Abstract
Ombrotrophic bogs accumulate large stores of soil carbon that eventually decompose to carbon dioxide and methane. Carbon accumulates because Sphagnum mosses slow microbial carbon decomposition processes, leading to the production of labile intermediate compounds. Acetate is a major product of Sphagnum degradation, yet rates of hydrogenotrophic methanogenesis far exceed rates of aceticlastic methanogenesis, suggesting that alternative acetate mineralization processes exist. Two possible explanations are aerobic respiration and anaerobic respiration via humic acids as electron acceptors. While these processes have been widely observed, microbial community interactions linking Sphagnum degradation and acetate mineralization remain cryptic. In this work, we use ordination and network analysis of functional genes from 110 globally distributed peatland metagenomes to identify conserved metabolic pathways in Sphagnum bogs. We then use metagenome-assembled genomes (MAGs) from McLean Bog, a Sphagnum bog in New York State, as a local case study to reconstruct pathways of Sphagnum degradation and acetate mineralization. We describe metabolically flexible Acidobacteriota MAGs that contain all genes to completely degrade Sphagnum cell wall sugars under both aerobic and anaerobic conditions. Finally, we propose a hypothetical model of acetate oxidation driven by changes in peat redox potential that explain how bogs may circumvent aceticlastic methanogenesis through aerobic and humics-driven respiration.
Similar content being viewed by others
Introduction
Peatlands are a type of wetland ecosystem where net primary production exceeds organic matter decomposition, leading to the accumulation of thick layers of organic peat soil, which store over 1000 Gt of carbon globally [1]. Long-term accumulation of peat carbon is facilitated by peat mosses, primarily of the genus Sphagnum, which eco-engineer nutrient-poor peatlands through peat acidification and the production of antimicrobial compounds that inhibit high rates of Sphagnum decay, resulting in a buildup of Sphagnum biomass that impedes the flow of mineral nutrients in groundwater so that their only source of water is rain [2]. In ombrotrophic (“rain-fed”) bogs, Sphagnum mosses are the major contributors to the botanical composition of the peat, and Sphagnum-derived cell wall polysaccharides contribute to the bulk soil organic matter in acidic peat bogs [2,3,4]. These cell wall polysaccharides include a hemicellulose of xyloglucomannans, cellulose, and pectin-like rhamnogalacturonan I-type polysaccharides (termed “sphagnan”), which contain the sugars arabinose, fucose, galactose, glucose, mannose, rhamnose, and xylose, in addition to galacturonic and glucuronic acids [3,4,5,6,7].
Recently, studies have implicated Acidobacteriota populations as primary degraders of Sphagnum polysaccharides in acidic peat bogs, mainly through the activity of glycoside hydrolases (GHs) and fermentation reactions that result in acetate as a dominant metabolic end product [8,9,10]. The mode through which acetate is mineralized in acidic bogs is driven by a variety of factors including temperature, pH, moisture content, and availability of terminal electron acceptors (TEAs) [2, 11,12,13,14,15]. In aerobic regions of peat, aerobic oxidation of acetate to carbon dioxide (CO2) is the primary acetate mineralization pathway [10, 12]. Under anaerobic conditions, TEA availability drives competition for acetate between anaerobic respiratory processes and aceticlastic methanogenesis [15].
In most ombrotrophic bogs, canonical TEAs such as nitrate, sulfate, or iron are in low micromolar concentrations and likely do not contribute substantially to acetate mineralization [15, 16]. Instead, solid-state humic peat substances are potential TEAs for acetate mineralization [16,17,18,19]. The humics are rich in quinone moieties, which can accept electrons and reduce their carbonyl groups to hydroxyl groups [14]. Quinone reduction has been linked to extracellular electron transfer (EET) via heme-containing cytochromes typically seen in iron reducers [20,21,22]. Reduced quinone moieties can be abiotically reoxidized when oxygen is present, creating a renewable redox cycle, which can be linked to seasonal and localized water table fluctuations [14, 23].
After available humic substances have been reduced, aceticlastic methanogenesis may proceed. Methanogenesis from acetate fuels only a fraction of acetate mineralization in many acidic bog soils, and the process may be severely limited by low temperature, low pH, and a smaller pool of readily decomposable carbon compounds under anaerobic conditions [11, 16, 19]. Inhibition of aceticlastic methanogenesis varies by site, with some sites reporting no aceticlastic methanogenesis and an accumulation of acetate under prolonged anaerobic conditions [10, 16] and other sites reporting activity of aceticlastic methanogens under anaerobic conditions [24,25,26]. These differences in dominant methanogenic pathway are likely attributable to differences in microbial community composition across sites, as sites with high relative abundance of aceticlastic methanogens have been reported to produce methane primarily through an aceticlastic mechanism [26] and sites with low relative abundance of aceticlastic methanogens have been reported to produce methane primarily through a hydrogenotrophic mechanism [25].
While microbial populations catalyzing Sphagnum degradation and aceticlastic methanogenesis are well known, there remains great uncertainty regarding populations catalyzing respiratory acetate oxidation in peat soils and the ecological relationships between these populations, Sphagnum degraders, and methanogens. Studies show notoriously variable rates of methanogenesis and emission of methane into the atmosphere from different ombrotrophic bogs [27], which has challenged our ability to incorporate bogs into global models that forecast future rates [28]. Having a better understanding of carbon mineralization pathways and the microbial players that carry out carbon mineralization will help parameterize these models.
In this study, we sought to elucidate relationships between these populations through a global analysis of publicly available peatland metagenomes and a local analysis of metagenome-assembled genomes (MAGs) from McLean Bog (MB), an acidic kettle-hole bog in New York State. The global analysis was used to identify networks of functional genes related to Sphagnum degradation. Genes in this network were then used for a targeted analysis of MAGs recovered from MB to reconstruct carbon degradation pathways in dominant MAGs within a characteristic Sphagnum bog. We hypothesized that (i) carbon degradation genes characteristic of Sphagnum bogs would be specialized for degradation of Sphagnum biomass, (ii) Sphagnum-degrading populations in MB would be dominated by Acidobacteriota, (iii) aerobic and humics-respiring populations (not aceticlastic methanogens) would be the most abundant acetate-mineralizing populations, and (iv) acetate-mineralizing populations would be metabolically flexible, capable of switching between aerobic and anaerobic metabolism.
Materials and methods
Comparative analysis of global peatland metagenomes
Genome clustering analysis based on metagenome scaffold functional profiles via enzyme classification (EC) system annotations of annotated genes from 110 publicly available peatland metagenomes from 12 sites was performed using the principal component analysis (PCA) feature on the Joint Genome Institute (JGI) Integrated Microbial Genomes System (IMG) platform [29]. A description of the metagenomes (including accession numbers) is provided in Table S1 and site descriptions for each of the sites are provided in Table S2. The 110 metagenomes were selected by searching for high throughput-sequenced peat soil metagenomes using the IMG Genome Browser. Any metagenome within the IMG database for which a publication already existed was automatically included in the study along with any unpublished metagenomes for which permission to use was obtained. Weighted gene correlation network analysis (WGCNA) was performed to find networks of statistically correlated functional genes (annotated with EC values) across all 110 metagenome samples and then relate those networks to sample positioning along the principal component axes [30, 31]. Networks of genes associated with the PCA positioning of Sphagnum bogs were queried to identify potential gene networks associated with Sphagnum degradation and acetate mineralization. These gene networks were then used for a genome-centric case study in a kettle-hole bog to assay community functioning with respect to Sphagnum degradation and acetate mineralization.
Site description, peat sampling, and microcosm incubations
MB is an acidic kettle-hole bog located in McLean, NY, USA (42°32′55.7″ N, 76°15′58.4″ W). The site is 0.04 km2 across with a peat depth of 8 m. Vegetation is dominated by a dense lawn of Sphagnum mosses (S. angustifolium and S. magellanicum) with varying cover of ericaceous shrubs (Chamaedaphne calyculata and Vaccinium corymbosum), sedges (Eriophorum vaginatum and Dolichium spp.), and pitcher plants (Sarracenia purpurea). Water table level is above the peat surface following spring snowmelt and recedes to a maximum depth of about 25 cm below the peat surface by mid-summer [32]. Additional site characteristics are described by Osvald [33].
Raw peat was collected from the 10–40 cm depth interval of peat in August 2018 and stored in an air-tight glass jar for immediate transport to the lab. In the lab, the jar was opened in an anaerobic glovebox within 2 h post sampling and peat was homogenized using sterile scissors. A sample of raw peat was frozen and stored at −80 °C for later nucleic acid extraction. Six microcosm incubations for a related project studying the effects of sulfate availability on the peat microbiome (James, unpublished manuscript; Fig. S1) were set up in 160 mL glass vials sealed with butyl rubber stoppers and a N2 headspace. Sequences from DNA extracted from the microcosm incubations were used here to increase genomic coverage for downstream assembly of MAGs using a co-assembly of the seven metagenomes. Each of the six microcosms contained 10 g homogenized raw peat and 90 mL distilled water. Microcosms were incubated at room temperature without shaking for 45 days with mixtures of electron donors (e.g., acetate, formate, lactate) with and without exogenously supplied sulfate. A description of the microcosm conditions can be found in Table S3. One of the microcosm incubations, which received 200 µM 13C-acetate, was used for targeted stable isotope probing analysis to assess acetate metabolism in MB. At the end of the incubations, peat was recovered from the vials and was frozen and stored at −80 °C for nucleic acid extraction.
Peat samples, nucleic acid extraction, sequencing, and assembly of metagenomes
DNA was extracted from raw peat and microcosm incubations using a DNeasy PowerSoil Kit (Qiagen; Hilden, Germany) according to the manufacturer’s instructions. For the subset of microcosms receiving either 13C-acetate or 13C-formate, the heavy DNA fraction was separated via CsCl centrifugation according to the protocol by [34], with the heavy fraction defined by a refractive index of 1.72–1.77. DNA was diluted to 25 ng/µL and sent to the DOE JGI for library sequencing and assembly using ~300 ng DNA. DNA was sequenced by JGI using 2 × 150 bp Illumina Regular Fragments on an Illumina NovaSeq. Raw reads were subjected to quality control and filtration using standard JGI protocols and reads were assembled into contigs using the metaSPAdes assembler (v.3.13.0) [35]. Assembled reads were annotated by JGI using the IMG annotation pipeline (v.5.0.0) [29]. To aid in recovery of high- and medium-quality MAGs, JGI performed a co-assembly of all metagenomic libraries within the site (7 metagenomes total) using the metaSPAdes assembler (v.3.13.0) [35]. To ensure metagenomes were suitable for co-assembly, mash distances were computed for pairwise comparisons between each metagenome [36].
Recovery and analysis of MAGs
MAGs were recovered from the metagenome co-assembly using MetaBat2 (v.2.12.2) [37] and subjected to quality control using CheckM (v.1.0.6) [38]. MAGs were retained if they were classified as high-quality (>95% complete, <5% contamination) or medium-quality (>50% complete, <10% contamination) according to the guidelines reported by Bowers et al. [39]. Phylogenetic relatedness of the MAGs was determined using the PhyloPhlAn2 [40] pipeline by aligning MAGs to a database of 400 marker genes via DIAMOND (v.0.9.24) [41] while taxonomic identification was performed in KBase, an open source predictive biology platform developed by the United States Department of Energy, using the GTDB-Tk classify application, which assigns objective taxonomic classification by aligning MAGs to the Genome Taxonomy Database (GTDB) [42]. Relative abundances of MAGs within the raw peat and 13C-acetate metagenomes were calculated using a method developed by [43]. In this method, reads from the raw peat and 13C-acetate metagenomes were aligned to the MAGs using Bowtie2 [44], and TAD80 (truncated average sequencing depth) scores were calculated per MAG per metagenome by estimating sequencing depth per base using BEDTools [45] and truncating to the central 80% values using the BedGraph.tad.rb script in the enveomics collection [46]. Total genome equivalents per metagenome were predicted using MicrobeCensus [47], and the relative abundance of each MAG was calculated by normalizing TAD80 scores (MAG genome equivalents) to the total estimated number of genome equivalents. To identify MAGs that significantly changed in abundance between the raw peat and 13C-acetate metagenomes, we computed Z-scores and p values based on TAD80 values for MAGs in each metagenome.
For identification of putative EET enzymes that may play a role in quinone respiration, we adopted methods used by He et al. to query MAGs for homologs of Cyc2, a monoheme c-type cytochrome used for EET in Fe(II) oxidizers; extracellular or outer-membrane associated multiheme c-type cytochromes common in Fe(III) reducers; and porin-cytochrome c protein complexes, also common in Fe(III) reducers [20]. For identification of other specific genes/pathways of interest (e.g., GHs, multiheme cytochromes, tricarboxylic acid (TCA) cycle), the BLASTp feature in DIAMOND was used [41]. All annotated genes for a specific function in the IMG database for the MB metagenome were used as references against our MAG database, with an e-value cutoff of 1e−100.
Results
Functional comparison of global peatland metagenomes and identification of Sphagnum-degrading GHs
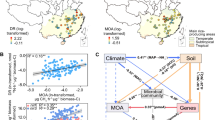
Functional clustering of 110 publicly available peatland metagenomes by PCA based on functional gene abundance by annotation with the EC system revealed positioning of metagenomes along PC1 by site description. MB positioned at the periphery near other Sphagnum-dominated acidic peat bogs (Fig. 1). Clustering along PC1 explained 84.5% of the variance while clustering along PC2 explained 7.2% of the variance. Given the large variance explained by PC1 and the positioning of MB and other acidic peat bogs on the negative PC1 axis, we next focused on modules significantly correlated with a negative positioning along PC1. WGCNA resulted in eight modules based on co-occurrence of functional genes across the 110 metagenomes (Fig. S2).Two modules significantly correlated with a negative position on PC1—one module (hereafter termed “primary bog module”) had a slightly stronger correlation (Pearson cor. = −0.63, p < 0.001) and smaller set of functional genes than the other (Fig. S2, Table S4).
Of the 45 genes in the primary bog module with a strong correlation between module membership and positioning on PC1 (Pearson cor. > 0.5), 18 encoded GHs involved in the endohydrolysis of plant polysaccharides and terminal hydrolysis of short oligosaccharides (Table 1), which result in the release of sugars and acids characteristic of Sphagnum hemicellulose [3,4,5,6,7]. These GHs are hereafter termed “Sphagnum-degrading GHs.” In addition to Sphagnum-degrading GHs, the primary bog module also contained genes for acetate activation and aldolases, epimerases, isomerases, and phosphotransferases specialized for acting on the sugars and organic acids released from the GH reactions (Table S5). The second module contained 347 genes, including some genes for acting on sugars and organic acids released from GH reactions. Downstream analyses focused primarily on analyzing the genes in the primary bog module, however, due to their strong association with Sphagnum degradation.
Quality filtering and assembly of metagenomes and MAGs
The raw peat metagenome (“MB metagenome”) contained 135,499,480 reads after quality filtering and error correction. Contig assembly by metaSPAdes resulted in 1,787,899 contigs with an N50 of 191,316 bp. The metagenome from the 13C-acetate incubation (“13C-acetate metagenome”) contained 97,177,238 reads after quality filtering and error correction. Contig assembly by metaSPAdes resulted in 1,631,687 contigs with an N50 of 217,080 bp. Mash distances between the raw peat metagenome and the six microcosm metagenomes were all small (<0.1, Table S6), indicating a high similarity of average nucleotide identity across all metagenomes [36] and a suitability for co-assembly to maximize recovery of MAGs. Thus, a co-assembly of the MB metagenome with the six metagenomes from microcosm incubations of MB peat was performed. The input to the MB co-assembly was 1,531,758,052 reads. The co-assembly resulted in 7,374,036 contigs with an N50 of 485,808 bp. The co-assembly was used to bin contigs into MAGs, resulting in the recovery of 272 draft MAGs (97 high-quality and 175 medium-quality). Characteristics of draft MAGs are provided in Table S7.
Taxonomic composition and relative abundance of MAGs in raw peat and acetate microcosms
The taxonomic composition and relative abundance of MAGs in the MB metagenome is shown in Fig. 2. Only MAGs with >10% genome coverage in the raw peat metagenome were considered in the subsequent analyses (250 of 272 MAGs). Based on an analysis of raw reads using MicrobeCensus [47], we predicted the metagenome contained ~5780 genome equivalents, and based on our analysis of TAD80 scores, our MAGs constituted ~3831 genome equivalents. The bacterial community MAGs were dominated by phylum Acidobacteriota (76 MAGs with 1267 total genome equivalents), with lower abundance of phyla Desulfobacterota (17 MAGs, 293 genome equivalents), Proteobacteria (43 MAGs, 166 genome equivalents), Verrucomicrobiota (18 MAGs, 156 genome equivalents), and Actinobacteriota (20 MAGs, 259 genome equivalents). The archaeal community was dominated by phyla Crenarchaeota (14 MAGs, 659 genome equivalents), Thermoplasmatota (10 MAGs, 457 genome equivalents), and Halobacterota (9 MAGs, 247 genome equivalents). MAGs from low abundance phyla accounted for the remaining 327 genome equivalents that mapped to 43 MAGs from 15 phyla (Table S7).
Tree construction was performed using PhyloPhlAn and phylogenetic inference was inferred using GTDB. The size of the node tip corresponds to relative abundance of the MAG in the MB metagenome. Inner circle: phylum or superphylum classification of MAGs. Green: number of Sphagnum-degrading GHs. Red: presence of complete WL pathway of acetate oxidation. Blue: presence of complete TCA cycle of acetate oxidation. Orange: acetate-oxidizing MAG with presence of terminal oxidases used in aerobic respiration. Purple: acetate-oxidizing MAG with presence of EET systems. Brown: acetate-oxidizing MAG with presence of terminal reductases used in nitrate/nitrite reduction. Black: acetate-oxidizing MAG with presence of terminal reductases used in sulfate/sulfite reduction.
In the 13C-acetate metagenome, we predicted ~3719 genome equivalents, with MAGs constituting ~2320 genome equivalents. There was an increase in the relative abundance of reads for MAGs associated with Actinobacteria (+216%), mostly explained by a significant increase in six Acidimicrobiia MAGs (Table S7). There was also a large relative increase in Proteobacteria (+80%), mostly explained by a significant increase in four Rhizobiales MAGs (Table S7). Desulfobacterota, Thermoplasmatota, and Crenarchaeota all saw large decreases in relative abundance (−64%, −58%, and −94%, respectively), the last largely explained by significant decreases in eight Bathyarchaeia MAGs (Table S7). Small changes were observed in Acidobacteriota (+13%), Verrucomicrobiota (−21%), and Halobacterota (+5.9%), (Fig. S3).
Sphagnum-degrading MAGs in MB
To identify MAGs that may play roles in Sphagnum degradation in MB, we surveyed our library of MAGs and identified those containing all, or nearly all, of the 18 Sphagnum-degrading GHs identified in the WGCNA. We identified 66 MAGs (989 genome equivalents) with at least 16 out of 18 of the relevant GHs (Fig. 2). The most abundant taxonomic groups represented were the phyla Acidobacteriota (50 MAGs, 884 genome equivalents) and Verrucomicrobiota (6 MAGs, 76 genome equivalents). The remaining ten MAGs accounted for 45 genome equivalents.
We performed targeted metabolic reconstructions of these MAGs using DIAMOND BLAST to identify which metabolic pathways were present that could utilize products of the Sphagnum-degrading GH reactions. Pathways for converting all sugars and acids released by these reactions into intermediates of glycolysis were identified. Figure 3 shows an overview of the metabolic reconstructions of the 66 Sphagnum-degrading MAGs. We observed complete pathways for the liberation of galacturonic and glucuronic acids from rhamnogalacturonan polysaccharides and downstream reactions for conversion of these acids into glyceraldehyde-3P (G3P) and pyruvate in only a small fraction of MAGs, though genes for the first step in the conversion of glucuronic and galacturonic acids were present in most MAGs. G3P also appeared to be a key intermediate for feeding xylose and arabinose into central carbon metabolism via xylulose 5-phosphate and, to a lesser extent, for fucose and rhamnose via glycerone phosphate. Pathways for converting mannose to fructose 6-phosphate for use in glycolysis and for converting glucose and galactose to glucose 6-phosphate were also identified in most MAGs. Genes for lactic acid fermentations were present in a small subset of MAGs while genes for acetate fermentation were present in most MAGs. A large fraction of MAGs also contained phosphoketolases, which catalyze the conversions of xylulose 5-phosphate and fructose 6-phosphate to acetyl-phosphate, which could further fuel acetic acid fermentation.
Acetate metabolism in MB
To identify MAGs that may play a role in the oxidation of acetate to CO2, we identified, from the total library of MAGs, those MAGs with (i) the acetate kinase (ack) and phosphotransacetylase (pta) acetate oxidation pathway and (ii) either the complete Wood–Ljungdahl (WL) pathway or the complete TCA cycle. We identified 59 MAGs with putative acetate oxidation capabilities (535 genome equivalents), including 6 MAGs with only the complete WL, 45 MAGs with only the complete TCA cycle, and 8 MAGs with both pathways.
To identify potential energy generating mechanisms for respiratory acetate oxidation, we queried these MAGs for terminal reductases involved in aerobic respiration, nitrate/nitrite reduction, sulfate/sulfite reduction, and putative EET complexes—monoheme cytochrome c (Cyc2) homologs, extracellular multiheme c-type cytochromes (MHC), and porin-cytochrome c protein complexes—which are believed to be involved in iron reduction and/or terminal reduction of quinone moieties in peat humus [20]. Among the putative acetate-oxidizing MAGs, terminal reductases for putative EET and aerobic respiration were present in nearly all MAGs, while terminal reductases for nitrate/nitrite and sulfate/sulfite reduction were present in few MAGs (Fig. 2). There was substantial overlap in MAGs with both aerobic cytochrome c oxidases and putative EET complexes. Acidobacteriota were the most abundant of these MAGs, with a low abundance of Alphaproteobacteria, Desulfuromonadata, and Desulfobacterota (Fig. 2). MAGs with extracellular multiheme c-type MHCs were dominated by Acidobacteriota, as were MAGs with porin-cytochrome complexes (PCCs), while MAGs with Cyc2 were dominated by both Acidobateriota and Alphaproteobacteria (Fig. S4). Among the 17 Acidobacteriota MAGs identified, 10 also contained all or nearly all the Sphagnum-degrading GHs.
Methanogenesis in MB
Within our MAG library, we recovered two low abundant Methanosarcinaceae MAGs (three genome equivalents) that contain genes for aceticlastic and hydrogenotrophic methanogenesis. We also recovered six MAGs within the hydrogenotrophic methanogen family Methanoregulaceae (220 genome equivalents) and 1 MAG for an unclassified hydrogenotrophic methanogen, also within phylum Halobacterota (24 genome equivalents). While the Methanoregulaceae MAGs did not substantially change in abundance upon anaerobic acetate incubation (+12%), the relative abundance of the aceticlastic Methanosarcinaceae MAG increased by 166% (Table S7).
Discussion
Although acidic peat bogs across the globe exhibit functional differences with respect to macro- and microtopography [48], the global PCA analysis shows they retain some similarities in their microbial communities’ functionality (Fig. 1), notably with respect to the abundance of specific, Sphagnum-polysaccharide degrading enzymes (Table 1). While Sphagnum bogs also contain vascular plants, Sphagnum mosses are the most abundant plant tissue below the peat surface [49]. In MB, Sphagnum-degrading enzymes are linked primarily to Acidobacteriota (Fig. 2). The high abundance of Acidobacteriota MAGs is consistent with their role as ubiquitous large polysaccharide degraders in other acidic peatlands [8, 9, 50]. Recent studies have identified specific classes of GHs responsible for Acidobacteriota-driven biopolymer degradation in bogs, including cellulases, xylanases, glucosidases, galactosidases, and xylosidases [8, 9]. Our global peatland analysis supports these previous findings, extends the list of relevant GHs (e.g., to include glucuronidases, mannosidases, rhamnosidases), and directly links these specific GHs to the chemical structure of Sphagnum cell wall polysaccharides [3,4,5,6,7].
To explore the capacity for Sphagnum degradation in MB, we performed metabolic reconstructions of 66 putative Sphagnum-degrading prokaryote MAGs. We identified an abundant subset of these MAGs with all or nearly all the enzymes required for incomplete oxidation of Sphagnum cell wall sugars to acetate, and to a lesser extent lactate, and for complete oxidation of acetate to CO2 via the TCA cycle or oxidative WL pathway (Fig. 3).
Degradation of Sphagnum cell wall polysaccharides is likely dominated by aerobic oxidation near the peat surface, though there may be a contribution from anaerobic degradation when the water table rises above the peat surface. Low levels of anaerobic degradation likely occur in the deeper, anaerobic region of the peat. Biochemical data from MB show a strong decline in hemicellulose concentrations in the top 15 cm of peat (from ~26% hemicellulose to ~11% hemicellulose) and a small decline between 15 and 145 cm (to ~8% hemicellulose) (Pipes and Yavitt, unpublished manuscript). These data support the claims that anaerobic degradation of Sphagnum occurs. The presence of both fermentative and respiratory pathways in Sphagnum-degrading MAGs suggests metabolic flexibility is an important feature of Sphagnum-degrading species in the 10–40 cm fraction of MB peat, particularly within Acidobacteriota, and that acetate is a likely major end product of fermentative Sphagnum degradation under anaerobic conditions.
Acetate as a major product of anaerobic decomposition in bogs is consistent with previous studies [10, 11], and the work here expands on these studies by linking acetate production with Sphagnum degradation. Given the speculative nature of genomic datasets, we used a highly conservative e-value cutoff (1e−100) when determining gene functional annotations. Thus, the data presented likely underestimate the actual percentage of MAGs with various carbohydrate metabolism pathways as novel enzymes or more distant homologs could potentially catalyze reactions further.
The metabolic reconstructions and clustering of acetate kinase in the primary bog module of functional genes in our WGCNA analysis further support the idea that acetate metabolism is prevalent in acidic peat bogs, even though free acetate levels are very low. Based on historically documented low levels of acetate in MB (<50 µM) [25], we explored the possibility that acetate oxidation pathways could be prevalent in MAGs from the site. We focused on oxidation through the TCA cycle and the oxidative WL pathway. Syntrophic acetate oxidation is another mechanism through which acetate could be oxidized; however, the low temperatures seen in MB should be inhibitory to this process [51], therefore our analysis did not focus on this process.
In our analysis of MAGs from MB, we identified complete acetate oxidation pathways in a subset of MAGs accounting for 535 genome equivalents (Fig. 2). While these pathways could be used acetogenically or acetotrophically, the high abundance of MAGs with terminal respiratory reductases for aerobic respiration or EET suggest the presence of an oxidative acetotrophic metabolism fueled by TEA reduction in MB, as has been observed in other peat soils, particularly at the oxic–anoxic water table interface [12].
Our analysis of terminal reductases in putative acetotroph MAGs suggests two primary mechanisms of respiratory acetate oxidation—aerobic respiration using cytochrome c oxidases and EET using Cyc2 homologs, extracellular MHCs, and PCCs. Based on phylogenetic analyses of these MAGs, we suggest that respiratory acetate oxidation would primarily be catalyzed by Acidobacteriota and to a lesser extent Alphaproteobacteria, Desulfuromonadata, and Desulfobacterota (Fig. 2). Among the 17 putative acetate-oxidizing Acidobacteriota MAGs, 10 also contained all or nearly all the Sphagnum-degrading GHs identified in the WGCNA analysis (Fig. 2). This finding suggests these populations may be specialized for complete mineralization of Sphagnum cell wall sugars to CO2 under both aerobic and anaerobic conditions. For humic substances to serve as TEAs, an oxidized pool of humic substances would need to be provided. One hypothesis is that this could occur through cyclic redox of humic substances driven by seasonal oxygenation during water table drawdown in late summer [14, 23]. Once the water table rises and humic substances are reduced, acetate may accumulate or fuel aceticlastic methanogenesis [10, 11, 13].
Unlike most anaerobic environments, in many ombrotrophic bogs, hydrogenotrophic methanogenesis typically predominates over aceticlastic methanogenesis [52, 53], which is true for methanogenesis in MB—seen here by the high relative abundance of Methanoregulaceae MAGs compared to Methanosarcinaceae and in previous surveys of MB [25, 32]. This phenomenon could be attributable to a number of factors, including competitive interactions between methanogens and acetate-oxidizing populations [15, 54, 55] and inhibition of aceticlastic methanogenesis by low temperature [56] or pH [11, 57]. Historically, anaerobic incubations of peat from MB amended with low levels of acetate have stimulated growth of aceticlastic methanogens, specifically Methanosaetaceae and Methanosarcinaceae [24, 26, 32]. Natively, Methanosarcinaceae species are at their lowest levels in the summer months when the water table is below the peat surface and thus oxygen is available to fuel aerobic respiration or oxidize quinone moieties to later serve as humic TEAs. However, the higher abundance of Methanosarcinaceae during spring and winter months when the water table is above the peat surface and the peat is anaerobic suggests that anaerobic conditions fuel aceticlastic methanogenesis over the long term of annual cycles. The increase in relative abundance of the Methanosarcinaceae MAGs in the 13C-acetate metagenome further suggests acetate supports a role for aceticlastic methanogenesis under anaerobic conditions after alternative TEAs have been reduced. This is in line with findings from other peat bogs which demonstrated acetate as a major end product of anaerobic decomposition [10, 52, 58].
The increased abundance of Acidimicrobiia (phylum Actinobacteriota) MAGs in the 13C-acetate metagenome suggests these MAGs represent acetate assimilators which were activated by the higher levels of acetate in the microcosm. While the MAGs do not have the Ack/Pta pathway for acetate activation, they do have AMP-forming acetyl-CoA transferases, which represent an alternative mode of acetate activation which is particularly important for acetate assimilation [59, 60]. Actinobacteriota have been shown to play important roles in aerobic organic matter decomposition in some Sphagnum-dominated peatlands [61]. In an experiment in a Fennoscandian tundra heath, Acidimicrobiia were identified as copiotrophic taxa in the soil [62]. In another experiment, Actinobacteriota increased in abundance in response to higher dissolved organic carbon concentrations during recovery from acidification [63]. In the context of these previous studies, we suggest the increase in Acidimicrobiia seen in the 13C-acetate metagenome is a result of a shift from a native oligotrophic environment in the bog to a copiotrophic environment in the microcosm incubation supplemented with 1 mM acetate as a carbon source. Likewise, the increase in abundance of Rhizobiales MAGs, specifically Beijerinckieaceae, may indicate a shift toward copiotrophy in the incubations as these generalist chemoorganotrophs are associated with growth on simple carbon substrates [50, 64].
Conclusion
In our global analysis of peatland metagenomes, we show that GHs with specific activity for Sphagnum cell wall sugar components are characteristic of acidic peat bogs, as are genes for acetate metabolism. Further, our work expands on previous studies characterizing Acidobacteriota-driven degradation of Sphagnum mosses by linking Sphagnum degradation and terminal acetate mineralization in single populations. We show that a distinct subset of Sphagnum-degrading populations in MB have the genomic capability for complete oxidation of Sphagnum cell wall sugars through versatile coupling of acetate oxidation to aerobic respiration or anaerobic reduction of humic substances (e.g., quinones) as TEAs via EET. Finally, we show that aceticlastic methanogens in MB are stimulated under strictly anaerobic conditions when humic TEAs are theoretically fully reduced and we hypothesize TEA reduction in acidic peat soils may be linked to changes in oxygen availability driven by water table fluctuations. We suggest future studies test this hypothesis directly through controlled manipulations of water table levels coupled to functional transcriptomic studies of Sphagnum-degrading and acetate-mineralizing MAGs.
References
Amesbury MJ, Gallego-Sala A, Loisel J. Peatlands as prolific carbon sinks. Nat Geosci. 2019;12:880–1.
Rydin H, Jeglum J. The biology of peatlands. 2nd ed. New York: Oxford University Press; 2013.
Kremer C, Pettolino F, Bacic A, Drinnan A. Distribution of cell wall components in Sphagnum hyaline cells and in liverwort and hornwort elaters. Planta. 2004;219:1023–35.
Theander O. Studies on Sphagnum peat. III. A quantitative study on the carbohydrate constituents of Sphagnum mosses and Sphagnum peat. Acta Chem Scand. 1954;8:989–1000.
Ballance S, Borsheim KY, Inngjerdingen K, Paulsen BS, Christensen BE. A re-examination and partial characterisation of polysaccharides released by mild acid hydrolysis from the chlorite-treated leaves of Sphagnum papillosum. Carbohydr Polym. 2007;67:104–15.
Painter TJ. Residues of D-lyxo-5-hexosulopyranuronic acid in Sphagnum holocellulose, and their role in cross-linking. Carbohydr Res. 1983;124:C18–C21.
Bartels D, Baumann A, Maeder M, Geske T, Heise EM, von Schwartzenberg K, et al. Evolution of plant cell wall: arabinogalactan-proteins from three moss genera show structural differences compared to seed plants. Carbohydr Polym. 2017;163:227–35.
Woodcroft BJ, Singleton CM, Boyd JA, Evans PN, Emerson JB, Zayed AAF, et al. Genome-centric view of carbon processing in thawing permafrost. Nature. 2018;560:49–54.
Ivanova AA, Wegner C-E, Kim Y, Liesack W, Dedysh SN. Identification of microbial populations driving biopolymer degradation in acidic peatlands by metatranscriptomic analysis. Mol Ecol. 2016;25:4818–35.
Duddleston KN, Kinney MA, Kiene RP, Hines ME. Anaerobic microbial biogeochemistry in a northern bog: acetate as a dominant metabolic end product. Glob Biogeochem Cycles. 2002;16:11–1-11–9.
Ye R, Jin Q, Bohannan B, Keller JK, McAllister SA, Bridgham SD. pH controls over anaerobic carbon mineralization, the efficiency of methane production, and methanogenic pathways in peatlands across an ombrotrophic-minerotrophic gradient. Soil Biol Biochem. 2012;54:36–47.
van Beelen P, Wouterse MJ, Masselink MJ, Spijker J, Mesman M. The application of a simplified method to map the aerobic acetate mineralization rates at the groundwater table of the Netherlands. J Contam Hydrol. 2011;122:86–95.
Conrad R. Importance of hydrogenotrophic, aceticlastic and methylotrophic methanogenesis for methane production in terrestrial, aquatic and other anoxic environments: a mini review. Pedosphere. 2020;30:25–39.
Walpen N, Getzinger GJ, Schroth MH, Sander M. Electron-donating phenolic and electron-accepting quinone moieties in peat dissolved organic matter: quantities and redox transformations in the context of peat biogeochemistry. Environ Sci Technol. 2018;52:5236–45.
Dettling MD, Yavitt J, Zinder S. Control of organic carbon mineralization by alternative electron acceptors in four peatlands, central New York State, USA. Wetlands. 2006;26:917–27.
Keller JK, Bridgham SD. Pathways of anaerobic carbon cycling across an ombrotrophic-minerotrophic peatland gradient. Limnol Oceanogr. 2007;52:96–107.
Keller JK, Takagi KK. Solid-phase organic matter reduction regulates anaerobic decomposition in bog soil. Ecosphere. 2013;4:1–12.
Keller JK, Weisenhorn PB, Megonigal JP. Humic acids as electron acceptors in wetland decomposition. Soil Biol Biochem. 2009;41:1518–22.
Yavitt JB, Seidman-Zager M. Methanogenic conditions in northern peat soils. Geomicrobiol J. 2006;23:119–27.
He S, Lau MP, Linz AM, Roden EE, McMahon KD. Extracellular electron transfer may be an overlooked contribution to pelagic respiration in humic-rich freshwater lakes. mSphere. 2019;4:1–8.
Lovley DR, Coates JD, Blunt-Harris EL, Phillips EJ, Woodward JC. Humic substances as electron acceptors for microbial respiration. Nature. 1996;382:445–8.
Stams AJM, De Bok FAM, Plugge CM, Van Eekert MHA, Dolfing J, Schraa G. Exocellular electron transfer in anaerobic microbial communities. Environ Microbiol. 2006;8:371–82.
Klupfel L, Piepenbrock A, Kappler A, Sander M. Humic substances as fully regenerable electron acceptors in recurrently anoxic environments. Nat Geosci. 2014;7:195–200.
Bräuer SL, Yavitt JB, Zinder SH. Methanogenesis in McLean Bog, an acidic peat bog in upstate New York: stimulation by H2/CO2 in the presence of rifampicin, or by low concentrations of acetate. Geomicrobiol J. 2004;21:433–43.
Cadillo-Quiroz H, Brauer S, Yashiro E, Sun C, Yavitt J, Zinder S. Vertical profiles of methanogenesis and methanogens in two contrasting acidic peatlands in central New York State, USA. Environ Microbiol. 2006;8:1428–40.
Kotsyurbenko O, Chin K, Glagolev M, Stubner S, Simankova M, Nozhevnikova A, et al. Acetoclastic and hydrogenotrophic methane production and methanogenic populations in an acidic West-Siberian pear bog. Environ Microbiol. 2004;6:1159–73.
Lai DYF. Methane dynamics in northern peatlands: a review. Pedosphere. 2009;19:409–21.
Xu XF, Elias DA, Graham DE, Phelps TJ, Carroll SL, Wullschleger SD, et al. A microbial functional group-based module for simulating methane production and consumption: application to an inbucated permafrost soil. J Geophys Res Biogeosci. 2015;120:1315–33.
Chen I-MA, Markowitz VM, Chu K, Palaniappan K, Szeto E, Pillay M, et al. IMG/M: integrated genome and metagenome comparative data analysis system. Nucleic Acids Res. 2017;45:D507–16.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformat. 2008;9:559.
Langfelder P, Horvath S. Eigengene networks for studying the relationships between co-expression modules. BMC Syst Biol. 2007;1:54.
Sun CL, Brauer SL, Cadillo-Quiroz H, Zinder SH, Yavitt JB. Seasonal changes in methanogenesis and methanogenic community in three peatlands, New York State. Front Microbiol. 2012;3:81.
Osvald H. Vegetation and stratigraphy of peatlands in North America. Uppsala: Acta Universitatis Upsaliensis; 1970.
Pepe-Ranney C, Campbell AN, Koechli CN, Berthrong S, Buckley DH. Unearthing the ecology of soil microorganisms using a high resolution DNA-SIP approach to explore cellulose and xylose metabolism in soil. Front Microbiol. 2016;7:703.
Nurk S, Meleshko D, Korobeynikov A, Pevzner PA. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 2017;27:824–34.
Ondov BD, Treangen TJ, Melsted P, Mallonee AB, Bergman NH, Koren S, et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 2016;17:132.
Kang D, Froula J, Egan R, Wang Z. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. PeerJ. 2015;3:e1165.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043–55.
Bowers RM, Kyrpides NC, Stepanauskas R, Harmon-Smith M, Doud D, Reddy TBK, et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat Biotechnol. 2017;35:725.
Segata N, Börnigen D, Morgan XC, Huttenhower C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat Commun. 2013;4:2304.
Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND. Nat Methods. 2014;12:59.
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil P-A, et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36:996–1004.
Rodriguez-R LM, Tsementzi D, Luo C, Konstantinidis KT. Iterative substractive binning of freshwater chronoseries metagenomes identifies over 400 novel species and their ecologic preferences. Environ Microbiol. 2020;22:3394–412.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–2.
Rodriguez-R LM, Konstantinidis KT. The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Prepr. 2016;4:e1900v1.
Nayfach S, Pollard KS. Average genome size estimation improves comparative metagenomics and sheds light on the functional ecology of the human microbiome. Genome Biol. 2015;16:51.
Zalman C, Keller JK, Tfaily M, Kolton M, Pfeifer-Meister L, Wilson RM, et al. Small differences in ombrotrophy control regional-scale variation in methane cycling among Sphagnum-dominated peatlands. Biogeochemistry. 2018;139:155–77.
Williams CJ, Yavitt JB. Botanical composition of peat and degree of peat decomposition in three temperate peatlands. Ecoscience. 2003;10:85–95.
Dedysh SN. Cultivating uncultured bacteria from northern wetlands: knowledge gained and remaining gaps. Front Microbiol. 2011;2:184.
Hattori S. Syntrophic acetate-oxidizing microbes in methanogenic environments. Microbes Environ. 2008;23:118–27.
Hines ME, Duddleson KN, Kiene RP. Carbon flow to acetate and C1 compounds in northern wetlands. Geophys Res Lett. 2001;28:4251–4.
Karakashev D, Batstone DJ, Trably E, Angelidaki I. Acetate oxidation is the dominant methanogenic pathway from acetate in the absence of Methanosaetaceae. Appl Environ Microbiol. 2006;72:5138–41.
Cervantes FJ, van der Velde S, Lettinga G, Field JA. Competition between methanogenesis and quinone respiration for ecologically important substrates in anaerobic consortia. FEMS Microbiol Ecol. 2000;34:161–71.
Cervantes FJ, Gutierrez CH, Lopez KY, Estrada-Alvarado MI, Meza-Escalante AC, Texier AC, et al. Contribution of quinone-reducing microorganisms to the anaerobic biodegradation of organic compounds under different redox conditions. Biodegradation. 2008;19:235–46.
Lokshina LY, Vavilin VA, Kettunen RH, Rintala JA, Holliger C, Nozhevnikova AN. Evaluation of kinetic coefficients using integrated monod and haldane models for low-temperature acetoclastic methanogenesis. Water Res. 2001;35:2913–22.
Kotsyurbenko OR, Friedrich MW, Simankova MV, Nozhevnikova AN, Golyshin PN, Timmis KN, et al. Shift from acetoclastic to H2-dependent methanogenesis in a West Siberain peat bog at low pH values and isolation of an acidophilic Methanobacterium strain. Appl Environ Microbiol. 2007;73:2344–8.
Schmidt O, Hink L, Horn M, Drake H. Peat: home to novel syntrophic species that feed acetate- and hydrogen-scavenging methanogens. ISME J. 2016;10:1954–66.
Wolfe AJ. The acetate switch. Microbiol Mol Biol Rev. 2005;69:12–50.
Starai VJ, Escalante-Semerena JC. Acetyl-coenzyme A synthetase (AMP forming). Cell Mol Life Sci. 2004;61:2020–30.
Pankratov TA, Dedysh SN, Zavarzin GA. The leading role of Actinobacteria in aerobic cellulose degradation in Sphagnum peat bogs. Dokl Biol Sci. 2006;410:428–30.
Mannisto M, Ganzert L, Tjirola M, Haggblom MM, Stark S. Do shifts in life strategies explain microbial community responses to increasing nitrogen in tundra soil? Soil Biol Biochem. 2016;96:216–28.
Kang H, Kwon MJ, Kim S, Lee S, Jones TG, Johncock AC, et al. Biologically driven DOC release from peatlands during recovery from acidification. Nat Commun. 2018;9:3807.
Dedysh SN, Dunfield PF. Beijerinckiaceae. In: Whitman WB, editor. Bergey’s manual of systematics of archaea and bacteria. Hoboken, NJ: John Wiley & Sons, Inc.; 2016. p. 1–4.
Acknowledgements
The authors would like to thank Tamar Barkay, Hinsby Cadillo-Quiroz, and Janet Jansson for their permission to include their unpublished metagenomes in this study. This work was supported by a microbial/metagenome project (CSP 503730) through the U.S. Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, which is supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC02–05CH11231.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
St. James, A.R., Yavitt, J.B., Zinder, S.H. et al. Linking microbial Sphagnum degradation and acetate mineralization in acidic peat bogs: from global insights to a genome-centric case study. ISME J 15, 293–303 (2021). https://doi.org/10.1038/s41396-020-00782-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00782-0
This article is cited by
-
Integrating Decomposers, Methane-Cycling Microbes and Ecosystem Carbon Fluxes Along a Peatland Successional Gradient in a Land Uplift Region
Ecosystems (2022)
-
Grazing by Tibetan Sheep Enhances Soil Bacterial and Fungal Diversity in Cold Season Pastures of Alpine Meadows on the Northern Qinghai–Tibetan Plateau
Journal of Soil Science and Plant Nutrition (2022)
-
Relationship Between Peat Type and Microbial Ecology in Sphagnum-Containing Peatlands of the Adirondack Mountains, NY, USA
Microbial Ecology (2021)