent, with the specific Zwf and GntZ activities increasing by 4.7- and 9.2-fold under oxidative stress, respectively. The in vitro activities of two enzymes of the ED pathway, in contrast, remained essentially unchanged within the experimental error in both control and H2O2-stressed cells. Although we cannot rule out that gene expression (and thus, protein levels) could have been affected in the time frame of the experiments, the results of the in vitro enzyme determinations are fully consistent with the previous observations in terms of metabolic flux distribution. Taken together, these results underline the importance of PP pathway for NADPH production when cells are challenged with an oxidative stress agent. We next explored the connection between catabolic NADPH overproduction and mechanisms that counteract oxidative stress such as the glutathione-dependent redox system.
a Enzyme activity ratios were calculated from the specific activity for each of the indicated reactions assessed under H2O2-induced oxidative stress and control (Ctrl.) conditions. Each bar represents the mean value of the corresponding ratio ± standard deviations of triplicate measurements from at least two independent experiments, and the horizontal dashed line indicates an activity ratio = 1 (i.e. no changes in enzymatic activities across experimental conditions). Statistical comparisons between enzyme activity ratios were assessed by the Student’s t test with Welch’s correction. Single (*) and double asterisks (**) identify significant differences at the p < 0.05 and p < 0.01 levels, respectively. Actual p values for the Glk and Gad activity ratios in the glucose conversion routes were p = 0.0019 and 0.0293, respectively. Actual p values for the Zwf and GntZ activity ratios in the PP pathway were p = 0.0087 and 0.0096, respectively. Circled numbers identify the enzymes in the biochemical network of Fig. S1. b Glutathione metabolism in P. putida KT2440. The key activities involved in biosynthesis and recycling of the reduced (GSH) and oxidized (GS–SG) forms of glutathione are indicated along with the corresponding PP identifiers. c Enzymatic determination of total glutathione and the fraction of the oxidized and reduced form. Bars represent the mean value of the corresponding parameter ± standard deviations of duplicate measurements from at least five independent experiments, with individual measurements indicated as empty circles, and the asterisk (*) identifies significant differences between stressed cells and control conditions at P < 0.05 as assessed by the Student’s t test with the Bonferroni correction. Actual p values for the total glutathione content and the GSH/GS–SG ratio between H2O2-treated and control cultures were p = 0.0361 and 0.0117, respectively. CDW, cell dry weight. d Impact of the carbon substrate on the growth of P. putida KT2440 upon an oxidative challenge. Normalized growth coefficients, representing the fraction of the specific growth rate (μ) in the presence of 3 mM H2O2 as compared with that of control (Ctrl.) conditions, were calculated in cultures using glucose, α-ketoglutarate (α-KG) or glycerol as the carbon source. Each bar represents the mean value of the normalized growth coefficients ± standard deviations of triplicate independent experiments, while the arrows and the accompanying percentages indicate the relative reduction in the growth rate under oxidative stress.
The glutathione system of P. putida links H2O2-induced overproduction of NADPH to ROS quenching in a carbon source-dependent fashion
Thiols play several roles in bacteria, which includes maintaining the redox balance, quenching ROS and nitrogen reactive species, and detoxifying other toxins and stress-inducing factors [57]. In most organisms, the major thiol involved in such processes is the tripeptide glutathione (GSH, γ–L-glutamyl-L-cysteinylglycine). The formation of GSH from L-glutamate and L-cysteine in strain KT2440 is catalyzed by GshAB (glutamate-cysteine ligase and glutathione synthase, respectively; Fig. 5b). The glutathione cycle, connecting the reduced (GSH) and the oxidized (GS–SG) forms of the thiol, acts as a first-line reductant of ROS (Fig. 5b) via glutathione peroxidases (e.g., PP_0777 and PP_1686) and glutaredoxins. Glutathione reductase (Gor, PP_3819) then replenishes the GSH pool by using NADPH as the reducing currency. Since the glutathione system of P. putida has not been explored thus far from a biochemical point of view, the cellular content of both GSH and GS–SG was assessed by means of a biochemical assay in both control and H2O2-stressed cultures (Fig. 5c). Cells exposed to H2O2 had a 1.6-fold higher total content of glutathione (i.e. GSH and GS–SG) than control experiments, and the composition of the total pool changed significantly upon exposure to oxidative stress. The GSH/GS–SG molar ratio, which reflects the fraction of reduced thiol in the cells, increased by threefold in H2O2–stressed P. putida. This observation suggests that the reduced form of glutathione can be sufficiently replenished by the cellular NADPH sources upon an oxidative stress—directly linking the increased NADPH turnover with the pool of antioxidants.
Considering that there is a direct connection between the carbon source on the overall metabolic regime adopted by P. putida—and thus, the operation of the EDEMP cycle—we explored oxidative stress tolerance when using substrates processed through mainly glycolytic, gluconeogenic, and mixed biochemical routes (Fig. 5d). To this end, the bacterial growth of cultures developed on the presence of glucose, α-ketoglutarate or glycerol was monitored upon an oxidative challenge by adding H2O2 at 3 mM (i.e., twice the amount used in the experiments above, and thereby affecting growth rates). While the growth of P. putida with glycolytic substrates, which foster NADPH-forming routes as explained above, was not significantly affected under stressful conditions (with a reduction of μ of only 14%), the other substrates promoted higher levels of sensitivity. The use of a mostly gluconeogenic substrate (α-ketoglutarate) led to μ values in cultures added with H2O2 of around half those observed in control experiments. Glycerol, which is partially processed by gluconeogenic and glycolytic modules of central metabolism [58], promoted a level of tolerance to oxidative stress in between the two other substrates (Fig. 5d). Although growth phenotypes are the consequence of several layers of multi-component regulation, these results expose the connection between the operativity of different segments of the versatile metabolism of strain KT2440, the coarse NADPH supply and the level of tolerance to oxidative stress.
Discussion
Environmental microorganisms are exposed to a variety of physicochemical perturbations that require dynamic and concerted responses to ensure cell survival [59]. On this background, P. putida KT2440 provides a unique experimental system to explore the interplay between stress and metabolic adaption. Broadening this understanding has not only a fundamental interest, but also practical consequences in view of the growing demand of robust agents for in situ environmental catalysis [60] and metabolic engineering [61]. In actively growing bacterial cells, metabolic fluxes are adapted to meet energy and redox demands [62], and a trade-off between steady-state growth rates and physiological adaptability in fluctuating environments seems to be a conserved feature of growing bacteria [63]. The distribution of metabolic fluxes under conditions where growth is not affected, on the other hand, is a largely unexplored aspect. Environmental bacteria do not divide actively in the natural niches they colonize, yet the cells are exposed to stressful conditions under these near-zero growth state—as mimicked in our current experimental design. Moreover, exploring nongrowing conditions has a significant impact on the design of industrial biocatalysts and bioprocesses [64], where stressful conditions are prevalent—and P. putida KT2440 is naturally primed to endow such circumstances [65].
The results of the present study strengthen the notion that network-wide adaption of metabolic fluxes [62, 66] accompanies (and plausibly, enables) the phenomenon traditionally described—at a genetic and regulatory level—as the general stress response [52, 53, 67, 68]. Metabolic adaption to oxidative stress has been revisited by Christodoulou et al. [30], demonstrating that reserve flux capacity in the PP pathway empowers E. coli to rapidly respond to H2O2 by increased flux through Zwf and Gnd. It was postulated that inhibition of Zwf caused by NADPH (either by a competitive, allosteric or combined mechanism) is lifted under these conditions—releasing ‘reserve’ flux through the oxidative PP pathway. Moreover, the activity of glyceraldehyde-3-phosphate is lowered due to ROS-mediated damage, which causes a metabolic jam in the metabolite pools of the PP route, as previously shown in redox-stressed yeast [69]. Product inhibition of Zwf is relieved due to NADPH drainage to counteract oxidative stress [70], and the suddenly enlarged 6PG pool is now channeled via (NADPH-forming) Gnd. Recently, such an NADPH replenishing mechanism catalyzed by Zwf and Gnd has been recognized as a general strategy in enteric bacteria, yeast, and mammalian cells [71]. In this study, we asked how P. putida, which lacks a linear, EMP-based glycolysis and processes glucose mostly through the conversion of the sugar into gluconate, reacts to oxidative stress. Since glycolysis is not an alternative glucose degradation route (therefore, P. putida cannot alter the split ratio between the EMP and PP pathways), we hypothesized that the EDEMP cycle could contribute to increased NADPH biosynthesis under oxidative stress via the PP pathway branch.
Metabolite changes and flux analysis revealed that the lower (EMP) glycolysis and the PP pathway were affected by H2O2 as compared with untreated controls. In a simplified model of the upper pathways for sugar processing (Fig. 6a), the output of NADPH is largely governed by the fluxes through Zwf and GntZ (Gnd)—that, in turn, respond to the split between oxidative conversion of glucose into gluconate or phosphorylation into G6P, and the recycling activity of the EDEMP cycle. As such, the overall NADPH turnover is dependent on rs (Fig. 6b), further modulated by the fraction of glucose oxidized in the bacterial periplasm. Consequently, a low activity of the EDEMP cycle contributes to NADPH formation when cells grow in the absence of any oxidative stress. Upon an oxidative challenge, the backward flux from the PP and ED routes to F6P increases >3-fold, and the ED flux through lower glycolysis decreases due to re-arrangement. The boost in the activities through the cyclic PP pathway is most remarkable, as P. putida maintains very low fluxes within this metabolic block under normal growth conditions [72]. Such metabolic adaption mechanism entails an increase in the fluxes through the enzymes needed for cyclic operation that burn carbon into CO2 to generate NADPH. This alteration of activities that lead to NADPH formation could stem (at least partially) from de novo transcriptional rearrangements—although the expression level of genes encoding the main NADP+-dependent dehydrogenases in the central catabolism of P. putida KT2440 seem not be overly affected by the H2O2 treatment [52]. One way or the other, under these conditions the overall biomass yield is similar to that observed in control, non-stressed cultures because formation of CO2 by mechanisms connected to cellular respiration (i.e. pyruvate dehydrogenase and the TCA cycle) is reduced. P. putida deploys a survival strategy that seems to differ from that reported for the close relative species P. fluorescens, where production of α-ketoacids and ATP formation via converging metabolic mechanisms prevails under oxidative stress [68, 73]—yet the observed responses are dependent on the carbon substrates used for both species.
a Schematic representation of the upper metabolism of P. putida KT2440. Several biochemical reaction have been lumped to illustrate the main routes for carbon circulation (see Fig. S1 for details and abbreviations). Note that the total rate of carbon uptake (qS) is split between glucose phosphorylation and oxidation to gluconate, such that qS = rp + rox. The overall cycling flux of trioses phosphate towards hexoses phosphate is indicated as rc and the flux through the PP pathway shunt is termed rs. Cofactors other than NADPH have been omitted in the drawing for the sake of clarity. b Functional relationship between rNADPH, the rate of NADPH formation within the simplified metabolic network of (a), and the fluxes through the oxidative loop for glucose processing and the PP pathway shunt. All (arbitrary) values are given as a fraction of qS, and the experimental conditions tested in this work are indicated with red dots (Ctrl. control conditions). c General model for flux distribution in the upper metabolic domain of P. putida. Under normal growth conditions, glucose is processed mostly through its oxidative conversion to gluconate, and the EDEMP cycle provides intermediates for biomass, with a very low flux through the PP pathway. Upon oxidative stress conditions (exerted either by endogenous or external perturbations), a rapid increase of fluxes via the PP pathway shunt replenishes the intermediates within upper metabolism and provides a direct source of NADPH that can be coupled to anti-oxidant defense mechanisms against reactive oxygen species (ROS). Fluxes predominant under each condition are highlighted.
Increasing evidence points to glutathione peroxidase as a key survival mechanism in stressed bacteria [74,75,76]. Glutathione peroxidase-dependent reduction of ROS requires a continuous supply of NADPH for regeneration [77]. Upon oxidative stress, the glutathione-based detoxification of ROS drains a lot of NADPH that must be replenished via the mechanisms explained above. Importantly, the results of this study reflect a situation where the immediate, early metabolic response (which in E. coli takes place within the first 10 s after exposure to the stress agent) is blended with early transcriptional responses (e.g., activation of inducible defense mechanisms, which take several minutes to occur). Experimental evidence indicates that there is a multilayered array of defense strategies against oxidative stress in P. putida, including (i) several catalases and superoxide dismutases [78, 79], (ii) ferredoxin-NADP+ oxido-reductases [80], (iii) polyphosphate-dependent NAD+ and NADH kinases [81], and (iv) stress-induced transhydrogenation activities [44]. However important, these mechanisms can be classified as (relatively) late responses to oxidative insults as they mostly depend on transcriptional regulation, and the re-arrangement of metabolic fluxes to ensure NADPH supply appears to represent a first line of defense in several bacterial species [82].
Regardless of which carbon source is used, the EDEMP cycle forms the metabolic core of P. putida, and virtually every anabolic or catabolic transaction of the cell involves the operation of this metabolic architecture. The reallocation of metabolic resources for the sake of generating redox currency NADPH under stressful conditions is not an exclusive feature of glucose-dependent growth (Fig. S1 and Table S1). On the contrary, the EDEMP cycle is the major biochemical device that endows this species with the metabolic versatility needed to tolerate redox stress independently of the main nutrients at hand. Moreover, the data presented above accounts to a large extent for the ability of P. putida [83] to withstand a variety of environmental insults—not just oxidative conditions. Desiccation and osmotic pressure [84], organic solvents [85], starvation [86], radiation [87] (including sunlight [88]), high temperature [89], and other physicochemical and nutritional circumstances [90] that often prevail in natural niches typically converge toward producing ROS. This is a well-documented phenomenon in the case of antibiotic treatment: the ensuing, indirectly produced ROS seem to be the ultimate agent of bacterial death [91] beyond the primary action on basic cellular functions. Any mechanism that counteracts ROS may thus result in shifting the survival window of the bacterium towards a more hostile environment, as it seems to be the case for P. putida. In either circumstance, an abundant, conditional NADPH supply is a key feature to ensure cell survival and durability in changing environmental (and industrial) scenarios. Given the capability of P. putida to adapt to stressful conditions, it cannot come as a surprise that this species has received considerable attention in recent years as a biological agent of choice for in situ bioremediation [60] in a time of global pollution challenges [92].
Data availability
Metabolite levels and 13C-labeling data are available in the Supplementary Information files. Other datasets generated and analyzed in this study are available from the corresponding authors on request.
References
Belda E, van Heck RGA, López-Sánchez MJ, Cruveiller S, Barbe V, Fraser C, et al. The revisited genome of Pseudomonas putida KT2440 enlightens its value as a robust metabolic chassis. Environ Microbiol. 2016;18:3403–24.
Nogales J, Mueller J, Gudmundsson S, Canalejo FJ, Duque E, Monk J, et al. High-quality genome-scale metabolic modelling of Pseudomonas putida highlights its broad metabolic capabilities. Environ Microbiol. 2020;22:255–69.
Nelson KE, Weinel C, Paulsen IT, Dodson RJ, Hilbert H, Martins dos Santos VAP, et al. Complete genome sequence and comparative analysis of the metabolically versatile Pseudomonas putida KT2440. Environ Microbiol. 2002;4:799–808.
Volke DC, Calero P, Nikel PI. Pseudomonas putida. Trends Microbiol. 2020;28:512–3.
Nikel PI, Chavarría M, Danchin A, de Lorenzo V. From dirt to industrial applications: Pseudomonas putida as a synthetic biology chassis for hosting harsh biochemical reactions. Curr Opin Chem Biol. 2016;34:20–9.
Wasi S, Tabrez S, Ahmad M. Use of Pseudomonas spp. for the bioremediation of environmental pollutants: a review. Environ Monit Assess. 2013;185:8147–55.
Akkaya Ö, Pérez-Pantoja D, Calles B, Nikel PI, de Lorenzo V. The metabolic redox regime of Pseudomonas putida tunes its evolvability toward novel xenobiotic substrates. mBio. 2018;9:e01512–18.
Brune A, Frenzel P, Cypionka H. Life at the oxic–anoxic interface: microbial activities and adaptations. FEMS Microbiol Rev. 2000;24:691–710.
Asaeda T, Barnuevo A. Oxidative stress as an indicator of niche-width preference of mangrove Rhizophora stylosa. For Ecol Manag. 2019;432:73–82.
Hinsinger P, Bengough AG, Vetterlein D, Young IM. Rhizosphere: biophysics, biogeochemistry and ecological relevance. Plant Soil. 2009;321:117–52.
Borer B, Tecon R, Or D. Spatial organization of bacterial populations in response to oxygen and carbon counter-gradients in pore networks. Nat Commun. 2018;9:769.
Morya R, Salvachúa D, Thakur IS. Burkholderia: an untapped but promising bacterial genus for the conversion of aromatic compounds. Trends Biotechnol. 2020;38:963–75.
Moreno-Forero SK, van der Meer JR. Genome-wide analysis of Sphingomonas wittichii RW1 behaviour during inoculation and growth in contaminated sand. ISME J 2015;9:150–65.
Cadet J, Douki T, Gasparutto D, Ravanat JL. Oxidative damage to DNA: formation, measurement and biochemical features. Mut Res. 2003;531:5–23.
Imlay JA, Sethu R, Rohaun SK. Evolutionary adaptations that enable enzymes to tolerate oxidative stress. Free Radic Biol Med. 2019;140:4–13.
Arts IS, Gennaris A, Collet JF. Reducing systems protecting the bacterial cell envelope from oxidative damage. FEBS Lett. 2015;589:1559–68.
Dan Dunn J, Álvarez LAJ, Zhang X, Soldati T. Reactive oxygen species and mitochondria: a nexus of cellular homeostasis. Redox Biol. 2015;6:472–85.
Imlay JA. The molecular mechanisms and physiological consequences of oxidative stress: lessons from a model bacterium. Nat Rev Microbiol. 2013;11:443–54.
Jamieson DJ. The effect of oxidative stress on Saccharomyces cerevisiae. Redox Rep. 1995;1:89–95.
Eleutherio E, Brasil AA, França MB, de Almeida DSG, Rona GB, Magalhães RSS. Oxidative stress and aging: learning from yeast lessons. Fungal Biol. 2018;122:514–25.
Brown AJP, Cowen LE, di Pietro A, Quinn J. Stress adaptation. Microbiol Spectr. 2017;5. https://doi.org/10.1128/microbiolspec.FUNK-0048-2016.
Boronat S, Domènech A, Paulo E, Calvo IA, García-Santamarina S, García P, et al. Thiol-based H2O2 signalling in microbial systems. Redox Biol. 2014;2:395–9.
Doroshow JH. Glutathione peroxidase and oxidative stress. Toxicol Lett 1995;82-83:395–8.
Palma M, DeLuca D, Worgall S, Quadri LEN. Transcriptome analysis of the response of Pseudomonas aeruginosa to hydrogen peroxide. J Bacteriol. 2004;186:248.
Reva ON, Weinel C, Weinel M, Böhm K, Stjepandic D, Hoheisel JD, et al. Functional genomics of stress response in Pseudomonas putida KT2440. J Bacteriol. 2006;188:4079–92.
Chechik G, Oh E, Rando O, Weissman J, Regev A, Koller D. Activity motifs reveal principles of timing in transcriptional control of the yeast metabolic network. Nat Biotechnol. 2008;26:1251–9.
Fuhrer T, Sauer U. Different biochemical mechanisms ensure network-wide balancing of reducing equivalents in microbial metabolism. J Bacteriol. 2009;191:2112–21.
Rui B, Shen T, Zhou H, Liu J, Chen J, Pan X, et al. A systematic investigation of Escherichia coli central carbon metabolism in response to superoxide stress. BMC Syst Biol. 2010;4:122.
Kuehne A, Emmert H, Soehle J, Winnefeld M, Fischer F, Wenck H, et al. Acute activation of oxidative pentose phosphate pathway as first-line response to oxidative stress in human skin cells. Mol Cell. 2015;59:359–71.
Christodoulou D, Link H, Fuhrer T, Kochanowski K, Gerosa L, Sauer U. Reserve flux capacity in the pentose phosphate pathway enables Escherichia coli’s rapid response to oxidative stress. Cell Syst. 2018;6:569–78.
Nikel PI, Martínez-García E, de Lorenzo V. Biotechnological domestication of pseudomonads using synthetic biology. Nat Rev Microbiol. 2014;12:368–79.
del Castillo T, Ramos JL, Rodríguez-Herva JJ, Fuhrer T, Sauer U, Duque E. Convergent peripheral pathways catalyze initial glucose catabolism in Pseudomonas putida: genomic and flux analysis. J Bacteriol. 2007;189:5142–52.
Sudarsan S, Dethlefsen S, Blank LM, Siemann-Herzberg M, Schmid A. The functional structure of central carbon metabolism in Pseudomonas putida KT2440. Appl Environ Microbiol. 2014;80:5292–303.
Sánchez-Pascuala A, Fernández-Cabezón L, de Lorenzo V, Nikel PI. Functional implementation of a linear glycolysis for sugar catabolism in Pseudomonas putida. Metab Eng. 2019;54:200–11.
Nikel PI, Chavarría M, Fuhrer T, Sauer U, de Lorenzo V. Pseudomonas putida KT2440 strain metabolizes glucose through a cycle formed by enzymes of the Entner-Doudoroff, Embden-Meyerhof-Parnas, and pentose phosphate pathways. J Biol Chem. 2015;290:25920–32.
Kohlstedt M, Wittmann C. GC-MS-based 13C metabolic flux analysis resolves the parallel and cyclic glucose metabolism of Pseudomonas putida KT2440 and Pseudomonas aeruginosaPAO1. Metab Eng. 2019;54:35–53.
Kukurugya MA, Mendonca CM, Solhtalab M, Wilkes RA, Thannhauser TW, Aristilde L. Multi-omics analysis unravels a segregated metabolic flux network that tunes co-utilization of sugar and aromatic carbons in Pseudomonas putida. J Biol Chem. 2019;294:8464–79.
Mendonca CM, Yoshitake S, Wei H, Werner A, Sasnow SS, Thannhauser TW, et al. Hierarchical routing in carbon metabolism favors iron-scavenging strategy in iron-deficient soil Pseudomonas species. Proc Natl Acad Sci USA. 2020;117:32358–69.
Cabiscol E, Tamarit J, Ros J. Oxidative stress in bacteria and protein damage by reactive oxygen species. Intern Microbiol 2000;3:3–8.
Bagdasarian M, Lurz R, Rückert B, Franklin FCH, Bagdasarian MM, Frey J, et al. Specific purpose plasmid cloning vectors. II. Broad host range, high copy number, RSF1010-derived vectors, and a host-vector system for gene cloning in Pseudomonas. Gene 1981;16:237–47.
Sambrook J, Russell DW. Molecular cloning: a laboratory manual. 3rd edn. Cold Spring Harbor: Cold Spring Harbor Laboratory; 2001.
Nikel PI, Pettinari MJ, Ramírez MC, Galvagno MA, Méndez BS. Escherichia coli arc Amutants: metabolic profile characterization of microaerobic cultures using glycerol as a carbon source. J Mol Microbiol Biotechnol. 2008;15:48–54.
Nikel PI, Chavarría M. Quantitative physiology approaches to understand and optimize reducing power availability in environmental bacteria. In: McGenity TJ, Timmis KN, Nogales-Fernández B, editors. Hydrocarbon and Lipid Microbiology Protocols–Synthetic and Systems Biology—Tools. Heidelberg, Germany: Humana Press; 2016. pp 39–70. https://doi.org/10.1007/8623_2015_84.
Nikel PI, Pérez-Pantoja D, de Lorenzo V. Pyridine nucleotide transhydrogenases enable redox balance of Pseudomonas putida during biodegradation of aromatic compounds. Environ Microbiol. 2016;18:3565–82.
Nikel PI, Pérez-Pantoja D, de Lorenzo V. Why are chlorinated pollutants so difficult to degrade aerobically? Redox stress limits 1,3-dichloroprop-1-ene metabolism by Pseudomonas pavonaceae. Philos Trans R Soc Lond B Biol Sci. 2013;368:20120377.
Rühl M, Rupp B, Nöh K, Wiechert W, Sauer U, Zamboni N. Collisional fragmentation of central carbon metabolites in LC-MS/MS increases precision of ¹³C metabolic flux analysis. Biotechnol Bioeng. 2012;109:763–71.
Nanchen A, Fuhrer T, Sauer U. Determination of metabolic flux ratios from 13C-experiments and gas chromatography-mass spectrometry data: protocol and principles. Methods Mol Biol. 2007;358:177–97.
Wijker RS, Sessions AL, Fuhrer T, Phan M. 2H/1H variation in microbial lipids is controlled by NADPH metabolism. Proc Natl Acad Sci USA. 2019;116:12173–82.
Zamboni N, Fischer E, Sauer U. FiatFlux – a software for metabolic flux analysis from 13C-glucose experiments. BMC Bioinformatics. 2005;6:209.
Corona F, Martínez JL, Nikel PI. The global regulator Crc orchestrates the metabolic robustness underlying oxidative stress resistance in Pseudomonas aeruginosa. Environ Microbiol. 2019;21:898–912.
Michie KL, Dees JL, Fleming D, Moustafa DA, Goldberg JB, Rumbaugh KP, et al. The role of Pseudomonas aeruginosa glutathione biosynthesis in lung and soft tissue infection. Infect Immun. 2020;88:00116–20.
Bojanovič K, D’Arrigo I, Long KS. Global transcriptional responses to osmotic, oxidative, and imipenem stress conditions in Pseudomonas putida. Appl Environ Microbiol. 2017;83:e03236–16.
Kim J, Park W. Oxidative stress response in Pseudomonas putida. Appl Microbiol Biotechnol. 2014;98:6933–46.
Fuhrer T, Fischer E, Sauer U. Experimental identification and quantification of glucose metabolism in seven bacterial species. J Bacteriol. 2005;187:1581–90.
Chavarría M, Nikel PI, Pérez-Pantoja D, de Lorenzo V. The Entner-Doudoroff pathway empowers Pseudomonas putida KT2440 with a high tolerance to oxidative stress. Environ Microbiol. 2013;15:1772–85.
Sánchez-Pascuala A, de Lorenzo V, Nikel PI. Refactoring the Embden-Meyerhof-Parnas pathway as a whole of portable GlucoBricks for implantation of glycolytic modules in Gram-negative bacteria. ACS Synth Biol. 2017;6:793–805.
Fahey RC. Glutathione analogs in prokaryotes. Biochim Biophys Acta. 2013;1830:3182–98.
Nikel PI, Kim J, de Lorenzo V. Metabolic and regulatory rearrangements underlying glycerol metabolism in Pseudomonas putida KT2440. Environ Microbiol. 2014;16:239–54.
Mailloux RJ, Lemire J, Appanna VD. Metabolic networks to combat oxidative stress in Pseudomonas fluorescens. Antonie van Leeuwenhoek. 2011;99:433–42.
Dvořák P, Nikel PI, Damborský J, de Lorenzo V. Bioremediation 3.0: engineering pollutant-removing bacteria in the times of systemic biology. Biotechnol Adv. 2017;35:845–66.
Weimer A, Kohlstedt M, Volke DC, Nikel PI, Wittmann C. Industrial biotechnology of Pseudomonas putida: advances and prospects. Appl Microbiol Biotechnol. 2020;104:7745–66.
Shimizu K, Matsuoka Y. Redox rebalance against genetic perturbations and modulation of central carbon metabolism by the oxidative stress regulation. Biotechnol Adv. 2019;37:107441.
Basan M, Honda T, Christodoulou D, Hörl M, Chang YF, Leoncini E, et al. A universal trade-off between growth and lag in fluctuating environments. Nature. 2020;584:470–4.
Lempp M, Lubrano P, Bange G, Link H. Metabolism of non-growing bacteria. Biol Chem. 2020;401:1479–85.
Ankenbauer A, Schäfer RA, Viegas SC, Pobre V, Voß B, Arraiano CM, et al. Pseudomonas putida KT2440 is naturally endowed to withstand industrial-scale stress conditions. Microb Biotechnol. 2020;13:1145–61.
Chakraborty S, Liu L, Fitzsimmons L, Porwollik S, Kim JS, Desai P, et al. Glycolytic reprograming in Salmonella counters NOX2-mediated dissipation of ΔpH. Nat Commun. 2020;11:1783.
Gottesman S. Trouble is coming: signaling pathways that regulate general stress responses in bacteria. J Biol Chem. 2019;294:11685–700.
MacLean A, Bley AM, Appanna VP, Appanna VD. Metabolic manipulation by Pseudomonas fluorescens: a powerful stratagem against oxidative and metal stress. J Med Microbiol. 2020;69:339–46.
Ralser M, Wamelink MM, Kowald A, Gerisch B, Heeren G, Struys EA, et al. Dynamic rerouting of the carbohydrate flux is key to counteracting oxidative stress. J Biol. 2007;6:10.
Stincone A, Prigione A, Cramer T, Wamelink MMC, Campbell K, Cheung E, et al. The return of metabolism: biochemistry and physiology of the pentose phosphate pathway. Biol Rev. 2015;90:927–63.
Christodoulou D, Kuehne A, Estermann A, Fuhrer T, Lang P, Sauer U. Reserve flux capacity in the pentose phosphate pathway by NADPH binding is conserved across kingdoms. iScience. 2019;19:1133–44.
Chavarría M, Kleijn RJ, Sauer U, Pflüger-Grau K, de Lorenzo V. Regulatory tasks of the phosphoenolpyruvate-phosphotransferase system of Pseudomonas putida in central carbon metabolism. mBio. 2012;3:e00028–12.
Bignucolo A, Appanna VP, Thomas SC, Auger C, Han S, Omri A, et al. Hydrogen peroxide stress provokes a metabolic reprogramming in Pseudomonas fluorescens: enhanced production of pyruvate. J Biotechnol. 2013;167:309–15.
Zhou H, Wang M, Smalley NE, Kostylev M, Schaefer AL, Greenberg EP, et al. Modulation of Pseudomonas aeruginosa quorum sensing by glutathione. J Bacteriol. 2019;201:e00685–18.
Wongsaroj L, Saninjuk K, Romsang A, Duang-Nkern J, Trinachartvanit W, Vattanaviboon P, et al. Pseudomonas aeruginosa glutathione biosynthesis genes play multiple roles in stress protection, bacterial virulence and biofilm formation. PLoS One. 2018;13:e0205815.
Lemire J, Alhasawi A, Appanna VP, Tharmalingam S, Appanna VD. Metabolic defence against oxidative stress: the road less travelled so far. J Appl Microbiol. 2017;123:798–809.
Staerck C, Gastebois A, Vandeputte P, Calenda A, Larcher G, Gillmann L, et al. Microbial antioxidant defense enzymes. Micro Pathog. 2017;110:56–65.
Hishinuma S, Yuki M, Fujimura M, Fukumori F. OxyR regulated the expression of two major catalases, KatA and KatB, along with peroxiredoxin, AhpC in Pseudomonas putida. Environ Microbiol. 2006;8:2115–24.
Heim S, Ferrer M, Heuer H, Regenhardt D, Nimtz M, Timmis KN. Proteome reference map of Pseudomonas putida strain KT2440 for genome expression profiling: distinct responses of KT2440 and Pseudomonas aeruginosa strain PAO1 to iron deprivation and a new form of superoxide dismutase. Environ Microbiol. 2003;5:1257–69.
Yeom J, Jeon CO, Madsen EL, Park W. In vitro and in vivo interactions of ferredoxin-NADP+ reductases in Pseudomonas putida. J Biochem. 2009;145:481–91.
Nikel PI, Chavarría M, Martínez-García E, Taylor AC, de Lorenzo V. Accumulation of inorganic polyphosphate enables stress endurance and catalytic vigour in Pseudomonas putida KT2440. Microb Cell Fact. 2013;12:50.
Ralser M, Wamelink MMC, Latkolik S, Jansen EEW, Lehrach H, Jakobs C. Metabolic reconfiguration precedes transcriptional regulation in the antioxidant response. Nat Biotechnol. 2009;27:604–5.
Wu X, Monchy S, Taghavi S, Zhu W, Ramos JL, van der Lelie D. Comparative genomics and functional analysis of niche-specific adaptation in Pseudomonas putida. FEMS Microbiol Rev. 2011;35:299–323.
Svenningsen NB, Pérez-Pantoja D, Nikel PI, Nicolaisen MH, de Lorenzo V, Nybroe O. Pseudomonas putida mt-2 tolerates reactive oxygen species generated during matric stress by inducing a major oxidative defense response. BMC Microbiol. 2015;15:202.
Ramos JL, Sol Cuenca M, Molina-Santiago C, Segura A, Duque E, Gómez-García MR, et al. Mechanisms of solvent resistance mediated by interplay of cellular factors in Pseudomonas putida. FEMS Microbiol Rev. 2015;39:555–66.
Kivisaar M. Mechanisms of stationary-phase mutagenesis in bacteria: mutational processes in pseudomonads. FEMS Microbiol Lett. 2010;312:1–14.
Slade D, Radman M. Oxidative stress resistance in Deinococcus radiodurans. Microbiol Mol Biol Rev. 2011;75:133–91.
Castro-Alférez M, Polo-López MI, Fernández-Ibáñez P. Intracellular mechanisms of solar water disinfection. Sci Rep. 2016;6:38145.
Marcén M, Ruiz V, Serrano MJ, Condón S, Mañas P. Oxidative stress in E. coli cells upon exposure to heat treatments. Int J Food Microbiol. 2017;241:198–205.
Imlay JA. Where in the world do bacteria experience oxidative stress? Environ Microbiol. 2019;21:521–30.
Dwyer DJ, Belenky PA, Yang JH, MacDonald IC, Martell JD, Takahashi N, et al. Antibiotics induce redox-related physiological alterations as part of their lethality. Proc Natl Acad Sci USA. 2014;111:E2100–9.
de Lorenzo V, Marlière P, Solé R. Bioremediation at a global scale: from the test tube to planet Earth. Micro Biotechnol. 2016;9:618–25.
Acknowledgements
This work was funded by The Novo Nordisk Foundation (individual grant NNF10CC1016517, and LiFe, NNF18OC0034818), the European Union’s Horizon 2020 Research and Innovation Programme under grant agreement No. 814418 (SinFonia) and the Danish Council for Independent Research (SWEET, DFF-Research Project 8021-00039B) to PIN. This work was also funded by the MADONNA (H2020-FET-OPEN-RIA-2017-1-766975), BioRoboost (H2020-NMBP-BIO-CSA-2018), SYNBIO4FLAV (H2020-NMBP/0500) and MIX-UP (H2020-Grant 870294) Contracts of the European Union and the S2017/BMD-3691 InGEMICS-CM Project of the Comunidad Autónoma de Madrid (European Structural and Investment Funds) to VDL.
Author information
Authors and Affiliations
Contributions
PIN and VDL conceived the whole study, analyzed the data, and wrote the manuscript with input from all other authors. PIN, TF, MC, and ASP carried out the quantitative physiology and tracer experiments. TF executed all metabolomics and flux analysis measurements and associated data curation and analysis. PIN and ASP measured in vitro enzymatic activities. TF and US discussed and drafted the analysis of metabolic flux distributions. All authors contributed to the discussion of the experimental results and conclusions of the study.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nikel, P.I., Fuhrer, T., Chavarría, M. et al. Reconfiguration of metabolic fluxes in Pseudomonas putida as a response to sub-lethal oxidative stress. ISME J 15, 1751–1766 (2021). https://doi.org/10.1038/s41396-020-00884-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00884-9
This article is cited by
-
The long-chain flavodoxin FldX1 improves the biodegradation of 4-hydroxyphenylacetate and 3-hydroxyphenylacetate and counteracts the oxidative stress associated to aromatic catabolism in Paraburkholderia xenovorans
Biological Research (2024)
-
Synthetically-primed adaptation of Pseudomonas putida to a non-native substrate D-xylose
Nature Communications (2024)
-
Recent advances on the systems metabolically engineered Pseudomonas species as versatile biosynthetic platforms for the production of polyhydroxyalkanoates
Systems Microbiology and Biomanufacturing (2024)
-
Increasing cellular fitness and product yields in Pseudomonas putida through an engineered phosphoketolase shunt
Microbial Cell Factories (2023)
-
GC/MS-based 13C metabolic flux analysis resolves the parallel and cyclic photomixotrophic metabolism of Synechocystis sp. PCC 6803 and selected deletion mutants including the Entner-Doudoroff and phosphoketolase pathways
Microbial Cell Factories (2022)