Abstract
The water-borne bacterium Legionella pneumophila is the causative agent of Legionnaires’ disease. In the environment, the opportunistic pathogen colonizes different niches, including free-living protozoa and biofilms. The physiological state(s) of sessile Legionella in biofilms and their functional consequences are not well understood. Using single-cell techniques and fluorescent growth rate probes as well as promoter reporters, we show here that sessile L. pneumophila exhibits phenotypic heterogeneity and adopts growing and nongrowing (“dormant”) states in biofilms and microcolonies. Phenotypic heterogeneity is controlled by the Legionella quorum sensing (Lqs) system, the transcription factor LvbR, and the temperature. The Lqs system and LvbR determine the ratio between growing and nongrowing sessile subpopulations, as well as the frequency of growth resumption (“resuscitation”) and microcolony formation of individual bacteria. Nongrowing L. pneumophila cells are metabolically active, express virulence genes and show tolerance toward antibiotics. Therefore, these sessile nongrowers are persisters. Taken together, the Lqs system, LvbR and the temperature control the phenotypic heterogeneity of sessile L. pneumophila, and these factors regulate the formation of a distinct subpopulation of nongrowing, antibiotic tolerant, virulent persisters. Hence, the biofilm niche of L. pneumophila has a profound impact on the ecology and virulence of this opportunistic pathogen.
Similar content being viewed by others
Introduction
The Gram-negative bacterium Legionella pneumophila ubiquitously occurs in natural as well as anthropogenic water systems and upon inhalation causes a severe pneumonia termed Legionnaires’ disease [1, 2]. In the environment, the opportunistic pathogenic bacterium infects and replicates within free-living protozoa, including amoebae and ciliates [3,4,5,6,7]. The mechanism of intracellular replication is evolutionarily conserved, requires the L. pneumophila Icm/Dot type IV secretion system (T4SS) and centers on the formation of a unique membrane-bound replication niche, the Legionella-containing vacuole (LCV) [8,9,10,11,12,13]. L. pneumophila is a facultative intracellular bacterium, which as an alternative niche also colonizes and forms biofilms in the environment [14,15,16,17], as well as under laboratory conditions [18,19,20].
Environmental bacteria face nutrient shortage and substantial physicochemical fluctuations, which they can respond to by arresting growth [21]. Growth arrest has a profound impact on bacterial physiology and traits, including stress resistance and an augmented tolerance toward antibiotics, called persistence [22, 23]. Pathogenic bacterial persisters are responsible for antibiotic treatment failures in clinical settings and cause relapsing infections [24,25,26]. However, bacterial persistence predates the clinical use of antibiotics, as L. pneumophila reversibly forms highly virulent nonreplicating persisters upon infection of primordial protozoan phagocytes, e.g., Acanthamoeba castellanii or Dictyostelium discoideum [27].
L. pneumophila adopts a biphasic lifestyle and cycles between a transmissive (growth-arrested, virulent and motile) and a replicative form [28]. The reversible switch between the transmissive and the replicative phase is controlled by the “stringent response” and the second messenger guanosine 3,5-bispyrophosphate (ppGpp) [29, 30], as well as by the Legionella quorum sensing (Lqs) system [31, 32]. Components of the Lqs system comprise the autoinducer synthase LqsA, which produces the α-hydroxyketone signaling molecule Legionella autoinducer-1 (LAI-1, 3-hydroxypentadecane-4-one) [33], the membrane-bound sensor histidine kinases LqsS [34] and LqsT [35], as well as the prototypic response regulator LqsR [36, 37], which dimerizes upon phosphorylation [38, 39]. While L. pneumophila lacking the autoinducer synthase gene lqsA (∆lqsA) is barely defective for virulence, the ∆lqsR mutant strain shows severe virulence and other phenotypes [32]. The Lqs system is linked to the cyclic-di-GMP signaling network through the transcription factor LvbR [40, 41]. LvbR is a pleiotropic transcription factor, which is negatively regulated by the sensor kinase LqsS, and directly controls the production of proteins involved in c-di-GMP metabolism as well as the architecture of L. pneumophila biofilms and pathogen–host cell interactions.
The Lqs system is a major regulator of various L. pneumophila traits, including the switch from the transmissive/virulent to the replicative phase [36], pathogen–host cell interactions [37], bacterial and host cell motility [38, 42], natural competence for DNA uptake [35], as well as extracellular filament formation and expression of a chromosomal “fitness island” [34]. Moreover, the Lqs system and in particular the autoinducer LAI-1 strongly induces the expression of the noncoding small regulatory RNA (sRNA) 6S RNA [38]. The conserved 6S RNA is a global regulator of transcription, which in L. pneumophila is abundantly produced in the postexponential growth phase and regulates the expression of icm/dot T4SS genes, as well as factors implicated in stress response or nutrient acquisition [43, 44].
While intracellular growth of L. pneumophila and LCV formation has been intensely studied, the physiology and ecological significance of extracellular, sessile bacteria are poorly understood. In this study, we investigate on a single-cell level the physiology and biphasic lifestyle of sessile L. pneumophila. We show that L. pneumophila exhibits phenotypic heterogeneity in biofilms and microcolonies, and—controlled by the Lqs system, the transcription factor LvbR and the temperature—forms subpopulations of growing and nongrowing bacteria. Nongrowing individuals are metabolically active, highly virulent, and tolerant toward antibiotics. Therefore, these sessile nongrowers are virulent persisters, which likely have a profound impact on the ecology and pathogenicity of L. pneumophila.
Results
The Lqs system regulates the ratio of nongrowers in L. pneumophila biofilms
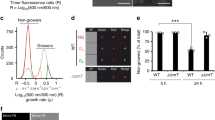
L. pneumophila forms biofilms upon undisturbed growth over several days [18, 40]. The physiology and functional features of sessile L. pneumophila in biofilms are poorly characterized. To investigate the traits of sessile L. pneumophila, we employed the Timer reporter, which allows to monitor the bacterial growth rate and to discriminate growing from nongrowing bacteria [24, 27]. Timer is a stable DsRed variant, which slowly changes its fluorescence from green (500 nm) to red (600 nm), and thus, the fluorescence ratio [500 nm (green)/600 nm (red)] accurately reflects the growth rate of L. pneumophila at a single-cell level [27]. Stationary phase Timer-producing L. pneumophila uniformly shows the characteristic red fluorescence of nongrowing bacteria (Fig. S1a, b).
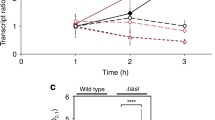
To assess growth rate heterogeneity within L. pneumophila biofilms and a possible role of the Lqs system for the process, we used Timer-producing L. pneumophila wild-type, ΔlqsA and a complemented ΔlqsA mutant strain (Fig. 1). Stationary phase bacteria were diluted in medium, and biofilm formation as well as bacterial growth rate (Timer color ratio, 500 nm/600 nm) was monitored by confocal microscopy after 24, 48 and 96 h growth at 25 °C. After 24 h, wild-type L. pneumophila/Timer formed a biofilm comprising interspersed growing (green) and nongrowing (red/orange) bacteria. The size of the initially substantial nongrowing subpopulation was considerably smaller in biofilms after 48 and 96 h. Interestingly, in biofilms formed by the ΔlqsA mutant strain, the portion of nongrowing bacteria was dramatically increased, and this phenotype was reverted upon providing the lqsA gene on a plasmid (Fig. 1a–c).
a–c L. pneumophila JR32 or ΔlqsA producing Timer (pNP107) or the complemented strain ΔlqsA(lqsA) producing Timer and LqsA (pNP120) were grown to stationary phase, diluted in AYE broth, and allowed to form biofilms at 25 °C. At the given time points, the Timer color ratio (500 nm/600 nm) was visualized by confocal microscopy. a 3D reconstruction (x = 200 μm, y = 200 μm, z = 30 μm), b orthogonal view, and c magnification are shown. d Biofilms formed after 24 h by Timer-producing L. pneumophila JR32, ΔlqsA or ΔlqsA(lqsA) were homogenized and analyzed by flow cytometry, and the subpopulation of nongrowing L. pneumophila was quantified. Data represent the mean ± SEM of three biological replicates (two-tailed Student’s t test; ***P < 0.001). Source data are provided as a Source Data file.
To quantify the percentage of nongrowing sessile L. pneumophila, the biofilms formed by Timer-producing wild-type L. pneumophila, ΔlqsA or complemented ΔlqsA, were homogenized by pipetting and analyzed by flow cytometry (Fig. 1d). This approach revealed for wild-type L. pneumophila ca. 15% nongrowers in the biofilm after 24 h of growth, which decreased to ca. 5% after 48 h of growth. Strikingly, biofilms formed by ΔlqsA mutant bacteria comprised more than 80% nongrowing bacteria after 24 h, which decreased to ca. 20% after 48 h (Fig. 1d). The phenotype was reverted to almost wild-type levels upon providing the lqsA gene on a plasmid. Taken together, the findings demonstrate that growing and nongrowing individuals co-exist in L. pneumophila biofilms at different ratios over time, and the Lqs system contributes to the regulation of the subpopulation ratio.
Nongrowing L. pneumophila in biofilms are persisters
Nongrowing bacteria can show increased tolerance toward antibiotics, a phenomenon termed persistence [23,24,25, 27, 45]. Accordingly, we assessed whether persistence occurs in L. pneumophila biofilms and how the formation of sessile L. pneumophila persisters is controlled. To this end, stationary phase Timer-producing L. pneumophila wild-type or ∆lqsA, ∆lqsS, ∆lqsT, ∆lqsS-∆lqsT, or ∆lqsR mutants were diluted in medium, and biofilm formation (Timer color ratio, 500 nm/600 nm) was monitored by confocal microscopy after 36 h at 25 °C (Figs. 2a and S2a). At this time point, L. pneumophila wild-type and the ∆lqsS-∆lqsT as well as the ∆lqsR mutant strains formed biofilms comprising mostly growing (green fluorescent = magenta) bacteria. In contrast, the ∆lqsA, ∆lqsS, and ∆lqsT mutant strains formed biofilms with a high ratio of nongrowing (orange/red fluorescent = cyan) L. pneumophila.
a 3D reconstruction of biofilms and b quantification of nongrowers by flow cytometry of Timer-producing L. pneumophila JR32, ΔlqsA, ΔlqsS, ΔlqsT, ΔlqsS ΔlqsT (ΔΔ), ΔlqsR, or ΔlvbR grown for 36 h (green = magenta, red = cyan). After 36 h, biofilms were treated with ampicillin (100 μg mL−1, 24 h), washed and regrown for 36 h. Biofilms formed for 48 h by Timer-producing L. pneumophila were treated for the time indicated with erythromycin (60 μg mL−1; orange), ampicillin (100 μg mL−1; magenta), or ofloxacin (30 μg mL−1; black) using c the strain JR32, or d ofloxacin and the strains JR32 (black), ΔlqsA (red), or complemented ΔlqsA (cyan). Bacteria were incubated at 25 °C and plated at defined time points to quantify CFU. Data represent the mean ± SEM of three biological replicates (two-tailed Student’s t test; *P < 0.05, ***P < 0.001).
Furthermore, we tested an L. pneumophila strain lacking the transcription factor LvbR, which forms a more “mat”-like biofilm compared to the more patchy biofilm architecture of the parental strain [40]. Interestingly, the ∆lvbR mutant strain formed biofilms comprising a largely increased population of nongrowing L. pneumophila (Fig. 2a), indicating that the transcription factor also regulates heterogeneity. Taken together, the formation of sessile L. pneumophila nongrowers is controlled by the Lqs system as well as by the transcription factor LvbR.
To assess the antibiotic tolerance of the L. pneumophila lqs mutant strains and correlate this feature to the growing and nongrowing subpopulations, the biofilms were treated with ampicillin (100× MIC) for 24 h, washed and let regrow in AYE medium for another 36 h (Figs. 2a and S2a). This approach indicated that ampicillin killed a large portion of wild-type L. pneumophila and ∆lqsS-∆lqsT as well as ∆lqsR mutant bacteria, and affected to a smaller extent ∆lqsA, ∆lqsS, ∆lqsT, and ∆lvbR mutant bacteria, which produced a large nongrowing subpopulation in biofilms. The nongrowing bacteria spared by ampicillin regrew upon removal of the antibiotic, and therefore, the nongrowing subpopulation is viable and culturable.
Quantification by flow cytometry validated these findings (Fig. 2b). Biofilms formed by L. pneumophila wild-type or the ∆lqsS-∆lqsT or ∆lqsR mutant strains comprised ca. 25–35% nongrowing bacteria, while biofilms formed by the ∆lqsA, ∆lqsS or ∆lqsT mutant strains comprised ca. 55–80% nongrowing bacteria. Strikingly, biofilms formed by the ∆lvbR mutant strain comprised almost only nongrowing bacteria. The treatment with ampicillin (100× MIC) for 24 h left only nongrowing L. pneumophila, as documented by a homogenous low Timer color ratio (orange/red fluorescence = cyan). Following the antibiotic treatment, the L. pneumophila strains resumed growth, as evidenced by a high Timer color ratio (green fluorescence = magenta), and the ratio of nongrowers formed by the different L. pneumophila strains correlated with the ratio observed prior to the antibiotic treatment. Biofilm formation was also assessed by CFU, which indicated that compared to wild-type L. pneumophila the ∆lqsA mutant produced less biofilm at 36 h (Fig. S2b). Therefore, the fraction of persisters is even higher for the ∆lqsA strain when normalized for the number of bacteria present before drug treatment. The final biomass of the biofilms produced by wild-type or lqs mutant L. pneumophila was comparable.
Antibiotic tolerance and persistence (as opposed to resistance) is typically characterized by biphasic killing curves [45, 46]. To test whether sessile L. pneumophila strains show antibiotic tolerance and persistence, stationary phase Timer-producing wild-type L. pneumophila was diluted in medium, allowed to form a biofilm, and exposed to antibiotics (Fig. 2c). After 2 days at 25 °C, sessile bacteria were treated with different classes of antibiotics (e.g., ampicillin, ofloxacin, or erythromycin) at a concentration of 100× MIC and plated at given time points to quantify CFU. Under these conditions, biphasic killing kinetics of L. pneumophila were observed for all antibiotics tested, consistent with the notion that a subpopulation rapidly died, while another subpopulation persisted for a longer period of time.
In analogous assays, the effect of the Lqs system on the persistence of L. pneumophila was tested. To this end, stationary phase Timer-producing wild-type L. pneumophila, ΔlqsA or complemented ΔlqsA was diluted in medium, allowed to form a biofilm and exposed to ofloxacin (Fig. 2d), which among the antibiotics tested most potently killed L. pneumophila. The deletion of lqsA increased the persistence of L. pneumophila, and the complemented strain partially restored the sensitivity to the antibiotic. The degree of persistence was proportional to the size of the nongrowing subpopulation formed by these strains during biofilm formation.
In agreement with a role for the Lqs system in stationary growth phase of L. pneumophila (high cell density and LAI-1 concentration), exponentially growing cultures of wild-type or ΔlqsA mutant bacteria produced in biofilms a smaller subpopulation of nongrowers (Fig. S3a–c) and displayed a poor persistence toward ofloxacin (Fig. S3d). In summary, the Lqs system and the transcription factor LvbR modulate the ratio between sessile populations of replicating and nonreplicating L. pneumophila, the latter of which are persisters that are tolerant toward different classes of antibiotics. Persistence of L. pneumophila is proportional to the size of the nongrowing subpopulation.
The Lqs system and LvbR control resuscitation of sessile L. pneumophila
Having established that L. pneumophila adopts phenotypic heterogeneity in biofilms and forms a subpopulation of nonreplicating persisters, we next analyzed the growth resumption (“resuscitation”) of sessile nongrowers. To this end, we developed a protocol to immobilize individual L. pneumophila in imaging chambers by agarose embedment [47]. This approach allows assessing traits of sessile L. pneumophila and their control on a single-cell level with high spatial resolution (Fig. 3).
a Stationary phase Timer-producing L. pneumophila wild-type (WT) or ΔlqsA were immobilized in AYE/0.5% agarose, and microcolony formation was recorded by time lapse microscopy over 40 h. Arrow heads indicate individuals that did not resume growth. Scale bars, 20 μm. b Fluorescence micrographs of microcolonies (scale bars, 20 μm), and c quantification of growth resumption of Timer-producing L. pneumophila wild-type (WT), ΔlqsA, ΔlqsA/plqsA, ΔlqsS, ΔlqsT, ΔlqsS-ΔlqsT, or ΔlqsR immobilized in AYE/0.5% agarose for 24 h. Data represent the mean ± SEM of three biological replicates (two-tailed Student’s t test; ***P < 0.001). d Stationary phase Timer-producing L. pneumophila wild-type (WT) or isogenic mutant strains were immobilized in AYE/0.5% agarose, and let form microcolonies for 40 h. Microcolony growth was analyzed using ImageJ (particle analysis, segmentation and area measurement), and the microcolony growth index was defined as [microcolony area (µm2)/time (h)]. Growth index differences between WT (n = 50), ∆lqsA (n = 50), ∆lqsS (n = 18), ∆lqsT (n = 32), or ∆lvbR (n = 38) were statistically not significant (n.s.).
Stationary phase Timer-producing L. pneumophila wild-type or ∆lqsA mutant strains were immobilized in agarose, and microcolony formation was recorded by time lapse confocal microscopy over 40 h (Fig. 3a, Supplementary Movies 1 and 2). After 24 h, ~60% of the red fluorescent growth-arrested wild-type bacteria had resumed growth and formed green fluorescent microcolonies. In contrast, barely any ∆lqsA mutant bacteria had resumed growth at this time point, and the mutant phenotype was complemented by plasmid-borne lqsA (Fig. 3a, b). The growth resumption frequency (i.e., the resuscitation efficiency) was also much lower for the ΔlqsS and ΔlqsT mutant strains, while the ΔlqsR and ΔlqsS-ΔlqsT mutants resumed growth at an even higher frequency than the wild-type strain (Fig. 3b). Quantification of the growth resumption frequency revealed that the percentage of ΔlqsA, ΔlqsS, or ΔlqsT mutant bacteria that resumed growth was two to three times lower than that of the wild-type strain, and the growth resumption frequency of the ΔlqsR and ΔlqsS-ΔlqsT mutant strains was 15–20% higher than that of the wild-type strain (Fig. 3c). Judged from the similar overall size of the microcolonies formed by wild-type or lqs mutant bacteria (Fig. 3a), the Lqs system controls the growth resumption frequency, but not the growth rate per se. Indeed, quantification of the microcolony growth revealed that there were no significant differences in growth between wild-type L. pneumophila and the mutant strains, which show reduced growth resumption frequency (∆lqsA, ∆lqsS, ∆lqsT, or ∆lvbR) (Fig. 3d).
If growing bacteria were used as inoculum, the frequency of microcolony formation of wild-type and ΔlqsA mutant bacteria was similar (Fig. S4a, b). This result is in agreement with the notion that the Lqs system is operative only in stationary phase (high cell density and LAI-1 concentration). Taken together, stationary phase L. pneumophila shows an inherent heterogeneity to resuscitate and resume sessile growth, which is controlled by the Lqs system.
Finally, compared to wild-type L. pneumophila, ca. 50% fewer ∆lvbR mutant bacteria resumed growth under these conditions, which is similar to the ΔlqsA, ΔlqsS, and ΔlqsT strains and contrasts the ΔlqsR strain (Fig. 3b, c). Hence, the transcription factor LvbR is a positive regulator of growth resumption, while the response regulator LqsR is a negative regulator of the process. In summary, the Lqs system and the transcription factor LvbR contribute to control growth resumption (“resuscitation”) of sessile L. pneumophila.
Sessile L. pneumophila nongrowers are antibiotic tolerant and virulent
Using the agarose-embedment microcolony assay, we next assessed antibiotic tolerance of sessile L. pneumophila subpopulations. To this end, stationary phase wild-type bacteria producing Timer were immobilized in agarose, let form microcolonies for 24 h and treated with ampicillin. The killing of L. pneumophila was monitored by confocal microscopy for up to 20 h (Fig. 4a, Supplementary Movie 3). Under these conditions, the green fluorescent growing bacteria were quantitatively killed and only red fluorescent nongrowing bacteria survived. Hence, the nongrowing sessile bacteria are persisters.
Timer-producing L. pneumophila JR32 was immobilized in AYE/0.5% agarose and allowed to form microcolonies for 24 h. a Ampicillin (100 μg mL−1) was added, and bacterial killing was monitored by confocal microscopy for up to 20 h. Arrow heads indicate an antibiotic susceptible, growing microcolony (green), and an antibiotic tolerant nongrowing individual (red). b Ethidium bromide was added (1 μg mL−1), and accumulation of the fluorescent compound was monitored by confocal microscopy after 1 h. Arrow heads indicate a nongrower that did not accumulate ethidium bromide. Data represent the mean ± SEM of three biological replicates (two-tailed Student’s t test; ***P < 0.001). c, d Timer-producing L. pneumophila JR32 (Ptac-timer-PsidC-mCerulean) was immobilized in AYE/0.5% agarose and allowed to form microcolonies. After 24 h, the bacteria were assessed by confocal microscopy for mCerulean production (expression of the virulence gene sidC) (c) in absence of antibiotics or (d) upon treatment with ampicillin (100 μg mL−1; 10 h). Arrow heads indicate a growing microcolony (green) or a nongrowing individual (red), which are negative or positive for mCerulean (sidC expression), respectively. Scale bars, 10 µm (a, b, d) and 5 µm (c).
Possibly, the nongrowers show increased persistence upon treatment with antibiotics due to decreased antibiotic uptake or increased efflux. To assess drug accumulation by sessile bacteria, agarose-embedded wild-type L. pneumophila/Timer grown for 24 h was treated with ethidium bromide, and the retention of the fluorescent compound was assessed by confocal microscopy (Fig. 4b). One hour after treatment, almost all growing bacteria accumulated and retained ethidium bromide, while only a minor portion (ca. 25%) of the nongrowers did so. This result is in agreement with the notion that decreased drug uptake or increased efflux contributes to antibiotic tolerance of nongrowing L. pneumophila.
Next, we assessed the expression of the gene encoding the Icm/Dot substrate SidC by sessile L. pneumophila as a proxy for bacterial virulence. Using agarose-embedded L. pneumophila wild-type harboring the fluorescent reporter Ptac-timer-PsidC-mCerulean [27], we detected sidC expression exclusively in nongrowing but not in growing bacteria (Fig. 4c). This result indicates that sessile stationary phase L. pneumophila individuals that do not resume growth express the virulence gene sidC, and consequently, are likely virulent. In agreement with this notion, 24 h biofilms originating from stationary phase wild-type L. pneumophila harboring the reporter Ptac-timer-PsidC-mCerulean comprised a much larger subpopulation of nongrowers expressing sidC as compared to biofilms formed by exponentially growing individuals (Fig. S5). Treatment of the L. pneumophila Ptac-timer-PsidC-mCerulean reporter strain with ampicillin selectively killed the growing bacteria, and only the nongrowing, sidC-expressing bacteria survived (Fig. 4d, Supplementary Movies 4 and 5). Taken together, these findings indicate that the antibiotic treatment enriches virulent bacteria and L. pneumophila nongrowers are virulent persisters.
Sessile L. pneumophila shows temperature-dependent phenotypic heterogeneity
In order to further assess physiological traits of sessile L. pneumophila and their control, we constructed transcriptional reporters for the sRNA 6S RNA [43], or the flaA gene encoding the major flagellum constituent, flagellin [48]. The 6S RNA and the flaA gene serve as proxies for stress response or stationary phase motility, respectively, and are positively regulated by the Lqs system and synthetic LAI-1 [38].
L. pneumophila constitutively producing mCherry and an unstable GFP variant under control of the P6SRNA promoter (Ptac-mCherry-P6SRNA-gfp) or the PflaA promoter (Ptac-mCherry-PflaA-gfp) [27] were grown in AYE medium to stationary phase, and the expression of the reporter constructs was assessed by confocal fluorescence microscopy and flow cytometry (Fig. S1a, b). Under these conditions, all bacteria expressed the PflaA reporter, while only approximately half of the bacteria expressed the P6SRNA reporter.
Using the agarose-embedment microcolony assay, we assessed the growth of sessile L. pneumophila bacteria expressing the P6SRNA reporter or not. To this end, stationary phase agarose-embedded bacteria were grown for 12 h, and single-cell bacterial growth was monitored by fluorescence microscopy (Fig. 5a, Supplementary Movie 6). During this period, ~50% bacteria started to divide, while the others remained growth-arrested. Interestingly, more than 80% of the L. pneumophila expressing the P6SRNA reporter resumed growth, while only about 25% of the bacteria not expressing the P6SRNA reporter divided (Fig. 5b). Taken together, sessile L. pneumophila immobilized on a glass surface shows phenotypic heterogeneity, where the expression of the 6S RNA reporter is positively correlated with the frequency of growth resumption.
a, b Stationary phase mCherry-producing L. pneumophila JR32 (Ptac-mCherry-P6SRNA-gfp) was immobilized in AYE/0.5% agarose, and the percentage of sessile bacteria resuming growth was visualized and quantified by time lapse microscopy. Arrow heads indicate GFP-positive and GFP-negative bacteria expressing 6S RNA reporter or not, which resume growth or not, respectively. Scale bar, 10 µm. Data represent the mean ± SEM of three biological replicates (two-tailed Student’s t test; ***P < 0.001). c Stationary phase mCherry-producing L. pneumophila JR32 harboring Ptac-mCherry-P6SRNA-gfp (upper panel) or Ptac-mCherry-PflaA-gfp (lower panel) was embedded in AYE/0.5% agarose and allowed to form microcolonies at 25 °C (72 h), at 25 °C (48 h) followed by a shift to 30 °C for 24 h, or at 30 °C (72 h). Scale bars, 10 µm. d Biofilms were formed with stationary phase mCherry-producing L. pneumophila JR32 harboring Ptac-mCherry-P6SRNA-gfp or Ptac-mCherry-PflaA-gfp at 25 °C (72 h), at 25 °C (48 h) followed by a shift to 30 °C for 24 h, or at 30 °C (72 h). Bacteria were fixed with 4% PFA, and analyzed by flow cytometry and FlowJo. Data represent the mean ± SEM of three biological replicates (two-tailed Student’s t test; *P < 0.05, **P < 0.01, ***P < 0.001). e Microcolony formation of L. pneumophila JR32 harboring Ptac-mCherry-P6SRNA-gfp or Ptac-mCherry-PflaA-gfp at 30 °C (72 h) was analyzed by confocal microcopy and 3D reconstruction (left panels; scale bars, 10 µm), and reporter (GFP/green = magenta, mCherry/red = cyan) fluorescence ratio log10 500 nm/600 nm (right panels; mean and SEM, two-tailed Student’s t test, ***P < 0.001).
To further assess the traits of sessile L. pneumophila, we let microcolony formation proceed at 25 °C or 30 °C for a prolonged time (up to 72 h) and analyzed the temporal and spatial expression of the 6S RNA reporter by confocal microscopy (Figs. 5c and S6a). Approximately 75% of L. pneumophila harboring a plasmid containing Ptac-mCherry-P6SRNA-gfp expressed the 6S RNA reporter at 30 °C, but only ca. 20% were GFP-positive at 25 °C (Figs. 5d and S6a). The expression of the 6S RNA reporter was also induced by a shift to 30 °C for 24 h after growth at 25 °C for 48 h (Figs. 5c, d and S6b). L. pneumophila bacteria expressing the 6S RNA reporter localized preferentially and statistically significantly to the boundaries of the microcolonies (Fig. 5c, e, Supplementary Movie 7).
The temperature dependency of gene expression was even more striking for the flaA reporter, since ~80% of L. pneumophila harboring a plasmid containing Ptac-mCherry-PflaA-gfp expressed the flaA reporter at 30 °C, but less than 10% were GFP-positive at 25 °C (Fig. 5c, d, Supplementary Movie 8). The spatial distribution of the PflaA reporter-expressing L. pneumophila subpopulation was similar to the one observed for 6S RNA expression (Fig. 5c, e). In summary, the expression of the 6S RNA reporter and even more so, the flaA reporter, is temperature dependent and significantly upregulated at 30 °C. Moreover, the 6S RNA or flaA reporters are expressed by spatially distinct subpopulations of sessile L. pneumophila in microcolonies.
The Lqs system and LvbR control heterogeneous gene expression in L. pneumophila microcolonies
The Lqs system regulates 6S RNA and flaA in L. pneumophila on a population level in medium [38]. We wondered whether and how quorum sensing regulates 6S RNA expression and its spatial distribution in L. pneumophila on a single-cell level in microcolonies. To this end, agarose-embedded L. pneumophila wild-type or the isogenic mutant strains ∆lqsA, ∆lqsS, ∆lqsT, ∆lqsS-∆lqsT, or ∆lqsR harboring a plasmid containing Ptac-mCherry-P6SRNA-gfp were allowed to form microcolonies for 72 h at 25 or 30 °C.
Only a few wild-type L. pneumophila expressed 6S RNA in the microcolonies at 25 °C as described above (Figs. 5c, d and S6a). At 30 °C, however, a much larger subpopulation of wild-type L. pneumophila and a similar percentage of ∆lqsR and ∆lqsS-∆lqsT mutant bacteria expressed 6S RNA, in particular at the microcolony boundaries (Figs. 6a and S6c). In contrast, microcolonies formed by the ∆lqsA, ∆lqsS or ∆lqsT mutant strains did not express 6S RNA at any temperature, indicating that the Lqs system indeed tightly controls the expression of 6S RNA in sessile L. pneumophila. Similar results were obtained upon shifting the growth temperature from 25 to 30 °C for 24 h (Fig. S6b). The quantification of 6S RNA expression in sessile L. pneumophila revealed an ~4-fold increase of 6S RNA-positive wild-type or ∆lqsR and ∆lqsS-∆lqsT mutant L. pneumophila from growth at the nonpermissive temperature (25 °C) to growth at the permissive temperature (30 °C), while there was no 6S RNA expression at all in the ∆lqsA, ∆lqsS or ∆lqsT mutants (Fig. 6b).
Stationary phase mCherry-producing L. pneumophila JR32 or ∆lqsA, ∆lqsS, ∆lqsT, ∆lqsS-∆lqsT, ∆lqsR, or ∆lvbR harboring a Ptac-mCherry-P6SRNA-gfp or c Ptac-mCherry-PflaA-gfp was embedded in AYE/0.5% agarose and allowed to form microcolonies at 30 °C (72 h). Microcolony formation was analyzed by confocal microcopy and 3D reconstruction. Scale bars, 10 µm. Quantification by flow cytometry of mCherry-producing, 6S RNA-expressing L. pneumophila JR32 or ∆lqsA, ∆lqsS, ∆lqsT, ∆lqsS-∆lqsT, ∆lqsR, or ∆lvbR harboring b Ptac-mCherry-P6SRNA-gfp grown at 25 °C (72 h), 30 °C (72 h), or 25 °C (72 h) followed by a shift to 30 °C for another 24 h, or d Ptac-mCherry-PflaA-gfp grown at 30 °C (72 h) and fixed with 4% PFA. Data represent the mean ± SEM of three biological replicates (two-tailed Student’s t test; *P < 0.05, **P < 0.01).
The transcription factor LvbR regulates the formation of persisters in biofilms (Fig. 2) and growth resumption of sessile L. pneumophila (Fig. 3). Accordingly, we also tested a role for the transcription factor in heterogeneous expression of 6S RNA in microcolonies. At the permissive temperature (30 °C), a subpopulation of ∆lvbR mutant bacteria indeed expressed 6S RNA, in particular at the microcolony boundaries (Figs. 6a and S6c). The quantification of 6S RNA expression in sessile ΔlvbR mutant L. pneumophila revealed ~20% of 6S RNA-positive bacteria at the nonpermissive temperature (25 °C), 30% upon shifting the temperature from 25 to 30 °C for 24 h, and ca. 50% 6S RNA-positive bacteria grown for 72 h at the permissive temperature (30 °C) (Fig. 6b). Compared to wild-type L. pneumophila, ~25% fewer ∆lvbR mutant expressed 6S RNA at 30 ºC, indicating that the LvbR transcription factor regulates 6S RNA expression. Taken together, these results demonstrate that the Lqs system and to a lesser extent LvbR positively regulate heterogeneous 6S RNA expression in sessile L. pneumophila.
We also assessed the role of quorum sensing for flaA expression in L. pneumophila microcolonies. L. pneumophila wild-type and the above mutant strains harboring a plasmid containing Ptac-mCherry-PflaA-gfp were allowed to form microcolonies at 30 °C for 72 h (Figs. 6c and S6c). Microcolonies formed by L. pneumophila wild-type, lqs mutant or lvbR mutant strains produced a subpopulation of bacteria that expressed flaA. The expression pattern was similar to the expression of 6S RNA, with GFP-positive bacteria mainly at the boundaries of the microcolonies. The quantification of flaA expression in sessile L. pneumophila revealed ~65% flaA-positive wild-type L. pneumophila and ca. 25 or 40% fewer ∆lqsA or ∆lvbR mutant bacteria, respectively (Fig. 6d). The expression of flaA by the other lqs mutant strains was also slightly, but statistically not significantly, reduced. Taken together, the Lqs system (in particular lqsA) and LvbR regulate the fraction and spatial distribution of sessile PflaA-expressing L. pneumophila in microcolonies.
In stationary phase L. pneumophila 6S RNA is upregulated [43], and the 6S RNA reporter is positively regulated by the Lqs system (Fig. 6a, b). To test whether 6S RNA is not only a manifestation of phenotypic heterogeneity, but also a regulator of the process, we assessed biofilm formation of a strain lacking 6S RNA. Timer-producing L. pneumophila Δ6S RNA mutant bacteria or the parental strain ΔcomR [43] were grown to stationary phase, diluted in fresh medium and let form a biofilm for 24 h. The quantification by flow cytometry revealed a ratio of ca. 10% nongrowing L. pneumophila in biofilms formed either by the Δ6S RNA mutant or the parental strain (Fig. S7a). In addition, both strains showed similar growth resumption frequency after immobilization in agarose (Fig. S7b). Hence, the 6S RNA does neither regulate the formation of nongrowers in L. pneumophila biofilms nor the growth resumption heterogeneity.
In summary, the Lqs system and the transcription factor LvbR determine the fraction and spatial distribution of the L. pneumophila subpopulation, which in microcolonies expresses 6S RNA and flaA. While the Lqs-dependent upregulation of the 6S RNA is a feature of phenotypic heterogeneity, the 6S RNA itself does not control the formation of biofilms or phenotypic heterogeneity of sessile L. pneumophila.
Discussion
In this study, we assessed traits of sessile L. pneumophila and their control on a single-cell level in biofilms and microcolonies. We revealed that sessile L. pneumophila exhibits phenotypic heterogeneity and forms growing and nongrowing subpopulations. Nongrowing sessile L. pneumophila are metabolically active and virulent persisters. L. pneumophila nongrowers might contribute as a “dormant” form to the long-term survival of the bacteria in the environment, and therefore, this feature of the opportunistic pathogen is likely of broad ecological significance.
In most ecosystems, nongrowing bacterial cells dominate. In clinical settings these bacteria are a major cause of chronic or relapsing infections due to their intrinsic capacity to be resuscitated in susceptible hosts and due to their notorious antibiotic tolerance [49,50,51]. Prominent examples of environmentally transmitted opportunistic pathogens, which adopt a nongrowing (“dormant” or “viable but non culturable”) state, include Vibrio cholerae, Mycobacterium tuberculosis, and Pseudomonas aeruginosa, the causative agents of cholera, tuberculosis, and pneumonia, respectively [52,53,54].
A sessile lifestyle on abiotic or biotic (biofilm) surfaces is arguably the most predominant form of bacterial life in the environment. Here, we provide evidence that upon sessile growth in biofilms or on abiotic surfaces L. pneumophila forms a population of nongrowing, virulent persisters (Fig. 4). L. pneumophila is a ubiquitous environmental bacterium and upon inhalation might become an “accidental” pathogen. Hence, the virulent, antibiotic tolerant form of L. pneumophila is likely important in clinical settings. In fact, these nongrowing bacteria might be the cause for severe, relapsing and antibiotic-nonresponsive forms of Legionnaires’ disease. Accordingly, the biofilm niche comprising different L. pneumophila subpopulations has a profound impact on the pathogenesis of the opportunistic pathogen.
In the environment, the subpopulation of sessile nongrowing virulent persisters may efficiently infect protozoan predators, naturally present in microbial communities. Functionally distinct subpopulations of sessile L. pneumophila may therefore adopt a bet-hedging strategy [55], which prepares for either extracellular growth (surface colonization) or intracellular growth. Thus, overall the sessile L. pneumophila population has the potential to colonize a range of vastly different environmental niches.
Nongrowing forms of environmental bacteria not only ensure long-term survival, but also need to retain the capacity to resume growth and be resuscitated. Resuscitation can happen at the onset of favorable conditions or by stochastic re-initiation of growth. Indeed nongrowing individual bacteria have been shown to stochastically re-initiate proliferation at a relatively low rate, regardless of the environmental conditions [56, 57]. Here, we show that the growth resumption frequency of sessile L. pneumophila under favorable conditions (rich medium) is genetically controlled. The Lqs system positively regulates growth resumption, and stationary phase ΔlqsA, ΔlqsS or ΔlqsT mutant strains resumed growth on a abiotic surface with a two to three times lower frequency compared to the parental strain (Fig. 3). Similarly, L. pneumophila lacking the transcription factor LvbR showed an impaired growth resumption frequency, and thus, LvbR also positively regulates the process. The mechanism underlying growth resumption frequency is unclear. The regulators might indeed control a switch from the nongrowing to the growing state. Alternatively, the regulators might control cell survival during stationary phase stress, allowing efficient growth resumption upon the encounter of more favorable conditions.
Quorum sensing is a major regulator of phenotypic heterogeneity of sessile L. pneumophila. The Lqs system controls the ratio between nongrowing and growing L. pneumophila in biofilms (Figs. 1 and 2), growth resumption (resuscitation) of sessile bacteria forming microcolonies (Fig. 3), antibiotic tolerance of L. pneumophila in biofilms (Figs. 2 and S2), and microcolonies (Fig. 4), as well as heterogeneous gene expression (Figs. 5 and 6). Phenotypic heterogeneity was primarily observed with stationary phase bacteria, and exponentially growing L. pneumophila cultures produced a much smaller subpopulation of nongrowers and antibiotic tolerant bacteria in biofilms (Fig. S3). These observations are in agreement with a role for the Lqs system in stationary growth phase (high cell density and LAI-1 concentration). Given that L. pneumophila predominantly exists in a nongrowing form in the environment, the Lqs system, which is operative in stationary phase, is likely also relevant for the control of phenotypic variation under natural conditions. Interestingly, the Lqs system negatively regulates the occurrence of nongrowing L. pneumophila in biofilms (Figs. 1 and 2a) and positively regulates the occurrence of nongrowers in phagocytes [27]. Hence, quorum sensing oppositely regulates the ratio between nongrowing and growing L. pneumophila in sessile or intracellular niches.
In addition to the Lqs system, the transcription factor LvbR controls phenotypic heterogeneity and the emergence of nongrowing, virulent L. pneumophila persisters (Figs. 2 and 3). LvbR is a pleiotropic regulator, which determines various L. pneumophila traits, such as biofilm architecture, natural competence for DNA uptake, and pathogen–host cell interactions [40, 41]. The transcription factor directly binds to the promoter of hnox1/lpg1056, which is divergently transcribed from the lvbR gene. Hnox1 is a nitric oxide sensor, which inhibits the diguanylate cyclase activity of Lpg1057 and reduces c-di-GMP production [58]. Thus, by negatively regulating the inhibitor Hnox1, LvbR positively regulates the diguanylate cyclase activity of Lpg1057 and, consequently, c-di-GMP levels are increased. It remains to be determined, whether LvbR controls phenotypic heterogeneity of L. pneumophila through the c-di-GMP network, and/or whether other regulatory circuits are involved.
Phenotypic heterogeneity of sessile L. pneumophila is not only controlled by endogenous circuits (Lqs, LvbR), but also by external cues such as the temperature. Compared to 25 °C the transcriptional reporters for 6S RNA or flaA expression were upregulated at 30 ºC ca. 4- or 8-fold, respectively, and a shift from 25 to 30 °C for 24 h already led to an induction of the reporters (Figs. 5, 6, and S6). Temperature is an important environmental cue sensed by many bacteria to adapt their behavior to changing environmental conditions [59]. We reveal in this study that L. pneumophila senses and responds to small changes in the environmental temperature. To our knowledge, this is also the first description of a thermo-regulated sRNA in L. pneumophila.
At the permissive temperature of 30 °C, the Lqs system and LvbR positively regulate 6S RNA expression. The Lqs system apparently overruled the regulation by temperature, since the ∆lqsA, ∆lqsS, or ∆lqsT mutant strains failed to express 6S RNA at any of the temperatures tested. Moreover, neither the ∆lqsS-∆lqsT double sensor kinase mutant nor the ∆lqsR response regulator mutant seemed to be involved in the regulation of 6S RNA expression (Fig. 6). These observations might be explained by reciprocal (and thus neutralizing) regulatory roles of the LqsS and LqsT sensor histidine kinases [35], and the presence of (a) regulatory element(s) other than the response regulator LqsR implicated in 6S RNA expression downstream of the sensor kinases.
The expression of 6S RNA as well as of flaA occurred with pronounced spatial heterogeneity, and the genes were preferentially expressed at the microcolony boundaries (Figs. 5 and 6). The functional significance of this heterogeneous expression pattern is unclear. The 6S RNA and flaA genes serve as proxies for stationary phase stress response or motility, respectively. Perhaps, peripheral bacteria that are preferentially exposed to stressors and predators might selectively upregulate stress response and virulence pathways. To facilitate the spread of sessile L. pneumophila, peripheral bacteria might preferentially become motile, since they are less firmly embedded in a biofilm (or microcolony). Alternatively, the spatial heterogeneity of P6SRNA and PflaA reporter induction might be the result of stochastic gene expression, which broadens the bacterial response repertoire in the context of a bet-hedging strategy [55].
The spatial heterogeneity of 6S RNA and flaA reporter expression is controlled by the Lqs system (Fig. 6). The Lqs system and the cognate signaling molecule LAI-1 are required (but might not be sufficient) to regulate gene expression. Accordingly, the 6S RNA and flaA promoter activities and corresponding expression reporters might function as (indirect) LAI-1 biosensors. It is tempting to speculate that the bacterial cells in a L. pneumophila microcolony or biofilm respond in a heterogeneous manner to LAI-1, and/or that the production of LAI-1 proceeds with spatial heterogeneity. However, other factor(s) likely (co-)determine the pattern of spatial heterogeneity of gene expression in sessile L. pneumophila.
In the current study, we identified endogenous genetic (lqs, lvbR) and exogenous physical (temperature) determinants of phenotypic heterogeneity of sessile L. pneumophila. The consequences of phenotypic heterogeneity are distinct subpopulations of growing and nongrowing bacteria, the latter of which are virulent and antibiotic tolerant persisters. This work paves the way for future studies addressing mechanistic aspects of cues and consequences of the phenotypic heterogeneity of sessile L. pneumophila in biofilms and on abiotic surfaces.
Methods
For details see Supplementary Information.
Bacterial strains and plasmids
Bacterial strains and plasmids used in this study are listed in Table S1. The Timer reporter is a stable DsRed variant, which slowly changes its fluorescence from green (500 nm) to red (600 nm) [60]. Accordingly, the Timer fluorescence ratio [500 nm (green)/600 nm (red)] reflects the growth rate of L. pneumophila at a single-cell level [27]. The dual fluorescence reporters (Ptac-mCherry-PflaA-gfp, Ptac-mCherry-P6SRNA-gfp) allow an assessment of L. pneumophila promoter activity.
Biofilm and microcolony analysis by confocal microscopy and flow cytometry
For biofilm formation, planktonic exponential or stationary phase grown L. pneumophila strains were diluted in fresh AYE/chloramphenicol at an OD600 of 0.1, placed in a multiwell plate or an ibidi imaging chamber, and incubated at 25 or 30 °C for the indicated time while avoiding mechanical disturbance [18, 40]. The formation and analysis of microcolonies is detailed in [47].
Biofilms and microcolonies were analyzed by confocal laser scanning microscopy (Leica TCS SP8 X CLSM). Flow cytometry was performed with a FACS-Fortessa II using homogenized and fixed biofilm samples. The gating for the Timer reporter was performed as described [27].
Biphasic kill curves
L. pneumophila biofilms grown for 48 h were treated or not with different classes of antibiotics at >100× MIC (ampicillin 100 μg mL−1, erythromycin 60 μg mL−1, or ofloxacin 30 μg mL−1) at 25 °C, washed three times, diluted, and plated on CYE agar plates to quantify CFU.
Data availability
All data is available in the main text or the Supplementary Material and provided as source data files.
References
Newton HJ, Ang DK, van Driel IR, Hartland EL. Molecular pathogenesis of infections caused by Legionella pneumophila. Clin Microbiol Rev. 2010;23:274–98.
Hilbi H, Hoffmann C, Harrison CF. Legionella spp. outdoors: colonization, communication and persistence. Environ Microbiol Rep. 2011;3:286–96.
Fields BS. The molecular ecology of Legionella. Trends Microbiol. 1996;4:286–90.
Greub G, Raoult D. Microorganisms resistant to free-living amoebae. Clin Microbiol Rev. 2004;17:413–33.
Hoffmann C, Harrison CF, Hilbi H. The natural alternative: protozoa as cellular models for Legionella infection. Cell Microbiol. 2014;16:15–26.
Boamah DK, Zhou G, Ensminger AW, O’Connor TJ. From many hosts, one accidental pathogen: The diverse protozoan hosts of Legionella. Front Cell Infect Microbiol. 2017;7:477.
Swart AL, Harrison CF, Eichinger L, Steinert M, Hilbi H. Acanthamoeba and Dictyostelium as cellular models for Legionella infection. Front Cell Infect Microbiol. 2018;8:61.
Sherwood RK, Roy CR. A Rab-centric perspective of bacterial pathogen-occupied vacuoles. Cell Host Microbe. 2013;14:256–68.
Asrat S, de Jesus DA, Hempstead AD, Ramabhadran V, Isberg RR. Bacterial pathogen manipulation of host membrane trafficking. Annu Rev Cell Dev Biol. 2014;30:79–109.
Finsel I, Hilbi H. Formation of a pathogen vacuole according to Legionella pneumophila: how to kill one bird with many stones. Cell Microbiol. 2015;17:935–50.
Qiu J, Luo ZQ. Legionella and Coxiella effectors: strength in diversity and activity. Nat Rev Microbiol. 2017;15:591–605.
Personnic N, Bärlocher K, Finsel I, Hilbi H. Subversion of retrograde trafficking by translocated pathogen effectors. Trends Microbiol. 2016;24:450–62.
Steiner B, Weber S, Hilbi H. Formation of the Legionella-containing vacuole: phosphoinositide conversion, GTPase modulation and ER dynamics. Int J Med Microbiol. 2018;308:49–57.
Declerck P. Biofilms: the environmental playground of Legionella pneumophila. Environ Microbiol. 2010;12:557–66.
Abdel-Nour M, Duncan C, Low DE, Guyard C. Biofilms: the stronghold of Legionella pneumophila. Int J Mol Sci. 2013;14:21660–75.
Pécastaings S, Allombert J, Lajoie B, Doublet P, Roques C, Vianney A. New insights into Legionella pneumophila biofilm regulation by c-di-GMP signaling. Biofouling. 2016;32:935–48.
Hochstrasser R, Hilbi H. Intra-species and inter-kingdom signaling of Legionella pneumophila. Front Microbiol. 2017;8:79.
Mampel J, Spirig T, Weber SS, Haagensen JAJ, Molin S, Hilbi H. Planktonic replication is essential for biofilm formation by Legionella pneumophila in a complex medium under static and dynamic flow conditions. Appl Environ Microbiol. 2006;72:2885–95.
Hindré T, Brüggemann H, Buchrieser C, Héchard Y. Transcriptional profiling of Legionella pneumophila biofilm cells and the influence of iron on biofilm formation. Microbiology. 2008;154:30–41.
Pécastaings S, Berge M, Dubourg KM, Roques C. Sessile Legionella pneumophila is able to grow on surfaces and generate structured monospecies biofilms. Biofouling. 2010;26:809–19.
Wai SN, Mizunoe Y, Yoshida S. How Vibrio cholerae survive during starvation. FEMS Microbiol Lett. 1999;180:123–31.
Balaban NQ, Merrin J, Chait R, Kowalik L, Leibler S. Bacterial persistence as a phenotypic switch. Science. 2004;305:1622–5.
Harms A, Maisonneuve E, Gerdes K. Mechanisms of bacterial persistence during stress and antibiotic exposure. Science. 2016;354:aaf4268.
Claudi B, Sprote P, Chirkova A, Personnic N, Zankl J, Schurmann N, et al. Phenotypic variation of Salmonella in host tissues delays eradication by antimicrobial chemotherapy. Cell. 2014;158:722–33.
Hélaine S, Cheverton AM, Watson KG, Faure LM, Matthews SA, Holden DW. Internalization of Salmonella by macrophages induces formation of nonreplicating persisters. Science. 2014;343:204–8.
Conlon BP, Rowe SE, Gandt AB, Nuxoll AS, Donegan NP, Zalis EA, et al. Persister formation in Staphylococcus aureus is associated with ATP depletion. Nat Microbiol. 2016;1:16051.
Personnic N, Striednig B, Lezan E, Manske C, Welin A, Schmidt A, et al. Quorum sensing modulates the formation of virulent Legionella persisters within infected cells. Nat Commun. 2019;10:5216.
Molofsky AB, Swanson MS. Differentiate to thrive: lessons from the Legionella pneumophila life cycle. Mol Microbiol. 2004;53:29–40.
Hammer BK, Swanson MS. Co-ordination of Legionella pneumophila virulence with entry into stationary phase by ppGpp. Mol Microbiol. 1999;33:721–31.
Dalebroux ZD, Yagi BF, Sahr T, Buchrieser C, Swanson MS. Distinct roles of ppGpp and DksA in Legionella pneumophila differentiation. Mol Microbiol. 2010;76:200–19.
Tiaden A, Spirig T, Hilbi H. Bacterial gene regulation by α-hydroxyketone signaling. Trends Microbiol. 2010;18:288–97.
Personnic N, Striednig B, Hilbi H. Legionella quorum sensing and its role in pathogen-host interactions. Curr Opin Microbiol. 2018;41:29–35.
Spirig T, Tiaden A, Kiefer P, Buchrieser C, Vorholt JA, Hilbi H. The Legionella autoinducer synthase LqsA produces an α-hydroxyketone signaling molecule. J Biol Chem. 2008;283:18113–23.
Tiaden A, Spirig T, Sahr T, Wälti MA, Boucke K, Buchrieser C, et al. The autoinducer synthase LqsA and putative sensor kinase LqsS regulate phagocyte interactions, extracellular filaments and a genomic island of Legionella pneumophila. Environ Microbiol. 2010;12:1243–59.
Kessler A, Schell U, Sahr T, Tiaden A, Harrison C, Buchrieser C, et al. The Legionella pneumophila orphan sensor kinase LqsT regulates competence and pathogen-host interactions as a component of the LAI-1 circuit. Environ Microbiol. 2013;15:646–62.
Tiaden A, Spirig T, Weber SS, Brüggemann H, Bosshard R, Buchrieser C, et al. The Legionella pneumophila response regulator LqsR promotes host cell interactions as an element of the virulence regulatory network controlled by RpoS and LetA. Cell Microbiol. 2007;9:2903–20.
Tiaden A, Spirig T, Carranza P, Brüggemann H, Riedel K, Eberl L, et al. Synergistic contribution of the Legionella pneumophila lqs genes to pathogen-host interactions. J Bacteriol. 2008;190:7532–47.
Schell U, Simon S, Sahr T, Hager D, Albers MF, Kessler A, et al. The α-hydroxyketone LAI-1 regulates motility, Lqs-dependent phosphorylation signalling and gene expression of Legionella pneumophila. Mol Microbiol. 2016;99:778–93.
Hochstrasser R, Hutter CAJ, Arnold FM, Bärlocher K, Seeger MA, Hilbi H. The structure of the Legionella response regulator LqsR reveals amino acids critical for phosphorylation and dimerization. Mol Microbiol. 2020;113:1070–84.
Hochstrasser R, Kessler A, Sahr T, Simon S, Schell U, Gomez-Valero L, et al. The pleiotropic Legionella transcription factor LvbR links the Lqs and c-di-GMP regulatory networks to control biofilm architecture and virulence. Environ Microbiol. 2019;21:1035–53.
Hochstrasser R, Hilbi H. Legionella quorum sensing meets cyclic-di-GMP signaling. Curr Opin Microbiol. 2020;55:9–16.
Simon S, Schell U, Heuer N, Hager D, Albers MF, Matthias J, et al. Inter-kingdom signaling by the Legionella quorum sensing molecule LAI-1 modulates cell migration through an IQGAP1-Cdc42-ARHGEF9-dependent pathway. PLoS Pathog. 2015;11:e1005307.
Faucher SP, Friedlander G, Livny J, Margalit H, Shuman HA. Legionella pneumophila 6S RNA optimizes intracellular multiplication. Proc Natl Acad Sci USA. 2010;107:7533–8.
Faucher SP, Shuman HA. Small regulatory RNA and Legionella pneumophila. Front Microbiol. 2011;2:98.
Balaban NQ, Hélaine S, Lewis K, Ackermann M, Aldridge B, Andersson DI, et al. Definitions and guidelines for research on antibiotic persistence. Nat Rev Microbiol. 2019;17:441–8.
Brauner A, Fridman O, Gefen O, Balaban NQ. Distinguishing between resistance, tolerance and persistence to antibiotic treatment. Nat Rev Microbiol. 2016;14:320–30.
Personnic N, Striednig B, Hilbi H. Single cell analysis of Legionella and Legionella-infected Acanthamoeba by agarose embedment. Methods Mol Biol. 2019;1921:191–204.
Byrne B, Swanson MS. Expression of Legionella pneumophila virulence traits in response to growth conditions. Infect Immun. 1998;66:3029–34.
Lewis K. Persister cells, dormancy and infectious disease. Nat Rev Microbiol. 2007;5:48–56.
Lewis K. Multidrug tolerance of biofilms and persister cells. Curr Top Microbiol Immunol. 2008;322:107–31.
Hall CW, Mah TF. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol Rev. 2017;41:276–301.
Brenzinger S, van der Aart LT, van Wezel GP, Lacroix JM, Glatter T, Briegel A. Structural and proteomic changes in viable but non-culturable Vibrio cholerae. Front Microbiol. 2019;10:793.
Defraine V, Fauvart M, Michiels J. Fighting bacterial persistence: current and emerging anti-persister strategies and therapeutics. Drug Resist Updates. 2018;38:12–26.
Wu B, Liang W, Kan B. Growth phase, oxygen, temperature, and starvation affect the development of viable but non-culturable state of Vibrio cholerae. Front Microbiol. 2016;7:404.
Ackermann M. A functional perspective on phenotypic heterogeneity in microorganisms. Nat Rev Microbiol. 2015;13:497–508.
Epstein SS. Microbial awakenings. Nature 2009;457:1083.
Sturm A, Dworkin J. Phenotypic diversity as a mechanism to exit cellular dormancy. Curr Biol. 2015;25:2272–7.
Carlson HK, Vance RE, Marletta MA. H-NOX regulation of c-di-GMP metabolism and biofilm formation in Legionella pneumophila. Mol Microbiol. 2010;77:930–42.
Loh E, Righetti F, Eichner H, Twittenhoff C, Narberhaus F. RNA thermometers in bacterial pathogens. Microbiol Spectr. 2018;6.
Terskikh A, Fradkov A, Ermakova G, Zaraisky A, Tan P, Kajava AV, et al. “Fluorescent timer”: protein that changes color with time. Science. 2000;290:1585–8.
Acknowledgements
We thank Selina Niggli for initial cloning and Ramon Hochstrasser for comments and discussions. We would also like to thank Sébastien Faucher (McGill University, Canada) for providing the Δ6S RNA mutant and parental strains. Research of NP in the laboratory of HH was supported by the Swiss National Science Foundation (SNF) Ambizione program (PZ00P3_161492 & PZ00P3_185529) awarded to NP. Research in the laboratory of HH was supported by the SNF (31003A_153200, 31003A_175557), the Novartis Foundation for Medical-Biological Research, the OPO Foundation, and the German Research Foundation (DFG; SPP 1617). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
NP conceived the study with input from HH. NP designed the experiments. NP and BS performed the experiments. NP and HH wrote the paper with input from BS.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Personnic, N., Striednig, B. & Hilbi, H. Quorum sensing controls persistence, resuscitation, and virulence of Legionella subpopulations in biofilms. ISME J 15, 196–210 (2021). https://doi.org/10.1038/s41396-020-00774-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00774-0
This article is cited by
-
Phagotrophic protists preserve antibiotic-resistant opportunistic human pathogens in the vegetable phyllosphere
ISME Communications (2023)
-
A clash of quorum sensing vs quorum sensing inhibitors: an overview and risk of resistance
Archives of Microbiology (2023)
-
Regulatory and innovative mechanisms of bacterial quorum sensing–mediated pathogenicity: a review
Environmental Monitoring and Assessment (2023)
-
Legionellosis risk—an overview of Legionella spp. habitats in Europe
Environmental Science and Pollution Research (2022)
-
Microbial warfare in the wild—the impact of protists on the evolution and virulence of bacterial pathogens
International Microbiology (2021)