Abstract
Nitrite is a pivotal component of the marine nitrogen cycle. The fate of nitrite determines the loss or retention of fixed nitrogen, an essential nutrient for all organisms. Loss occurs via anaerobic nitrite reduction to gases during denitrification and anammox, while retention occurs via nitrite oxidation to nitrate. Nitrite oxidation is usually represented in biogeochemical models by one kinetic parameter and one oxygen threshold, below which nitrite oxidation is set to zero. Here we find that the responses of nitrite oxidation to nitrite and oxygen concentrations vary along a redox gradient in a Pacific Ocean oxygen minimum zone, indicating niche differentiation of nitrite-oxidizing assemblages. Notably, we observe the full inhibition of nitrite oxidation by oxygen addition and nitrite oxidation coupled with nitrogen loss in the absence of oxygen consumption in samples collected from anoxic waters. Nitrite-oxidizing bacteria, including novel clades with high relative abundance in anoxic depths, were also detected in the same samples. Mechanisms corresponding to niche differentiation of nitrite-oxidizing bacteria across the redox gradient are considered. Implementing these mechanisms in biogeochemical models has a significant effect on the estimated fixed nitrogen budget.
Similar content being viewed by others
Introduction
Nitrogen is required for all life and its availability in natural systems affects global climate through control of biological carbon cycling and nitrous oxide fluxes. The loss and retention of fixed inorganic nitrogen are thought to be spatially separated: loss is restricted to anoxic environments, while the oxidation of nitrite (NO2−) to nitrate (NO3−) by nitrite-oxidizing bacteria (NOB) is considered to be an aerobic process. Anaerobic NO2− oxidation in anoxic layers of marine oxygen minimum zones (OMZs) has been suggested [1,2,3], but has not been experimentally proven. It could not be shown that NO2− oxidation is independent of O2 because O2 concentrations were not measured in the experimental samples. Unavoidable O2 contamination below previous analytical detection limits during sampling cannot be quantified but may be enough to support measured NO2− oxidation rates. NO2− oxidation by microaerophilic NOB has been recognized [4, 5] and may explain some apparently anaerobic NO2− oxidation rates in oxic–anoxic interfaces of OMZs. Whether measured NO2− oxidation rates in anoxic seawaters were also fueled by O2 introduced by sampling and incubation manipulations or were actually anaerobic rates remains unknown. Contradictory results in O2 manipulation experiments complicate the story: additions of O2 to samples from oxic–anoxic interfaces caused decreased [3, 6] or increased [5] NO2− oxidation, suggesting that these dynamic environments contain diverse NOB with different O2 preferences.
Diverse NOB were recently detected in both oxic [7] and anoxic seawater [8] via single-cell sequencing and metagenomics. Among the seven genera of NOB, Nitrococcus and Nitrospina-like NOB or Candidatus Nitromaritima were found in hypoxic and anoxic seawater [6, 8,9,10]. Dominant OMZ NOB were absent in oxic oceanic regions, and NOB from oxic seawater were rare in OMZs [8]. The difference in NOB communities under different conditions suggests niche differentiation among marine NOB. It also implies that different NOB may have different O2 sensitivities and substrate affinities.
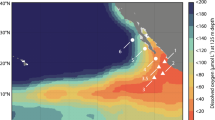
To explore whether the composition of the NOB assemblage could be responsible for the previous contradictory results, we investigated substrate kinetics and effects of O2 on NO2− oxidation at depths with distinct redox features. We sampled the oxycline (oxic), the top of the anoxic oxygen deficient zone (ODZ, the oxic–anoxic interface) and the anoxic ODZ core at two stations in the Eastern Tropical North Pacific (ETNP) OMZ to capture different responses by the diverse NOB communities occupying distinct niches. ODZ, also known as anoxic marine zones (AMZs) [11], refers to the anoxic zone of the OMZ and OMZ refers to an oceanic region that includes an ODZ and oxic water layers. In addition, to determine whether NO2− oxidation could occur without O2, we measured NO2− oxidation rates in 15N stable isotope tracer experiments with samples collected from anoxic ODZs while monitoring O2 using highly sensitive LUMOS sensors (detection limit ≈1 nM) [12, 13].
Results and discussion
Substrate kinetics and the effects of O2 addition on NO2 − oxidation
NO2− oxidation was detected at several depths under various oxygen concentrations from an open ocean station (PS2) and a coastal station (PS3) (Fig. S1a). In most samples from the oxycline and top of the ODZ, NO2− oxidation rates increased with increasing NO2− concentration, consistent with Michaelis–Menten kinetics (Fig. 1). Half-saturation constants (Km) for these samples were similar to or slightly higher than those previously determined in low-nitrite OMZ [3] and non-OMZ [14] seawater, but much lower than those determined in cultures or other environments [14], confirming that the in situ assemblage of marine NOB is adapted to low-nitrite conditions. NO2− oxidation rate from the ODZ core did not respond to NO2− addition, suggesting saturation of the rate at ambient NO2− concentrations (~2 µM, Fig. S1b and Table S1). O2 addition significantly increased the Km of NO2− oxidation at the ODZ top at station PS2 (Fig. 1f), indicating the co-existence of NOB with significantly different affinities for nitrite, as well as different oxygen preferences in this interface environment.
Samples were from oxycline (a, b, c, d), ODZ top (e, f, g, h), and ODZ core (i, j, k, l), respectively. Closed circles indicate incubation under low O2 conditions (0.3–0.7 μM), and open circles indicate high O2 conditions (2.3–8.7 μM). Solid lines indicate a Km significantly larger than zero. The adjusted R square (r2) for each fitting curve is shown. Error bars around each point show standard errors of linear regression slope calculated from time course incubations with five bottles. Sampling depths for station PS2: Oxycline (90 m), ODZ top (120 m), and ODZ core (250 m); for station PS3: Oxycline (33 m), ODZ top (80 m), and ODZ core (160 m).
Unlike NO2−, O2 did not elicit simple Michaelis–Menten-like responses in NO2− oxidation rate. O2 addition stimulated NO2− oxidation rates in samples collected at oxycline depths (Fig. 2a, b), but had no effect on NO2− oxidation rates at the top of the ODZ at either station (Fig. 2c, d). Notably, NO2− oxidation rate at the ODZ core at station PS2 (Fig. 2e) appeared to be stimulated by 0.6 µM O2 but decreased to zero at higher O2 concentrations. To our knowledge, this is the first observation of full inhibition of NO2− oxidation by O2 addition, and it occurred only in samples collected from the anoxic ODZ core. There was no significant response to added O2 at the ODZ core at station PS3 (Fig. 2f). Different responses to O2 of NO2− oxidation at different depths imply different O2 tolerances as the basis of niche differentiation among OMZ NOB [8]. In particular, inhibition of NO2− oxidation by O2 addition (>0.6 µM) at the ODZ core at station PS2 implies that the dominant NOB in these particular samples are adapted to anoxic environments. The lack of response to O2 at the top (Fig. 2c, d) or core of the ODZ (Fig. 2f) implies that diverse NOB with different O2 preferences co-existed and at least part of the in situ assemblage is something other than conventional obligate aerobic NOB.
Samples were from oxycline (a, b), ODZ top (c, d), and ODZ core (e, f), respectively. Original O2 in all samples was purged with helium, and O2 concentrations shown on the X-axis were measured O2 concentrations in sample bottles after O2 additions. Dashed line means that Km (half-saturation constant; see “Methods”) was not significantly different from zero. The adjusted R square (r2) for the fitting curve is shown. Error bars around each point show standard errors of the linear regression slope calculated from time course incubations with five bottles. Sampling depths for station PS2: Oxycline (93 m), ODZ top (120 m), and ODZ core (250 m); for station PS3: Oxycline (50 m), ODZ top (60 m), and ODZ core (160 m).
NO2 − oxidation and nitrite-oxidizing communities in the absence of O2 consumption
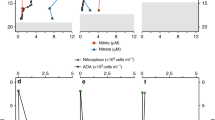
The lowest O2 concentrations in the above incubations (0.3 µM at station PS2 and 2.6 µM at station PS3) with samples from the anoxic ODZ core (Fig. 2e, f) might still be high enough to support aerobic NO2− oxidation. Therefore, week-long incubation experiments with samples from the anoxic ODZ cores were performed to determine whether NO2− oxidation could occur independently of the O2 concentration. NO2− oxidation occurred in samples collected from the anoxic ODZ core at both stations, where the measured O2 consumption rate was too slow to support the observed oxidation rates (Fig. 3a–d). O2 concentration increased during the first few days (Figs. 3a, b, S2a, b), probably due to release of trace O2 remaining in the rubber septa. Aerobic NO2− oxidation occurred at day 2.5 at station PS2 and at 1.5 and 2 days at station PS3 (Fig. 3c, d). Then net O2 consumption commenced, which we attribute to microbial respiration and the complete deoxygenation of the septa. NO2− oxidation occurred again (Fig. 3c, d) when ambient O2 was very low (i.e., 3 nM at station PS2 and 12 nM at station PS3, Fig. S2a, b) and O2 concentration stabilized after 4 days at station PS2 and 6 days at station PS3 (Fig. 3a, b). These later oxidation rates were much higher than could be accounted for by the contemporaneous O2 consumption rate (Fig. 3a, b) based on the stoichiometry of aerobic nitrite oxidation (Table S2). NO2− oxidation occurred with and without O2, indicating either the co-existence of aerobic and anaerobic NOB at both stations, or a switch between metabolisms by the same microbes. Abiotic reactions can be ruled out (killed controls, see “Methods”). The maximum NO2− oxidation rate (31.7 ± 6.3 nM d−1) at station PS2 was detected in the absence of O2 consumption and the maximum rate (38.64 ± 12.4 nM d−1) at station PS3 was aerobic. Together with evidence of O2 inhibition at station PS2 but not station PS3 (Fig. 2e, f), these results suggest that a larger proportion of NOB in the anoxic ODZ core at station PS2 were adapted to anaerobic conditions.
All rates are in nM/d or nM-N/d. a, b O2 increasing (positive values) and consumption (negative values) rates (black line) (nM/d) and estimated O2 consumption rates from nitrite oxidation if the nitrite oxidation had been aerobic (blue circles) (nM/d); (c, d) nitrite oxidation rates (blue) (NO2− ox, nM-N/d) and estimated anaerobic nitrite oxidation rates from anammox (red); (e, f) denitrification rates (Denit, nM-N/d) (black stars) and estimated N2 production rates from nitrite disproportionation (blue circles); (g) anammox rates (Amx, nM-N/d). Error bars around each point show standard errors of linear regression slope calculated from time course incubations. Pink shade indicates the occurrence of nitrite oxidation (rates with error bars overlapping with zero line of Y-axis were not considered significant). O2 concentrations and excess nitrate or N2 used to calculate these rates are shown in Fig. S2.
To explore NOB communities in the anoxic ODZ cores at the two ETNP stations, we obtained metagenomes from the two ODZ core depths where the week-long incubation samples were collected. Two OMZ NOB “species” and several other putative NOB, recently identified in samples from the OMZ of the Eastern Tropical South Pacific (ETSP) [8] recruited ETNP metagenomic reads from this study, indicating that very similar NOB likely reside in the ETNP ODZ core samples where NO2− oxidation was detected. NOB MAGs constructed here (ETNP PS2 MAG-11 and ETNP PS3 MAG-54) belong to the same species as MAG-2 from the ETSP based on their high average nucleotide identity (ANI) (99.5 and 99.1%, Table S3). We used the most complete draft genomes of the two novel OMZ NOB “species”, MAG-1 and MAG-2 from the ETSP (Table S4), to estimate their relative abundance in the total microbial communities in the ODZ cores at our two ETNP stations by mapping new sequencing reads from the ETNP to the two ETSP MAGs. Relative abundance of NOB, expressed as RPKG (reads per kilobase per genome equivalent) [15], varied between the two stations (Fig. 4), but MAG-2 was always more abundant than MAG-1 (Table S5). Based on the relative enrichment of MAG-1 at oxic–anoxic interfaces and MAG-2 at anoxic ODZ cores, MAG-1 was proposed as a microaerophile while MAG-2 might be an anaerobic specialist [8]. A terminal oxidase with high affinity for O2 was encoded by MAG-1 but was missing in MAG-2 [8]. Although anaerobic metabolisms were not confirmed in MAG-2, MAG-1 and MAG-2 may have different O2 sensitivities. The relative abundance of MAG-2 was 9.8-fold higher than MAG-1 at station PS2, but only 1.9-fold higher at station PS3 (Table S5), consistent with the greater relative dominance of apparently anaerobic NO2− oxidation (Fig. 3c) and the inhibition of NO2− oxidation by O2 (Fig. 2e) at station PS2. We also estimated the relative abundance (RPKG) of anammox and other (putative) NOB by mapping ETNP metagenomic reads obtained in this study to previously identified OMZ nxrB (nitrite oxidoreductase) gene sequences (Fig. 4). The two most abundant nxrB genes at both stations belonged to the putative nxrB cluster and they cannot be linked to any known NOB or anammox. The abundance of these unknown genes, representing potentially unknown metabolisms, suggests that other undiscovered microbes could be even more important than the known species and MAGs in the metabolism of nitrite in the ODZ.
Y-axis shows IDs of nxrB genes. The presence of nxrB genes from clusters defined previously [8] shown in different colors indicates the presence of diverse nitrite oxidizers. Putative nxrB clustered between Nitrospina nxrB and Anammox nxrB, and Unknown nxrB did not fall into any known nxrB clusters.
Potential mechanisms corresponding to NOB niche differentiation along redox gradients
NO2− oxidation in distinct redox layers (i.e., the top of the ODZ and the core of the ODZ) can be explained by different mechanisms (Fig. S3). NOB respire O2 in order to oxidize nitrite in the oxic layer. At the top of the ODZ, the interface where O2 is usually undetectable, transient intrusions [16] could support NO2− oxidation. When O2 is undetectable, cryptic oxygen cycling (i.e., supported by in situ O2 production in the deep chlorophyll maximum, attributed to abundant Prochlorococcus) [12] could support the in situ rate, even though all incubations in this study were in the dark. Some of the NOB at the top of the ODZ were found to have very high affinity for O2, higher than their nitrification partner ammonia oxidizers in the same environment [5]. Thus, these NOB could use the trace amount of O2 which is available at this oxic–anoxic interface. Alternatively, iodate is proposed as a possible oxidant for NO2− [17] and iodate flux from the oxic layer is possible at the ODZ top.
In the ODZ core, several mechanisms must be considered: (a) Neither O2 nor iodate is available (Fig. S3) in the ODZ core. Iodate and iodide concentration measurements made in the same month or year that these rates were measured show that iodate was usually depleted in the ETNP ODZ core [18]. Iodate concentrations are nearly constant near the secondary nitrite maximum inside ODZs [18]; thus iodate flux is not sufficient to support observed NO2− oxidation rates. In addition, iodate reduction was detected in oxyclines but was usually below detection limit in anoxic ODZs when iodate was added in the presence of nitrite [19], which indicates that microbes in those samples could not use iodate to oxidize nitrite.
(b) Transient intrusion of O2 has been reported [20] but is very rare in the core [16]. Anaerobic NO2− oxidation observed in incubation experiments above suggests that this reaction could occur without O2.
(c) Chlorophyll concentration was very low (Fig. S1b) in ODZ samples and the ODZ core was dark, so photosynthesis could not support an O2 flux for nitrite oxidation in situ or in the vials.
(d) The oxidation of NO2− by Mn4+ or Fe3+ is thermodynamically favored only at very low pH (<6) [21]. Even if enzymatic oxidation by these metals is possible at higher pH, total dissolved Mn (0–5 nM) and Fe (0–1 nM) concentrations are too low in OMZs [22, 23] to support the observed nitrite oxidation rates.
(e) Anammox bacteria anaerobically oxidize NO2− to NO3− while simultaneously using NH4+ to reduce NO2− to N2 [24]. Based on known reaction stoichiometry (Table S2) [24], the anammox rates measured in the same bottles where we detected NO2− oxidation rates at station PS2 (Fig. 3g) accounted for 7% and 2% of the measured anaerobic NO2− oxidation rates at days 5 and 5.5, respectively (Fig. 3c). At station PS3, anammox was not detected. Thus, anammox alone could not explain anaerobic NO2− oxidation.
(f) The reversible nitrite oxidoreductase enzyme [25] has been proposed to catalyze the bidirectional exchange between NO2− and NO3−, independent of net concentration change [26, 27]. This “futile” cycle could lead to NO2− and NO3− isotopic fractionation without generating energy to support cell growth. Thus, this process alone might explain the isotope enrichment in NO3− from labeled 15NO2− (Fig. S2c, d), but cannot explain inhibition of NO2− oxidation by O2 at station PS2 (Fig. 2e), the presence of NOB at both stations (Fig. 4), transcriptionally active NOB in the ODZs of ETNP and ETSP OMZs (Fig. S4), or the increasing number of nitrite oxidoreductase enzymes with decreasing O2 in the OMZ [28]. This suggests that in situ NO2− oxidation at both stations occurs by some energy generating mechanism, at least some of the time, to support NOB growth (Fig. S3).
(g) NO2– disproportionation
was suggested as a potential explanation for nitrite oxidation carried out by NOB adapted to anoxic environments [6, 29]. This reaction is thermodynamically favorable [30, 31] although it has not been detected in nature. Disproportionation, if confirmed, constitutes a third pathway of nitrogen loss in addition to anammox and denitrification. Because the two N atoms in N2 must come from NO2− during both NO2− disproportionation and denitrification, the isotope signatures of N2 produced from both pathways in the tracer incubations would be identical. Thus, previously measured “denitrification” using stable isotope tracer experiments might actually be a combination of canonical denitrification plus NO2− disproportionation. Based on stoichiometry (Eq. (1)), N2 produced from NO2− disproportionation could be responsible for 20–100% of the measured rate of N2 production from NO2− based on measured N2 production and NO2− oxidation rates here (Fig. 3e, f).
OMZ NOB may have the potential to carry out NO2− disproportionation by coupling NO2− oxidation and NO2− reduction (Table S6). The only gene required for this process that is missing from the ODZ MAGs is a NO dismutase gene (nod). No sequences related to the nod of Candidatus Methylomirabilis oxyfera [32] were found in the two ETSP NOB MAGs [8] or the two ETNP MAGs. Unidentified nod distinct from that of Ca. Methylomirabilis oxyfera might be present but cannot be identified due to the lack of nod references. Alternatively, chlorite dismutation following chlorate reduction to chlorite by NXR [33] could supply internal O2, although chlorate and chlorite concentrations in the ocean are vanishingly small. Chlorite dismutase (Cld) was found to be functional in Nitrobacter NOB and Nitrospira NOB [34, 35], and was proposed to provide O2 for NOB in hypoxic conditions [34]. DNA sequences encoding Cld-like proteins were present in NOB MAGs from the ETSP OMZ [8] and ETNP PS2 MAG-11 and ETNP PS3 MAG-54 despite the low MAG quality of ETNP MAGs. However, arginine173, which was proposed as a marker for Cld activity [36], was replaced by leucine in MAG-1 and MAG-2 (Fig. S5a). Although the same position of Cld-like proteins was not included in the short Cld-like protein sequences (only 120 aa) encoded by ETNP MAGs, their high similarity to the Cld-like protein of MAG-2 (Fig. S5b) suggests that the chance of finding arginine173 in Cld-like proteins of ETNP NOB MAGs was low. Thus, the primary function of Cld-like proteins in all Nitrospina-like OMZ NOB needs to be further confirmed by experiments. NOB from other genera have surprisingly versatile metabolic capabilities [6, 9, 25]. OMZ NOB may also use other yet to be discovered metabolisms to survive in anoxic waters.
Implications for the marine nitrogen budget from a biogeochemical model
We added NO2− disproportionation to an established inverse model [37] to quantitatively test the importance to the nitrogen budget of this newly proposed pathway (Fig. 5a, b). This model simulated nitrogen cycling rates in ODZs of the ETSP OMZ, where measured concentrations [38] and natural abundance isotope data [37] of nitrogen compounds were available (Table S7). All rates simulated in the revised model (Fig. 5c–j), especially NO2− reduction and NO2− oxidation through NO2− disproportionation (up to 36 nM-NO3− d−1), were a better match (i.e., the same magnitude) for the measured rates (at the same time and location of the collection of model input data, shown in Fig. 8 in Peters et al. 2016 [37]) than the rates simulated in the original model [37], which were an order of magnitude larger than the measured rates. The simulated total nitrogen loss was similar or much lower (by up to 62%) when NO2− disproportionation is included in anoxic ODZs (Fig. 5f, j) because the light NO2− and heavy NO3− in the ODZ can be attributed to the inverse isotopic fractionation of NO2− oxidation [39, 40] in addition to the fractionation of conventional nitrogen loss processes. Although the mechanisms for NO2− oxidation in anoxic waters remain to be experimentally verified, the modeling results here indicate the necessity to consider microbial niche differentiation when simulating reactions in distinct redox conditions.
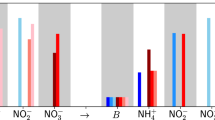
a, b A schematic of physical processes and net biochemical production or consumption (R) that affect concentrations of inorganic nitrogen compounds (a), and nitrogen cycle processes resulting in net production or consumption (R) in the anoxic core of the ODZ represented in the model (b). Black arrows: nitrate reduction and denitrification (nitrite reduction to N2). Red arrows: anammox. Blue arrows: nitrite oxidation through disproportionation. c–j Model results: (c, g) Modeled rates of nitrate reduction; (d, h) Nitrate production from nitrite disproportionation; (e, i) Denitrification (black), anammox (red), N2 production from nitrite disproportionation (blue); (f, j) Total N2 production when including nitrite oxidation through disproportionation (blue) and excluding nitrite oxidation in the model (black) at stations BB1 (c–f) and BB2 (g–j) in the ETSP OMZ. Shaded areas indicate the location of the anoxic ODZ.
NO2− oxidation is a critical pivot in the nitrogen cycle, constraining the proportion of nitrogen retention and affecting total estimates of nitrogen loss. Most current models use a fixed positive O2 threshold to determine when and where NO2− oxidation occurs in OMZs, and one parameterization to characterize its kinetics. Our results, however, demonstrate that NO2− oxidation occurred throughout the redox gradient of OMZs, and that substrate kinetics, O2 responses and potential mechanisms of NO2− oxidation differed among depths, potentially due to diverse NOB occupying distinct niches. Marine suboxic zones are predicted to expand over the next 130 years due to global climate change [41]. Dynamic regions such as coastal upwelling zones are subject to seasonal nutrient and oxygen fluctuations. Thus, responses of NO2− oxidation to NO2− and O2 determined here should be applied to other systems and could improve predictions of both long- and short-term nitrogen budgets in a changing ocean. The finding of nitrite oxidation occurring without O2 but inhibited by O2 addition in ODZs calls for follow-up research, especially the isolation of novel NOB from anoxic environments.
Methods
Experimental site
Seawater samples were collected from two stations (Fig. S1a, offshore OMZ station PS2 and coastal OMZ station PS3) in the ETNP in March and April 2018 on board R/V Sally Ride (Cruise ID: SR 1805). NO2− kinetics and O2 effects experiments (≤1 day incubation) were performed at the oxycline, the oxic/anoxic interface (top) of the ODZ and the core of the ODZ at each station (Fig. S1b and Table S1). Long incubations (6–7 days) were performed at the core of the ODZ at both stations to investigate nitrogen reaction rates at very low O2 concentrations.
Sampling and incubation experiments for NO2 − kinetics and O2 effects
Twelve 30-L Niskin bottles on a rosette with a conductivity–temperature–depth (CTD) profiler were used to collect seawater while recording in situ O2 concentration, temperature, pressure, salinity, and chlorophyll fluorescence. O2 concentration at selected ODZ depths was measured by STOX sensor (on the CTD profiler) with detection limit of 10 nM [42]. NO2− concentrations were measured by standard spectrophotometric methods onboard. O2 and NO2− concentrations were used to select sampling depths. NO3− and NO2− concentrations in incubation samples were measured on a chemiluminescence NO/NOx Analyzer (Teledyne API, San Diego, CA, USA) in the lab.
Seawater was collected from Niskin bottles into 60 mL air-tight serum bottles after overflowing three times in order to minimize O2 contamination. Serum bottles were sealed with rubber septa and aluminum seals ensuring the absence of bubbles inside bottles. Septa were deoxygenated in anaerobic chambers with three cycles of vacuum/helium flushing over a period of 1 month prior to the cruise. A helium headspace was created for each sample collected from anoxic depths (the top and the core of the ODZ), and then samples were flushed with helium for at least 15 min to remove O2 that might have been introduced during sampling.
To determine a single NO2− oxidation rate, a set of five serum bottles amended with 15NO2− tracer was incubated on board at 12 °C in the dark. Incubations were terminated in time series (one bottle at day zero (T0), two at 0.5 day and two at 1 day) by adding 0.2 mL of saturated ZnCl2. The T0 bottles served as killed controls for both tracer contamination and abiotic reactions. The observed temporal changes in isotopic signals that occurred in the live samples over a few days in the time courses would not be detected if abiotic reactions were taking place during the >3 months that elapsed before all samples were measured on the mass spec.
In the lab, NO2− was removed from samples using sulfamic acid, and NO3− in serum bottles was converted into N2O using the denitrifier method [43, 44]. Both concentration and isotopic composition of N2O were measured on a mass spectrometer (Delta Vplus, Thermo Fisher Scientific, Waltham, MA, USA) for calculating nitrite oxidation rate from the linear regression of the five nitrate concentrations at the three time points as previously described [3]. NO2− kinetics of NO2− oxidation were determined by measuring NO2− oxidation rates under different 15NO2− tracer concentrations (0.5–13.8 μM). For responses of NO2− oxidation to O2, different amounts (0, 0.2, 0.5, 1, 2, and 5 mL) of O2 saturated seawater collected from the same Niskin bottle were added into serum bottles to achieve different O2 concentrations. O2 concentrations in serum bottles were monitored by optical oxygen sensors with a detection limit of 62.5 nM (PyroScience GmbH, Aachen, Germany).
Half-saturation constant
Half-saturation constant (Km) is the nitrite concentration at which nitrite oxidation rate (V) equals half of the potential maximum rate (Vm). The curve fitting tool in MATLAB_R2015a was used to fit the Michaelis–Menten model (Eq. (2)) to determine Km.
Long incubation experiments
Longer (≥6 days) incubations were performed in 12-mL exetainers. Seawater from the core of the ODZ (250 m at station PS2, 160 m at station PS3) was sampled into 320 mL ground glass-stoppered bottles, which were immediately transferred into a N2 flushed glove bag. 15NO2− was added into these bottles to reach final concentrations of 7.24 and 8.01 µM at stations PS2 and PS3, respectively. 15NO2− labeled seawater was aliquoted into exetainers and capped within the glove bag. The septa had been stored under helium for at least 1 month. Exetainers were purged with helium for 5 min. Every 12 h, microbial activity in triplicate exetainers was terminated by adding 0.05 mL of 50% w/v ZnCl2. In the lab, N2 produced in exetainers was measured on a mass spectrometer (Europa Scientific 20–20, Crewe, UK) [45]. Denitrification and anammox rates were computed from linear regression of N2 produced at three time points. 15NO3− was measured (as described above) in the same exetainers, and NO2– oxidation rates were computed from linear regression of NO3− produced at three time points. O2 was monitored throughout the incubations in parallel exetainer vials using LUMOS sensors [12, 13]. Each O2 production or consumption rate was determined by linear regression of O2 concentrations at 32 time points. One sensing spot, glued inside an exetainer vial, allowed the optode to measure O2 concentrations every 5 min from the outside. Detection limit and resolution of LUMOS sensors was ≈1 nM.
DNA sampling, extraction, sequencing, metagenomics, and metatranscriptomics analysis
Particulate DNA samples were collected by filtration onto 0.22 µm Sterivex filters from the ODZ core samples (250 m at station PS2, 160 m at station PS3). DNA was extracted using the modified plant tissue protocol (All Prep DNA/RNA Mini Kit, Qiagen, Valencia, CA, USA), and subjected to paired-end sequencing on an Illumina MiSeq to generate over 10 million read pairs for each sample by the Genomics Core Facility of Lewis-Sigler Institute for Integrative Genomics at Princeton University. Quality control of raw reads was performed by BBDuk (DOE Joint Genome Institute, Walnut Creek, CA, USA), and assembled into contigs using metaSPAdes v3.12.0 [46] with specified options (-k 21, 33, 55, 77, 99, 127 -m 500). Metagenome-assembled genomes (MAGs) were constructed using MetaBAT v2.12.1 [47] with default (sensitive) mode and contigs longer than 1500 bp. The quality of MAGs was determined by checkM [48]. Taxonomy of MAGs was predicted by GTDB-tk [49]. ETNP PS2 MAG-11 from the ODZ core at station PS2 and ETNP PS3 MAG-54 from the ODZ core at station PS3 were identified as Nitrospina-like NOB. The taxonomy of these two NOB MAGs (ETNP PS2 MAG-11 and ETNP PS3 MAG-54) was further confirmed by comparing them with known OMZ NOB MAGs (MAG-1 and MAG-2) [8] using ANI. ANI between MAGs was assessed using enveomics [50]. MAG-2 [8], ETNP PS2 MAG-11 and ETNP PS3 MAG-54 belong to the same species (threshold for species: ANI ≥95%) based on their ANI values (Table S3). Considering the low completeness of MAG-11 and the high contamination of MAG-54 constructed from the two new metagenomes here (Table S4), we decided to use MAG-2 as the representative for this NOB species.
We estimated the relative abundance and the transcriptional activity of the two known OMZ NOB species in different oceanic regions (including the two ETNP stations in this study) by mapping metagenomes and metatranscriptomes from this and other studies to MAG-1 and MAG-2. The relative abundance of MAG-1 or MAG-2 at stations PS2 and PS3 in the ETNP was calculated as RPKG (the number of metagenomic reads obtained in this study mapped to a MAG per MAG length (kb) per genome equivalents) [15]. Genome equivalents were estimated using MicrobeCensus v1.1.1 [15]. The transcriptional activity of MAG-1 or MAG-2 in ETNP and ETSP OMZs was assessed by mapping published metatranscriptomic reads from the ETNP (PRJNA263621) [51] and the ETSP (SRA023632.1) [52] to MAG-1 and MAG-2. The relative abundance of RNA in Fig. S4 was calculated as the number of metatranscriptomic reads mapped to a MAG divided by the number of total reads. Mapping was performed by Bowtie2 [53] using “very-sensitive” mode, and only reads with a mapping quality above 20 were included as mapped reads.
In order to explore the possibility of the presence of other NOB in the ODZ core at stations PS2 and PS3, we also estimated the relative abundance of other (putative) NOB using their marker gene, nxrB (nitrite oxidoreductase). First, we downloaded previously identified (putative) nxrB sequences from the ETSP OMZ [8]. Then, we color coded the genes in Fig. 4 based on previously defined clusters in a phylogenetic tree (see Fig. 3 in [8]): the Nitrospina cluster (blue) contains nxrB grouped with cultured marine NOB, Nitrospina gracilis. Based on BLASTp search results, amino acid sequence identities between all the OMZ nxrB in this cluster and that of Nitrospina gracilis were 96.71%, 96.71%, and 96.87% for NODE_69234, NODE_114897, and NODE_102582, respectively. Only this cluster is associated with known NOB, and the nitrite oxidation capacity of all the other genes associated with nxrB needs to be confirmed. The anammox cluster (red) contains anammox nxrB sequences. The putative cluster (black) contains nxrB grouped with microbes in which nitrite oxidation capacity has not been proven. This putative cluster fell between known NOB and anammox, and was implied to represent unidentified NOB [54]. The last cluster is called unknown nxrB (gray) because neither their phylogeny nor function can be determined based on their distant relationship with known NOB or anammox nxrB. Finally, we mapped the ETNP metagenomic reads obtained in this study to each nxrB gene. Relative abundance of nxrB genes was also expressed as RPKG: relative abundance of nxrB gene = (number of mapped reads to a certain nxrB gene)/(length of this nxrB in kb)/(genome equivalents).
To explore the potential metabolisms of NOB in anoxic ODZ cores, we searched for chlorite dismutase and NO dismutase (nod) genes in the NOB MAGs. First, protein-coding sequences in the two new NOB MAGs obtained here (ETNP PS2 MAG-11 and ETNP PS3 MAG-54) were predicted by Prodigal v2.6.3 [55]. The protein-coding sequences were annotated by the best BLASTp hits against the nr protein database. DNA sequences encoding Cld were identified in both ETNP NOB MAGs. Predicted Cld amino acid sequences encoded by ETNP MAGs were too short (120 aa) to be compared to Cld of Candidatus Nitrospira defluvii, but they only had one mismatch with Cld amino acid sequences of MAG-2 from the ETSP OMZ based on MUSCLE alignment using MEGA 7 software (Fig. S5b). Thus, we looked for the arginine173, the Cld activity marker, in longer Cld sequences of MAG-1 and MAG-2 by aligning their Cld with the Cld of Candidatus Nitrospira defluvii using MUSCLE in MEGA 7 software. Since nod was not found in the ETNP MAGs via annotation and was not reported in MAG-1 and MAG-2 in the previous study [8], gene search (i.e., searching nod against NOB MAGs) was performed using Hidden Markov Models by HMMER3 [56]. Reference sequences of the search included a nod sequence retrieved from Candidatus Methylomirabilis oxyfera [32] genome (accession numbers FP565575.1), and three environmental nod sequences (accession numbers: KX364450.1, KX364454.1, and KU933965.1). Search queries were the two ETNP MAGs and two ETSP MAGs in Table S3.
Estimation of NO2 − oxidation through disproportionation using an inverse isotope model
We simulated the distribution of NO2− oxidation rates via disproportionation using a 1-D inverse isotope model [37] in the anoxic ODZ core from two stations in the ETSP OMZ (Fig. S6), where the complete suite of isotope and rate data have been previously published (Table S7). Briefly, the net biochemical production or consumption rate of each nitrogen compound (R14Ammonium, R14Nitrite, R14Nitrate, R15Nitrite, and R15Nitrate) was balanced by vertical diffusion and advection at steady state, and then five equations were used to balance measured 14NH4+, 14NO2−, 15NO2−, 14NO3−, and 15NO3− concentrations (Eqs. (3–7)). The exclusion of horizontal processes was justified [37] since the model was not run all the way to the surface, and using a constant vertical advection term does not violate continuity of the model. F is the rate of each nitrogen cycle process represented in this model (Fig. 5a, b). We modified the previous model by replacing canonical NO2− oxidation with NO2− disproportionation and using the recently determined stoichiometry of nitrate production by anammox from 0.3 to 0.16 [24]. Since OMZ NOB genomes encode nitrite oxidoreductase (catalyzes NO2−→NO3−) and nitrite reductase (catalyzes NO2−→NO), we assumed that the fractionation factor of nitrate production from nitrite by disproportionation is the same as by canonical nitrite oxidation (αDis = αNxr) based on the close clustering between OMZ NOB nxr and aerobic NOB (Nitrospina gracilis) nxr [8], and the fractionation factor of N2 produced by disproportionation is the same as by denitrification (αDisN2 = αNir). Rates of NO2− disproportionation (FDis), NO3− reduction (FNar), denitrification (FNir), and anammox (FAmx) were solved from the equations by the nonnegative least squares optimization routine (lsqnonneg) in MATLAB_R2015a as described previously [37].
Data availability
NOB MAG-1 and MAG-2 from ETSP metagenomes were submitted to NCBI under BioSample accession numbers SAMN10411459 (MAG-1) and SAMN10411419 (MAG-2). Metagenomic reads from the core of the ODZ at stations PS2 and PS3 were deposited to NCBI under the SRA accession number PRJNA505148.
Code availability
Code for the 1-D model is available at https://github.com/xinsun12/AnaerobicNitriteOxidation.
Change history
06 July 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41396-021-01032-7
References
Lipschultz F, Wofsy SC, Ward BB, Codispoti LA, Friedrich G, Elkins JW. Bacterial transformations of inorganic nitrogen in the oxygen-deficient waters of the Eastern Tropical South Pacific Ocean. Deep Sea Res A Oceanogr Res Pap. 1990;37:1513–41.
Füssel J, Lam P, Lavik G, Jensen MM, Holtappels M, Gunter M, et al. Nitrite oxidation in the Namibian oxygen minimum zone. ISME J. 2012;6:1200–9.
Sun X, Ji Q, Jayakumar A, Ward BB. Dependence of nitrite oxidation on nitrite and oxygen in low oxygen seawater. Geophys Res Lett. 2017;44:7883–91.
Penn J, Weber T, Deutsch C. Microbial functional diversity alters the structure and sensitivity of oxygen deficient zones. Geophys Res Lett. 2016;43:9773–80.
Bristow LA, Dalsgaard T, Tiano L, Mills DB, Bertagnolli AD, Wright JJ, et al. Ammonium and nitrite oxidation at nanomolar oxygen concentrations in oxygen minimum zone waters. Proc Natl Acad Sci USA. 2016;113:10601–6.
Füssel J, Lücker S, Yilmaz P, Nowka B, Van KesselMAHJ, Bourceau P, et al. Adaptability as the key to success for the ubiquitous marine nitrite oxidizer Nitrococcus. Sci Adv. 2017;3:e1700807.
Pachiadaki MG, Sintes E, Bergauer K, Brown JM, Record NR, Swan BK, et al. Major role of nitrite-oxidizing bacteria in dark ocean carbon fixation. Science. 2017;358:1046–51.
Sun X, Kop LFM, Lau MCY, Frank J, Jayakumar A, Lücker S, et al. Uncultured Nitrospina-like species are major nitrite oxidizing bacteria in oxygen minimum zones. ISME J. 2019;13:2391–402.
Daims H, Lücker S, Wagner M. A new perspective on microbes formerly known as nitrite-oxidizing bacteria. Trends Microbiol. 2016;24:699–712.
Ngugi DK, Blom J, Stepanauskas R, Stingl U. Diversification and niche adaptations of nitrospina-like bacteria in the polyextreme interfaces of Red Sea brines. ISME J. 2016;10:1383–99.
Ulloa O, Canfield DE, DeLong EF, Letelier RM, Stewart FJ. Microbial oceanography of anoxic oxygen minimum zones. Proc Natl Acad Sci USA. 2012;109:15996–6003.
Garcia-Robledo E, Padilla CC, Aldunate M, Stewart FJ, Ulloa O, Paulmier A, et al. Cryptic oxygen cycling in anoxic marine zones. Proc Natl Acad Sci USA. 2017;114:8319–24.
Lehner P, Larndorfer C, Garcia-Robledo E, Larsen M, Borisov SM, Revsbech NP, et al. LUMOS—a sensitive and reliable optode system for measuring dissolved oxygen in the nanomolar range. PLoS ONE. 2015;10:1–15.
Zhang Y, Qin W, Hou L, Zakem EJ, Wan X, Zhao Z, et al. Nitrifier adaptation to low energy flux controls inventory of reduced nitrogen in the dark ocean. Proc Natl Acad Sci USA. 2020;117:4823–30.
Nayfach S, Pollard KS. Average genome size estimation improves comparative metagenomics and sheds light on the functional ecology of the human microbiome. Genome Biol. 2015;16:1–18.
Thamdrup B, Dalsgaard T, Revsbech NP. Widespread functional anoxia in the oxygen minimum zone of the Eastern South Pacific. Deep Res Part I Oceanogr Res Pap. 2012;65:36–45.
Babbin AR, Peters BD, Mordy CW, Widner B, Casciotti KL, Ward BB. Multiple metabolisms constrain the anaerobic nitrite budget in the Eastern Tropical South Pacific. Global Biogeochem Cycles. 2017;31:258–71.
Moriyasu R, Evans N, Bolster KM, Hardisty DS, Moffett JW. The distribution and redox speciation of iodine in the Eastern Tropical North Pacific Ocean. Global Biogeochem Cycles. 2020;34:1–23.
Hardisty DS, Horner TJ, Evans N, Moriyasu R, Babbin AR, Wankel SD, et al. Limited iodate reduction in shipboard seawater incubations from the Eastern Tropical North Pacific oxygen deficient zone. Earth Planet Sci Lett. 2021;554:116676.
Tiano L, Garcia-Robledo E, Dalsgaard T, Devol AH, Ward BB, Ulloa O, et al. Oxygen distribution and aerobic respiration in the north and south eastern tropical Pacific oxygen minimum zones. Deep Res Part I Oceanogr Res Pap. 2014;94:173–83.
Luther GW. The role of one- and two-electron transfer reactions in forming thermodynamically unstable intermediates as barriers in multi-electron redox reactions. Aquat Geochem. 2010;16:395–420.
Vedamati J, Chan C, Moffett JW. Distribution of dissolved manganese in the Peruvian Upwelling and Oxygen Minimum Zone. Geochim Cosmochim Acta. 2015;156:222–40.
Kondo Y, Moffett JW. Iron redox cycling and subsurface offshore transport in the eastern tropical South Pacific oxygen minimum zone. Mar Chem. 2015;168:95–103.
Oshiki M, Satoh H, Okabe S. Ecology and physiology of anaerobic ammonium oxidizing bacteria. Environ Microbiol. 2016;18:2784–96.
Koch H, Lücker S, Albertsen M, Kitzinger K, Herbold C, Spieck E, et al. Expanded metabolic versatility of ubiquitous nitrite-oxidizing bacteria from the genus Nitrospira. Proc Natl Acad Sci USA. 2015;112:11371–6.
Wunderlich A, Meckenstock RU, Einsiedl F. A mixture of nitrite-oxidizing and denitrifying microorganisms affects the δ18O of dissolved nitrate during anaerobic microbial denitrification depending on the δ18O of ambient water. Geochim Cosmochim Acta. 2013;119:31–45.
Kemeny PC, Weigand MA, Zhang R, Carter BR, Karsh KL, Fawcett SE, et al. Enzyme-level interconversion of nitrate and nitrite in the fall mixed layer of the Antarctic Ocean. Global Biogeochem Cycles. 2016;30:1069–85.
Saito MA, McIlvin MR, Moran DM, Santoro AE, Dupont CL, Rafter PA, et al. Abundant nitrite-oxidizing metalloenzymes in the mesopelagic zone of the tropical Pacific Ocean. Nat Geosci. 2020;13:355–62.
Babbin AR, Buchwald C, Morel FMM, Wankel SD, Ward BB. Nitrite oxidation exceeds reduction and fixed nitrogen loss in anoxic Pacific waters. Mar Chem. 2020;224:103814.
Strohm TO, Griffin B, Zumft WG, Schink B. Growth yields in bacterial denitrification and nitrate ammonification. Appl Environ Microbiol. 2007;73:1420–4.
van de Leemput IA, Veraart AJ, Dakos V, De Klein JJM, Strous M, Scheffer M. Predicting microbial nitrogen pathways from basic principles. Environ Microbiol. 2011;13:1477–87.
Ettwig KF, Butler MK, Le Paslier D, Pelletier E, Mangenot S, Kuypers MMM, et al. Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature. 2010;464:543–8.
Youngblut MD, Wang O, Barnum TP, Coates JD. (Per)chlorate in Biology on Earth and Beyond. Annu Rev Microbiol. 2016;70:435–57.
Maixner F, Wagner M, Lücker S, Pelletier E, Schmitz-Esser S, Hace K, et al. Environmental genomics reveals a functional chlorite dismutase in the nitrite-oxidizing bacterium ‘Candidatus Nitrospira defluvii’. Environ Microbiol. 2008;10:3043–56.
Mlynek G, Sjöblom B, Kostan J, Füreder S, Maixner F, Gysel K, et al. Unexpected diversity of chlorite dismutases: a catalytically efficient dimeric enzyme from Nitrobacter winogradskyi. J Bacteriol. 2011;193:2408–17.
Kostan J, Sjöblom B, Maixner F, Mlynek G, Furtmüller PG, Obinger C, et al. Structural and functional characterisation of the chlorite dismutase from the nitrite-oxidizing bacterium ‘ Candidatus Nitrospira defluvii’: Identification of a catalytically important amino acid residue. J Struct Biol. 2010;172:331–42.
Peters BD, Babbin AR, Lettmann KA, Mordy CW, Ulloa O, Ward BB, et al. Vertical modeling of the nitrogen cycle in the eastern tropical South Pacific oxygen deficient zone using high-resolution concentration and isotope measurements. Global Biogeochem Cycles. 2016;30:1661–81.
Sun X, Jayakumar A, Ward BB. Community composition of nitrous oxide consuming bacteria in the oxygen minimum zone of the Eastern Tropical South Pacific. Front Microbiol. 2017;8:1–11.
Casciotti KL, Buchwald C, McIlvin M. Implications of nitrate and nitrite isotopic measurements for the mechanisms of nitrogen cycling in the Peru oxygen deficient zone. Deep Res Part I Oceanogr Res Pap. 2013;80:78–93.
Martin TS, Primeau F, Casciotti KL. Assessing marine nitrogen cycle rates and process sensitivities with a global 3D inverse model. Global Biogeochem Cycles. 2019;33:1026–47.
Resplandy L. Will ocean zones with low oxygen levels expand or shrink? Nature. 2018;557:314–5.
Revsbech NP, Larsen LH, Gundersen J, Dalsgaard T, Ulloa O, Thamdrup B. Determination of ultra-low oxygen concentrations in oxygen minimum zones by the STOX sensor. Limnol Oceanogr Methods. 2009;7:371–81.
Sigman DM, Casciotti KL, Andreani M, Barford C, Galanter M, Böhlke JK. A bacterial method for the nitrogen isotopic analysis of nitrate in seawater and freshwater. Anal Chem. 2001;73:4145–53.
Weigand MA, Foriel J, Barnett B, Oleynik S, Sigman DM. Updates to instrumentation and protocols for isotopic analysis of nitrate by the denitrifier method. Rapid Commun Mass Spectrom. 2016;30:1365–83.
Babbin AR, Keil RG, Devol AH, Ward BB. Organic matter stoichiometry, flux, and oxygen control nitrogen loss in the ocean. Science. 2014;344:406–8.
Nurk S, Meleshko D, Korobeynikov APP. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 2017;1:30–47.
Kang DD, Froula J, Egan R, Wang Z. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. PeerJ. 2015;3:e1165.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043–55.
Chaumeil P-A, Mussig AJ, Hugenholtz P, Parks DH. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics. 2019;36:1925–7.
Rodriguez-R LM, Konstantinidis KT. The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. Peer J Prepr. 2016;4:e1900v1.
Ganesh S, Bristow LA, Larsen M, Sarode N, Thamdrup B, Stewart FJ. Size-fraction partitioning of community gene transcription and nitrogen metabolism in a marine oxygen minimum zone. ISME J. 2015;9:2682–96.
Stewart FJ, Ulloa O, DeLong EF. Microbial metatranscriptomics in a permanent marine oxygen minimum zone. Environ Microbiol. 2012;14:23–40.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–60.
Lüke C, Speth DR, Kox MAR, Villanueva L, Jetten MSM. Metagenomic analysis of nitrogen and methane cycling in the Arabian Sea oxygen minimum zone. PeerJ. 2016;4:e1924.
Hyatt D, Chen G, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010;11:1–11.
Finn RD, Clements J, Eddy SR. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 2011;39:29–37.
Acknowledgements
We thank all the crew and scientists on R/V Sally Ride for assistance. In particular, we thank Nicole Travis for providing nitrite concentration data. We thank Erika Lee-Sanchez for sampling assistance. We thank Keith Shadle for helping with CTD deployment. We thank Sergey Oleynik for maintaining mass spectrometers. We thank Niels Peter Revsbech for supplying STOX sensors. We thank Yi Zhang for her suggestions on the model and Elizabeth Wallace for additional nutrient data from the incubation bottles. The cruise was supported by an NSF grant to BBW. EGR was supported by Poul Due Jensen Foundation.
Author information
Authors and Affiliations
Contributions
XS and BBW conceptualized the study. XS carried out long incubation experiments, and EGR provided oxygen concentration measurements with LUMOS in these experiments and performed in situ STOX measurements. XS and CF performed incubations in serum bottles and measured oxygen concentrations in these experiments. AJ sampled and extracted DNA. XS measured all samples, analyzed data, modeled rates, and performed metagenomic analysis. XS and BBW wrote the paper. All authors proofread the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised. The authors realized that proteins encoded by the OMZ NOB MAGs were incorrectly annotated as chlorite dismutase (Cld).
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, X., Frey, C., Garcia-Robledo, E. et al. Microbial niche differentiation explains nitrite oxidation in marine oxygen minimum zones. ISME J 15, 1317–1329 (2021). https://doi.org/10.1038/s41396-020-00852-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00852-3
This article is cited by
-
An abundant bacterial phylum with nitrite-oxidizing potential in oligotrophic marine sediments
Communications Biology (2024)
-
Microscale dynamics promote segregated denitrification in diatom aggregates sinking slowly in bulk oxygenated seawater
Communications Earth & Environment (2023)
-
Network analysis of 16S rRNA sequences suggests microbial keystone taxa contribute to marine N2O cycling
Communications Biology (2023)
-
The marine nitrogen cycle: new developments and global change
Nature Reviews Microbiology (2022)
-
Novel metagenome-assembled genomes involved in the nitrogen cycle from a Pacific oxygen minimum zone
ISME Communications (2021)