Abstract
Most aerobic bacteria exist in dormant states within natural environments. In these states, they endure adverse environmental conditions such as nutrient starvation by decreasing metabolic expenditure and using alternative energy sources. In this study, we investigated the energy sources that support persistence of two aerobic thermophilic strains of the environmentally widespread but understudied phylum Chloroflexi. A transcriptome study revealed that Thermomicrobium roseum (class Chloroflexia) extensively remodels its respiratory chain upon entry into stationary phase due to nutrient limitation. Whereas primary dehydrogenases associated with heterotrophic respiration were downregulated, putative operons encoding enzymes involved in molecular hydrogen (H2), carbon monoxide (CO), and sulfur compound oxidation were significantly upregulated. Gas chromatography and microsensor experiments showed that T. roseum aerobically respires H2 and CO at a range of environmentally relevant concentrations to sub-atmospheric levels. Phylogenetic analysis suggests that the hydrogenases and carbon monoxide dehydrogenases mediating these processes are widely distributed in Chloroflexi genomes and have probably been horizontally acquired on more than one occasion. Consistently, we confirmed that the sporulating isolate Thermogemmatispora sp. T81 (class Ktedonobacteria) also oxidises atmospheric H2 and CO during persistence, though further studies are required to determine if these findings extend to mesophilic strains. This study provides axenic culture evidence that atmospheric CO supports bacterial persistence and reports the third phylum, following Actinobacteria and Acidobacteria, to be experimentally shown to mediate the biogeochemically and ecologically important process of atmospheric H2 oxidation. This adds to the growing body of evidence that atmospheric trace gases are dependable energy sources for bacterial persistence.
Similar content being viewed by others
Introduction
Bacteria from the phylum Chloroflexi are widespread and abundant in free-living microbial communities [1,2,3,4]. One reason for their success is their metabolic diversity; cultured strains from the phylum include heterotrophs, lithotrophs, and phototrophs adapted to both oxic and anoxic environments [5]. Cultured representatives of the phylum are classified into four classes by the genome taxonomy database [6], the primarily aerobic Chloroflexia and Ktedonobacteria and the anaerobic Anaerolineae and Dehalococcoidia [5]. Studies have provided insight into the metabolic strategies that anaerobic classes within the phylum use to adapt to oligotrophic niches [7, 8]. However, surprisingly little is known about how aerobic heterotrophic Chloroflexi colonise oxic environments. Global surveys have reported that Chloroflexi comprise 4.3% of soil bacteria [2] and 3.2% of marine bacteria [3]. However, the most dominant lineages within these ecosystems (notably Ellin6529 and SAR202) have not been cultivated [2, 6, 9]. Instead, most of our knowledge about the ecophysiological strategies of aerobic heterotrophic Chloroflexi is derived from studies on thermophilic isolates. Various strains from the classes Chloroflexia and Ktedonobacteria have been isolated and characterised from hot springs and geothermal soils [10,11,12,13,14].
Within geothermal environments, Chloroflexi strains are likely to encounter temporal and spatial variations in the availability of organic carbon compounds and other nutrients [15]. It is currently unknown how members of this phylum stay energised in response to these environmental perturbations. Carbon monoxide (CO) and molecular hydrogen (H2) of both geothermal and atmospheric origin are available in such environments and may be particularly important energy sources for sustaining growth and persistence [16,17,18,19,20]. Consistently, genomic, and metagenomic studies have revealed that Chloroflexi encode carbon monoxide dehydrogenases [21,22,23] and hydrogenases [21, 24,25,26] known to mediate aerobic respiration of these gases. Chloroflexi isolates have been shown to aerobically oxidise CO at a range of environmentally significant concentrations: Thermomicrobium roseum consumes high concentrations of CO when available during growth [21] and multiple Thermogemmatispora isolates have been shown to oxidise CO, including T. carboxidovorans to atmospheric concentrations (0.10 ppmv) [12]. While H2 oxidation has yet to be reported in aerobic heterotrophic Chloroflexi, strains of the phylum are known to encode the high-affinity group 1h [NiFe]-hydrogenase [24, 27]. This enzyme class has been shown to support bacterial persistence by mediating oxidation of atmospheric H2 (0.53 ppmv) [26, 28,29,30,31,32,33,34,35,36]. To date, atmospheric H2 oxidation has only been experimentally confirmed in Actinobacteria [28, 30,31,32, 34, 37] and two acidobacterial isolates [33, 38].
In this study, we investigated the persistence strategies of thermophilic isolates from two classes of Chloroflexi. We focused primarily on Thermomicrobium roseum (class Chloroflexia, formerly class Thermomicrobia [6]), a strain originally isolated from Toadstool Spring of Yellowstone National Park, USA [10]. This obligately aerobic bacterium is known to grow heterotrophically on a variety of carbohydrates, organic acids, and proteinaceous substrates [10, 13, 21]. Previous analyses have shown T. roseum encodes a type I carbon monoxide dehydrogenase and a group 1h [NiFe]-hydrogenase [21, 24], and can oxidise high concentrations of CO. However, the bacterium appears to be incapable of supporting chemolithoautotrophic growth and lacks key enzymes for the Calvin-Benson cycle [14, 21]. A combination of transcriptome sequencing and targeted activity assays were used to holistically determine the metabolic basis of persistence in this organism, including demonstrating that CO and H2 are oxidised by this strain during nutrient limitation. To help generalise these findings, we also investigated Thermogemmatispora sp. T81 (class Ktedonobacteria), a cellulolytic thermophilic strain which we previously isolated from geothermal soils in Tikitere, New Zealand [11, 39, 40]. Collectively, our results demonstrate that atmospheric H2 and CO serve as important energy sources that support the persistence of members of this phylum.
Materials and methods
Bacterial strains
Thermomicrobium roseum DSM 5159 [10, 13] and Thermogemmatispora sp. T81 [11, 39] were imported from the Extremophiles Research Group (GNS Science, Wairakei, New Zealand) culture collection in February 2017. Cultures of both bacterial isolates were routinely maintained in 120 mL serum vials sealed with treated lab-grade butyl rubber stoppers. Cultures of T. roseum contained 30 mL Castenholz media [41] supplemented with 1 g L−1 yeast extract and 1 g L−1 tryptone, whereas Thermogemmatispora sp. T81 cultures were maintained in 30 mL 10% R2A media [42]. Unless otherwise stated, both strains were incubated at 60 °C at an agitation speed of 150 rpm in an Eppendorf 40 Incubator. For T. roseum, cultures were inoculated to an OD600 of 0.03 and cells entered stationary phase within 48 h (ODmax = 0.75 to 1.0); growth curves confirmed that the strain entered stationary phase as a result of nutrient limitation, likely because of exhausting organic carbon supplies (Figure S1). For Thermogemmatispora sp. T81, 1 mL of stationary-phase cells were inoculated into 29 mL medium and cultures were incubated for 294 h before gas consumption assays were performed. Sporulation of Thermogemmatispora sp. T81 at this timepoint was verified by light microscopy of Gram-stained cultures. Gram staining and 16S rRNA gene amplicon sequencing also confirmed that both cultures were axenic.
Transcriptomics
Full transcriptome sequencing (RNA-Seq) was used to compare gene expression in T. roseum cultures under nutrient-rich (exponential phase; 10 mL, OD600 of 0.3) and nutrient-exhausted (stationary phase; 10 mL, OD600 of 0.75, 48 h post ODmax) conditions. Biological triplicate samples for each condition were harvested by centrifugation (21,000 × g, 15 min, 4 °C), the supernatants were removed, and the cell pellets were resuspended in 1 mL of RNAlater Stabilisation Solution (ThermoFisher Scientific) prior to freezing at −20 °C. Extraction and sequencing of RNA was performed by Macrogen Inc., Seoul, Korea. Briefly, RNA was extracted using the RNeasy Plant Mini Kit (Qiagen), libraries were constructed using a TruSeq RNA v2 Sample Prep Kit (Illumina), and rRNA was removed using the Ribo-Zero rRNA Removal Kit (Illumina). The resultant complementary DNA was sequenced on an Illumina HiSeq4000 platform using a paired-end, 100 bp high-throughput protocol. Sequence analysis was performed using the automated cloud-based transcriptomics pipeline, AIR (Sequentia Biotech). Briefly, the steps performed were read trimming, read quality analysis using FastQC [43], and read mapping against the T. roseum reference genome (NCBI ID: NC_011959.1 [21]) using an intrinsic platform read aligner with default parameters. Aligned reads were checked for quality prior to data normalisation using the trimmed mean of M-values method [44] and the ‘normalizaData’ command within the R package HTSFilter [45]. A principle component analysis was then performed on the normalised data prior to statistical analysis using edgeR [46] to obtain differential gene expression counts.
Gas chromatography
Gas chromatography measurements were performed to determine whether the two Chloroflexi strains could use atmospheric levels of CO and H2. Briefly, sealed serum vials containing stationary-phase cultures of T. roseum (72 hr post ODmax / 120 h post-inoculation) and sporulating cultures of Thermogemmatispora sp. T81 (294 h post-inoculation) were opened, equilibrated with ambient air (1 h), and resealed. These vials were then amended with H2 (via 1% v/v H2 in N2 gas cylinder, 99.999% pure) or CO (via 1% v/v CO in N2 gas cylinder, 99.999% pure) to achieve headspace concentrations of ~14 ppmv for each gas. The first headspace samples were collected within minutes after closure in order to measure the initial gas concentrations. The vials were maintained at the growth temperature (60 °C) and agitated (150 rpm) for the entire incubation period (75 h) to enhance H2, CO, and O2 transfer to the cultures. Six to nine headspace samples (1 mL) were collected at different time intervals using a gas-tight syringe to measure H2 and CO. Concomitantly, headspace gas concentrations in heat-killed negative controls (autoclaved; 30 mL) were measured to confirm that observed rates of gas consumption occurred due to a biotic process. Headspace gas concentrations were determined by gas chromatography using a pulsed discharge helium ionization detector [47, 48]. This customized trace gas analyser (model TGA-6791-W-4U-2, Valco Instruments Company Inc.) is designed to analyse a suite of atmospheric gases across six orders of magnitude of concentrations. Briefly, the system is configured to use two valves as injectors/backflushers, and two valves to front flush or heart cut from the precolumns (Mole Sieve 5A, set at 140 °C). Gases are then separated on the main columns (5′ X 1/8” HayseSep Db, set at 55 °C). The fifth valve is used as a sample loop selector to accommodate a larger range of gas concentrations. Concentrations of H2 and CO in each sample were regularly calibrated against ultra-pure H2 and CO gas standards of known concentrations. With the standards used, the limit of detection was 42 ppbv H2 and 9 ppbv CO.
Kinetic measurements
The whole-cell kinetic parameters of H2 and CO oxidation in T. roseum were measured by comparing rates of gas consumption at different substrate concentrations. Briefly, the headspace of stationary-phase cultures (72 h post ODmax) were amended with 100, 1000, or 4000 ppmv H2 or CO; whereas 100 ppmv mixing ratios were attained as described above, 100% H2 and 100% CO cylinders (99.999% pure) were used to attain mixing ratios of 1000 and 4000 ppmv. Cultures were incubated at 60 °C at an agitation speed of 300 rpm in an Eppendorf 40 Incubator. Headspace gas samples were measured at various time intervals (0, 0.5, 1, 2, 3, 4, and 5 h after substrate addition) by gas chromatography as described above. Reaction velocity relative to the gas concentration was measured at each timepoint and plotted on a Michaelis–Menten graph. Curves of best fit, Vmax app values, and Km app values were initially calculated in GraphPad Prism (version 7.01) using non-linear regression models (enzyme kinetics – substrate vs. velocity, Michaelis–Menten, least squares fit). Michaelis–Menten parameters were also derived using linear regressions based on Lineweaver-Burk, Eadie-Hofstee, and Hanes-Woolf plots [49].
Activity staining
Hydrogenase and carbon monoxide dehydrogenase activity was stained using whole-cell lysates of stationary-phase cultures of T. roseum. Five hundred mL of culture was harvested by centrifugation (10,000 × g, 10 min, 4 °C), washed in phosphate-buffered saline solution (PBS; 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4 and 2 mM KH2PO4, pH 7.4), and resuspended in 16 mL lysis buffer (50 mM Tris-Cl, pH 8.0, 1 mM PMSF, 2 mM MgCl2, 5 mg ml−1 lysozyme, 40 µg ml−1 DNase, 10% glycerol). The resultant suspension was then lysed by passage through a Constant Systems cell disruptor (40,000 psi, four times), with unbroken cells removed by centrifugation (10,000 × g, 20 min, 4 °C). Protein concentration was calculated using the bicinchoninic acid assay [50] against bovine serum albumin standards. Next, 20 µg protein was loaded onto a native 7.5% (w/v) Bis-Tris polyacrylamide gel prepared as described elsewhere [51] and run alongside a protein standard (NativeMark Unstained Protein Standard, Thermo Fisher Scientific) at 25 mA for 1.5 h. The gel was cut into three sections that were stained either for total protein, hydrogenase activity, or carbon monoxide dehydrogenase activity. For total protein staining, the gel section was incubated in AcquaStain Protein Gel Stain (Bulldog Bio) at room temperature for 3 h. For hydrogenase staining [28], the gel section was incubated in 50 mM potassium phosphate buffer (pH 7.0) supplemented with 500 µM nitroblue tetrazolium chloride (NBT) in an anaerobic jar (5% H2, 10% CO2, 85% N2 v/v) at 60 °C for 1 h. For carbon monoxide dehydrogenase staining [52], the gel section was incubated in 50 mM Tris-HCl buffer (pH 7.5) containing 50 µM NBT and 100 µM phenazine methosulfate in an anaerobic jar (100% CO v/v atmosphere) maintained at 60 °C for 1 h.
Electrode measurements
For T. roseum cultures, rates of H2 oxidation with and without treatment of respiratory chain uncouplers were measured amperometrically, following previously established protocols [53, 54]. Prior to the start of measurement, a Unisense H2 microsensor electrode was polarised at +800 mV for 1 h using a Unisense multimeter and calibrated with standards of known H2 concentration. Gas-saturated PBS was prepared by bubbling the solution with 100% (v/v) of either H2 or O2 for 5 min. For untreated cells, 1.1 mL microrespiration assay chambers were sequentially amended with stationary-phase T. roseum cell suspensions (OD600 = 1; 0.9 mL), H2-saturated PBS (0.1 mL), and O2-saturated PBS (0.1 mL) stirred at 250 rpm, 37 °C. Following measurements of untreated cells, the assay mixtures were treated with 100 µM carbonyl cyanide m-chlorophenyl hydrazine (CCCP), 10 µM nigericin, or 10 µM valinomycin. Changes in H2 concentrations were recorded using Unisense Logger Software. Upon observing a linear change in H2 concentration, initial rates of consumption were calculated over a period of 20 s and normalised against total protein concentration.
Phylogenetic analyses
Phylogenetic trees were constructed to investigate the evolutionary history and distribution of uptake hydrogenases and carbon monoxide dehydrogenases within the Chloroflexi phylum. Specifically, the catalytic subunits of [NiFe]-hydrogenases (HhyL and homologues) and type I carbon monoxide dehydrogenases (CoxL) were retrieved from Chloroflexi genomes and metagenome-assembled genomes (MAGs) in the NCBI RefSeq database via protein BLAST [55] in October 2018. Using MEGA7 [56], the amino acid sequences were aligned with reference sequences [23, 24] with ClustalW and evolutionary relationships were visualised by constructing a maximum-likelihood phylogenetic tree; specifically, initial trees for the heuristic search were obtained automatically by applying Neighbour-Join and BioNJ algorithms to a matrix of pairwise distances estimated using a JTT model, and then selecting the topology with superior log likelihood value. Gaps were treated with partial deletion and trees were bootstrapped with 100 replicates.
Results and discussion
Thermomicrobium roseum upregulates hydrogenase and carbon monoxide dehydrogenase expression during a coordinated response to nutrient starvation
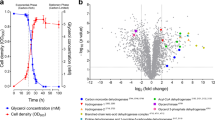
We compared the transcriptomes of triplicate T. roseum cultures under nutrient-rich (exponential growing) and nutrient-limited (stationary phase) conditions. A total of 401 genes were significantly upregulated and 539 genes were significantly downregulated by at least two-fold (p < 10−6) in response to nutrient limitation (Fig. 1a; Table S1). Three major trends were observed with respect to energy acquisition and utilisation. Firstly, genes associated with energetically expensive processes were downregulated, including those encoding ribosomal proteins, cytochrome c and menaquinone biosynthesis enzymes, and the megaplasmid-encoded chemotactic and flagellar apparatus (Table S1). Secondly, there was evidence of mobilisation of internal carbon stores, including an acetoin dehydrogenase complex and an electron transfer flavoprotein complex (ETF). Thirdly, the expression profiles indicate there is extensive remodelling of the respiratory chain. Two primary respiratory dehydrogenases involved in heterotrophic growth (type I and II NADH dehydrogenases) were downregulated, whereas complexes involved in lithotrophic energy generation and a succinate dehydrogenase were upregulated (Fig. 1a; Table S1). In both conditions, the terminal oxidases that mediate aerobic respiration were highly expressed and there was no evidence of the use of other electron acceptors; the cytochrome aa3 oxidase was expressed in both phases and the alternative cytochrome bo3 oxidase was upregulated during stationary phase. In contrast, the F1Fo-ATPase (ATP synthase) was downregulated, a finding consistent with an expected decrease in the availability of respiratory electron donors during nutrient limitation (Table S1).
Differential gene expression of nutrient-rich (exponential phase) and nutrient-limited (stationary phase) cultures of Thermomicrobium roseum. a Volcano plot showing relative expression change of genes following nutrient limitation. The fold-change shows the ratio of normalised transcript abundance of three stationary phase cultures divided by three exponential phase cultures (biological replicates). Each gene is represented by a grey dot and respiratory genes are highlighted as per the legend. b, c Heat maps of normalised abundance of the putative operons encoding the structural subunits of the group 1h [NiFe]-hydrogenase (hhyLS; b) and type I carbon monoxide dehydrogenase (coxLSM; c). The read counts per kilobase million (RPKM) are shown for three exponentially growing and three stationary phase biological replicates. HP = hypothetical protein. d Differential regulation of the respiratory complexes mediating aerobic respiration of organic and inorganic compounds. Complexes are differentially shaded depending on whether they are significantly upregulated (green), downregulated (orange), or unchanged (grey) in nutrient-limited compared to nutrient-rich cultures. Gene names, loci numbers, and average fold changes in transcriptome abundance are shown for each complex. Shown are the structural subunits of type I NADH dehydrogenase (nuoA-E,H-N), type II NADH dehydrogenase (ndh), succinate dehydrogenase (sdhA-D), group 1h [NiFe]-hydrogenase (hhyLS), type I carbon monoxide dehydrogenase (coxLMS), heterodisulfide reductase (hdrABC), electron transfer flavoprotein (etfAB), sulfur-carrier protein (tusA), cytochrome aa3 oxidase (coxABC), cytochrome bo3 oxidase (cyoAB), and ATP synthase (atpA-H). Note that the physiological role of the highly upregulated hdrABC, etfAB, and tusA genes is yet to be experimentally validated in T. roseum
Thermomicrobium roseum upregulates genes associated with H2 and CO metabolism under nutrient-limiting conditions. The genes encoding the structural subunits of a group 1h [NiFe]-hydrogenase (hhyLS; trd_1878–1877) [24, 25, 57], which are a class of oxygen-tolerant enzymes known to mediate atmospheric H2 oxidation [28, 31, 33, 58, 59], were upregulated by an average of 12.6-fold (Fig. 1b). Also upregulated were the conserved hypothetical proteins hhaABC (trd_1876–1874; 5.5-fold) [27], encoded on the same putative operon as the structural subunits, as well as a separate putative operon of maturation factors (trd_1873–1863; 3.1-fold) (Figure S2; Table S1). The structural (trd_1206–1208) and maturation (trd_1209–1215) subunits encoding a type I carbon monoxide dehydrogenase were upregulated by an average of two-fold (Fig. 1c & S2) in response to nutrient limitation. Consistent with previous reports of CO utilisation during growth in this organism [21], carbon monoxide dehydrogenase genes were highly expressed in both exponential and stationary-phase cultures. (Fig. 1c; Table S1). This suggests that T. roseum uses CO to supplement available organic carbon during growth (mixotrophy) and persistence. These findings are broadly similar to observations made in other phyla, notably Actinobacteria and Proteobacteria, that hydrogenase and carbon monoxide dehydrogenase expression are induced by organic carbon limitation [28, 31, 60,61,62,63,64].
Overall, the greatest differential in gene expression involved a 19-gene cluster (trd_0160–0142) putatively involved with the oxidation of sulfur compounds. The cluster contains gene encoding a putative soluble heterodisulfide reductase (hdrABC), an electron transfer flavoprotein complex (etfAB), three sulfur-carrier proteins (tusA, dsrE1, dsrE2), three lipoate-binding proteins (lbpA), and various hypothetical proteins, which are upregulated by an average of 45-fold during persistence. Most of these components have homologues in a system recently shown to mediate the oxidation of diverse organic and inorganic sulfur compounds in Hyphomicrobium denitrificans [65, 66]. One role of this cluster may be to mediate the activation and oxidation of endogenous or exogenous thiol-containing compounds. To achieve this, we predict that the Hdr complex catalyses disulfide bond formation between the thiol compound and a sulfur-carrier protein (e.g., TusA); the Hdr complex then transfers the liberated electrons into the respiratory chain, possibly via the ETF complex. Supporting this notion, thiol oxidation to disulfide is exergonic with oxygen as the terminal electron acceptor. While Hdr complexes are best-characterized for their roles in heterodisulfide reduction in methanogenic archaea [67], they have also been studied in sulfur-oxidizing and sulfate-reducing bacteria, where they have been predicted to be physiologically reversible [68, 69]. Consistently, the Hdr complex of T. roseum is most closely related to those of sulfur-oxidising Sulfobacillus, Hyphomicrobium, and Acidithiobacillus strains [65, 70, 71]. It seems plausible that T. roseum would benefit from a survival advantage if it can harness reduced sulfur compounds available in geothermal springs. However, further work is needed to verify the activity, substrates, and physiological role of this system.
Collectively, these findings show that T. roseum is more metabolically flexible than previously thought. Fig. 1d illustrates the predicted remodelling of the respiratory chain that occurs during the transition from nutrient-rich to nutrient-limited conditions. The upregulation of enzymes involved in harnessing inorganic compounds, in conjunction with the downregulation of gene clusters involved in NADH oxidation, suggests that T. roseum has evolved mechanisms to maintain aerobic respiration despite nutrient fluctuations and deprivation within its environment.
T. roseum aerobically oxidises H2 and CO at a wide range of concentrations, including sub-atmospheric levels, during persistence
The high expression levels for genes encoding the group 1h [NiFe]-hydrogenase and type I carbon monoxide dehydrogenase suggested that T. roseum may support persistence by oxidising atmospheric H2 and CO. To test this, we incubated nutrient-limited cultures of T. roseum in an ambient air headspace supplemented with ~14 ppmv of either H2 or CO and monitored their consumption using gas chromatography. In agreement with our hypothesis, cultures aerobically oxidised both gases in a first-order kinetic process; within 71 h, mixing ratios of these gases (103 ppbv H2, 22 ppbv CO) were five times below atmospheric levels (Fig. 2a, b). This constitutes the first observation of both aerobic H2 respiration and atmospheric H2 oxidation within the phylum Chloroflexi.
Hydrogenase and carbon monoxide dehydrogenase activity of Thermomicrobium roseum cultures during nutrient limitation. a, b Oxidation of molecular hydrogen (H2; a) and carbon monoxide (CO; b) to sub-atmospheric levels by T. roseum cultures. Error bars show standard deviations of three biological replicates, with heat-killed cells monitored as a negative control (grey dashed lines). Mixing ratios of H2 and CO are displayed on a logarithmic scale and dotted lines show the average atmospheric mixing ratios of H2 (0.53 ppmv) and CO (0.10 ppmv). c, d Apparent kinetic parameters of H2 (c) and CO (d) oxidation by T. roseum whole cells. Curves of best fit and kinetic parameters were calculated based on a Michaelis–Menten non-linear regression model. Values calculated based on Lineweaver-Burk, Hanes-Woolf, and Eadie-Hofstee plots are shown in Table S2. e Zymographic observation of hydrogenase and carbon monoxide dehydrogenase activity in T. roseum whole-cell lysates. The first two lanes show protein ladder and whole protein stained with Coomassie Blue. The third and fourth lanes show hydrogenase and carbon monoxide dehydrogenase activity stained with the artificial electron acceptor nitroblue tetrazolium in a H2-rich and CO-rich atmosphere respectively. f Amperometric measurements of hydrogenase activity in T. roseum whole cells. The rate of H2 oxidation was measured with a hydrogen electrode before and after treatment with the respiratory uncouplers and ionophores carbonyl cyanide m-chlorophenyl hydrazine (CCCP), nigericin, and valinomycin
Whole-cell kinetic measurements revealed that T. roseum efficiently oxidises H2 and CO across a wide range of concentrations through hydrogenase and carbon monoxide dehydrogenase activity. In cultures, the enzymes display a moderate apparent velocity (Vmax app of 376 nmol H2 and 149 nmol CO g−1 of protein min−1) and moderate apparent affinity (Km app of 569 nM H2 and 285 nM CO) for these substrates (Fig. 2c, d; Table S2). With respect to carbon monoxide dehydrogenase, these observations are consistent with the organism being able to utilise CO at elevated concentrations for growth [21] and atmospheric concentrations for persistence. The apparent kinetic parameters of the group 1h [NiFe]-hydrogenase are more similar to those recently described for the verrucomicrobial methanotroph Methylacidiphilum fumariolicum (Km = 600 nM) [72] than to the high-affinity, low-activity hydrogenases of previously described atmospheric H2 scavengers (Km < 50 nM) [28, 31, 33]. Altogether, these findings suggest that T. roseum can take advantage of the elevated H2 and CO concentrations when available through geothermal activity and subsist on atmospheric concentrations of these gases otherwise.
Consistent with the observed whole-cell activities, cell-lysates run on native polyacrylamide gels strongly stained for hydrogenase and carbon monoxide dehydrogenase activity (Fig. 2e). The molecular weight of the major bands were, respectively, at the expected molecular weight for a carbon monoxide dehydrogenase dimer [266 kDa, (CoxLMS)2] and slightly below the expected molecular weight of a hydrogenase dimer [210 kDa, (HhyLS)2]. This is compatible with biochemical studies in other organisms that have shown type I carbon monoxide dehydrogenases and group 1h [NiFe]-hydrogenases form homodimers [58, 59, 73]. We next verified that the hydrogenase was coupled to the respiratory chain by measuring H2 oxidation using a H2 electrode under aerobic conditions. Untreated cells oxidised H2 at a rapid rate. This activity decreased by 2.5-fold upon addition of the respiratory uncoupler CCCP and ceased upon addition of the ionophore valinomycin, whereas no significant change in H2 oxidation rate was observable with the protonophore nigericin (Fig. 2f). The combination of these results suggests that the oxidation of hydrogen is tightly coupled to the respiratory chain and this interaction may be linked to the electrical gradient (Δψ), but not pH gradient (ΔpH), of the membrane.
Findings from the transcriptome analysis and activity studies therefore suggest that T. roseum persists through oxidation of atmospheric H2 and CO. We propose that the group 1h [NiFe]-hydrogenase and type I carbon monoxide dehydrogenase directly use electrons derived from atmospheric H2 and CO to support aerobic respiration (Fig. 1d). It is probable that these electrons are relayed via electron carriers into the menaquinone pool and are subsequently transferred to the terminal oxidases. However, further studies are needed to confirm how these proteins functionally and physically interact with the respiratory chain, including their localisation and which electron carriers they interact with. Due to the genetic intractability of Chloroflexi and the lack of specific hydrogenase or carbon monoxide dehydrogenase inhibitors, we were also unable to determine the necessity of either H2 or CO oxidation for prolonged survival for this organism. However, previous studies have demonstrated that genetic deletion of the group 1h [NiFe]-hydrogenase reduces longevity of M. smegmatis cells [29, 30, 60] and Streptomyces avermitilis exospores [31, 32].
Scavenging of atmospheric gases is potentially a common persistence strategy within the aerobic heterotrophic Chloroflexi
Having demonstrated that T. roseum oxidises atmospheric trace gases during persistence, we subsequently investigated whether this is a common strategy employed by the Chloroflexi. We first analysed the respiratory capabilities of Thermogemmatispora sp. T81, a heterotrophic cellulolytic and sporulating thermophile, which we previously isolated from geothermal soils from Tikitere, New Zealand [11, 40]. Analysis of the organism’s genome (Assembly ID: GCA_003268475.1) indicated that it encodes core respiratory chain components similar to T. roseum, including primary dehydrogenases (nuo, ndh, sdh), terminal oxidases (cox, cyo), and ATP synthase (atp). The genome also encodes putative operons for the structural subunits of a group 1h [NiFe]-hydrogenase, the maturation factors of this hydrogenase, and structural subunits of a type I carbon monoxide dehydrogenase (Figure S3). However, homologues of the putative heterodisulfide reductase and ETF complexes encoded by T. roseum are absent from the Thermogemmatispora sp. T81 genome.
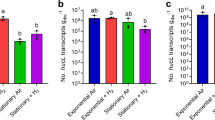
We verified that sporulating cultures of Thermogemmatispora sp. T81 actively consume H2 and CO. The organism slowly oxidised available H2 and CO in the headspace to sub-atmospheric levels (120 ppbv H2, 70 ppbv CO) over ~320 h (Fig. 3a, b). Although this strain has previously been shown to oxidise carbon monoxide [12], this is the first observation that it can do so to sub-atmospheric concentrations and during persistence. These results suggest that, despite their distinct evolutionary histories and ecological niches, Thermogemmatispora sp. T81 and T. roseum have both evolved similar metabolic strategies to survive nutrient limitation.
Hydrogenase and carbon monoxide dehydrogenase activity of Thermogemmatispora sp. T81 during sporulation. Oxidation of molecular hydrogen (H2; a) and carbon monoxide (CO; b) to sub-atmospheric levels by Thermogemmatispora sp. T81 cultures. Error bars show standard deviations of three biological replicates, with heat-killed cells monitored as a negative control (grey dashed lines). Mixing ratios of H2 and CO are displayed on a logarithmic scale and dotted lines show the average atmospheric mixing ratios of H2 (0.53 ppmv) and CO (0.10 ppmv)
Analysis of the distribution of hydrogenases and carbon monoxide dehydrogenases within publicly available reference genomes showed that genetic capacity for trace gas scavenging is a common trait among aerobic Chloroflexi. Specifically, group 1h [NiFe]-hydrogenases and type I carbon monoxide dehydrogenases were encoded in three of the four reference genomes within the Thermomicrobiales (class Chloroflexia) and four of the five reference genomes within the Ktedonobacteriales (class Ktedonobacteria) (Fig. 4a, b). The latter includes the genomes of the heterotrophic soil bacterium Ktedonobacter racemifer [74] and the nitrite-oxidising bioreactor isolate Nitrolancea hollandica [75]. In addition, seven strains within the photosynthetic order Chloroflexales encoded group 1f and/or group 2a [NiFe]-hydrogenases (Figure S4). These hydrogenase classes have been shown to mediate aerobic H2 oxidation in a range of bacteria, including to sub-atmospheric concentrations in Acidobacterium ailaaui and M. smegmatis respectively [28, 38]. Moreover, a metatranscriptome study revealed that homologs of the group 1f [NiFe]-hydrogenase of Roseiflexus species are highly expressed in geothermal microbial mats at night [76]. Hence, it is likely that the traits of aerobic H2 respiration and possibly atmospheric H2 oxidation extends to the photosynthetic strains of this phylum. A range of metagenome-assembled genomes, including from the abundant candidate class Ellin6529 [2, 26], also encoded genes for aerobic H2 and CO oxidation (Figure S4 & S5). Consistent with previous reports, Dehalococcoidia encode group 1a [NiFe]-hydrogenases known to facilitate dehalorespiration [77,78,79].
Evolutionary history of the group 1h [NiFe]-hydrogenase and type I carbon monoxide dehydrogenase. Phylogenetic trees showing the distribution and evolutionary history of the catalytic (large) subunits of the group 1h [NiFe]-hydrogenase (hhyL; a) and type I carbon monoxide dehydrogenase (coxL; b) in the phylum Chloroflexi. Chloroflexi sequences (labelled by class) are shown in bold against reference sequences (labelled by phylum). Trees were constructed using amino acid sequences through the maximum-likelihood method (gaps treated with partial deletion) and were bootstrapped with 100 replicates. The trees were respectively rooted with group 1g [NiFe]-hydrogenase sequences (WP_011761956.1, WP_048100713.1) and type II carbon monoxide dehydrogenase sequences (WP_011388721.1, WP_012893108.1). The distribution of other respiratory uptake hydrogenases within genomes and metagenome-assembled genomes (MAGs) in the phylum Chloroflexi is shown in Figure S4. The distribution of type I carbon monoxide dehydrogenases within metagenome-assembled genomes (MAGs) in the phylum Chloroflexi is shown in Figure S5
Our analyses suggest that the capacity for atmospheric H2 and CO oxidation may have evolved on two or more occasions within the Chloroflexi. Phylogenetic trees show that the group 1h [NiFe]-hydrogenases from Chloroflexia and Ktedonobacteria are divergent and fall into two distinct, robustly supported branches (Fig. 4a). It is therefore more likely that Chloroflexia and Ktedonobacteria independently acquired these enzymes, for example as a result of horizontal gene transfer events from other Terrabacteria, rather than vertically inheriting them from a common ancestor. Phylogenetic analysis also suggests that the type I carbon monoxide dehydrogenase may have also been acquired on two or three occasions in this phylum (Fig. 4b). In line with their probable independent acquisition, the putative operons encoding the hydrogenase and carbon monoxide dehydrogenase in T. roseum (Figure S2) and Thermogemmatispora sp. T81 (Figure S3) are distinctly organized. For example, the structural and accessory factors of carbon monoxide dehydrogenase are encoded in a single putative operon in Thermogemmatispora sp. T81 (coxMSLIG), but are separated into a structural operon (coxGSLM) and accessory operon (including coxG and coxE) in T. roseum. These findings agree with previous inferences of horizontal dissemination of hhyL and coxL genes [23, 24, 27] and suggest there is strong selective pressure for the acquisition of metabolic enzymes that support persistence. However, other explanations for their observations cannot be ruled out and further analysis is required to unravel the complex evolutionary histories of hydrogenases and carbon monoxide dehydrogenases.
Ecological and biogeochemical significance of metabolic flexibility and trace gas oxidation in Chloroflexi
Aerobic heterotrophic bacteria from the phylum Chloroflexi are more metabolically versatile than previously thought. The transcriptome analyses clearly show that T. roseum regulates its metabolism in response to nutrient limitation, enabling persistence on a combination of exogenous inorganic compounds and likely endogenous carbon reserves. In support of this, gas chromatography measurements showed that the bacterium efficiently oxidises H2 and CO down to sub-atmospheric concentrations during persistence through an aerobic respiratory process. We made similar findings for the ktedonobacterial isolate Thermogemmatispora sp. T81, suggesting that trace gas scavenging might be a common persistence strategy employed by aerobic Chloroflexi. Analyses of primary sequence phylogeny and operon structure indicate that the group 1 h [NiFe]-hydrogenases and carbon monoxide dehydrogenases within these organisms fall into different clades and are relatively divergent. Hence, it is probable that these organisms have horizontally acquired the capacity to oxidise atmospheric H2 and CO via separate events, though other explanations are possible. The apparent convergence in persistence strategies is notable given the distinct evolutionary histories, persistence morphologies (i.e., sporulation in T81), and ecological niches of these bacteria. Resource generalism is therefore likely to be a common ecological strategy for the survival of Chloroflexi in environments where organic carbon and other nutrients may be periodically scarce.
More broadly, these findings provides pure culture support for the hypothesis that atmospheric carbon monoxide serves as an energy source for persistence [26]. Our findings suggest that the expression and activity of carbon monoxide dehydrogenase is linked to persistence, and provide evidence that atmospheric CO may serve as an electron donor for the aerobic respiratory chain in this condition. Indeed, as with atmospheric H2, atmospheric CO is likely to be a dependable energy source for microbial survival given its ubiquity, diffusibility, and energy density. Integrating these findings with the wider literature, it is probable that atmospheric CO oxidation is a general strategy supporting long-term survival of aerobic heterotrophic bacteria. Indeed, various heterotrophic bacteria have previously been inferred to be capable of oxidising atmospheric CO, including Proteobacteria [80,81,82,83], Actinobacteria [84, 85], and a Thermogemmatispora strain [12]. Moreover, other datasets have shown that carbon monoxide dehydrogenase expression is activated during nutrient limitation in other aerobic organisms [60,61,62,63,64]. However, in contrast to atmospheric H2 [29, 32], it remains to be validated through genetic and biochemical studies that atmospheric CO oxidation can enhance survival of bacteria during persistence. In line with previous activity-based measurements [21], the transcriptome analysis shows that T. roseum expresses carbon monoxide dehydrogenase at high levels during growth. Unlike carboxydotrophs such as Oligotropha carboxidovorans [62, 73, 86], T. roseum as a carboxydovore cannot grow chemolithoautotrophically [14, 21] and instead appears to use CO as an additional energy source during heterotrophic growth. The wide kinetic range of the T. roseum carbon monoxide dehydrogenase in whole cells likely enables this isolate to both persist on ubiquitously available atmospheric CO [87] and grow mixotrophically in microenvironments where CO is available at elevated concentrations (up to 6000 ppmv) through geothermal activity [17].
Finally, this study establishes Chloroflexi as the third phylum experimentally shown to scavenge atmospheric H2, following the Actinobacteria [28, 31, 34, 37, 57] and Acidobacteria [33, 38]. The findings made here are similar to those previously reported for the actinobacterium Mycobacterium smegmatis [28, 60] and acidobacterium Pyrinomonas methylaliphatogenes [33], both of which also shift from heterotrophic respiration to atmospheric H2 oxidation in response to energy limitation, including through expressing group 1h [NiFe]-hydrogenases. Given at least four other cultured phyla (Fig. 4a) and two candidate phyla [26] also encode group 1h [NiFe]-hydrogenases, it seems increasingly likely that atmospheric H2 serves as a general energy source for aerobic heterotrophic bacteria. This observation is also potentially biogeochemically significant, given aerobic soil bacteria are known to be the main sink in the global hydrogen cycle [88]. Further work, however, is needed to test these whether these principles extend to the still enigmatic Chloroflexi species inhabiting mesophilic soil environments.
References
Thompson LR, Sanders JG, McDonald D, Amir A, Ladau J, Locey KJ, et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature. 2017;551:457–63.
Delgado-Baquerizo M, Oliverio AM, Brewer TE, Benavent-González A, Eldridge DJ, Bardgett RD, et al. A global atlas of the dominant bacteria found in soil. Science. 2018;359:320–5.
Sunagawa S, Coelho LP, Chaffron S, Kultima JR, Labadie K, Salazar G, et al. Structure and function of the global ocean microbiome. Science. 2015;348:1261359.
Mehrshad M, Salcher MM, Okazaki Y, Nakano S, Šimek K, Andrei A-S, et al. Hidden in plain sight—highly abundant and diverse planktonic freshwater Chloroflexi. Microbiome. 2018;6:176.
Whitman WB. Bergey’s manual of systematics of Archaea and Bacteria. Wiley Online Library; New York, United States, 2015.
Parks DH, Chuvochina M, Waite DW, Rinke C, Skarshewski A, Chaumeil P-A, et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat Biotechnol. 2018;36:996–1004.
Hug LA, Castelle CJ, Wrighton KC, Thomas BC, Sharon I, Frischkorn KR, et al. Community genomic analyses constrain the distribution of metabolic traits across the Chloroflexi phylum and indicate roles in sediment carbon cycling. Microbiome. 2013;1:22.
Dong X, Greening C, Rattray JE, Chakraborty A, Chuvochina M, Mayumi D, et al. Potential for microbial anaerobic hydrocarbon degradation in naturally petroleum-associated deep-sea sediments. bioRxiv. 2018; 400804. https://www.biorxiv.org/content/10.1101/400804v2.
Mehrshad M, Rodriguez-Valera F, Amoozegar MA, López-García P, Ghai R. The enigmatic SAR202 cluster up close: shedding light on a globally distributed dark ocean lineage involved in sulfur cycling. ISME J. 2018;12:655.
Jackson TJ, Ramaley RF, Meinschein WG. Thermomicrobium, a new genus of extremely thermophilic bacteria. Int J Syst Evol Microbiol. 1973;23:28–36.
Stott MB, Crowe MA, Mountain BW, Smirnova AV, Hou S, Alam M, et al. Isolation of novel bacteria, including a candidate division, from geothermal soils in New Zealand. Environ Microbiol. 2008;10:2030–41.
King CE, King GM. Description of Thermogemmatispora carboxidivorans sp. nov., a carbon-monoxide-oxidizing member of the class Ktedonobacteria isolated from a geothermally heated biofilm, and analysis of carbon monoxide oxidation by members of the class Ktedonobacter. Int J Syst Evol Microbiol. 2014;64:1244–51.
Houghton KM, Morgan XC, Lagutin K, MacKenzie AD, Vyssotski M, Mitchell KA, et al. Thermorudis pharmacophila WKT50.2T sp. nov., a novel isolate of class Thermomicrobia isolated from geothermal soil. Int J Syst Evol Microbiol. 2015;65:4479–87.
King CE, King GM. Thermomicrobium carboxidum sp. nov., and Thermorudis peleae gen. nov., sp. nov., carbon monoxide-oxidizing bacteria isolated from geothermally heated biofilms. Int J Syst Evol Microbiol. 2014;64:2586–92.
Power JF, Carere CR, Lee CK, Wakerley GLJ, Evans DW, Button M, et al. Microbial biogeography of 925 geothermal springs in New Zealand. Nat Commun. 2018;9:2876.
Spear JR, Walker JJ, McCollom TM, Pace NR. Hydrogen and bioenergetics in the Yellowstone geothermal ecosystem. Proc Natl Acad Sci USA. 2005;102:2555–60.
Shock EL, Holland M, Amend JP, Osburn GR, Fischer TP. Quantifying inorganic sources of geochemical energy in hydrothermal ecosystems, Yellowstone National Park, USA. Geochim Cosmochim Acta. 2010;74:4005–43.
King GM, Weber CF. Interactions between bacterial carbon monoxide and hydrogen consumption and plant development on recent volcanic deposits. ISME J. 2008;2:195–203.
King GM. Contributions of atmospheric CO and hydrogen uptake to microbial dynamics on recent Hawaiian volcanic deposits. Appl Environ Microbiol. 2003;69:4067–75.
Yang J, Zhou E, Jiang H, Li W, Wu G, Huang L, et al. Distribution and diversity of aerobic carbon monoxide-oxidizing bacteria in geothermal springs of China, the Philippines, and the United States. Geomicrobiol J. 2015;32:903–13.
Wu D, Raymond J, Wu M, Chatterji S, Ren Q, Graham JE, et al. Complete genome sequence of the aerobic CO-oxidizing thermophile Thermomicrobium roseum. PLoS ONE 2009;4:e4207.
King GM, Weber CF. Distribution, diversity and ecology of aerobic CO-oxidizing bacteria. Nat Rev Microbiol. 2007;5:107–18.
Quiza L, Lalonde I, Guertin C, Constant P. Land-use influences the distribution and activity of high affinity CO-oxidizing bacteria Associated to type I-coxL genotype in soil. Front Microbiol. 2014;5:271.
Greening C, Biswas A, Carere CR, Jackson CJ, Taylor MC, Stott MB, et al. Genomic and metagenomic surveys of hydrogenase distribution indicate H2 is a widely utilised energy source for microbial growth and survival. ISME J. 2016;10:761–77.
Søndergaard D, Pedersen CNS, Greening C. HydDB: a web tool for hydrogenase classification and analysis. Sci Rep. 2016;6:34212.
Ji M, Greening C, Vanwonterghem I, Carere CR, Bay SK, Steen JA, et al. Atmospheric trace gases support primary production in Antarctic desert surface soil. Nature. 2017;552:400–3.
Greening C, Constant P, Hards K, Morales SE, Oakeshott JG, Russell RJ, et al. Atmospheric hydrogen scavenging: from enzymes to ecosystems. Appl Environ Microbiol. 2015;81:1190–9.
Greening C, Berney M, Hards K, Cook GM, Conrad R. A soil actinobacterium scavenges atmospheric H2 using two membrane-associated, oxygen-dependent [NiFe] hydrogenases. Proc Natl Acad Sci USA. 2014;111:4257–61.
Greening C, Villas-Bôas SG, Robson JR, Berney M, Cook GM. The growth and survival of Mycobacterium smegmatis is enhanced by co-metabolism of atmospheric H2. PLoS ONE 2014;9:e103034.
Berney M, Greening C, Conrad R, Jacobs WR, Cook GM. An obligately aerobic soil bacterium activates fermentative hydrogen production to survive reductive stress during hypoxia. Proc Natl Acad Sci USA. 2014;111:11479–84.
Constant P, Chowdhury SP, Pratscher J, Conrad R. Streptomycetes contributing to atmospheric molecular hydrogen soil uptake are widespread and encode a putative high-affinity [NiFe]-hydrogenase. Environ Microbiol. 2010;12:821–9.
Liot Q, Constant P. Breathing air to save energy—new insights into the ecophysiological role of high-affinity [NiFe]-hydrogenase in Streptomyces avermitilis. MicrobiologyOpen. 2016;5:47–59.
Greening C, Carere CR, Rushton-Green R, Harold LK, Hards K, Taylor MC, et al. Persistence of the dominant soil phylum Acidobacteria by trace gas scavenging. Proc Natl Acad Sci USA. 2015;112:10497–502.
Meredith LK, Rao D, Bosak T, Klepac-Ceraj V, Tada KR, Hansel CM, et al. Consumption of atmospheric hydrogen during the life cycle of soil-dwelling actinobacteria. Environ Microbiol Rep. 2014;6:226–38.
Kanno M, Constant P, Tamaki H, Kamagata Y. Detection and isolation of plant-associated bacteria scavenging atmospheric molecular hydrogen. Environ Microbiol. 2015;18:2495–506.
Khdhiri M, Hesse L, Popa ME, Quiza L, Lalonde I, Meredith LK, et al. Soil carbon content and relative abundance of high affinity H2-oxidizing bacteria predict atmospheric H2 soil uptake activity better than soil microbial community composition. Soil Biol Biochem. 2015;85:1–9.
Constant P, Poissant L, Villemur R. Isolation of Streptomyces sp. PCB7, the first microorganism demonstrating high-affinity uptake of tropospheric H2. ISME J. 2008;2:1066–76.
Myers MR, King GM. Isolation and characterization of Acidobacterium ailaaui sp. nov., a novel member of Acidobacteria subdivision 1, from a geothermally heated Hawaiian microbial mat. Int J Syst Evol Microbiol. 2016;66:5328–35.
Vyssotski M, Ryan J, Lagutin K, Wong H, Morgan X, Stott M. A novel fatty acid, 12,17-dimethyloctadecanoic acid, from the extremophile Thermogemmatispora sp. (Strain T81). Lipids. 2012;47:601–11.
Tomazini A, Lal S, Munir R, Stott M, Henrissat B, Polikarpov I, et al. Analysis of carbohydrate-active enzymes in Thermogemmatispora sp. strain T81 reveals carbohydrate degradation ability. Can J Microbiol. 2018. https://doi.org/10.1139/cjm-2018-0336.
Ramaley RF, Hixson J. Isolation of a nonpigmented, thermophilic bacterium similar to Thermus aquaticus. J Bacteriol. 1970;103:527.
Reasoner DJ, Geldreich EE. A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microbiol. 1985;49:1–7.
Andrews S. FastQC: a quality control tool for high throughput sequence data. 2010. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
Robinson MD, Oshlack A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010;11:R25.
Rau A, Gallopin M, Celeux G, Jaffrézic F. Data-based filtering for replicated high-throughput transcriptome sequencing experiments. Bioinformatics. 2013;29:2146–52.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Wentworth WE, Vasnin SV, Stearns SD, Meyer CJ. Pulsed discharge helium ionization detector. Chromatographia. 1992;34:219–25.
Novelli PC, Crotwell AM, Hall BD. Application of gas chromatography with a pulsed discharge helium ionization detector for measurements of molecular hydrogen in the atmosphere. Environ Sci Technol. 2009;43:2431–6.
Price NC, Dwek RA, Wormald M, Ratcliffe RG. Principles and Problems in Physical Chemistry for Biochemists. 3rd edn. Oxford: Oxford University Press; 2002.
Smith PK, Il KrohnR, Hermanson GT, Mallia AK, Gartner FH, Provenzano Md, et al. Measurement of protein using bicinchoninic acid. Anal Biochem. 1985;150:76–85.
Walker JM (ed.). Nondenaturing polyacrylamide gel electrophoresis of proteins. The protein protocols handbook. Springer; Totowa, NJ, United States, 2009, p. 171–6.
Lorite MJ, Tachil J, Sanjuán J, Meyer O, Bedmar EJ. Carbon monoxide dehydrogenase activity in Bradyrhizobium japonicum. Appl Environ Microbiol. 2000;66:1871–6.
Berney M, Greening C, Hards K, Collins D, Cook GM. Three different [NiFe] hydrogenases confer metabolic flexibility in the obligate aerobe Mycobacterium smegmatis. Environ Microbiol. 2014;16:318–30.
Carere CR, Hards K, Houghton KM, Power JF, McDonald B, Collet C, et al. Mixotrophy drives niche expansion of verrucomicrobial methanotrophs. ISME J. 2017;11:2599–610.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Kumar S, Stecher G, Tamura K MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4.
Constant P, Chowdhury SP, Hesse L, Pratscher J, Conrad R. Genome data mining and soil survey for the novel Group 5 [NiFe]-hydrogenase to explore the diversity and ecological importance of presumptive high-affinity H2-oxidizing bacteria. Appl Environ Microbiol. 2011;77:6027–35.
Schäfer C, Friedrich B, Lenz O. Novel, oxygen-insensitive group 5 [NiFe]-hydrogenase in Ralstonia eutropha. Appl Environ Microbiol. 2013;79:5137–45.
Schäfer C, Bommer M, Hennig SE, Jeoung J-H, Dobbek H, Lenz O, et al. Structure of an actinobacterial-type [NiFe]-hydrogenase reveals insight into O2-tolerant H2 oxidation. Structure. 2016;24:285–92.
Berney M, Cook GM. Unique flexibility in energy metabolism allows mycobacteria to combat starvation and hypoxia. PLoS ONE. 2010;5:e8614.
Patrauchan MA, Miyazawa D, LeBlanc JC, Aiga C, Florizone C, Dosanjh M, et al. Proteomic analysis of survival of Rhodococcus jostii RHA1 during carbon starvation. Appl Environ Microbiol. 2012;78:6714–25.
Santiago B, Schübel U, Egelseer C, Meyer O. Sequence analysis, characterization and CO-specific transcription of the cox gene cluster on the megaplasmid pHCG3 of Oligotropha carboxidovorans. Gene. 1999;236:115–24.
Christie-Oleza JA, Fernandez B, Nogales B, Bosch R, Armengaud J. Proteomic insights into the lifestyle of an environmentally relevant marine bacterium. ISME J. 2012;6:124.
Muthusamy S, Lundin D, Mamede Branca RM, Baltar F, González JM, Lehtiö J, et al. Comparative proteomics reveals signature metabolisms of exponentially growing and stationary phase marine bacteria. Environ Microbiol. 2017;19:2301–19.
Koch T, Dahl C. A novel bacterial sulfur oxidation pathway provides a new link between the cycles of organic and inorganic sulfur compounds. ISME J. 2018;12:2479–91.
Cao X, Koch T, Steffens L, Finkensieper J, Zigann R, Cronan JE, et al. Lipoate-binding proteins and specific lipoate-protein ligases in microbial sulfur oxidation reveal an atpyical role for an old cofactor. eLife. 2018;7:e37439.
Wagner T, Koch J, Ermler U, Shima S. Methanogenic heterodisulfide reductase (HdrABC-MvhAGD) uses two noncubane [4Fe-4S] clusters for reduction. Science. 2017;357:699–703.
Ramos AR, Grein F, Oliveira GP, Venceslau SS, Keller KL, Wall JD, et al. The FlxABCD‐HdrABC proteins correspond to a novel NADH dehydrogenase /heterodisulfide reductase widespread in anaerobic bacteria and involved in ethanol metabolism in Desulfovibrio vulgaris Hildenborough. Environ Microbiol. 2015;17:2288–305.
Boughanemi S, Lyonnet J, Infossi P, Bauzan M, Kosta A, Lignon S, et al. Microbial oxidative sulfur metabolism: biochemical evidence of the membrane-bound heterodisulfide reductase-like complex of the bacterium Aquifex aeolicus. FEMS Microbiol Lett. 2016;363:1–8.
Justice NB, Norman A, Brown CT, Singh A, Thomas BC, Banfield JF. Comparison of environmental and isolate Sulfobacillus genomes reveals diverse carbon, sulfur, nitrogen, and hydrogen metabolisms. BMC Genom. 2014;15:1107.
Quatrini R, Appia-Ayme C, Denis Y, Jedlicki E, Holmes DS, Bonnefoy V. Extending the models for iron and sulfur oxidation in the extreme acidophile Acidithiobacillus ferrooxidans. BMC Genom. 2009;10:394.
Mohammadi S, Pol A, van Alen TA, Jetten MSM, Op den Camp HJM. Methylacidiphilum fumariolicum SolV, a thermoacidophilic ‘Knallgas’ methanotroph with both an oxygen-sensitive and -insensitive hydrogenase. ISME J. 2017;11:945–58.
Dobbek H, Gremer L, Meyer O, Huber R. Crystal structure and mechanism of CO dehydrogenase, a molybdo iron-sulfur flavoprotein containing S-selanylcysteine. Proc Natl Acad Sci. 1999;96:8884–9.
Cavaletti L, Monciardini P, Bamonte R, Schumann P, Rohde M, Sosio M, et al. New lineage of filamentous, spore-forming, gram-positive bacteria from soil. Appl Environ Microbiol. 2006;72:4360–9.
Sorokin DY, Vejmelkova D, Lücker S, Streshinskaya GM, Rijpstra WIC, Damste JSS, et al. Nitrolancea hollandica gen. nov., sp. nov., a chemolithoautotrophic nitrite-oxidizing bacterium isolated from a bioreactor belonging to the phylum Chloroflexi. Int J Syst Evol Microbiol. 2014;64:1859–65.
Klatt CG, Liu Z, Ludwig M, Kühl M, Jensen SI, Bryant DA, et al. Temporal metatranscriptomic patterning in phototrophic Chloroflexi inhabiting a microbial mat in a geothermal spring. ISME J. 2013;7:1775.
Jayachandran G, Görisch H, Adrian L. Studies on hydrogenase activity and chlorobenzene respiration in Dehalococcoides sp. strain CBDB1. Arch Microbiol. 2004;182:498–504.
Nijenhuis I, Zinder SH. Characterization of hydrogenase and reductive dehalogenase activities of Dehalococcoides ethenogenes strain 195. Appl Environ Microbiol. 2005;71:1664–7.
Hartwig S, Dragomirova N, Kublik A, Türkowsky D, von Bergen M, Lechner U, et al. A H2‐oxidizing, 1, 2, 3‐trichlorobenzene‐reducing multienzyme complex isolated from the obligately organohalide‐respiring bacterium Dehalococcoides mccartyi strain CBDB1. Environ Microbiol Rep. 2017;9:618–25.
King GM. Molecular and culture-based analyses of aerobic carbon monoxide oxidizer diversity. Appl Environ Microbiol. 2003;69:7257–65.
Weber CF, King GM. The phylogenetic distribution and ecological role of carbon monoxide oxidation in the genus Burkholderia. FEMS Microbiol Ecol. 2012;79:167–75.
King GM. Carbon monoxide as a metabolic energy source for extremely halophilic microbes: implications for microbial activity in Mars regolith. Proc Natl Acad Sci USA. 2015;112:4465–70.
Weber CF, King GM. Volcanic soils as sources of novel CO-oxidizing Paraburkholderia and Burkholderia: Paraburkholderia hiiakae sp. nov., Paraburkholderia metrosideri sp. nov., Paraburkholderia paradisi sp. nov., Paraburkholderia peleae sp. nov., and Burkholderia alpina sp. nov., a member of the Burkholderia cepacia complex. Front Microbiol. 2017;8:207.
King GM. Uptake of carbon monoxide and hydrogen at environmentally relevant concentrations by Mycobacteria. Appl Environ Microbiol. 2003;69:7266–72.
Gadkari D, Schricker K, Acker G, Kroppenstedt RM, Meyer O. Streptomyces thermoautotrophicus sp. nov., a thermophilic CO-and H2-oxidizing obligate chemolithoautotroph. Appl Environ Microbiol. 1990;56:3727–34.
Meyer O, Schlegel HG. Reisolation of the carbon monoxide utilizing hydrogen bacterium Pseudomonas carboxydovorans (Kistner) comb. nov. Arch Microbiol. 1978;118:35–43.
Khalil MAK, Rasmussen RA. The global cycle of carbon monoxide: Trends and mass balance. Chemosphere. 1990;20:227–42.
Ehhalt DH, Rohrer F. The tropospheric cycle of H2: a critical review. Tellus B. 2009;61:500–35.
Acknowledgements
This work was supported by an ARC DECRA Fellowship (DE170100310; awarded to CG), an ARC Discovery Project (DP180101762; awarded to CG), a Marsden Grant (16-GNS-035; awarded to CRC and CG), an SNF Early Mobility Postdoctoral Fellowship (P2EZP3_178421; awarded to EC), an Australian Government Research Training Program Stipend (awarded to ZFI), and Monash University Doctoral Scholarships (awarded to PRFC, Y-JC, and SB). We thank Prof Craig White and Prof Gary King for their helpful comments.
Author information
Authors and Affiliations
Contributions
CG conceived this study, designed experiments, and analysed data. ZFI, PRFC, and JF performed experiments and analysed data. CG, RMG, and ZFI supervised students. YJ-C, SKB, TJ, CRC, MBS, and EC contributed to methods development and data analysis. CG, ZFI, CRC, MBS, and EC wrote and edited the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Islam, Z.F., Cordero, P.R.F., Feng, J. et al. Two Chloroflexi classes independently evolved the ability to persist on atmospheric hydrogen and carbon monoxide. ISME J 13, 1801–1813 (2019). https://doi.org/10.1038/s41396-019-0393-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0393-0
This article is cited by
-
Survival and rapid resuscitation permit limited productivity in desert microbial communities
Nature Communications (2024)
-
Trace gas oxidation sustains energy needs of a thermophilic archaeon at suboptimal temperatures
Nature Communications (2024)
-
Current status of carbon monoxide dehydrogenases (CODH) and their potential for electrochemical applications
Bioresources and Bioprocessing (2023)
-
Structural basis for bacterial energy extraction from atmospheric hydrogen
Nature (2023)
-
Molecular hydrogen in seawater supports growth of diverse marine bacteria
Nature Microbiology (2023)