Abstract
Up to half of marine N losses occur in oxygen-deficient zones (ODZs). Organic matter flux from productive surface waters is considered a primary control on N2 production. Here we investigate the offshore Eastern Tropical North Pacific (ETNP) where a secondary chlorophyll a maximum resides within the ODZ. Rates of primary production and carbon export from the mixed layer and productivity in the primary chlorophyll a maximum were consistent with oligotrophic waters. However, sediment trap carbon and nitrogen fluxes increased between 105 and 150 m, indicating organic matter production within the ODZ. Metagenomic and metaproteomic characterization indicated that the secondary chlorophyll a maximum was attributable to the cyanobacterium Prochlorococcus, and numerous photosynthesis and carbon fixation proteins were detected. The presence of chemoautotrophic ammonia-oxidizing archaea and the nitrite oxidizer Nitrospina and detection of nitrate oxidoreductase was consistent with cyanobacterial oxygen production within the ODZ. Cyanobacteria and cyanophage were also present on large (>30 μm) particles and in sediment trap material. Particle cyanophage-to-host ratio exceeded 50, suggesting that viruses help convert cyanobacteria into sinking organic matter. Nitrate reduction and anammox proteins were detected, congruent with previously reported N2 production. We suggest that autochthonous organic matter production within the ODZ contributes to N2 production in the offshore ETNP.
Similar content being viewed by others
Introduction
The oxygen content of the Pacific Ocean has been decreasing since the 1980s in response to increased oxygen utilization [1]. Model predictions suggest a 1% reduction in the Pacific’s O2 inventory will double the size of the Pacific oxygen-deficient zones (ODZs) [2]. Indeed, repeat measurements indicate that the Eastern Tropical North Pacific (ETNP) ODZ has thickened over the past 40 years and intruded into shallower waters [3]. In the absence of oxygen as an electron acceptor, oxidized nitrogen species can be reduced to N2 via two main pathways: heterotrophic denitrification and autotrophic anaerobic ammonia oxidation (anammox) [4, 5]. Both processes depend on organic matter, directly in the case of denitrification and indirectly in the case of anammox, which can use ammonium released during organic matter degradation as its reductant [6]. Organic matter flux has been correlated with N2 production in the environment [7,8,9,10,11], and indeed, the addition of sterilized sediment trap material significantly increased both denitrification and anammox rates in all three marine ODZs: the Arabian Sea, the Eastern Tropical South Pacific (ETSP), and the ETNP [5, 12]. Up to half of marine N losses from N2 production occur in marine ODZs [13], so an increase in the volume of ODZs may lead to an increase in N loss and stronger N limitation on primary producers globally.
Due to sluggish circulation [14], most of the volume of the ETNP ODZs is offshore, where productivity is 10× lower than in coastal regions [15]. In general, the phytoplankton communities responsible for organic matter production also differ between coastal and offshore regions, with larger eukaryotic phytoplankton more abundant in coastal regions while cyanobacteria are the dominant primary producers offshore [15]. Given this combination of low primary productivity rates and the lower export efficiency of communities dominated by smaller-celled phytoplankton [16], export fluxes are expected to be lower in the offshore ETNP. However, in addition to production in surface waters, populations of the cyanobacterium Prochlorococcus have been found at depth within all three major ODZs [17, 18] at a secondary chlorophyll a maximum where the 1% of the deeply penetrating blue irradiance (490 nm) overlaps with the ODZ [19]. Genetic evidence using the 16S–23S ribosomal RNA (rRNA) internal transcribed spacer (ITS) region suggests that this population may be composed of novel lineages of Prochlorococcus that are closely related to cultured isolates of the low light IV (LLIV) ecotype [20]. At five stations in the ETNP, primary production was measured at the Prochlorococcus-dominated secondary chlorophyll a maximum [21]. It has been suggested that in situ production in the ODZ could significantly increase the organic C supply to N2-producing heterotrophs [21]. Here we show productivity and export measurements along with metaproteomic and metagenomic analyses that strongly supports this hypothesis. The pathway to deliver organic carbon fixed by Prochlorococcus cells to heterotrophic denitrifiers has not been established in the ODZ. Viruses have been implicated in organic matter cycling in the ocean [22], and abundance of cyanobacterial viruses have been shown to strongly correlate with organic matter flux in the oxic oligotrophic ocean [23]. From the abundance and composition of cyanobacterial viruses on particles, we suggest a role for these viruses in the conversion of cells into sinking organic matter in the ODZ.
Methods
Sampling
Samples were collected in April 2012 aboard the R/V Thompson during cruise TN278 using 10 L Niskin bottles attached to a 24 bottle CTD (conductivity–temperature–depth) rosette. A Seabird 911 Conductivity Temperature Density meter, a Seabird SBE 43 Dissolved Oxygen Sensor, a WETLabs ECO Chlorophyll Fluorometer, and a Biospherical/Licor PAR/Irradiance Sensor were attached to the CTD rosette. Hydrographic and nutrient data from the cruise are deposited at http://data.nodc.noaa.gov/accession/0109846.
Three free-floating surface-tethered sediment net traps [24] were deployed at station BB2 at 105, 150, and 750 m. Sediment trap carbon and nitrogen fluxes were measured as in Babbin et al [5]. Logistics precluded duplicate measurements of sediment trap-derived flux estimates. Carbon flux from 105 m was previously published [5], but carbon fluxes from 150 and 750 m and nitrogen fluxes have not been previously published.
Productivity measurements
Samples for triple oxygen isotope (17Δ; TOI) and O2/Ar dissolved gas analysis were collected in duplicate from the surface mixed layer on a coastal to open ocean transect (station locations shown in Fig. 1 and station information listed in Table S1) and analyzed following the laboratory procedures described in Palevsky et al. [25]. Gross primary production (GPP) rates can be calculated from mixed layer TOI by leveraging the unique isotopic signature of atmospheric-derived oxygen, which is depleted in 17O as compared to photosynthetically derived oxygen due to mass-independent fractionation in a stratospheric photochemical reaction. The TOI system is insensitive to mass-dependent fractionation as respiration removes oxygen, isolating the influence of photosynthesis from respiration and enabling calculation of mixed layer GPP by combining TOI measurements with air–sea gas exchange rates [26]. Net community production (NCP) rates, which represent organic carbon export from the mixed layer under steady-state conditions, can similarly be calculated by combining biological oxygen supersaturation determined from measured O2/Ar with air–sea gas exchange rates.
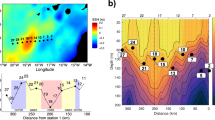
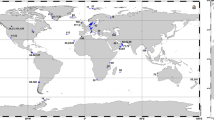
a Map of 8-day averaged satellite chlorophyll over the time period of sampling. Red dots indicate productivity sampling stations where mixed layer triple oxygen isotopes and oxygen/argon dissolved gas ratios were measured and small pink dots indicate stations where 13C incubations took place. Dark gray line indicates 10 µM O2 oxygen-deficient zones (ODZ) boundary as determined from World Ocean Atlas. A box indicates stations 136 and BB2. b Sediment trap data from station BB2. Dashed lines indicate ODZ boundary. c Transect of chlorophyll a. Station numbers are listed above the graph and correspond to triple oxygen isotope (TOI) productivity measurements (NPP; mmol C m−2 day−1) and export efficiency. Black dashes indicate mixed layer depth, and red dashes indicate the top of the ODZ
Mixed layer GPP and NCP rates were calculated from estimated air–sea gas transfer velocity and measured TOIs ([27]; Eq. 7) and O2/Ar ([28]; Eq. 3), respectively. These equations assume steady-state conditions and negligible influence of mixing and advection in the mixed layer dissolved gas budget, which model results show to be a reasonable approximation for the ETNP in April [29]. Daily wind speed data from National Oceanic and Atmospheric Administration National Climatic Data Center’s multiple-satellite Blended Sea Winds product (https://www.ncdc.noaa.gov/oa/rsad/air-sea/seawinds.html) were used to calculate air–sea gas transfer velocity [30] with the Reuer et al. [28] weighting scheme. Uncertainty in GPP and NCP rates were determined using a Monte Carlo approach (as in [25]). Mixed layer GPP rates were converted to rates equivalent to incubation-based net primary production (NPP) using a gross O2:net C ratio of 2.7 [31]. NCP rates were converted to carbon units using a O2:C ratio of 1.1 appropriate for production using a recycled nitrogen source [32].
Volumetric NPP was measured on 13 CTD casts at 10 stations (multiple casts at stations BB1 and BB2; Table S2) using stable isotope highly labeled (13C; 99%) bicarbonate in 24 h incubations. Whole water was transferred directly from Niskin bottles to 1 L polyethylene terephthalate glycol bottles using inert tubing. Incubations in triplicate light and duplicate dark bottles were initiated before dawn with the addition of a 10% enrichment of 13C-sodium bicarbonate, and bottles were incubated in shipboard incubators equipped with flow thru surface seawater to maintain temperature and neutral density screens to simulate in situ light levels where samples were collected. After 24 h, incubations were terminated by filtration (≤127 mmHg) of the entire bottle onto a pre-combusted (2 h at 450 °C) GF/F filter, and filters were stored at −20 °C. Upon return to the lab, filters were dried at 40 °C, pelletized in tin capsules, and analyzed on a Europa 20/20 isotope ratio mass spectrometer equipped with an automated nitrogen and carbon analyzer. Independent measurements of the initial particular carbon concentration and particle isotopic enrichment were used to calculate net primary production. Rates of carbon fixation were calculated as in Mulholland and Capone [33], and rates in the dark were subtracted from rates measured in light incubations.
Satellite chlorophyll data for 6–13 April 2012 (MODIS Aqua R2014, available from http://www.science.oregonstate.edu/ocean.productivity) was extracted from a 1/3° box around each station.
Metagenomics
Metagenomic libraries constructed, sequenced, and assembled using 125 and 150 bp paired end sequences at station 136 (−106.543°W 17.043°N) and station BB2 (107.148°W 16.527°N; cast 141) were previously published in Fuchsman et al [34]. DNA samples for the depth profile at station 136 were filtered onto a 0.2 μm SUPOR filters. At station BB2, DNA samples were filtered through 30 μm filters at 100, 120, and 150 m depths and subsequently filtered onto a 0.2 μm SUPOR filter. At 120 m, this prefiltered sample (30–0.2 μm fraction) was also sequenced. Depth and pore size of filters used for metagenomic analysis are found in Table S1. Hydrographic conditions were very similar at these two stations that were only 83 km apart [34]. Metagenomic sequences were assembled independently from each sample into larger contigs using the VELVET (1.2.10) assembler [35], as described in Fuchsman et al. [34] and the MEGAHIT assembler [36] as described in Widner et al [37]. Assembled contigs were functional annotation by PROKKA [38]. ETNP 2012 metagenomic reads and assembled contigs can be found at GenBank bioproject PRJNA350692.
Although the metagenomic sequences have been previously published, the genes of interest for this paper (psbD and DNA polymerase A and B) have not been previously examined. As in Fuchsman et al. [34], for each gene, a reference phylogenetic tree was constructed using both published full-length and full or nearly full-length sequences assembled from the metagenomes themselves. Sequences of interest from the metagenomes were found by searching a custom blast database [39] of all our assembled open reading frames as called by Prodigal [40] and annotated with Prokka [38], using representative published sequences as query sequences. All sequences recruited from blast were combined with the previously published full-length gene sequences and aligned in amino acid space with MUSCLE v. 3.8.1551 [41]. Maximum likelihood phylogenetic trees were constructed with the reference sequence alignments of the genes of interest using the program RAxML v. 8.1.20 [42]. The trees were constructed with a gamma model of rate heterogeneity, and appropriate amino acid substitution models were determined for each tree. Bootstrap analyses (n = 100) were performed on each tree. For the rRNA ITS tree a replicate of the Lavin et al. [20] phylogenetic tree was used.
A phylogenetic placement approach [43] was used to characterize short metagenomic reads related to the targeted genes of interest (psbD, viral DNA polymerase, and the ITS region of rRNA), in a semi-quantitative and phylogenetically specific manner [34, 44]. This allows us to both confirm the accuracy of a read assignment to a gene and also to differentiate between different taxa that might have been sources for the read. For read placement, the short metagenomic reads were recruited via tblastn search with an e value cutoff of <−5 [39]. The recruited reads were trimmed to the edge of the gene of interest to remove any overhang of up- or down-stream sequence, trimmed to the proper reading frame of the blast results. DNA polymerase and psbD reads were converted to amino acid space, while ITS reads remained in nucleotide space. Any sequence ambiguities and stop codons were removed. Only sequences longer than 100 bp (33 amino acids) after quality trimming were used. These reads were aligned to the reference sequences using PaPaRa: Parsimony-based Phylogeny-Aware Read Alignment program [45]. Paired end reads were combined into the same alignment and placed as one on the tree using the EPA: Evolutionary Placement Algorithm portion of RAxML [42]. Each read has a number, or “branchlength,” which corresponds to the similarity between the read and the sequence to which it is placed. Reads placed with a read “branchlength” longer than 2.0 were removed as erroneous. Spot testing indicated that these reads belonged to different genes than the one examined. Only a small percentage of reads were thus removed (0.1–2.5%). Reads are normalized by the number of sequences in each metagenome, divided by gene length, and then multiplied by 100 to make the numbers easier to visualize. For the ITS region, read abundance for Synechococcus, Prochlorococcus LLIV/V, and NC1 were divided by two since those organisms contain two copies of the ITS region.
Metaproteomics
At station BB2 metaproteomic analyses were conducted on both water column and sediment trap samples. Water was filtered in situ at 55, 100, 145, 160, and 250 m using a McLane pump (McLane Research Laboratories, Falmouth, MA, USA) fitted with a 4 mm mesh screen and combusted 142 mm diameter GF/F filters (0.7 µm) that were double-stacked. Volumes filtered were between 300 and 1000 L. Depth and filter pore size used for metaproteomic analysis are found in Table S3. Materials from two 11 mm circular punches of the GF/F filter were used for protein extraction. Additionally proteins were extracted from two 0.2 µm Sterivex filters from 100 and 150 m (~6 L of water filtered from the CTD) to compare to the results from the larger GF/F pore size from the McLane pumps. The sediment trap samples were allowed to settle and then decanted to a small volume and split between carbon and proteomics analyses. Only 105 and 750 m sediment trap samples were analyzed for proteomics. Filters or sediment trap material were extracted into ammonium bicarbonate following a protocol adapted from Bridoux et al [46]. Briefly, the chilled suspension was lysed with the help of a high-power sonicator, resulting in a whole-debris lysate. The lysate was enzymatically digested with trypsin, following a protocol adapted from Nunn et al. [47]. The resulting tryptic peptides were desalted using a macro-spin C18 column (NestGroup), resuspended in 5% acetonitrile with 0.1% formic acid (FA), and 1 µg of protein was introduced to a Thermo Q Exactive Plus mass spectrometer using a Waters nanoAcquity UPLC. Peptides were separated on a home-packed analytical column consisting of a 37 cm log, 75-μm i.d. fused-silica capillary column packed with C18 particles (Magic C18AQ, 100 Å, 5 mm; Michrom) coupled to a 4 cm long, 100 μm i.d. precolumn (Magic C18AQ, 200 Å, 5 mm; Michrom). Solvents of 100% water with 0.1% FA (A) and 100% acetonitrile with 0.1% FA (B) were used to elute peptides over a 90 min gradient from 5 to 35% solvent B. A data-dependent acquisition strategy was utilized, where tandem mass spectrometry spectra were collected on the ten most intense ions observed in the MS1 precursor scan.
The resulting mass spectra were analyzed using PEAKS (version 8.5; Bioinformatics Solutions, Waterloo, Canada; [48]). Proteins were identified using PEAKS DB, a database matching search platform augmented by de novo peptide sequencing, using a database consisting of assembled proteins from ETNP metagenomes [34, 49, 50] as well as all proteins translated from the Marine Microbial Eukaryote Transcript Project [51] and a selection of genomes from relevant cultures (Table S4). A contaminant database downloaded from http://www.thegpm.org/crap/ was included in the database search. A target-decoy search approach was used to estimate the false discovery rate of peptide-spectrum matches. All MS2 spectra were searched against the database with the following parameters: enzyme type, trypsin; allowed missed cleavages, 3; peptide confidence cutoff: ≥15 −log P; protein confidence cutoff: ≥20 –log P; unique peptides: ≥1; protein false discovery rate: 4%. Fixed modification of cysteine residues by 57.021 Da, due to iodoacetamide modification, and variable modification of methionine by 15.995 Da, due to oxidation, were allowed. Protein identifications were accepted if they could be established at >95.0% probability.
When the identified protein was a protein from assembled ETNP metagenomes, in an attempt to identify the organism the protein derived from, the protein was blasted against a custom database composed of the proteins from genomes used for the proteomics database (Table S4). The best blast hit was recorded if the e value was <−20.
The phylogenetic composition of the proteins identified from the GF/F and 0.2 μm Sterivex filters from the same depths are fundamentally similar with some variations in the abundance of taxa (Fig. S3). The percent of cyanobacteria decrease and the amount of Marine Group A/SAR406 increase in 0.2 μm samples compared to GF/F (Fig. S3).
Results and discussion
Productivity and export
In April 2012, we occupied 10 stations along an 830 km transect from the coast (18.8°N 104°W) to offshore waters (14.0°N 110°W) through the heart of the ETNP ODZ (Fig. 1a). We use the triple oxygen isotope composition of mixed layer dissolved oxygen to calculate mixed layer productivity across the spatial extent of the transect and 13C enrichment incubations to expand our productivity measurements below the mixed layer at a few stations. Satellite-derived surface chlorophyll a concentrations were 20 μg L−1 at the station nearest the coast, decreasing greatly to 0.1–0.3 μg L−1 at offshore stations (Fig. 1, Table S1). These offshore chlorophyll a values are typical of the region [15]. Similarly, gross primary production in the mixed layer, determined by the incubation-independent triple oxygen isotope (TOI) technique, was highest near the coast (St 133: 182 ± 29 mmol O2 m−2 d−1) and lower (41 ± 7 to 94 ± 12 mmol O2 m−2 d−1) at the more offshore stations (Table S1). Using a gross O2:net C ratio of 2.7 [31], these values are equivalent to net primary production (NPP) offshore of 15 ± 2 to 34 ± 5 mmol C m−2 d−1; Table S1 and Fig. 1). Volumetric NPP rates measured using 13C incubations were approximately ten times greater in surface waters at the coastal stations (5–11 µM C d−1 or 71–130 mmol C m−2 d−1 integrated across the mixed layer; Figure S1 & Table S2) than in the offshore region (0.5–1 µM C d−1 or 16–29 mmol C m−2 d−1 integrated across the mixed layer) (Fig. 2 & Table S2). In the literature, the average surface productivity in this offshore region is 0.2 µM C d−1 [15], even smaller than our measured surface values. TOI and 13C NPP estimates agree well given that TOI measurements represent an average over the mixed layer over several weeks while the incubation experiments were 24 h.
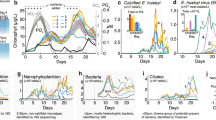
Primary production, autotrophic bacteria, and water column properties offshore. a %PAR (photosynthetically active radiation) and density profiles at station BB2, b Chlorophyll a fluorescence, STOX (Switchable Trace Oxygen) oxygen data from repeat casts at BB2 [54], net primary productivity (NPP) from 13C experiments from three offshore stations. c Metagenomic rpoB reads for cyanobacteria, and oxygen-utilizing ammonia-oxidizing Thaumarchaea (AOA) and nitrite oxidizer Nitrospina at station 136 [34]
The majority of the offshore stations exhibited vertical profiles with two chlorophyll a maxima; both were below the surface mixed layer and the secondary chlorophyll a maximum was within in the upper ODZ (Fig. 1c). Mixed layer TOI samples only provide estimates of productivity within the mixed layer, but 13C incubations were conducted from the surface to 1% PAR at five stations to determine sub-mixed layer primary production rates (Table S2). At the offshore stations where 13C-NPP incubations were conducted, volumetric rates at the primary chlorophyll a maximum varied between 0.7 and 1.8 µM C day−1 (Fig. 2; Table S2). Thus, at the offshore stations volumetric NPP rates were similar from the surface to 1% PAR (Fig. 2). At station BB2, volumetric NPP integrated from the mixed layer down to the 1% PAR level at 75 m (Fig. 2) for 13C-based NPP measurements was 49 mmol C m−2 day−1, only slightly higher than the mean euphotic zone NPP at oligotrophic Station ALOHA (42 ± 7 mmol C m−2 day−1) [52].
At the offshore stations, carbon export from the mixed layer, as determined from dissolved oxygen/argon ratios, ranged from 0.2 to 3.2 mmol C m−2 day−1, or 1–13% of NPP (Table S1 and Fig. 1). Thus, the majority of photosynthetically fixed carbon was recycled within the mixed layer, with only a small fraction sinking out of the surface mixed layer. These export rates are lower than mean export from the mixed layer determined by dissolved oxygen/argon ratios at Station ALOHA near Hawaii (10 ± 3 mmol C m−2 day−1; [52]). Export rates were also lower than offshore stations off the coast of Southern California (9 ± 3 mmol C m−2 day−1) where mixed layer net productivity from TOI was 31 ± 14 mmol C m−2 day−1 [53]. However, the rates reported here in the ETNP do not include contributions to export from below the mixed layer, which need to be considered since the mixed layer was shallower than the euphotic depth throughout the study region and both chlorophyll a maxima were substantially below the surface mixed layer. Sediment trap flux estimates from traps deployed at 105 m at station BB2 (the top of the ODZ as determined by a STOX (Switchable Trace Oxygen) oxygen sensor; [54]) were lower (0.39 mmol C m2 day−1 [5] and 38 μmol N m2 day−1; Fig. 1b) than those estimated at the bottom of the mixed later (42 m; Fig. 2) by the TOI method (Table S1), indicating that carbon export 60 m below the mixed layer was also low. These flux rates were lower than sediment trap-based flux estimates at a similar depth from the persistently oligotrophic regime at Station ALOHA (1.7–3.3 mmol C m−2 day−1; [55]).
Although organic matter fluxes generally attenuate with depth in the ocean, at our station in 2012, sediment trap fluxes at 150 m (0.66 mmol C m−2 day−1 and 64 μmol N m2 day−1) were higher than fluxes measured at 105 m (Fig. 1b). The 750 m sediment trap did measure the lowest fluxes (0.34 mmol C m−2 day−1 and 6 μmol N m−2 day−1), however. If we assume the difference between fluxes at 105 and 150 m (0.26 mmol C m−2 day−1 and 25 μmol N m−2 day−1) are due to in situ productivity in the ODZ, this productivity would make up 40% of the particle flux at 150 m. Although no primary production measurements were made within the secondary chlorophyll a maximum (110–125 m) in the ODZ during the 2012 cruise, rates of NCP integrated across the secondary chlorophyll a maximum of 0.13– 0.93 mmol C m−2 day−1 were measured by 13C incubations at five stations in a similar area of the ETNP in 2014 [21], demonstrating the potential for in situ productivity within the ODZ. Although our calculation of a 40% contribution to flux involves a comparison of only two sediment trap samples, combined with previously measured productivity rates in the ODZ, these data suggest a significant amount of primary production and carbon export at the secondary chlorophyll a maximum.
Phytoplankton in and above the ODZ
To characterize the functional and taxonomic diversity of microbes within this oligotrophic ODZ community, we examined metagenomes from this 2012 cruise [34] from nine depths (60–180 m) at the offshore station 136 (Fig. 1a) with particular attention to the secondary chlorophyll a maximum. We identified short sequences by both function and phylotype by placing them on reference trees constructed from full-length genes, allowing us to use the short-read data in a semi-quantitative manner.
We used genes for the photosystem I reaction center core protein PsbD to identify photoautotrophs (Fig. S2). The primary chlorophyll a maximum contained both cyanobacteria and eukaryotes, including green algae (an unidentified prasinophyte and Mamiellales related to Ostreococcus and Bathycoccus) and heterokonts (Pelagophytes and diatoms) (Fig. 3a, b). In contrast, the secondary chlorophyll a maximum was entirely cyanobacteria with Prochlorococcus dominating the photoautotrophic community (Fig. 3a, b), which is consistent with previous flow cytometry results [17, 21]. The proportion of cyanobacterial reads declined with depth from 18% of the total microbial community in oxic waters, as determined by rpoB, to about 5% in the upper ODZ (Fig. 2c). We used the rRNA ITS to further identify the Prochlorococcus genotypes in each region of the water column. Prochlorococcus clade LLI was dominant at 60 m (Fig. 3), which corresponds to the primary chlorophyll a maximum (Fig. 2), and Prochlorococcus clades LLII and the uncultured clade NC1 [56] were also found in oxic waters (Fig. 3b). Prochlorococcus clade LLIV/LLV was the dominant ecotype in the anoxic secondary chlorophyll a maximum (Fig. 3b).
Diversity of photoautotrophs in the water column and on particles. a Depth profile of normalized reads for eukaryotes as determined by psbD in water column metagenomes. b Depth profile of normalized reads for cyanobacterial genotypes based on internal transcribed spacer (ITS) sequences in water column metagenomes, including Synechococcus (Syn) and Prochlorococcus high light (HL) and low light (LL) clades and the uncultured NC1 clade. c Eukaryotic psbD normalized reads found in 100, 120, and 150 m particle metagenomes. Colors indicate psbD phylotypes as indicated in a. d Cyanobacterial ITS genotypes from normalized reads found in 100, 120, and 150 m particle metagenomes. Colors indicate ITS genotypes as indicated in b. Dashed lines indicate the top of the oxygen-deficient zones (ODZ)
Additionally, we performed metaproteomics on water column samples (McClane pump) at five depths at station BB2 (55, 100, 145, 160, and 250 m), Cyanobacteria dominated the metaproteome at 100 m composing 50% of the total identified proteins (Fig. S3; Table S5). The number of cyanobacterial proteins decreased with depth, composing 22% of identified proteins at 145 m and 10% at 160 m, and 4% at 250 m (Fig. S3; Table S5). Prochlorococcus-derived proteins exceeded those from Synechococcus in the water column (Fig. S3; Table S5), probably due to the higher abundance of Prochlorococcus in the community (Fig. 3). Although our analysis was not quantitative and many proteins present in the sample were likely not identified due to the database-dependent limitations of the metaproteomics technique [57], the presence of a protein in metaproteomic sample, particularly one that might be expected to have a short half life, such as an enzyme, suggests it is being actively expressed.
Identified proteins indicate that cyanobacteria were actively photosynthesizing and fixing carbon in the water column at 100 m, a depth with 0.8 μM oxygen [54]. Proteins identified included those used in light capture, cyanobacterial enzymes for photosynthesis (28 photosystem proteins, including all 3 extrinsic proteins of the oxygen evolving complex), electron transport (cytochrome c6 and ferredoxin), ATP synthesis (4 subunits of ATP synthase), bicarbonate transporters, carbon concentration (carboxysome shell protein and CO2 concentrating protein CcmK), and carbon fixation (both subunits of RubisCo), as well as replication (DNA-binding protein HU), transcription (2 subunits of DNA directed RNA polymerase), translation (Elongation Factors Tu, Ts, and G), and cell division (FtsZ) were all detected (Fig. 4a, Table S5). Additionally, Prochlorococcus were actively attempting to acquire nitrogen compounds for assimilation as evidenced by detection of nitrogen regulatory protein P-II, a urea ABC transporter, a nitrite transporter, and an assimilatory nitrite reductase. Since ammonium in the ODZ is generally below detection limits [37], Prochlorococcus populations may need to use alternative sources of N for assimilation. Thus, Prochlorococcus was clearly active at the secondary chlorophyll a maximum in the ODZ.
Metaproteomic data indicating active cyanobacterial production within the oxygen-deficient zones (ODZ) and evidence for the ODZ cyanobacterial population contribution to particles. a Cyanobacterial proteins involved in primary production identified at 100 m. The Oxygen evolving complex (OEC) and ferredoxin NADP+ reductase (FNR) are abbreviated on the figure. b A comparison of total peptides identified between the 105 m sediment trap, 50 m oxic water column, and 100 m ODZ water column samples. c A comparison of only the cyanobacterial peptides identified between the 105 m sediment trap, 50 m oxic water column, and 100 m ODZ water column samples
Cyanobacteria on particles
To investigate whether ODZ cyanobacteria can contribute to the particles that make up the sinking organic matter in the ODZ, we examined metagenomes from the >30 μm size fraction collected from Niskin bottles at 100, 120, and 150 m, and metaproteomes from sediment trap material at 105 and 750 m, both at offshore station BB2. Cyanobacteria were present in >30 μm particles, especially at 100 m, where, when normalized by the single copy core gene rpoB, they made up 7.5% of the prokaryotic community [34]. In contrast, we note that the % of community of SAR11, the most abundant bacteria in the ODZ, decreased dramatically on the >30 μm particles compared to the water column [34], indicating that the 30 μm filters were not clogged. The cyanobacteria in the particle fraction were predominately LLIV/LLV Prochlorococcus, prevalent in the water column in the ODZ, and Synechococcus group CRD1, which was present in the oxic water column but not within the ODZ water column (Fig. 3d). The difference in Synechococcus types between particles and the ODZ water column indicates that Synechococcus on particles came from surface waters. Synechococcus has been found on particles in other systems [58]. Notably, the fraction of LLI Prochlorococcus, dominant at the primary chlorophyll a maximum, was miniscule at the secondary chlorophyll a maximum and in the particle fraction (Fig. 3d), implying that in contrast to Synechococcus, Prochlorococcus cells were not sinking into the ODZ from the primary chlorophyll a maximum.
In the 105 m sediment trap metaproteomes, cyanobacterial proteins composed 17% of the identified proteins (Fig. S3). The cyanobacterial proteins consisted of structural proteins used in light capture, proteins used in photosynthesis, ATP synthesis, and transporters (Table S5). When cyanobacterial peptides were compared between the oxic 50 m water column sample, the 100 m ODZ water column sample and the 105 m sediment trap, 67% of the cyanobacterial peptides in the 105 m trap were shared with the 100 m water column sample (Fig. 4c). Thus, our particulate metaproteomic data and metagenomic data both suggest that there is transfer of some organic material from cyanobacteria living in the ODZ to sinking particles.
Viruses as source of cyanobacterial mortality
We evaluated cyanophages (viruses that infect cyanobacteria) as a potential contributor to cyanobacteria mortality and particle formation. Cyanophages were detected in both free-living and particle metagenomes using DNA polymerase A and B, forms of DNA polymerase found in all known T7-like cyanopodoviruses and T4-like cyanomyoviruses (Fig. 5). Genes for cyanophage DNA polymerase can be split into phylotypes (Fig. 5 and Figs. S4, S5; following [59]). Cyanomyoviruses were more abundant than cyanopodoviruses in the ODZ with cyanomyovirus clade II as the dominant type of virus (Fig. 5A). While cyanomyovirus clade III decreases in the ODZ, cyanomyovirus clade I and cyanopodovirus A increase in the ODZ (Fig. 5A). Thus, the relative proportions of cyanophage types differ between the oxic waters and the ODZ. The virus/host ratio (as calculated using cyanophage DNA polymerase/cyanobacteria rpoB) was high (30–40) in the free-living metagenomes and even higher (>100) in the >30 μm particulate metagenomes, which is consistent with active infection (Fig. 5B). Viral infection has been shown to cause bacteria to clump and form particles [60]. Although sinking particles may also scavenge viruses from the water column [61], the cyanophage phylotypes on 100 and 120 m particles are found in the same proportions as found in the ODZ water column, but differed in proportion compared to those in the oxic overlying water (Fig. 5C, D and Fig. S6). This is consistent with the idea that cyanobacteria and their viruses were not sinking into the ODZ from particles formed in oxic water above, but rather were creating particles in situ.
Cyanophage prevalence in the water column and on particles. a Depth profiles of normalized reads for DNA polymerase phylotypes. b Virus to host ratio in cells (>0.2 μm) and on particles (>30 μm) and one free-living microbial sample (<30 μm). Ratios are calculated using cyanophage DNA polymerase divided by cyanobacteria RNA polymerase B (rpoB). Dashed line indicates the top of the ODZ. c Viral DNA polymerase normalized reads found in 100, 120, and 150 m particle metagenomes. Colors indicate DNA polymerase phylotypes as indicated in a and delineated in Figs. S4, S5. d Comparison of the proportion of DNA polymerase phylotypes between 100 m particulate (>30 μm) and >0.2 μm water column samples from the ODZ (100, 110, and 120 m). Each symbol represents the proportion of a phylotype of DNA polymerase. The black line represents the 1:1 line
Viral proteins were also detected in the metaproteomes from the water column and the sediment trap. At 100 m many structural capsid proteins for cyanophage were identified in the proteome (Table S4). Cyanophage composed 2% of total identified proteins at 100 m and 1% in the 105 m sediment trap sample (Fig. S3). These structural proteins could be indicative of detection of free viruses. However, we also detected proteins expected to be expressed only during the process of viral infection including a viral DNA maturase and a host-derived viral photosystem I P700 chlorophyll a protein (PsaB), both detected at 100 m [62, 63]. This is additional evidence for ongoing cyanophage infection in these waters. Viral lysis of cyanobacteria would inject organic matter into the ODZ. Our viral results are consistent with the hypothesis that in situ production by cyanobacteria in the ODZ leads to release of organic matter into the system.
Impacts of in situ productivity on nitrogen cycling
Consistent with observations of active cyanobacterial oxygenic photosynthesis within the ODZ, combined STOX sensor depth profiles from five casts from BB2 indicate transitory nanomolar oxygen concentrations in the depth range of the secondary chlorophyll a maximum (Fig. 2) [54]. The creation of oxygen by photosynthesis may enable oxygen-requiring organisms to exist and thrive within the nominally anoxic ODZ. In particular, both ammonia and nitrite oxidization can occur at very low oxygen concentrations. The Km for nitrite oxidizers at low oxygen is 0.5 ± 4.0 nM O2 (from two component fitting) and for ammonia oxidizers is 333 ± 130 nM [64]. Metagenomes indicate the aerobic chemoautotrophic nitrite oxidizer, Nitrospina, co-occurred with cyanobacteria in the ODZ (Fig. 2), and the aerobic ammonia-oxidizing Thaumarchaea were also found at lower abundances in the same depth range (Fig. 2) [34]. We also detected the nitrite-oxidizing protein nitrate oxidoreductase from Nitrospina at 100 m (Table S5). Thus, the presence of Prochlorococcus may contribute to nitrite oxidation in this ODZ. Measured potential oxidation rates from this same cruise are consistent with this idea [65]. Nitrite oxidation rates had their maximum of 120 nM day−1 at 105 m at BB2 and were undetectable at 150 m [65], below the LLIV/LLV Prochlorococcus maximum. Ammonia oxidation rates were 9 nM day−1 at 105 m (1/3 of their maximum) and were undetectable at 150 m, but were not measured in between [65]. Similarly, transcripts indicated activity of nitrite oxidizer Nitrospina and ammonia-oxidizing archaea at the secondary chlorophyll a maximum in 2014 [21]. Transcript evidence also supports the possibility of heterotrophic use of oxygen in anoxic waters at the secondary chlorophyll a maximum [21]. Our results are consistent with the idea that oxygen production from photosynthesis during the day by Prochlorococcus in the ODZ may fuel an oxygen-utilizing community under anoxic conditions, and thus the activity of bacteria and archaea at these depths may vary over a diel cycle.
Several lines of evidence indicate active production of N2 in the offshore region. We have previously reported the diverse community of denitrifiers identified at this station with metagenomics [34]. Here, in the metaproteomes at least six types of nitrate reductase proteins, including both types found in the ODZ SAR11 [50], were found in 145, 160, and 250 m samples (Table S5). Respiratory nitrate reductase proteins were also found in 105 m sediment trap material (Table S5). Anammox activity was also detected in the metaproteome, consistent with the previously described presence of Candidatus Scalindua in the water [34]. Proteins detected for the anammox process included hydrazine synthase (both subunits), hydroxylamine oxidoreductase, nitrite oxidoreductase, and an ammonium transporter, found at 145, 160, and 250 m in the water column (Table S5). Thus, the proteins detected here are consistent with in situ occurrence of both denitrification and anammox. Previously reported denitrification and anammox rates were 2.1 and 0.9 nM day−1, respectively, at 150 m at offshore station BB2 [5, 66]. Rates increased to 15.1 and 6.0 nM day−1, respectively, when excess sterile sediment trap material was added [5].
Carbon flux and offshore production of N2
Despite low surface productivity and export, in the following calculation, we show that measured N2 gas concentrations and N2 production rates can be supported by the amount of organic C in the offshore ODZ without the need for transport of N2 from coastal waters. Since sediment trap organic C was 40% greater at 150 m than 105 m, in situ production may contribute to N2 production rates. We can estimate N2 production rates from organic matter fluxes using an equation for denitrification [67]:
and assuming that anammox bacteria can use the ammonia produced in this organic matter degradation in a 1:1 ratio with nitrite to make N2. We use the difference between sinking C fluxes at 150 m (0.66 mmol C m−2 day−1) and 750 m (0.34 mmol C m−2 day−1), at the bottom of the ODZ, to estimate the amount of organic C consumed in the ODZ. However, sediment traps do not quantify all types of C fluxes. Both export of dissolved organic C through mixing and active C export by zooplankton migration could be important factors increasing the organic C flux [68]. Total organic C fluxes, including dissolved organics and active transport, are estimated to be 2–3 times larger than sediment trap fluxes in the North Pacific [68]. For our calculation, we double the amount of organic C consumed in the ODZ from sediment trap fluxes (0.32 mmol C m−2 day−1) to estimate total C fluxes (0.64 mmol C m−2 day−1). N2 production is much higher in the upper part of the ODZ compared to the lower [5]. If the majority of organic matter consumption occurred in the top 30% of the ODZ (200 m), our calculated N2 production rate in N units would be 4–6 nM N day−1, which is similar to measured rates (1.5–5 nM N day−1; [5]). The residence time of water in the ETNP ODZ in a 3-D model was 3.9 ± 0.8 years [69], allowing 6–9 μM N as N2 to build up in the top 200 m from consumption of organic C fluxes. Measured biological N2 was 10 μM N as N2 at 150 m at this station [34]. Organic matter fluxes like those seen here would allow offshore N2 production to create measured values of N2 gas. Thus, we have evidence from rate measurements and proteomics that N2 production occurs locally offshore and our calculations indicate that the majority of the N2 gas measured offshore could be produced in situ. The increase in sediment trap fluxes from 105 to 150 m suggests that in situ primary production accounted for 40% of these fluxes and thus of 40% of the N2 production. Given the extent of the secondary chlorophyll a maximum in the offshore ETNP ODZ (Fig. 1), the impact of in situ primary production in the ODZ could be significant. In corroboration, among three offshore stations examined in the ETSP, active anammox cells were only detected at the station with a deep cyanobacterial maximum within the ODZ [70]. Additionally, no N2 production was detected at stations in the offshore Southern ETNP where the top of the ODZ was at 340 m [71], a depth too deep for cyanobacteria to obtain light.
Conclusions
We propose that offshore N loss processes were at least partly due to the influx of organic matter created by primary production inside the ODZ. Together, our metagenomic and metaproteomic data indicates that cyanobacteria were abundant in the upper ODZ and were actively photosynthesizing and dividing. According to metagenomic and metaproteomic data, cyanobacteria comprised 7% of prokaryotes and 17% of identified proteins on 100 m particles. Proteins derived specifically from this ODZ population were found in sediment traps, suggesting an in situ cyanobacterial contribution to particles. Detection of cyanophage proteins indicative of active infection and high corresponding cyanophage-to-cyanobacterial ratios suggest that cyanobacteria may have been experiencing viral-induced death. The release of organic matter from this process could have been a source of carbon fueling microbial activity within the ODZ. In support of this concept, the sediment trap-derived particle flux estimates at 150 m, right below the secondary chlorophyll maximum (100–140 m) were higher than those at 105 m, suggesting that cyanobacteria were actively contributing to particle flux there.
The thickness of the ETNP ODZ has increased over the past 40 years and the upper oxycline has become shallower [3]. Further shoaling of the ODZ into sunlit waters could increase the habitat of cyanobacteria and thus the amount of in situ primary production in the ODZ. The presence of significant primary production within the N-replete ODZ may contribute organic carbon used for N2 production, allowing N loss from oceanic ODZs to continue even with increasing N limitation in surface waters.
References
Ito T, Minobe S, Long MC, Deutsch C. Upper ocean O2 trends: 1958–2015. Geophys Res Lett. 2017;44:4214–23.
Deutsch C, Brix H, Ito T, Frenzel H, Thompson L. Climate-forced variability of ocean hypoxia. Science. 2011;333:336–9.
Horak REA, Ruef W, Ward BB, Devol AH. Expansion of denitrification and anoxia in the eastern tropical North Pacific from 1972 to 2012. Geophys Res Lett. 2016;43:5252–60.
Dalsgaard T, Thamdrup B, Farías L, Peter Revsbech N. Anammox and denitrification in the oxygen minimum zone of the eastern South Pacific. Limnol Oceanogr. 2012;57:1331–46.
Babbin AR, Keil RG, Devol AH, Ward BB. Organic matter stoichiometry, flux, and oxygen control nitrogen loss in the ocean. Science. 2014;344:406.
Devol AH. Solution to a marine mystery. Nature. 2003;422:575.
Fuchsman CA, Murray JW, Konovalov SK. Concentration and natural stable isotope profiles of nitrogen species in the Black Sea. Mar Chem. 2008;111:90–105.
Kalvelage T, Lavik G, Lam P, Contreras S, Arteaga L, Löscher CR, et al. Nitrogen cycling driven by organic matter export in the South Pacific oxygen minimum zone. Nat Geosci. 2013;6:228–34.
Kirkpatrick JB, Fuchsman CA, Yakushev E, Staley JT, Murray JW. Concurrent activity of anammox and denitrifying bacteria in the Black Sea. Front Microbiol. 2012;3:Article 256.
Montes E, Altabet MA, Scranton MI, Thunell RC, Lorenzoni L. Biogenic nitrogen gas production at the oxic– anoxic interface in the Cariaco Basin, Venezuela. Biogeosciences. 2013;10:267–79.
Ward BB, Devol AH, Rich JJ, Chang BX, Bulow SE, Naik H, et al. Denitrification as the dominant nitrogen loss process in the Arabian Sea. Nature. 2009;461:78.
Chang BX, Rich JR, Jayakumar A, Naik H, Pratihary AK, Keil RG, et al. The effect of organic carbon on fixed nitrogen loss in the eastern tropical South Pacific and Arabian Sea oxygen deficient zones. Limnol Oceanogr. 2014;59:1267–74.
DeVries T, Deutsch C, Rafter PA, Primeau F. Marine denitrification rates determined from a global 3-D inverse model. Biogeosciences. 2013;10:2481–96.
Kessler WS. The circulation of the eastern tropical Pacific: a review. Prog Oceanogr. 2006;69:181–217.
Pennington JT, Mahoney KL, Kuwahara VS, Kolber DD, Calienes R, Chavez FP. Primary production in the eastern tropical Pacific: a review. Prog Oceanogr. 2006;69:285–317.
Boyd PW, Newton PP. Does planktonic community structure determine downward particulate organic carbon flux in different oceanic provinces? Deep Sea Res Part I. 1999;46:63–91.
Goericke R, Olson RJ, Shalapyonok A. A novel niche for Prochlorococcus sp. in low-light suboxic environments in the Arabian Sea and the Eastern Tropical North Pacific. Deep Sea Res Part I. 2000;47:1183–1205.
Johnson Z, Landry ML, Bidigare RR, Brown SL, Campbell L, Gunderson J, et al. Energetics and growth kinetics of a deep Prochlorococcus spp. population in the Arabian Sea. Deep Sea Res Part II. 1999;46:1719–43.
Cepeda-Morales J, Beier J, Gaxiola-Castro G, Lavin MF, Godinez VM. Effect of the oxygen minimum zone on the second chlorophyll maximum in the Eastern Tropical Pacific off Mexico Universidad Autónoma de Baja California Effect of the oxygen minimum zone on the second chlorophyll maximum in the Eastern Tropical Pacific off. Ciencias Mar. 2009;35:389–403.
Lavin P, González B, Santibáñez JF, Scanlan DJ, Ulloa O. Novel lineages of Prochlorococcus thrive within. Environ Microbiol Rep. 2010;2:728–38.
Garcia-Robledo E, Padilla CC, Aldunate M, Stewart FJ, Ulloa O, Paulmier A, et al. Cryptic oxygen cycling in anoxic marine zones. Proc Natl Acad Sci USA. 2017;114:8319–24.
Weitz J, Wilhelm S. Ocean viruses and their effects on microbial communities and biogeochemical cycles. F1000 Biol Rep. 2012;4:17.
Guidi L, Chaffron S, Bittner L, Eveillard D, Larhlimi A, Roux S, et al. Plankton networks driving carbon export in the oligotrophic ocean. Nature. 2016;532:465–70.
Peterson ML, Wakeham SG, Lee C, Askea MA, Miquel JC. Novel techniques for collection of sinking particles in the ocean and determining their settling rates. Limnol Oceanogr. 2005;3:520–32.
Palevsky HI, Quay PD, Lockwood DE, Nicholson DP. The annual cycle of gross primary production, net community production, and export efficiency across the North Pacific Ocean. Glob Biogeochem Cycles. 2016;30:361–80.
Juranek LW, Quay PD. Using triple isotopes of dissolved oxygen to evaluate global marine productivity. Annu Rev Mar Sci. 2013;5:503–24.
Prokopenko MG, Pauluis OM, Granger J, Yeung LY. Exact evaluation of gross photosynthetic production from the oxygen triple-isotope composition of O2: implications for the net-to-gross primary production ratios. Geophys Res Lett. 2011;38:L14603.
Reuer MK, Barnett Ba, Bender ML, Falkowski PG, Hendricks MB. New estimates of Southern Ocean biological production rates from O2/Ar ratios and the triple isotope composition of O2. Deep Sea Res Part I. 2007;54:951–74.
Nicholson D, Stanley RHR, Doney SC. The triple oxygen isotope tracer of primary productivity in a dynamic ocean model. Glob Biogeochem Cycles. 2014;28:538–52.
Nightingale PD, Malin G, Law CS, Watson AJ, Liss PS, Liddicoat MI, et al. In situ evaluation of air–sea gas exchange parameterizations using novel conservative and volatile tracers. Glob Biogeochem Cycles. 2000;14:373.
Marra J. Approaches to the measurement of plankton production. In: Williams PJ, Thomas DR, Reynolds CS, editors. Productivity and carbon assimilation in marine and freshwater ecosystems. London: Blackwell; 2002. p. 78–108.
Laws EA. Photosynthetic quotients, new production and net community production in the open ocean. Deep Sea Res Part A. 1991;38:143–67.
Mulholland MR, Capone DG. Stoichiometry of nitrogen and carbon utilization in cultured populations of Trichodesmium IMS101: implications for growth. Limnol Oceanogr. 2001;46:436–43.
Fuchsman CA, Devol AH, Saunders JK, McKay C, Rocap G. Niche partitioning of the N cycling microbial community of an offshore oxygen deficient zone. Front Microbiol. 2017;8:2384.
Zerbino DR. Using the Velvet de novo assembler for short-read sequencing technologies. Curr Protoc Bioinformatics. 2010;Unit 11.5:1–13.
Li D, Liu CM, Luo R, Sadakane K, Lam TW. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics. 2015;31:1674–6.
Widner B, Fuchsman CA, Chang BX, Rocap G, Mulholland MR. Utilization of urea and cyanate in waters overlying and within the eastern tropical north Pacific oxygen deficient zone. FEMS Microbiol Ecol. 2018;94:fiy138.
Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30:2068–9.
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–402.
Hyatt D, Chen G-L, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics. 2010;11:119.
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7.
Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3.
Berger SA, Krompass D, Stamatakis A. Performance, accuracy, and web server for evolutionary placement of short sequence reads under maximum likelihood. Syst Biol. 2011;60:291–302.
Saunders JK, Rocap G. Genomic potential for arsenic efflux and methylation varies among global Prochlorococcus populations. ISME J. 2016;10:197–209.
Berger SA, Stamatakis A. Aligning short reads to reference alignments and trees. Bioinformatics. 2011;27:2068–75.
Bridoux MC, Neibauer J, Ingalls AE, Nunn BL, Keil RG. Suspended marine particulate proteins in coastal and oligotrophic waters. J Mar Syst. 2015;143:39–48.
Nunn BL, Ting YS, Malmstrom L, Tsai YS, Squier A, Goodlett DR, et al. The path to preservation: using proteomics to decipher the fate of diatom proteins during microbial degradation. Limnol Oceanogr. 2010;55:1790–804.
Zhang J, Xin L, Shan B, Chen W, Xie M, Yuen D, et al. PEAKS DB: de novo sequencing assisted database search for sensitive and accurate peptide identification. Mol Cell Proteomics. 2012;11:M111.010587.
Glass JB, Kretz CB, Ganesh S, Ranjan P, Seston SL, Buck KN, et al. Meta-omic signatures of microbial metal and nitrogen cycling in marine oxygen minimum zones. Front Microbiol. 2015;6:Article 998.
Tsementzi D, Wu J, Deutsch S, Nath S, Rodriguez-r LM, Burns AS, et al. SAR11 bacteria linked to ocean anoxia and nitrogen loss. Nature. 2016;536:179–83.
Keeling PJ, Burki F, Wilcox HM, Allam B, Allen EE, Amaral-Zettler LA, et al. The Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP): illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol. 2014;12:e1001889.
Quay PD, Peacock C, Björkman K, Karl DM. Measuring primary production rates in the ocean: enigmatic results between incubation and non-incubation methods at Station ALOHA. Glob Biogeochem Cycles. 2010;24:GB3014.
Munro DR, Quay PD, Juranek LW, Goericke R. Biological production rates off the Southern California coast estimated from triple O2 isotopes and O2: Ar gas ratios. Limnol Oceanogr. 2013;58:1312–28.
Tiano L, Garcia-Robledo E, Dalsgaard T, Devol AH, Ward BB, Ulloa O, et al. Oxygen distribution and aerobic respiration in the north and south eastern tropical Pacific oxygen minimum zones. Deep Sea Res Part I. 2014;94:173–83.
Church MJ, Lomas MW, Muller-Karger F. Sea change: charting the course for biogeochemical ocean time-series research in a new millennium. Deep Sea Res Part II. 2013;93:2–15.
Martiny AC, Tai APK, Veneziano D, Primeau F, Chisholm SW. Taxonomic resolution, ecotypes and the biogeography of Prochlorococcus. Environ Microbiol. 2009;11:823–32.
Saito MA, Bertrand EM, Duffy ME, Gaylord DA, Held NA, Hervey WJ. et al. Progress and challenges in ocean metaproteomics and proposed best practices for data sharing. J Proteome Res. 2019. https://doi.org/10.1021/acs.jproteome.8b00761.
Thiele S, Fuchs BM, Amann R, Iversen H. Colonization in the photic zone and subsequent changes during sinking determine bacterial community composition in marine snow. Appl Environ Microbiol. 2015;81:1463–71.
Huang S, Zhang S, Jiao N, Chen F. Marine cyanophages demonstrate biogeographic patterns throughout the global ocean. Appl Environ Microbiol. 2015;81:441–52.
Shibata A, Kogure K, Koike I, Ohwada K. Formation of submicron colloidal particles from marine bacteria by viral infection. Mar Ecol Prog Ser. 1997;155:303–7.
Weinbauer MG, Bettarel Y, Cattaneo R, Luef B, Maier C, Motegi C, et al. Viral ecology of organic and inorganic particles in aquatic systems: avenues for further research. Aquat Microb Ecol. 2009;57:321–41.
Lindell D, Jaffe JD, Coleman ML, Futschik ME, Axmann IM, Rector T, et al. Genome-wide expression dynamics of a marine virus and host reveal features of co-evolution. Nature. 2007;449:83–6.
Sullivan MB, Lindell D, Lee JA, Thompson LR, Bielawski JP, Chisholm SW. Prevalence and evolution of core photosystem II genes in marine cyanobacterial viruses and their hosts. PLoS Biol. 2006;4:e234.
Bristow LA, Dalsgaard T, Tiano L, Mills DB, Bertagnolli AD, Wright JJ, et al. Ammonium and nitrite oxidation at nanomolar oxygen concentrations in oxygen minimum zone waters. Proc Natl Acad Sci USA. 2016;113:10601–6.
Peng X, Fuchsman CA, Jayakumar A, Oleynik S, Martens-habbena W. Ammonia and nitrite oxidation in the Eastern Tropical North Pacific. Glob Biogeochem Cycles. 2015;29:2034–49.
Babbin AR, Bianchi D, Jayakumar A, Ward BB. Rapid nitrous oxide cycling in the suboxic ocean. Science. 2015;348:1127–9.
Fuchsman CA, Devol AH, Chase Z, Reimers CE, Hales B. Benthic fluxes on the Oregon shelf. Estuar Coast Shelf Sci. 2015;163:156–66.
Emerson S. Annual net community production and biological carbon flux in the ocean. Glob Biogeochem Cycles. 2014;28:14–28.
DeVries T, Deutsch C, Primeau F, Chang B, Devol A. Global rates of water-column denitrification derived from nitrogen gas measurements. Nat Geosci. 2012;5:547–50.
Galan A, Molina V, Thamdrup B, Woebken D, Lavik G, Kuypers MMM, et al. Anammox bacteria and the anaerobic oxidation of ammonium in the oxygen minimum zone off northern Chile. Deep Sea Res Part II. 2009;56:1021–31.
Trimmer M, Chronopoulou P-M, Maanoja ST, Upstill-Goddard RC, Kitidis V, Purdy KJ. Nitrous oxide as a function of oxygen and archaeal gene abundance in the North Pacific. Nat Commun. 2016;7:13451.
Acknowledgements
We thank the Keil lab sediment trap team for sediment trap deployments, captain and crew of the R/V Thompson, Peter Bernhardt for assistance with 13C net productivity rates, and Johnny Stutsmann for assistance with TOI analyses. We also thank David Garcia Prieto for assistance with the metagenomic read placement analyses. This work is supported in part by the University of Washington’s Proteomics Resource (UWPR95794). This study was funded by NSF grants OCE-1046017 to A.H.D., OCE-1138368 to G.R., DEB-1542240 to G.R., R.G.K., and A.H.D., and OCE-1230051 and OCE-1155566 to M.R.M.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fuchsman, C.A., Palevsky, H.I., Widner, B. et al. Cyanobacteria and cyanophage contributions to carbon and nitrogen cycling in an oligotrophic oxygen-deficient zone. ISME J 13, 2714–2726 (2019). https://doi.org/10.1038/s41396-019-0452-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0452-6
This article is cited by
-
Viruses under the Antarctic Ice Shelf are active and potentially involved in global nutrient cycles
Nature Communications (2023)
-
Partitioning of the denitrification pathway and other nitrite metabolisms within global oxygen deficient zones
ISME Communications (2023)
-
The marine nitrogen cycle: new developments and global change
Nature Reviews Microbiology (2022)
-
Anoxic chlorophyll maximum enhances local organic matter remineralization and nitrogen loss in Lake Tanganyika
Nature Communications (2021)
-
Stable aerobic and anaerobic coexistence in anoxic marine zones
The ISME Journal (2020)