Abstract
Bioaerosol transport in the atmosphere disperses microbial species between continents, affects human and plant health, and may influence hydrologic cycling. However, there have been few quantitative observations of bioaerosols at altitudes more than a few kilometers above the surface. Lack of data on bioaerosol distributions in the atmosphere has impeded efforts to assess the aerial dissemination of microbes and their vertical extent in the biosphere. In this study, a helium balloon payload system was used to sample microbial cells and dust particles in air masses as high as 38 km above sea level over three locations in the southwestern United States. The cell concentrations at altitudes between 3 and 29 km were highly similar (2–5 × 105 cells m−3) and approximately threefold lower than those observed in the convective boundary layer (CBL; 1 × 106 cells m−3), decreasing to 8 × 104 cells m−3 at 35–38 km. The detection of adenosine triphosphate (ATP) and recovery of bacteria possessing extreme tolerance to desiccation and shortwave ultraviolet radiation confirmed that certain microorganisms have the capacity to persist at lower altitudes of the stratosphere. Our data and related calculations provide constraints on the upper altitudinal boundary for microbial habitability in the biosphere.
Similar content being viewed by others
Introduction
Microorganisms comprise a portion of the bioaerosols that are distributed and transported in Earth’s atmosphere [1,2,3]. However, logistical challenges have limited opportunities to examine the microorganisms disseminated in high-altitude air masses. The vast majority of bioaerosol data available have been obtained from near-surface air in the troposphere. Above the troposphere exists an atmospheric layer with a temperature inversion called the stratosphere (Fig. 1a). While there have been reports of viable microorganisms recovered from stratospheric altitudes [4,5,6,7], the dearth of quantitative bioaerosol data at high altitudes (Fig. S1) has hampered meaningful assessments of microbial abundances in the atmosphere. Given that microorganisms are capable of dispersing large distances as aerosols [1, 2], remain metabolically active under certain atmospheric conditions [8, 9], and may affect meteorological processes [10], there is a critical need for new observations that improve understanding of their distribution in the atmosphere.
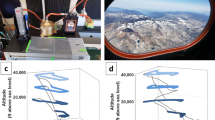
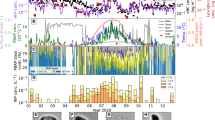
Temperature (a), microbiological (b), and particle (c) data with altitude. a Temperature profiles from Amarillo, TX were used to identify the CBL, free troposphere, tropopause, and stratosphere, the approximate boundaries of which are demarcated by the gray horizontal lines. b Cell (black circles) and cell carbon (gray squares) concentration estimates for the flights listed in Table 1. The values for “viable” cells (black triangles) were derived from the ATP data (Table S4). Open symbols are samples that did not exceed the procedural limits of detection. c Particle concentration (black circles) and mass (gray squares) for each flight in Table 1. Error values are obscured by symbols and provided in Table 1
Global dissemination of microorganisms via atmospheric transport is limited to those species tolerant of water loss and exposure to ultraviolet (UV) radiation, stressors that intensify with altitude. Survival of water deficit is a trait possessed by various insect, plant, yeast, and protozoa species [11], but in bacteria, most of what is known is based on work from a few unrepresentative strains (i.e., the cyanobacterial genera Nostoc and Chroococcidiopsis; e.g., ref. [12]). Oxygen and ozone in the stratosphere absorb incident shortwave UV radiation (UVC, 100–280 nm), strongly attenuating the flux of these mutagenic photons to the surface [13]. As such, bioaerosols transported into the stratosphere represent the only natural laboratories on Earth for examining the biological effects of UVC radiation [14].
We developed a high-altitude bioaerosol sampling approach for helium balloon platforms [15] to test the hypothesis that the concentration and viability of microorganisms incrementally decreases with altitude in the atmosphere due to low water availability, temperature, and pressure, and high UV radiation flux. In this study, we collected aerosol samples in air masses as high as 38 km, returned them to the surface, and conducted a comparative analysis of the data obtained with procedural controls that were included on each balloon mission [15]. In addition to providing the first quantitative profile of bioaerosol concentrations for the troposphere and stratosphere, a variety of bacteria were cultured from the aerosol samples and investigated for their tolerance of desiccation and UVC radiation. Here, we use these data to examine bacterial survival in the stratosphere and constrain the upper boundary of the biosphere.
Materials and methods
Balloon flights and sampling
Data from nine sounding balloon flights and one long-duration balloon flight (Table 1; Table S1) were collected at sites in proximity to LSU (Baton Rouge) and a NASA Columbia Science Balloon Facility (Palestine, Texas and Fort Sumner, New Mexico).
A 2-kg latex helium sounding balloon was used for short-duration (<3 h) flights at altitudes from 1.3 to 29 km [15] and the following missions: two in Louisiana [June 24, 2010 near Church Point, LA (30.40° N, 92.22° W, elevation 0.01 km above sea level, asl), November 21, 2010 near Denham Springs, LA (30.48° N, 90.95° W, elevation 0.01 km asl)]; one in Texas [May 22, 2013 near Palestine, TX (31.76° N, 95.64° W, elevation 0.15 km asl)]; and six in New Mexico [between August 17 and September 4, 2013 in the vicinity of Ft. Sumner, NM (34.47° N, 104.24° W, elevation 1.2 km asl)]. Rates of ascension for the sounding balloon flights ranged between 6.5 and 7.9 m s−1 (average of 7.1 m s−1).
Aerosol sampling was conducted using an autonomous payload system that was decontaminated using a combination of sodium hypochlorite, UVC irradiation, and ethylene oxide, as described by Bryan et al. [15]. To assess background microbial and particle contamination, each flight included a procedural control chamber that was identical to and decontaminated with the sample chambers; however, the chamber doors of the control remained sealed for the duration of the flight [15]. Particles and cells in the air masses were sampled by impaction on silicon-coated Rotorods® (IMS Health, Inc.; 40 rods per chamber) that were exposed during ascent by opening retractable doors with linear actuators (Supplementary Video 1). Sampling was terminated by commanding the linear actuators to extend, closing the doors and sealing the chamber. The volume of air sampled per rod (Table 1) was calculated by multiplying the altitude interval by the rod area (35 mm2). At altitudes of ~30 km, the payload string was separated from the sounding balloon, facilitating a controlled descent to the surface via a parachute. Upon payload recovery, the chambers were inspected to verify their seal during impact, and the doors were manually locked to prevent breach during transport. The chambers were opened in a laminar flow cabinet with high-efficiency particulate air filters, and all samples were processed within 6 h of recovery.
To sample at float altitude on the High Altitude Student Platform [16] (HASP), the payload was configured to rotate on a fixed axis, allowing the chambers to pass through the air at an average velocity of 2.1 m s−1 (ref. [15]; Supplementary Video 2). The modified payload was integrated on the HASP and launched September 2, 2013 (14:57 UTC) from Ft. Sumner, NM. The HASP support vehicle was tethered to a 3.11 × 105 m3 polyethylene, zero pressure helium-filled balloon and provided power, uplink commanding, and downlink telemetry for payload science operations. Upon reaching float altitude (~36 km), the chambers were warmed and commanded to open. All operations were confirmed in the downlinked telemetry as well as in the real-time video on-board HASP (Supplementary Video 2). Sampling was conducted for 8.4 h, the payload was commanded to stop rotation, and the chambers were closed. Based on a rotation rate of 92 rpm, the cross-sectional area and number of rods, and the distance from the rotation axis to the center of the chamber, the total volume of air sampled at float altitudes of 35–38 km was estimated at 2.2 m3 rod−1 (total volume of 177 m3). Flight termination ensued when HASP was ~1000 km west of the launch site. Payload touchdown occurred at 33.96° N, 112.98° W on September 3, 2013 (14:10 UTC). HASP was accessible ~12 h after touchdown, the payload was chilled during transport, and the samples were processed within 48 h.
Physical properties of the air masses
Temperature and relative humidity (RH) data collected by National Oceanic and Atmospheric Administration balloon sounding flights (http://weather.uwyo.edu/upperair/sounding.html) were examined from Amarillo, TX (35.22° N, 101.83° W) and Albuquerque, NM (35.09° N, 106.61° W). Temperature profiles and air mass origins for both locations (at 12:00 UTC) showed similar characteristics during each flight; therefore, only data from Amarillo are shown (Fig. 1a; Fig. S2). Temperature and RH in the free troposphere over Lake Charles, LA (30.23° N, 93.22°W) for the flight dates in 2010 (Table 1) were similar to those observed at Amarillo, TX during 2013 (Fig. 1a). No temperature or RH data are available for the stratospheric altitudes sampled in the vicinity of Palestine, TX on May 22, 2013. Based on the Amarillo temperature profiles during flights in August and September 2013, the CBL (from the surface to ∼3 km), free troposphere (3–15 km), tropopause (15–18), and stratosphere (altitudes above 18 km) were identified (Fig. 1). The CBL is well mixed, and as shown in Fig. S2, its upper boundary with the free troposphere is indicated by linearly decreasing temperature with increasing altitude [17,18,19]. For flights targeting the lowest sampling altitudes (17 and 18 August 2013), the top of the CBL was at 2.1 and 2.5 km, respectively (Fig. S2). The troposphere and stratosphere are separated by the tropopause, which was at a height of 15–18 km during the 2013 flight campaign. The stratosphere is the layer of air immediately above the tropopause where air temperature increases with altitude (Fig. 1a).
Air mass trajectory and classification
The trajectories of air masses arriving at the launch site were forecast prior to each flight with the HYbrid Single Particle Lagrangian Integrated Trajectory (HYSPLIT) model [20]. The model output was used to predict the boundary altitudes for the CBL, free troposphere, and stratosphere based on temperature. The predicted air mass trajectories were verified post flight by generating 72-h backward trajectories with archived data using HYSPLIT. The trajectory history was used to categorize air masses as continental tropical (cT) or maritime tropical (mT) according to the method of Rauber et al. [21].
Characterization of cells and particles
Particles and SYBR® Gold-stained cells on the sampling rods were microscopically enumerated as previously described [15]. Particles were sized using ImageJ software and cell dimensions were estimated using a calibrated ocular scale and epifluorescence microscopy; diameter was based on the major axis length. Data for particles 1.0–1.9 µm in diameter were categorized into ten bins and particles 2.0–6.4 µm were categorized into five bins (e.g., Table S2). Size distributions were used to estimate particle mass (Table 1) by assuming a particle density of 1.0 g cm−3 (ref. [22]). The cell data were separated into 1 µm bins of 1.0 and 2.0 µm in diameter (Table S1). Cellular carbon was calculated based on representative volumes for the 1.0 (0.52 µm3) and 2.0 µm (4.2 µm3) cell size bins, a biomass density of 1.1 g cm−3 (ref. [23]), and assuming carbon represented 50% of the cell dry weight [24].
The mean number of particles or cells per field of view in the controls was subtracted from the mean number of particles or cells per field of view in each sample. Error was propagated by combining the standard errors of the control and sample following the method of Baker and Nissim [25]. Data from the samples and procedural controls were scaled to the sampling rod area, corrected for efficiency and the associated errors (Fig. S3; Supplemental Discussion), and divided by the air volume sampled (Table 1). The procedural level of detection for each altitude interval sampled in Table 1 is based on cell abundance in the controls (average of 460 cells rod−1, n = 21) and the volume of air sampled (range of 0.059–1.0 × 106 cells m3). Cell abundance data that were not significantly different from the procedural controls (Fig. 1b, open symbols, α > 0.05) are reported as limit of detection values [26].
For global bioaerosol estimates (Table S3), a rudimentary atmospheric structure was presumed to derive the volume of four concentric atmospheric layers: the CBL (0–3 km, volume of 1.5 × 109 km3), the free troposphere and tropopause (3–20 km, 8.7 × 109 km3), and two altitude ranges in the stratosphere [20–30 km (5.1 × 109 km3), and 30–40 km, (5.2 × 109 km3)]. The minimum and maximum concentration values available at altitudes <30 km (Table 1) were used for the calculations. The single data point at 36 km (7.7 × 104 cells m−3) was used for the estimate at altitudes of 30–40 km. Similarly, the biomass and cell carbon data (Table 1) were used to estimate their magnitude in the atmosphere (Table S3).
Aerosol collection efficiency
The aerosol collection efficiency was measured by rotating two sampling chambers at different velocities on the roof of a building at Louisiana State University (30.45° N, 91.14° W, elevation ~0.02 km; Supplemental Discussion). For each trial, the velocity of the payload sampling chambers was varied by adjusting the length of the rotational arm affixed to the sampling chambers. During the first efficiency measurement, the chamber velocity ranged from 2.8 to 3.1 m s−1, with an average velocity of 2.9 m s−1. Based on the radial distance from center, each sampling rod traveled from 3.8 to 4.2 m per revolution, sampling between 0.35 and 0.39 m3 of air after 1 h. In the second trial, the average chamber velocity was 4.7 m s−1 (ranging from 4.6 to 4.9 m s−1), the distance traveled by the rods was 5.9–6.3 m per revolution, and the total volume of air sampled per rod during 1 h ranged from 0.58 to 0.62 m3 (Supplemental Discussion).
Simultaneous with rotational aerosol collection, a six-stage (cutpoint aerodynamic diameters of 10, 2.5, 1.4, 0.77, 0.44, and 0.25 µm) Model 131 MOUDITM (MSP Corporation) cascade impactor was used to sample (100 L min−1; total of 12 m3 air sampled in 2 h) at the same height as the sampling chambers and within a horizontal distance of ~1 m. Prior to sampling, the foil substrates used as impaction surfaces were coated with silicone spray (CRC Industries) and allowed to dry for 15 min in a laminar flow hood. Particles impacted on the collection substrate (3.5 × 109 µm2 of sampling surface) from three stages of the MOUDI sampler were obtained: stage 1 (with a cutpoint diameter of 8.5 µm), stage 2 (cutpoint diameter 2.5 µm), and stage 3 (cutpoint diameter 1.4 µm). Based on the cell size distribution observed in the air masses sampled (Table S1), the data from stage 3 were selected for determination of the size and abundance of particles 0.95–6.4 µm (Table S2; Supplemental Discussion).
For each velocity test, the particles in 60 fields of view (total area of 7.7 × 105 µm2 per rod) for each sample rod (n = 3) were counted and measured. Particles on three replicate samples of the substrate from stage 3 were enumerated and sized by examining 30 fields of view per replicate (area of 9.6 × 104 µm2 per replicate). Identical collection substrates were prepared and processed in parallel to serve as procedural controls. The particles in each field of view were manually traced, counted, and sized [15], and the average number of particles per field of view for each size bin determined (Table S2; Supplemental Discussion). The resulting modeled efficiency values (Fig. S3, Table S2; Supplemental Discussion) were used to determine the total cell and particle concentrations at altitude (Table 1).
Quantification of cellular ATP
Free ATP was enzymatically eliminated and the concentration of cellular ATP was determined as described by Bryan et al. [15]. Procedural control rods from seven flights (an average of 12 rods per flight) were analyzed to determine the level of background and analytical detection limit for cellular ATP. The level of background associated with the procedural controls averaged 7.5 ± 0.85 × 102 amol ATP rod−1 (n = 96). The concentrations reported (Table S4) were adjusted for background using the same procedure described for the cell data. A 250:1 weight ratio of C:ATP [27] was used to estimate cell carbon from the ATP data. To make comparisons per volume of air sampled, cell carbon values derived from the ATP data were partitioned according to observed cell size (Table S1), converted to biovolume [24], and the number of 1 and 2 µm cells per rod was estimated (Table S4). The cell concentration inferred from the ATP data was corrected for efficiency (Supplemental Discussion) and divided by the sampling volume (Fig. 1b; Table S4).
Enrichment culturing and isolation of bacteria
Within 6 hr of payload recovery, the sealed sampling chambers were opened under class 100 conditions and individual rods were aseptically removed and placed into culture tubes containing 5 mL of sterile media previously shown successful for the recovery of heterotrophic microorganisms in the atmosphere [4, 5]. For the samples collected in November 2010 and May 2013, ten sample and ten control rods were placed into R2A broth (Difco cat. no.: 218262) amended with 100 mg mL−1 cycloheximide. To expand opportunities for resuscitating bacteria and encouraging their growth, a rich media (tryptic soy) and recipes of 1% concentration were also used during the 2013 New Mexico campaign. R2A broth, 1% (v/v) R2A broth, tryptic soy broth (Difco cat. no.: 236950), and 1% (v/v) tryptic soy broth [5] amended with 100 mg mL−1 cycloheximide were individually inoculated with five sample and five control rods.
The inoculated media was stored at 4 °C in the dark until returned to Louisiana State University (<3 weeks), whereupon the cultures were incubated aerobically at 22 °C. Microbial growth in the liquid cultures was monitored by visual inspection and biweekly spread plating 100 μL of each onto agar-solidified media of identical composition. Individual colonies that appeared on the agar media following aerobic incubation at 22 °C were isolated in pure culture using a standard three-phase streak technique.
Amplification and sequencing of 16S rRNA gene sequences from bacterial isolates
Liquid cultures of the bacterial isolates were harvested by centrifugation (17,000 × g; 10 min); lysed chemically (10% SDS w/v) and enzymatically (lysozyme, 10 mg mL−1); and genomic DNA was extracted and purified using phenol–chloroform followed by ethanol precipitation [28]. The DNA pellet was suspended in 0.22 µm-filtered and autoclaved deionized water, and quantified by absorbance at 260 nm using a Nanodrop ND 1000 spectrophotometer (Thermo Scientific, Inc.).
A portion of the 16S ribosomal RNA gene (~1.4 kb) was amplified in a 50 µL polymerase chain reaction (PCR) containing 1.5 U of Taq DNA polymerase (5PRIME, cat. no.: 2200000), 1× TaqMaster PCR enhancer, 1× MasterTaq buffer, 200 µm deoxynucleotide triphosphates, 20 µm of each primer (27F [29] and 1492R [30]), and genomic DNA template.
The purified PCR products were sequenced using BigDye® terminator cycle sequencing (Applied BiosystemsTM), an ABI 3130XL Genetic Analyzer (Applied BiosystemsTM), and the following primers: 27F, 515F [30], 907R [29], and 1492R. Individual sequences were aligned using BioEdit software [31] to create consensus sequences for positions 227–1257 (Escherichia coli 16S rRNA gene numbering). The nearest neighbors to the queried 16S rRNA gene sequences were identified with the SINA aligner using the SEED database [32]. Distance analyses were performed with MEGA7 (ref. [33]).
Desiccation survival
Cultures of Deinococcus radiodurans R1 (ATCC 13939), Escherichia coli MG1655, and the isolates in Table S5 were grown aerobically with shaking (175 rpm) at 30 °C in R2A broth. Optical density at 620 nm was used to measure culture turbidity, cells were harvested from stationary phase, and washed twice and resuspended in 10 mM MgSO4. Aliquots of the suspensions (10 µL) were deposited onto sterile glass coverslips and placed in a desiccation chamber containing Drierite® (≥98% CaSO4, W.A. Hammond Drierite Company, Ltd, cat. no.: 7778-18-9); the liquid evaporated in ~4 h. RH in the chamber was controlled at 25% and monitored using a dial hygrometer (Fisher ScientificTM, cat. no.: 11-661-6B) with an accuracy of ±2.5%.
Three replicates of each sample were tested during each desiccation survival experiment. The concentration of colony forming units (CFUs) recovered immediately after drying of the coverslip (~4 h) was designated as the t = 0 time point (N0). Samples were removed from the desiccation chamber after 1, 2, 4, 6, 8, 10, 12, and 14 days, serially diluted in 10 mM MgSO4, spread plated on R2A agar (DifcoTM, cat. no.: 218263), and incubated at 30 °C in the dark. After 3–7 days, the number of CFUs on a dilution plate series was used to calculate the surviving fraction, expressed as the ratio of N/N0. The average (± the standard deviation) of parameters describing the desiccation survival curves (the slope, and R2 values) were determined using the exponential equation, N = N0e−kt, where k is the inactivation rate (day−1) and t is the time of exposure in days (Table S6).
UVC radiation survival
Triplicate samples of desiccated bacteria, prepared as described above, were exposed to 254 nm UVC radiation using a model 3UV-38 lamp (Part no.: 95-0341-01, UVP LLC) at a dose rate of 10.6–11.4 W m−2 (Table S7). The populations were initially subjected to UVC exposures of 0.15–0.65 kJ m−2 (Table S7). For those isolates demonstrating higher levels of resistance, UVC radiation doses of 0.65–2.8 kJ m−2 were evaluated (Table S7). Serial dilutions of irradiated samples were prepared as described above, and cultures were incubated at 30 °C in the dark. The surviving fraction (N/N0) and survival curves were generated using the equation N = N0e−kF, where k is the inactivation rate (m2 kJ−1) and F is UV radiation fluence (kJ m−2; ref. [34]).
Statistical analysis
Statistical analysis was performed using SAS® University Edition [35]. Following tests for normality, the PROC GLIMMIX procedure for a negative binomial distribution, along with the LSMEANS statement and the DUNNETTU option, was used for one-sided significance testing to determine if the cell, particle, and ATP data were at concentrations above those observed in the procedural controls. The limit of detection for measured parameters was determined using the PROC GLIMMIX procedure with 95% confidence intervals. Linear regressions and ANOVA (for comparing tropospheric with stratospheric data) were performed using StatPlus:mac statistical analysis software [36].
The survival curves of individual isolates were compared by statistical analysis of k values using a one-way ANOVA followed by post hoc Tukey HSD analysis [36]. Pairwise comparisons with values of p < 0.05 were considered to be statistically significant.
Results
Helium balloon payloads developed for high-altitude aerosol sampling [15] were used to investigate the size, concentration, and viability of microorganisms in atmospheric parcels between 1.3 and 38 km asl (Fig. 1b, Table 1; hereafter referenced to the median sampling altitude). Twelve air masses were sampled during ten flights in Louisiana, Texas, and New Mexico between 2010 and 2013 (Table 1, Supplementary Discussion). Temperature and RH data from each flight were highly similar to the data obtained from Amarillo, TX (Fig. 1a). The CBL (Fig. S2), free troposphere, and stratosphere were identified based on temperature profiles (Fig. 1a). Air temperature in the CBL was ~25 oC (Fig. S2) with RH values of 58–76%, which contrasted to higher altitudes in the troposphere and stratosphere that had low RH (20–30%) and a minimum temperature of −74 oC (Fig. 1a).
According to 72-h back trajectories, the sampled CBL air masses (2.2 and 2.3 km) had traversed over the Chihuahuan Desert in the south and the High Plains region in the west before entering New Mexico, and therefore, were categorized as continental tropical (cT; n = 2) air masses [21]. In contrast, all other air masses sampled were classified as maritime tropical (mT; n = 10) [21]. Aerosols sampled in the free troposphere over New Mexico (8.1 km; n = 2) on consecutive days in 2013 and Louisiana during November 2010 (6.1 km; n = 1) were sourced from mT air masses that originated over the Pacific Ocean. Sampling in the free troposphere over Louisiana in June 2010 (6.1 km; n = 1) also occurred within an mT air mass, but it originated over the Gulf of Mexico. Despite differences in launch date and location, all of the stratospheric balloon missions sampled similar mT air masses that originated over the Atlantic Ocean and traveled west towards Texas and New Mexico (n = 6).
DNA-containing cells (Fig. 2a–f) and particles impacted onto the polystyrene collection rods were characterized using epifluorescence and scanning electron microscopy, respectively [15]. Comparison of the data with procedural blanks included on each flight indicated most samples contained cells with long-axis diameters ranging from 0.5 to 2.0 µm (Fig. 2a–f) that were at abundances statistically higher than background. Statistically significant numbers of cells >2.0 µm were not observed in any of the samples, so size bins of 1.0 and 2.0 µm were used to parse the cell data (Table S1). Abundances of particles >6.0 µm were statistically indistinguishable from those in the controls; therefore, the data for particles were limited to those 1.0–6.0 µm in diameter (Table 1, Table S2).
The efficiency at which the rods collected particles 1.0–6.0 µm was empirically determined (Fig. S3; Supplemental Discussion). The modeled efficiency values for each sampling velocity were used to correct for the number of particles counted (Table S2), and the coefficients derived were used to estimate the concentration of each size bin per volume of air mass sampled (Table 1, Fig. S3; Supplementary Discussion). Particle concentration and mass decreased exponentially with altitude (Fig. 1c), and values from the free troposphere were 16- and 10-fold lower, respectively, than those observed in the CBL. The lowest particle abundances were observed in samples from the stratosphere, which ranged in concentration from 1.2 to 2.5 × 106 particles m−3 (Table 1) and had highly similar size and mass distributions (Fig. S4).
In general, the concentration of microbial cells and associated cellular carbon decreased with increasing altitude from the CBL into the stratosphere (Fig. 1b; Table 1 and S1). Microbial abundances in the CBL (1.2 and 1.4 × 106 cells m−3) were approximately threefold higher than those in free tropospheric air masses (Fig. 1b). Although slightly higher cell concentrations were observed in the lower portion of the stratosphere (19–29 km, 3.8–5.3 × 105 cells m−3) when compared with values from the free troposphere (2.2–4.0 × 105 cells m−3), these differences were not statistically significant (ANOVA: n = 7, p = 0.11). Rods from the 8.1 and 21 km flights that sampled 0.18 and 0.16 m3 of air, respectively, had average cell numbers higher than the controls but are not statistically different from background and reported at the detection limit (Fig. 1b, open symbols). Sampling of larger air volumes reliably provided cell data that were statistically higher than those in the controls (Table 1). The highest altitude sampled (36 km) had the lowest cell and cell carbon concentrations (7.7 × 104 cells m−3 and 0.086 µg m−3, respectively). It is important to note that the epifluorescent microscopic method of cell enumeration requires that the dye diffuse through the cell membrane and bind to DNA. Since this process would not be expected to occur with endospores, their abundances are unlikely to be accounted for by this method and the data in Table 1 and Fig. 1.
Statistically significant levels of ATP, a proxy for viable microbial biomass, were measured in samples collected from 2.2, 24, and 36 km (Table S4). Consistent with the cell data (Fig. 1b), the highest ATP concentrations were detected at 2.2 km and lowest at 36 km. The number of “viable” cells in the CBL inferred from the ATP data (3.8 ± 0.10 × 105 cells m−3) suggests ~30% of the DNA-containing cells were potentially viable (Fig. 1b). In the stratosphere, the data from 24 and 36 km (1.1 ± 0.062 × 105 and 7.6 ± 0.25 × 103 cells m−3, respectively) implies that cell viability decreased (~20% and 10% of the cells, respectively) with altitude (Fig. 1b; Table S4). Given that the cell extraction method in this analysis would not release ATP from endospores, our data should be viewed as a conservative estimate of microbial viability in these samples.
Fifteen isolates were cultured from aerosol samples collected at 6, 21, and 26 km and identified as members of the bacterial phyla Actinobacteria, Firmicutes, and Proteobacteria (Table S5). Two of the isolates from Louisiana and New Mexico air masses (ML2-1 and L9-1, respectively; Table S5) were most closely related to the endospore-forming species Paenibacillus barengoltzii. Based on the low tolerance of vegetative cells from these isolates to water deficit, it is likely these taxa survive as endospores during atmospheric transport. Approximately three-fourths of the non-sporulating isolates are desiccation tolerant, with survival rates significantly greater than E. coli. Seven isolates exhibited moderate to high levels of desiccation resistance, with LD90 values ranging from 6 to 25 days at 25% RH (Table S6; Supplemental Discussion). Tolerance to UVC radiation was also documented among six of the desiccation tolerant isolates from the stratosphere (Table S7, Supplemental Discussion). Three isolates displayed moderate UVC radiation resistance by surviving doses that reduced E. coli populations by >99.9%, but they were significantly less tolerant than Deinococcus radiodurans R1 (Table S7). The betaproteobacterial isolate L7-7A and actinobacterial isolates L6-1 and L9-4 demonstrated extreme levels of UVC radiation tolerance, with survival rates equivalent to D. radiodurans R1 (Fig. 3b; Table S7, p > 0.05).
Extreme tolerance to desiccation and UVC radiation in bacteria isolated from the stratosphere. Survival of populations during desiccation (a; Table S6) and UVC radiation exposure at 254 nm (b; Table S7). The isolates are labeled as follows: L6-1 (Curtobacterium sp., black circles), L7-7A (Noviherbaspirillum sp., white circles), L9-4 (Modestobacter sp., gray squares), and D. radiodurans (gray triangles). c Conservative estimated timeframe to inactivate populations of L6-1 and L7-7A based on the combined effect of desiccation and the estimated flux of UVC radiation (data from panels a and b) with altitude in the stratosphere [13]. Some error bars in panel c are obscured by the symbols and provided in Table S8. *Assuming 12 h of sun day−1
Discussion
Bioaerosols are ubiquitous in the near-surface atmosphere (e.g., Jaenicke et al., ref. [3]), but there are very few quantitative observations outside of the CBL and no data have been available for altitudes above 10 km [37] (Fig. S1). To provide fundamental information on the extent of microbial life in the atmosphere, we investigated vertical profiles of bioaerosols in the southwestern US from altitudes in the CBL to the stratosphere (Fig. 1b; Table 1). The range of bioaerosol concentration data available have been determined using widely differing techniques, but nevertheless, microbial abundances in CBL air masses (1.2 and 1.4 × 106 cells m−3) were comparable with values from a mountain observatory in the western United States (Fig. S1), and those in the free troposphere were approximately twofold higher than those reported at 9–10 km above the Caribbean Sea [37]. While cell abundance decreased with altitude in the stratosphere (Fig. 1b), cell concentrations from 21 to 26 km were highly similar and statistically indistinguishable from sampled altitudes in the free troposphere. The particle concentration estimates for altitudes up to 20 km are within the several order of magnitude range of published values (Fig. S5). Above 20 km, the particle concentrations observed are higher than estimates for liquid aerosols [38, 39] and particles smaller than 1.0 µm [40].
A portion of the bioaerosols collected and characterized were viable bacteria that demonstrated extreme tolerance to desiccation and short-wavelength UV radiation (Fig. 3). Many of these bacteria are phylogenetically related to strains previously isolated from the atmosphere [8, 41, 42] or other arid soil environments [43], including the Atacama [44] and Antarctica [45]. Although resistance to desiccation and UV radiation has not been reported within the betaproteobacterial genus Noviherbaspirillum, it is a common trait of the Actinobacteria [46] and species of Curtobacterium [47] and Modestobacter [48]. Several of the isolates (L6-1, L7-7A, and L9-4) were as UVC radiation tolerant as D. radiodurans (Fig. 3b) and the most UVC radiation resistant endospore-forming species documented [49]. Interestingly, L6-1 belongs to the most frequently isolated genus from high-altitude samples and has 100% 16S rRNA gene identity to the agriculturally relevant plant pathogen Curtobacterium flaccumfaciens pv. flaccumfaciens LMG 3645 (ref. [50]; Table S5).
Based on a population size of 3.8 × 105 cells m−3 (minimum observed between 21 and 26 km; Table 1) and the range of cell inactivation rates for the most desiccation tolerant bacterial isolates (Fig. 3a), best case scenarios for survival at RH values relevant to the stratosphere (25% RH) range from timeframes of 35 to 140 days (Table S8). UVC radiation flux estimates in the stratosphere (Supplemental Discussion) imply a negligible germicidal effect at altitudes below 30 km compared with desiccation (Table S8). However, the modeled UVC radiation flux increases by three orders of magnitude between 30 and 40 km (Table S8) due to decreasing atmospheric ozone concentration at these altitudes [13]. Assuming that attachment to particles abundant at the stratospheric altitudes sampled (<6 μm; Table 1) would provide ineffective shielding from UV radiation [51], the combined effects of desiccation and UVC radiation at 36 and 40 km should inactivate a population of L6-1 (i.e., a highly desiccation and UVC radiation-resistant isolate) in 39 and 7.8 days, respectively (Fig. 3c, Table S8).
Given the capacity of the isolates to survive prolonged desiccation and high UVC radiation dosage (Fig. 3c), a fundamental question is whether or not microbes in the stratosphere can return to the surface in a viable state. Brewer–Dobson circulation (BDC) is a major pathway for injecting aerosols through the tropical tropopause and into the stratosphere [52, 53], and the mean age of air at the sampled latitudes is ~3 years at 20 km and ~7.5 years at 36 km [52]. Gravitational settling is the primary route for aerosol removal from the stratosphere, with sedimentation rates at 25 km for 1 and 2 µm particles estimated at 2 and 1 years, respectively [54]. Considering this, populations of even the most desiccation and UVC radiation resistant species described would not survive the timeframes necessary for stratospheric transport via BDC (Fig. 3c). However, tropospheric aerosols can be transported to altitudes of 20–30 km over timeframes shorter than necessary for BDC through quasi-horizontal eddy mixing [52, 53]. The most rapid route for aerosol removal from the stratosphere is through downward transport during stratospheric intrusions, which are common at high-elevation sites in the western United States and northern hemispheric midlatitudes during the westerly phase of the quasi-biennial oscillation [55, 56].
We used a novel approach to sample high-altitude bioaerosols [15] and generated the first quantitative profile of microbial abundances to altitudes of 38 km. A key observation from this study is that large quantities of microbial aerosols were inferred to be transported to and present in the tropospheric and stratospheric air masses sampled. Extrapolation of the cell and biomass concentration data in Table 1 implies the atmosphere (i.e., from the surface to 40 km) may contain ~1024 cells (Table S3). Clearly this value should be viewed as tentative and revised when more data are available, but this estimate is at least 100-fold higher than prior inventories [57, 58] and suggests the stratosphere could harbor ~40% of all bioaerosols (Table S3). However, there are acute impediments to microbial survival with altitude (Fig. 3c), the most significant of which are the ozone concentrations that attenuate the flux of UVC radiation [13] and allow lower altitudes of the stratosphere to exist within the boundary conditions for microbial life.
References
Smith DJ, Jaffe DA, Birmele MN, Griffin DW, Schuerger AC, Hee J, et al. Free tropospheric transport of microorganisms from Asia to North America. Microb Ecol. 2012;64:973–85.
Griffin DW, Westphal DL, Gray MA. Airborne microorganisms in the African desert dust corridor over the mid-Atlantic ridge, Ocean Drilling Program, Leg 209. Aerobiologia. 2006;22:211–26.
Jaenicke R, Matthias-Maser S, Gruber S. Omnipresence of biological material in the atmosphere. Environ Chem. 2007;4:217–20.
Griffin DW. Terrestrial microorganisms at an altitude of 20,000 m in Earth's atmosphere. Aerobiologia. 2004;20:135–40.
Smith DJ, Griffin DW, Schuerger AC. Stratospheric microbiology at 20 km over the Pacific Ocean. Aerobiologia. 2010;26:35–46.
Yang Y, Itahashi S, Yokobori SI, Yamagishi A. UV-resistant bacteria isolated from upper troposphere and lower stratosphere. Biol Sci Space. 2008;22:18–25.
Yang Y, Yokobori S, Kawaguchi J, Yamagami T, Iijima I, Izutsu N, et al. Investigation of cultivable microorganisms in the stratosphere collected by using a balloon in 2005. JAXA Research and Development Report, JAXA-RR-08-001; Ibaraki, 2009. p. 35–42. https://global.jaxa.jp
Vaïtilingom M, Attard E, Gaiani N, Sancelme M, Deguillaume L, Flossmann AI, et al. Long-term features of cloud microbiology at the puy de Dôme (France). Atmos Environ. 2012;56:88–100.
Sattler B, Puxbaum H, Psenner R. Bacterial growth in supercooled cloud droplets. Geophys Res Lett. 2001;28:239–42.
Phillips VT, Andronache C, Christner BC, Morris CE, Sands DC, Bansemer A, et al. Potential impacts from biological aerosols on ensembles of continental clouds simulated numerically. Biogeosciences. 2009;6:987–1014.
Potts M. Desiccation tolerance of prokaryotes. Microbiol Rev. 1994;58:755–805.
Billi D, Wright DJ, Helm RF, Prickett T, Potts M, Crowe JH. Engineering desiccation tolerance in Escherichia coli. Appl Environ Microbiol. 2000;66:1680–4.
Gueymard C. A simple radiative model of the atmospheric radiative transfer of sunshine: algorithms and performance assessment. Florida Solar Energy Center; Clear Lake Road, Cocoa, Florida, 1995, FSEC-PF-270-95.
Brash DE. UV mutagenic photoproducts in Escherichia coli and human cells: a molecular genetics perspective on human skin cancer. Photochem Photobiol. 1988;48:59–66.
Bryan NC, Stewart M, Granger D, Guzik TG, Christner BC. A method for sampling microbial aerosols using high altitude balloons. J Microbiol Methods. 2014;107:161–8.
Guzik TG. The high altitude student platform (HASP) as a model multi-payload balloon platform. In: Aerospace Conference, IEEE 2015 Mar 7. p. 1–10.
Stull R. The Atmospheric Boundary Layer. In: Wallace JM, Hobbs PV, editors. Atmospheric science: an introductory survey, 2nd ed. Boston, USA: Elsevier Academic Press; 2006. p.375–417.
Cushman-Roisin B. Atmospheric boundary layer. In: Environmental fluid dynamics. New York, USA: John Wiley & Sons, Inc.; 2014. p. 165–79.
Ouwersloot HG, Vilà-Guerau de Arellano J, Nölscher AC, Krol MC, Ganzeveld LN, Breitenberger C, et al. Characterization of a boreal convective boundary layer and its impact on atmospheric chemistry during HUMPPA-COPEC-2010. Atmos Chem Phys. 2012;12:9335–53.
Draxler RR, Rolph GD. HYSPLIT (HYbrid Single Particle Lagrangian Integrated Trajectory). NOAA Air Resources Laboratory READY website. College Park, MD: NOAA Air Resources Laboratory; 2003. http://www.arl.noaa.gov/HYSPLIT.php.
Rauber RM, Walsh JE, Chalevoix DJ. Airmasses and fronts. In: Severe and hazardous weather: an introduction to high impact meteorology, 3rd ed. Dubuque, USA: Kendall/Hunt Publishing Company; 2008. p.166–167.
Huffman JA, Sinha B, Garland RM, Snee-Pollmann A, Gunthe SS, Artaxo P, et al. Size distributions and temporal variations of biological aerosol particles in the Amazon rainforest characterized by microscopy and real-time UV-APS fluorescence techniques during AMAZE-08. Atmos Chem Phys. 2012;12:11997–2019.
Bratbak GU, Dundas IA. Bacterial dry matter content and biomass estimations. Appl Environ Microbiol. 1984;48:755–7.
West AW, Sparling GP, Grant WD. Correlation between four methods to estimate total microbial biomass in stored, air-dried, and glucose-amended soils. Soil Biol Biochem. 1986;18:569–76.
Baker RW, Nissim JA. Expressions for combining standard errors of two groups and for sequential standard error. Nature. 1963;198:1020.
Eaton AD, Clesceri LS, Greenberg AE, Franson MAH, editors Standard methods for the examination of water and wastewater. 20th ed. Washington, USA: American Public Health Association; 1998.
Karl DM. Cellular nucleotide measurements and applications in microbial ecology. Microbiol Rev. 1980;44:739.
Sambrook J, Russell DW, editors. Molecular cloning: a laboratory manual. Cold Spring Harbor Cold Spring Harbor, USA: Laboratory Press; 2001.
Lane DJ, Stackebrandt E, Goodfellow M. 16S/23S rRNA sequencing. Nucleic acid techniques in bacterial systematics. New York, USA: Wiley; 1991. p. 115–76.
Turner S, Pryer KM, Miao VP, Palmer JD. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis 1. J Eukaryot Microbiol. 1999;46:327–38.
BioEdit v. 7.2.5, Ibis Biosciences, Carlsbad, CA, copyright 1997–2013. http://www.mbio.ncsu.edu/bioedit/bioedit.html.
Pruesse E, Peplies J, Glöckner FO. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics. 2012;28:1823–9.
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4.
Lin CY, Li CS. Control effectiveness of ultraviolet germicidal irradiation on bioaerosols. Aerosol Sci Technol. 2002;36:474–8.
SAS® University Edition, SAS Insititude Inc., Cary, NC, copyright 2014. http://www.sas.com/en_us/software/university-edition.html.
AnalystSoft, StatPlus, AnalystSoft Inc.—statistical analysis program for Mac OS®. Version v6. http://www.analystsoft.com/en/.
DeLeon-Rodriguez N, Lathem TL, Rodriguez-R LM, Barazesh JM, Anderson BE, Beyersdorf AJ. et al. Microbiome of the upper troposphere: species composition and prevalence, effects of tropical storms, and atmospheric implications. Proc Natl Acad Sci USA. 2013;110:2575–80.
Renard J, Brogniez C, Berthet G, Bourgeois Q, Gaubicher B, Chartier M, et al. Vertical distribution of the different types of aerosols in the stratosphere: detection of solid particles and analysis of their spatial variability. J Geophys Res. 2008;113. https://doi.org/10.1029/2008JD010150
Renard J, Berthet G, Salazar V, Catoire V, Tagger M, Gaubicher B, et al. In situ detection of aerosol layers in the middle stratosphere. Geophys Res Lett. 2010;37. https://agupubs.onlinelibrary.wiley.com/doi/full/10.1029/2010GL044307
Xu L, Okada K, Iwasaka Y, Hara K, Okuhara Y, Tsutsumi Y, et al. The composition of individual aerosol particle in the troposphere and stratosphere over Xianghe (39.45°N, 117°E), China. Atmos Environ. 2001;35:3145–53.
Wei K, Zheng Y, Li J, Shen F, Zou Z, Fan H, et al. Microbial aerosol characteristics in highly polluted and near-pristine environments featuring different climatic conditions. Sci Bull. 2015;60:1439–47.
Weon HY, Kim BY, Hong SB, Jeon YA, Koo BS, Kwon SW, et al. Massilia niabensis sp. Nov. and Massilia niatensis sp. nov., isolated from air samples. Int J Syst Evol Microbiol. 2009;59:1656–60.
Reddy GS, Potrafka RM, Garcia-Pichel F. Modestobacter versicolor sp. nov., an actinobacterium from biological soil crusts that produces melanins under oligotrophy, with emended descriptions of the genus Modestobacter and Modestobacter multiseptatus Mevs et al. 2000. Int J Syst Evol Microbiol. 2007;57:2014–20.
Busarakam K, Bull AT, Trujillo ME, Riesco R, Sangal V, van Wezel GP, et al. Modestobacter caceresii sp. nov., novel actinobacteria with an insight into their adaptive mechanisms for survival in extreme hyper-arid Atacama Desert soils. Syst Appl Microbiol. 2016;39:243–51.
Mevs U, Stackebrandt E, Schumann P, Gallikowski CA, Hirsch P. Modestobacter multiseptatus gen. nov., sp. nov., a budding actinomycete from soils of the Asgard Range (Transantarctic Mountains). Int J Syst Evol Microbiol. 2000;50:337–46.
Bull AT. Actinobacteria of the extremobiosphere. In: Horikoshi K, Antranikian G, Bull A, Robb FT, Setter KO, editors. Extremophiles handbook. Tokyo, Japan: Springer; 2011. p. 1230–40.
Jacobs JL, Carroll TL, Sundin GW. The role of pigmentation, ultraviolet radiation tolerance, and leaf colonization strategies in the epiphytic survival of phyllosphere bacteria. Micro Ecol. 2005;49:104–13.
Gtari M, Essoussi I, Maaoui R, Sghaier H, Boujmil R, Gury J, et al. Contrasted resistance of stone-dwelling Geodermatophilaceae species to stresses known to give rise to reactive oxygen species. FEMS Microbiol Ecol. 2012;80:566–77.
Link L, Sawyer J, Venkateswaran K, Nicholson W. Extreme spore UV resistance of Bacillus pumilus isolates obtained from an ultraclean spacecraft assembly facility. Microb Ecol. 2004;47:159–63.
Harveson RM, Schwartz HF, Urrea CA, Yonts CD. Bacterial wilt of dry-edible beans in the central high plains of the US: past, present, and future. Plant Dis. 2015;99:1665–77.
Schuerger AC, Mancinelli RL, Kern RG, Rothschild LJ, McKay CP. Survival of endospores of Bacillus subtilis on spacecraft surfaces under simulated martian environments: implications for the forward contamination of Mars. Icarus. 2003;165:253–76.
Haenel FJ, Stiller GP, Von Clarmann T, Funke B, Eckert E, Glatthor N, et al. Reassessment of MIPAS age of air trends and variability. Atmos Chem Phys. 2015;15:13161–76.
Ploeger F, Riese M, Haenel F, Konopka P, Müller R, Stiller G. Variability of stratospheric mean age of air and of the local effects of residual circulation and eddy mixing. J Geophys Res Atmos. 2015;120:716–33.
Benduhn F, Lawrence MG. An investigation of the role of sedimentation for stratospheric solar radiation management. J Geophys Res Atmos. 2013;118:7905–21.
Lin M, Fiore AM, Horowitz LW, Langford AO, Oltmans SJ, Tarasick D, et al. Climate variability modulates western US ozone air quality in spring via deep stratospheric intrusions. Nat Commun. 2015a;6:7105.
Lin M, Horowitz LW, Cooper OR, Tarasick D, Conley S, Iraci LT, et al. Revisiting the evidence of increasing springtime ozone mixing ratios in the free troposphere over western North America. Geophys Res Lett. 2015b;42:8719–28.
Whitman WB, Coleman DC, Wiebe WJ. Prokaryotes: the unseen majority. Proc Natl Acad Sci USA. 1998;95:6578–83.
Burrows SM, Butler T, Jöckel P, Tost H, Kerkweg A, Pöschl U, et al. Bacteria in the global atmosphere—part 2: modeling of emissions and transport between different ecosystems. Atmos Chem Phys. 2009;9:9281–97.
Acknowledgements
This work was supported by a grant from the National Aeronautics and Space Administration (NNX10AN07A), Established Program to Stimulate Competitive Research (EPSCoR), and the Louisiana Board of Regents (to BC and TG). Partial support was also provided by a Graduate Student Research Assistance Award from the Louisiana Space Grant Consortium (to NB) and from NSF awards 1241161 and 1643288 (to BC). Flight operations in Texas and New Mexico were supported by the NASA Balloon Program Office (BPO), the Columbia Scientific Balloon Facility (CSBF), and the High Altitude Student Platform (HASP).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Bryan, N.C., Christner, B.C., Guzik, T.G. et al. Abundance and survival of microbial aerosols in the troposphere and stratosphere. ISME J 13, 2789–2799 (2019). https://doi.org/10.1038/s41396-019-0474-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0474-0
This article is cited by
-
The Habitability of Venus
Space Science Reviews (2023)
-
Routes and rates of bacterial dispersal impact surface soil microbiome composition and functioning
The ISME Journal (2022)
-
Wildland fire as an atmospheric source of viable microbial aerosols and biological ice nucleating particles
The ISME Journal (2021)
-
A rather dry subject; investigating the study of arid-associated microbial communities
Environmental Microbiome (2020)
-
Sierra Nevada sweep: metagenomic measurements of bioaerosols vertically distributed across the troposphere
Scientific Reports (2020)