Abstract
Ecological factors contributing to depth-related diversification of marine Thaumarchaeota populations remain largely unresolved. To investigate the role of potential microbial associations in shaping thaumarchaeal ecotype diversification, we examined co-occurrence relationships in a community composition dataset (16S rRNA V4-V5 region) collected as part of a 2-year time series in coastal Monterey Bay. Ecotype groups previously defined based on functional gene diversity—water column A (WCA), water column B (WCB) and Nitrosopumilus-like clusters—were recovered in the thaumarchaeal 16S rRNA gene phylogeny. Networks systematically reflected depth-related patterns in the abundances of ecotype populations, suggesting thaumarchaeal ecotypes as keystone members of the microbial community below the euphotic zone. Differential environmental controls on the ecotype populations were further evident in subnetwork modules showing preferential co-occurrence of OTUs belonging to the same ecotype cluster. Correlated abundances of Thaumarchaeota and heterotrophic bacteria (e.g., Bacteroidetes, Marinimicrobia and Gammaproteobacteria) indicated potential reciprocal interactions via dissolved organic matter transformations. Notably, the networks recovered ecotype-specific associations between thaumarchaeal and Nitrospina OTUs. Even at depths where WCB-like Thaumarchaeota dominated, Nitrospina OTUs were found to preferentially co-occur with WCA-like and Nitrosopumilus-like thaumarchaeal OTUs, highlighting the need to investigate the ecological implications of the composition of nitrifier assemblages in marine waters.
Similar content being viewed by others

Introduction
Thaumarchaeota are important contributors to carbon and nitrogen cycling in the ocean due to their roles in nitrogen remineralization, nitrous oxide production, and carbon fixation. Accounting for up to 20% of marine picoplankton communities, they are often the most abundant microbial organisms in marine waters, particularly below the euphotic zone [1, 2]. All cultivated members of the phylum Thaumarchaeota are capable of oxidizing ammonia to nitrite, the first and rate-limiting step in nitrification (e.g., refs. 3,4,5,6,7). Distribution and diversity patterns of Thaumarchaeota have been studied extensively in natural systems using the gene encoding the α-subunit of ammonia monooxygenase (amoA) as a molecular marker (e.g., refs. 8,9,10,11,12,13,14).
Phylogenetic analysis of amoA genes from marine waters suggests the existence of two distinct depth-related ‘ecotype’ populations of Thaumarchaeota in the ocean. First identified by Francis et al. [8], the water column “A” (WCA) and water column “B” (WCB) clades are suggested to represent vertically segregated “shallow” and “deep” ecotypes of pelagic Thaumarchaeota, respectively [8,9,10, 12, 15,16,17,18,19,20,21,22,23]. Distribution patterns of the two ecotypes along depth gradients suggest differential niche adaptations for the ecotype lineages: the “shallow” ecotype likely possesses a broader niche than the “deep” ecotype [24], as the former is often found throughout the water column, although more abundant at shallower depths, while the “deep” ecotype is more abundant below the euphotic zone [10, 12, 19, 20]. Genomic and physiological differences between the ecotype populations and environmental factors driving their niche separation, however, are not well resolved [22, 24]. Suggested depth-related physicochemical gradients influencing ecotype partitioning include: light levels [25], reactive oxygen species [26], and substrate (ammonia) concentrations [27].
Prior studies examining thaumarchaeal ecotype distributions in the water column have focused primarily on the role of physicochemical factors on niche partitioning [12, 20, 22, 23], and rarely on potential ecological interactions involving Thaumarchaeota and other co-occurring microbial populations. The latter is important because microbial composition and functional distribution in ecosystems are known to be significantly influenced by ecological interactions within microbiomes [28,29,30]. In recent years, analyses of microbial co-occurrence patterns have been applied to explore ecologically-meaningful interactions between taxa in both environmental and host-associated microbiomes, primarily through inferring co-occurrence networks from the relative abundances of taxa [30,31,32,33,34,35,36,37,38,39]. While most such studies have been limited to exploring interactions within bacterial communities, a few have examined ecological associations involving archaeal groups (e.g., refs. 32, 36, 39, 40). The two prior studies that directly incorporated thaumarchaeal phylotypes into association networks— one in grassland soils [40] and another in a coastal marine environment [39]—have both reported frequent associations between the two nitrifier guilds (i.e., thaumarchaeal ammonia oxidizers and nitrite-oxidizing bacteria). Examining the dynamics and co-occurrence patterns of archaeal populations at the San Pedro Ocean Time-Series (SPOT) station in Southern California, Parada and Fuhrman [39] suggested that distinct nitrifier assemblages may be responsible for nitrification at different spatiotemporal regimes, in agreement with the co-varying abundances of archaeal and bacterial nitrifiers observed in other marine systems [9, 13]. However, to date no study has specifically addressed any potential links between ecotype differentiation and co-occurrence patterns.
In this study, we seek to examine co-occurrence relationships involving Thaumarchaeota in the water column, in the context of depth-related ecotype diversification. We apply co-occurrence network analysis to community sequence data (16S rRNA gene) from coastal Monterey Bay to decipher patterns of association between thaumarchaeal ecotypes and other members of the microbial community. Specifically, we investigate: (i) whether depth-related changes in the abundance and diversity of bacteria and archaea are reflected in the co-occurrence relationships of thaumarchaeal ecotypes, and (ii) to what degree ecotype diversification matter for network structure and association patterns at different depth layers. We also examine the relative importance of individual thaumarchaeal phylotypes in the networks to ascertain indications of niche separation, hypothesizing that differences in association patterns would result from ecotypes adapting to divergent ecological niches.
While our approach is fundamentally different from inferring and interpreting ecological networks that entail direct causal relationships between taxa [41], we attempt to trace depth-related patterns of taxa co-occurrence in light of suggested ecological relationships between microbial groups. Unlike prior correlation-based studies, we also map phylogenetic information on the co-occurrence networks to analyze thaumarchaeal association patterns at a much greater taxonomic resolution to gain insights into ecotype diversification.
Materials and methods
Oceanographic setting and sample collection
In collaboration with the Monterey Bay Aquarium Research Institute (MBARI) 29-year oceanographic time series, we sampled two stations, M1 (36.747 N, −122.022 W) and M2 (36.697 N, −122.378 W), in Monterey Bay over a period of two years (from May 2014 to February 2016). Samples were collected nearly monthly (although no sampling was carried out between December-March in 2014-15 and January 2016) from up to 10 depths at each station, ranging from surface to 200 m (in 2014) or 500 m (in 2015 and 2016). Using Niskin (10 L volume) bottles mounted onto a conductivity-temperature-depth (CTD) rosette, duplicate water samples were collected from each depth. A volume of 1 L each of the collected water was filtered using a peristaltic pump through a 10 µm pore size polyester pre-filter (in-line; GE Osmonics, Minnesota, USA), followed by a 0.22 µm filter (Supor, Pall Inc, New York, USA). Both filters were placed into individual 2 ml bead tubes containing a mixture of 0.1 and 0.5 mm glass beads, which were flash-frozen in liquid nitrogen immediately after filtering and stored at −80 °C until processing. For every sample filtered for molecular analyses, we also obtained nutrient measurements (NH4+, NO3−, NO2−, PO43−, SiO42−) and other water column characteristics (chlorophyll a, dissolved oxygen, relative fluorescence, and total carbon), using previously described methods [42, 43].
Samples analyzed in this study include all time points and depths from both stations, except for September 2015 and February 2016 (these two profiles were not included in the first phase of 16S rRNA iTag sequencing runs due to logistical delays associated with processing the samples for nucleic acid extractions).
Nucleic acid extractions, 16S rRNA iTag sequencing and processing
Total DNA and RNA were extracted from all 0.2 micron filters using a previously described two-step co-extraction protocol [19, 43], with a few modifications - for each set of extraction, we added 10 μL of fresh β-mercaptoethanaol per 1 mL of the lysis buffer (mirVana miRNA isolation kit, Life Technologies, Carlsbad, CA, USA). To maximize yield, DNA and RNA were eluted twice from the capture columns with 50 μL each of the respective elution buffers heated to 95 0C. Total DNA yield for each sample was quantified using the Quant-iTTM High Sensitivity dsDNA assay with a Qubit Fluorometer (Invitrogen, NY, USA). Estimated concentrations ranged from 0.12–120 μg DNA L−1 of seawater filtered, with a mean yield of 19.85 μg L−1.
The 16S rRNA gene amplicons were prepared from each sample and sequenced using the MiSeq platform (Illumina; iTag amplicon sequencing), as part of a Community Science Program (CSP) project with the DOE Joint Genome Institute (JGI). Universal (Bacteria and Archaea) V4-V5 primers 515F-Y (5′- GTGCCAGCMGCCGCGGTAA) and 926 R (5′- CCGYCAATTYMTTTRAGTTT) [39] were used for amplification.
Demultiplexed, quality-filtered reads were processed through the MiSeq Standard Operating Procedure for the mothur software package [44]. Briefly, quality-filtered unique reads were aligned against the Silva v128 16S rRNA gene alignment [45]. Chimeric reads were detected using UCHIME [46] and removed from the alignment. Taxonomic assignments were carried out using the Silva v128 reference database [45]. Clustering was performed at 97% similarity using the cluster.split() command in mothur. At the completion of the pipeline, 10,151 Operational Taxonomic Units (OTUs) were retained for downstream analyses.
Statistical analyses
All statistical analyses on the OTU abundance data were carried out in R (v. 3.4.1; ref. 47), primarily using the Phyloseq (v 1.20.0; ref. [48]) and vegan (v 2.4.4; ref. [49]) packages. Samples with < 10,000 reads were excluded (only 1 sample out of 275 total; the 5 m sample from August 2014 had only 311 reads). OTU abundance tables were rarefied to even depth (35,927 OTUs per sample) prior to beta-diversity analyses. Community variability was evaluated by nonmetric multidimentional scaling (nMDS) using the Bray-Curtis dissimilarity matrix. PERMANOVA analyses [using adonis() in vegan] were performed to further confirm the ordination results. Constrained ordinations [Constrained Analysis of Principal Coordinates (CAP) and Canonical Correspondence Analysis (CCA)] were performed on the Bray-Curtis dissimilarity matrix to examine environmental controls on beta-diversity using Phyloseq functions. Hierarchical clustering was used to identify depth-based grouping patterns in overall community composition (at the phylum level; relative abundance cutoff: 2%). Phyloseq alpha-diversity command plot_richness was used to compute and visualize richness (# observed OTUs) and evenness (Inverse Simpson index).
Association network inference and analyses
Low-abundance OTUs were filtered out prior to network calculation, retaining only the OTUs that were present at least 10 times in 10% of the samples at each depth. In addition to reducing network complexity by removing poorly represented taxa, this filtering step also aided in avoiding spurious correlations between taxa resulting from skewed abundance distributions.
Correlation networks were calculated at the six chosen depths at each station (see Results and Discussion for details on depth selection) using the SpiecEasi R package (v. 0.1.2). Spiec-Easi (Sparse InversE Covariance estimation for Ecological Association and Statistical Inference) is a novel statistical method developed specifically for compositional data; network inference is done under the assumption of sparsity, using sparse neighborhood and inverse covariance selection algorithms [50]. We ran Spiec-Easi in the neighborhood selection (MB) model, and selected final networks for each depth-station combination by random subsampling and interaction re-estimation using the Stability Approach to Regularization Selection (StARS), using a variability threshold of 0.05%.
The resulting adjacency matrices were converted into network objects using the R igraph package (v 1.1.2). Various network summary statistics and node centrality measures were calculated using igraph functions to compare network topologies across stations and depths. Detailed explanations of computed network statistics are provided in the Supplementary Information accompanying this article. Cytoscape (v 3.5.1; ref. [51]) was used for visualizing the networks and identifying network modules.
In addition to network topology measures, we also computed the following node centrality measures to compare the relative importance of individual nodes within a network:
-
I.
Degree centrality refers to the number of connections that a node has, and is based on the idea that a central node is connected to many others in the network.
-
II.
Eigenvector centrality measures the importance of a node based on the number of ‘central’ nodes it is connected to (i.e., a central node would be connected to many other central nodes).
-
III.
Betweenness centrality measures the extent to which a node lies on the shortest paths between other nodes in the network. In other words, it is an indicator of a node’s influence on the ‘communication’ between parts of the network.
We further used degree and betweenness centrality scores as metrics to evaluate the degree of importance or ‘keystone-ness’ of a taxon (i.e., node) within a given network (discussed further in Results and Discussion).
Results and discussion
Depth-associated patterns in community structure
The dataset consisted of 44 distinct phyla, including 3 archaeal phyla (Fig. 1a). Proteobacteria and Bacteroidetes were the most abundant phyla, comprising 51.5 and 18.7% of the classified community, respectively. Alphaproteobacteria, specifically the SAR11 clade, was the most abundant class overall. A total of 14.3% of the reads were classified as Archaea, and Thaumarchaeota were the third most abundant phylum constituting 7.99% of the total classified reads (i.e., both Bacteria and Archaea). Depths > 60 m at both stations showed relative numerical dominance of thaumarchaeal OTUs over most bacterial and all other archaeal phyla, consistent with their reported distribution in the ocean’s interior [1, 2].
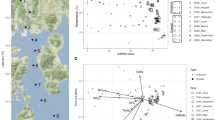
(a) Relative abundance of major bacterial and archaeal phyla in the 16 S rRNA iTag dataset. The data are from all 10 depths sampled across the 2-year period. Only phyla with overall relative abundance > 2% are shown. Note the increasing relative abundance of Thaumarchaeota with depth. (b) Principal Component Analysis (PCA) plot showing the spread of samples along a depth gradient (PC1), based on measured environmental variables. Shapes correspond to the two stations, and vectors are drawn to highlight significant environmental variables explaining the variation along PC1 and PC2. Percent variability explained by each component is given in brackets next to the axis labels
Relative abundance data also indicated Thaumarchaeota as the primary ammonia oxidizers in the system, since ammonia-oxidizing bacteria (AOB) were considerably less abundant (0.05%). All AOB in the dataset belonged to the betaproteobacterial class (Family: Nitrosomanadaceae). Across depth and time points, the most abundant thaumarchaeal OTU (#3) was 20 to 2700 times more abundant than the most abundant AOB OTU (#213).
Relative abundances of all major microbial groups showed clear spatiotemporal patterns at the two stations, M1 and M2. However, no significant difference in taxa composition was observed between the two stations (PERMANOVA and ANOSIM p-values > 0.05), which may reflect the fact that oceanographic conditions (i.e., measured macronutrient levels and physiochemical variables) did not change significantly between the stations during the sampling period (PERMANOVA p > 0.05; Fig. 1b). Both water column depth and sampling time significantly correlated to community structure with p < 0.01 (based on PERMANOVA on Bray-Curtis dissimilarities), also supported by analysis of similarities (ANOSIM) performed on sampled depths (p = 0.001, R-value: 0.54). Similar vertical and temporal structuring of the pelagic microbial community is commonly observed in coastal and open ocean systems (e.g., refs. 39, 52,53,54,55,56,57,58). While seasonality appeared to be a strong factor controlling beta-diversity in our dataset (a temporal gradient was evident along PC2 in Fig. 1b, presented in Fig. S1A), the most variability in community structure explained was along the spatial dimension (Fig. 1b). Therefore, we focus primarily on depth-related changes in community composition and co-occurrence patterns from this point forward.
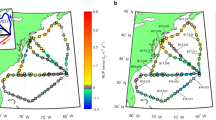
To examine the environmental variables significantly correlating with depth-based partitioning of community structure, we performed a Constrained Analysis of Principal coordinates (CAP) on the Bray-Curtis dissimilarities between samples (Fig. 2a). The first two constrained axes together explained 55.5% of the total compositional variation; spread of samples along the first ordination axis was suggestive of the strong depth gradient. The most significant environmental variables correlating with community composition along the CAP1 direction were nitrate and dissolved oxygen, closely followed by phosphate and silicate concentrations, temperature and salinity (Fig. 2a).
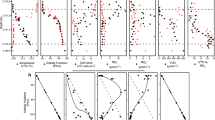
a Constrained Analysis of Principal Coordinates (CAP) plot of community dissimilarity (Bray-Curtis) between samples, color-coded by depth. Shapes correspond to the two stations, and vectors show significant environmental variables constraining the variability in community composition. Rarefied OTU abundances were used for the analysis. b Results of a hierarchical cluster analysis on the Bray-Curtis dissimilarity (based on OTU composition) between depths; OTUs were agglomerated at the Phylum level, and low-abundance OTUs (i.e., < 2% relative abundance) were filtered out prior to analysis. Each column in the heatmap shows the abundance pattern of a unique phylum while each row is a depth sampled from each station. c Alpha diversity measurements - richness and evenness - for (i) overall dataset prior to filtering and (ii) filtered dataset used for network inference. Details on filtering criteria explained in Materials and Methods and discussed further in the text
Results of a hierarchical clustering analysis on the Bray-Curtis dissimilarity between samples along the vertical dimension revealed three major clusters - depths ≤ 40 m formed one cluster (which we refer to as the ‘shallow’ cluster), while 80, 100 and 200 m depths clustered together (the ‘intermediate’ cluster). The 500 m depth samples were distinct from all other depths at both stations, suggestive of a significantly different community composition at this depth (Fig. 2b). These depth clusters broadly reflect the oceanographic features of our study system, representing the mixed layer, twilight waters and the zone below the seasonal thermocline, respectively. Alpha diversity analyses also supported the depth-related clustering patterns: estimates of richness (#observed OTUs) and evenness (Inverse-Simpson) were lowest at the shallow depths and peaked at intermediate depths (Fig. 2c[i]). Richness increased steadily with depth, while evenness peaked at 80–100 m at both stations (Fig. 2c[i]). Thus, since both alpha and beta diversity patterns changed significantly with depth, we hypothesized that the nature of interactions involving Thaumarchaeota would also reflect the change in community structure and therefore show depth-related patterns.
To compare the co-occurrence relationships of Thaumarchaeota at different depth regimes, we computed association networks for groups of samples belonging to the three depth clusters identified in Fig. 2b. Networks were inferred using OTU compositional data filtered to include only the most abundant OTUs, for all depths ≥80 m. From the ‘shallow’ cluster, we picked only the 20 m and 40 m depths for network inference because: (i) overall thaumarchaeal relative abundances were <6% at the shallow cluster depths (Fig. 3a) and (ii) the four uppermost depths in the ‘shallow’ cluster (i.e., 5, 10, 20, and 30 m) were not significantly different from each other in terms of beta-diversity (confirmed by PERMANOVA test, p > 0.05). The final filtered dataset had similar diversity patterns as that of the overall dataset (Fig. 2c[ii] and Fig. S1B), indicating that the filtering step did not substantially alter community structure to effectuate a marked change in inferred co-occurrence patterns.
a Relative abundance of the top 10 thaumarchaeal OTUs at the depths considered for network inference at M1 and M2. Also indicated in each plot window is the overall relative abundance of thaumarchaeal OTUs at the corresponding depths. b Distribution of centrality measures for all thaumarchaeal nodes in each network corresponding to the 12 station-depth combinations. Mean values are indicated by gray horizontal lines
Global network topologic features vary across stations and depths
To assess changes in network structure with depth, we computed global topological properties for each network, and compared these measures to corresponding random networks. Computed measures for all networks were significantly larger than those of their comparable random networks (one sample z-test p < 0.001; 1000 Erdos-Renyi random networks were created for each network). This suggested that the co-occurrence patterns we observed were not random, and that OTUs/taxa involved in these potential associations are highly likely to be found together.
As a summary measure of local network topologies, we computed graphlet correlation distance between the networks [59], and inferred a two-dimensional embedding of the distances using MDS (Fig. S1C). The resulting clustering patterns broadly corresponded to the degree distributions for individual networks depth (Fig. S1D), as well as to in-situ alpha diversity at each. Network complexity (as determined by clustering coefficient and average degree scores) generally increased with depth at both stations (Table 1), and was significantly greater than those of comparable Erods-Renyi random networks (p < 0.01). This is consistent with prior studies reporting ‘small-world’ properties for microbial ecological networks (e.g., refs. [32, 34])–each OTU is linked to many others and, therefore, randomly removing one should not significantly alter network/community structure [60]. Total number of connections in the networks increased near-linearly with the number of unique phylotypes at each depth (Pearson’s R2: 0.95 and 0.98, respectively for M1 and M2). Thus, networks from depths with the highest alpha diversity values also had the highest average degree (Table 1). Degree distributions for the individual networks also followed this trend as right-shifted distributions characterized depths with the largest alpha-diversity values, particularly at M1 (Fig. S1D). As noted earlier, degree distribution of the M2 20 m network was considerably shifted to the left compared to other networks from this station (Fig. S1D), consistent with the lowest average alpha diversity values observed at this depth (Fig. 2c[ii]). These observations imply that the density of network connections is directly affected by the diversity of interacting OTUs, as observed previously for human microbiome networks [28].
Role of thaumarchaeal OTUs in shaping network structure
Consistent with the depth profile observed for the relative abundances of Thaumarchaeota (Fig. 3a), the number of thaumarchaeal OTUs meeting the abundance filtering criteria and incorporated into the networks increased with depth-final networks from 20 m depth included only 2 thaumarchaeal OTUs, while the 500 m networks contained 18 and 17 OTUs each, at M1 and M2, respectively.
Distribution of node centrality scores computed for thaumarchaeal OTUs in the networks, however, did not follow the trend in their relative abundances, but closely tracked the vertical alpha diversity patterns (Fig. 3b). At both stations, thaumarchaeal nodes with the highest average centrality scores appeared in the 80 and 100 m networks, although overall relative abundances of Thaumarchaeota at these depths were 4–5% lower than at 500 m (Fig. 3a). These depths also had among the highest OTU richness (Fig. 2c), which likely correlates directly to the number of possible links between taxa. Likewise, the relative drop in centrality scores in the 200 m networks, particularly at M1 (Fig. 3b), appears to reflect the relatively lower overall richness at this depth (Fig. 2c).
Nodes with high centrality scores are predicted to be ‘keystone’ taxa in ecological networks - microbial members important for maintaining network connectivity, and potentially controlling the stability of the entire community [32, 35, 61,62,63]. Many of the thaumarchaeal nodes had high centrality scores, and thus appeared to be hub nodes in the networks, fitting the keystone species definition [29] (Fig. S2) in the depths considered. Particularly in the ‘intermediate’ depth layers, centrality of thaumarchaeal nodes were often comparable to the highest scoring nodes in the network (Fig. S2). Moreover, some thaumarchaeal nodes (e.g., OTUs 21, 211, and 373) had among the highest betweenness centrality scores among all nodes, despite having relatively fewer connections, suggesting that many of them were important for the flow of information between parts of the network by acting as ‘bridges’ between poorly linked subnetworks/modules.
Intriguingly, abundance of individual OTUs did not correlate to their importance in the network. Even though the overall relative abundance of Thaumarchaeota increased with increasing depth, there were substantial variations in the abundances of individual OTUs (Fig. 3a). While generally, the most abundant OTUs were also well-connected, many of the ‘hub’ nodes in the networks were relatively low-abundance members of the phylum. For instance, thaumarchaeal nodes with the highest centrality scores in the 500 m networks were OTU 812 and OTU 929, neither of which were among the top 10 most abundant thaumarchaeal OTUs at this depth (Fig. 3a). A similar pattern has previously been observed in human [38] and plant [64] microbiome networks, where highly abundant taxa often appeared to be poorly connected while low-abundance ones served as highly connected hub nodes.
In addition to comparing individual node centralities, we also examined whether Thaumarchaeota played a more important role in network structure compared to other phyla, by testing for group effect on betweenness centrality scores for all nodes agglomerated by phylum, individually for each network. Results suggested that Thaumarchaeota were significantly important to network structure for all but three of the station-depth combinations—the 20 m and 80 m networks from M1, and the 20 m network from M2 (Kruskal-Wallis P < 0.05 for all networks except the three specified). Other phyla identified as important for network structure included the following: Nitrospinae in the M1 40 m, 100 m and 200 m, and M2 80 m networks; Lentisphaerae and Planctomycetes in the 80 m networks; Bacteroidetes in the M1 80 m and 500 m networks; and Marinimicrobia in the M2 100 m network. Proteobacteria, despite being the most abundant phylum in the dataset, were significantly central to network structure only in the M1 80 m and M2 40 m networks (based on Kruskal-Wallis significance test; P > 0.05).
Co-occurrence links involving Thaumarchaeota
Across all depths at both stations, thaumarchaeal nodes had frequent co-occurrence links with Bacteroidetes, Verrucomicrobia, Planctomycetes, Euryarchaeota, and Alpha-Proteobacteria and Gamma-Proteobacteria (Fig. S3, S4). Many of these connections likely reflect changes at the community level, resulting from similar eco-physiological responses to varying environmental conditions with depth.
Another potential explanation for co-occurrence relationships is metabolic interactions between taxa. While it is not possible to parse out the underlying forces at play from network models alone, inferences can be made based on described physiologies and ecological niches of the microbial groups involved in co-occurrence relationships. For example, networks from the ‘intermediate’ and 500 m depth clusters showed many connections between thaumarchaeal and marinimicrobial nodes (Fig. S3, S4), and a positive correlation between their abundances was apparent in phylum-level relative abundance plots (Fig. S5). This is in agreement with a recently proposed co-metabolic interaction between Thaumarchaeota and Marinimicrobia in oxygenated marine waters, via the sharing of nitrous oxide (N2O) [65]. Thaumarchaeota are known to produce N2O, likely as a byproduct of ammonia oxidation [12, 66] though this may come from an abiotic reaction from NO, a known intermediate in ammonia oxidation [67, 68]. Examining a population genome bin resolved from metagenomic data, Hawley and colleagues noted that a Marinimicrobia clade abundant in oxic waters encoded the genetic machinery for N2O reduction to N2 (i.e., nosZ, nosL, nosD, and nosY genes). Based on gene expression patterns, they proposed metabolic coupling between Thaumarchaeota and Marinimicrobia, resulting in the net reduction of N2O produced during nitrification. Although yet to be validated in experimental studies, this offers a potential explanation for the co-occurrence patterns we observe in our networks.
The other frequent co-occurrence patterns observed between Thaumarchaeota, Bacteroidetes and Gammaproteobacteria potentially indicate grazing/remineralization relationships involving these microbial groups. Both Bacteriodetes and Gammaproteobacteria are major organic matter degraders in the ocean [69, 70]. Negative cross-correlation patterns were evident in phylum-level relative abundance plots of Thaumarchaeota and Bacteroidetes (predominantly Flavobacteria), especially in the epipelagic (Fig. S6). Temporal trends in thaumarchaeal and gammaproteobacterial abundances also appeared to be correlated, particularly ≤100 m (Fig. S7). These abundance patterns broadly reflect changes in community composition in response to seasonal changes in organic matter availability—abundances of both Bacteroidetes and Gammaproteobacteria peak during the post-upwelling summer months with greater overall productivity, followed by lower abundances in winter. These patterns point to thaumarchaeal blooms potentially fueling heterotrophic bacterioplankton communities in shallow waters where the latter is especially abundant (Fig. 1a).
Oceanospirillales and Salinisphaerales accounted for most gammaproteobacterial nodes linked to Thaumarchaeota in the networks. One exception was the 200 m networks where less well-characterized deep-water Orders such as E01-9C-26 and KI89A comprised considerable fractions of the total shared links between the two groups.
A total of 6 Orders accounted for all alphaproteobacterial nodes in the networks, even though the dataset consisted of 64 different Families. SAR11 was the most abundant Order at all depths and among the most highly connected in the networks, accounting for 40–60% of the alphaproteobacterial nodes connected to Thaumarchaeota. Rhodospirillales were also dominant in the networks, particularly at intermediate depths, sharing a relatively larger proportion of the total alphaproteobacterial links with Thaumarchaeota. While Rhodobacterales were the second most abundant Order in the dataset after SAR11, very few links were observed between this group of Alphaproteobacteria and Thaumarchaeota.
AOB were so low in abundance that all except the most abundant OTU (#213) were filtered out prior to network computation. Only the 80 and 100 m networks incorporated OTU 213. In these networks, direct links were present between this AOB OTU and other thaumarchaeal and/or Nitrospinae OTUs. We did not detect any preferential (ecotype-specific) association patterns in these co-occurrence links.
Associations with Nitrospina OTUs were detected in all networks except in the M2 200 m network–we expand upon the nature of these associations in later sections.
Ecotype-specific co-occurrence patterns vary by depth
Having established that thaumarchaeal OTUs were important for network composition, we next considered if taxonomic distribution of OTUs was relevant to co-occurrence patterns by examining whether the distribution of potential ‘ecotype’-specific thaumarchaeal OTUs correlated with their importance in network structure.
Thaumarchaeal OTUs incorporated into the networks formed three distinct groups on a phylogenetic tree (Fig. 4a), broadly corresponding to the ‘ecotype’ clusters generally observed for the amoA gene in marine waters [14, 71]. Most OTUs (n = 11) clustered with the “deep” ecotype (WCB-like) sequences, while several others (n = 6) clustered with Nitrosopelagicus brevis-like sequences (WCA clade). The Nitrosopumilus-like cluster was represented by the least number of OTUs (n = 2), although it included the most abundant thaumarchaeal OTU in our dataset (OTU 3).
a Phylogenetic tree of all thaumarchaeal OTUs included in the networks, together with 16S rRNA gene sequences from reference genomes. The OTUs fall in three distinct clusters potentially corresponding to Nitrosopumilus-like Thaumarchaeota and the two pelagic “ecotypes” (WCA and WCB). b Table showing the most abundant versus and the most important thaumarchaeal OTUs for network structure at each depth (the latter inferred based on number of connections and betweenness centrality scores). OTU names are colored according to the phylogenetic cluster they belong to as shown in panel a. c Relative abundances of thaumarchaeal ecotypes with depth at M1 and M2 (left); summary of co-occurrence patterns observed between thaumarchaeal OTUs belonging to the three ecotype groups and other major microbial Phyla (right)
As noted earlier, centrality measures for thaumarchaeal nodes did not follow the relative abundance trends for individual OTUs as many of the low-abundance OTUs had relatively higher centrality scores. Examining the genus-level taxonomic identity of the most connected and ‘most central’ OTUs (defined as the node with the highest degree and betweenness centrality scores, respectively in each network), we observed that the “deep” WCB-like OTUs became progressively more important with depth (Fig. 4b). The pattern was more conspicuous at M1; in the 20 m networks, Nitrosopumilus-like OTUs had the highest centrality scores, and for depths between 40 and 200 m, WCA-like OTUs became more central to network structure, and WCB-like OTUs gained prominence at 500 m (Fig. 4b). This is in complete agreement with the observed depth distributions of WCA and WCB ecotypes in the northeast Pacific ocean [20]. Our results thus point to a progressively greater ecological role of WCB ecotypes in the community with increasing depth, which is consistent with previous studies reporting dominance of the WCB ecotype below the euphotic zone [12, 19, 22, 43].
Networks also resolved many co-occurrence links among thaumarchaeal OTUs that appeared to suggest preferential interactions within ecotype populations. In a previous study, clade-independent associations were found between SAR11 OTU subtypes at the SPOT station, which led Fuhrman (2009) [72] to suggest that phylogenetic relatedness may not equate to ecological relatedness. Co-occurrence links connecting thaumarchaeal OTUs in our networks were generally consistent with this suggestion, as many frequent associations existed between OTUs of different ecotypes at both stations. However, we also observed many clade-specific interactions: within each network, OTUs belonging to the WCA-like and WCB-like ecotype clusters were found to be preferentially co-located together to different parts of the network (Fig. S3, S4)—suggesting that it is more likely for OTUs of the same ecotype to co-occur.
Partitioning the Thaumarchaeota-centered subnetworks into modules-densely connected clusters of OTUs, potentially representing shared ecological and/or functional niches between taxa—also recovered ecotype-specific modules. Figure 5 and S8 demonstrate these patterns—for each station-depth combination, most modules could be considered to represent separate subnetworks centered around WCA, WCB and Nitrosopumilus-like thaumarchaeal ecotypes. Whether there is an ecological or functional relationship underlying these connections has yet to be examined, although they likely represent shared environmental niches within each ecotype population.
Phylogenetic relatedness shaping co-occurrence patterns between Thaumarchaeota and nitrite-oxidizing bacteria
Previous studies have documented co-varying abundances of Thaumarchaeota and nitrite-oxidizing bacteria (NOB) in the marine water column [9, 13], as well as spatiotemporal patterns in co-occurrence relationships between the two nitrifier guilds [39]. While metabolic coupling between ammonia-oxidizing Thaumarchaeota and NOB predict co-varying abundances, the specifics of their interaction remain unknown and unexplored, including any favored connections between ecotype groups. To address this knowledge gap, we examined whether phylogenetic relatedness mattered for the co-occurrence relationships between Thaumarchaeota and Nitrospina OTUs (presumably NOB) in our networks.
All networks recovered links between thaumarchaeal and Nitrospina nodes, with the exception of the 200 m network from M2. To examine whether specific phylogenetic clusters of Thaumarchaeota and Nitrospina preferentially interacted with each other, we compared the nature of co-occurrence links between distinct phylogenetic clades within the two groups. Specifically, we asked what fraction of the co-occurrence links between Thaumarchaeota and Nitrospina could be associated with each of the three ecotype clades identified in Fig. 4a. The results suggested that OTUs in different thaumarchaeal ecotype clusters tend to co-occur preferentially with distinct Nitrospina OTUs (Fig. 6a). For example, the two clade 1 Nitrospina OTUs occupying relatively derived positions in the NOB phylogeny (i.e., OTUs 33 and 240) linked preferentially to OTUs in the WCA and WCB-like thaumarchaeal ecotype clades (Fig. 6a). In contrast, three other Nitrospina OTUs (OTUs 69, 235, and 470) were found to be linked to Nitrosopumilus and WCA-like OTUs, but not with WCB-like OTUs. Moreover, OTU 64 had connections only with Nitrosopumilus-like thaumarchaeal OTUs.
(a) Phylogenetic tree of all Nitrospina OTUs included in the networks, together with 16 S rRNA gene sequences from reference genomes. The pie-charts next to each OTU show the percent of links observed between that OTU and the thaumarchaeal OTUs belonging to each ecotype cluster identified in panel a. (b) Canonical correspondence analysis (CCA) plot of thaumarchaeal and NOB OTUs from the networks based on Bray-Curtis dissimilarity on relative abundances. Ellipses are drawn to highlight the three ecotype clusters. Vectors correspond to significant environmental variables controlling the distribution of OTUs in the ordination space. The two components together explain 52% of the total constrained inertia
We hypothesized that these patterns reflected similar environmental controls on the distributions of interacting nitrifiers, and thus explored these relationships further by performing a canonical correspondence analysis (CCA) on thaumarchaeal and Nitrospina relative abundances. Clustering patterns in the CCA were consistent with the observed co-occurrence relationships: the three prominent clusters (denoted as I, II, and III) placed Nitrospina OTUs close to the corresponding thaumarchaeal ecotype OTUs that they were found to be preferentially linked to in the networks (Fig. 6b). Relative positions of all Nitrospina OTUs in the ordination space agreed with the co-occurrence patterns depicted in Fig. 6a.
Nitrospina OTUs in our networks systematically co-occurred more frequently with WCA-like and Nitrosopumilus-like thaumarchaeal OTUs, even at deeper depths where the WCB-like ecotype was more abundant, resulting in a relatively distinct association pattern between the two nitrifier groups (Fig. 4c). Network modules presented in Fig. 5 and S8 further illustrate these preferential association patterns. This observation also concurs well with previous work in Monterey Bay by Smith et al. [19], implying a potentially more significant role of the WCA ecotype in mediating nitrification in the water column compared to WCB. Co-occurrence patterns inferred in the present study also imply that nitrification over the euphotic to mid-mesopelagic depths in Monterey Bay is likely mediated primarily by WCA and Nitrosopumilus-like thaumarchaeal ammonia-oxidizers and Nitrospina-like NOB. The relative contribution of the WCB ecotype to nitrification in these waters remains to be determined, especially given their high abundance below surface depths.
These results compare to the findings of Parada and Fuhrman [39], who interpreted the correlation between nitrifier OTUs as an indication of spatiotemporal shifts in populations mediating nitrification at the SPOT station. Here, we add a phylogenetic perspective to the suggested interactions, and find that thaumarchaeal ecotypes may associate differentially with co-occurring Nitrospina OTUs. Our dataset also included 7 Nitrospira OTUs, but their relative abundances were so low that they were not incorporated into the networks. Therefore, whether there exist clade-specific associations between Thaumarchaeota (or Nitrospina) and Nitrospira is impossible to determine based on networks alone. This also means we cannot definitively conclude that the specific co-occurring thaumarchaeal and Nitrospina OTUs in our networks are solely responsible for nitrification at the depths considered.
Unlike ammonia-oxidizers, NOB remain poorly characterized in the marine environment. Recent single amplified genome (SAG)-based exploration of the marine NOB genetic repertoire implicated nitrifiers in yet another type of co-metabolic relationship - several Nitrospinae and Nitrospirae SAGs encode genes for hydratases and lyases involved in the reduction of urea and cyanate to ammonium, suggesting a co-metabolic interaction between ammonia-oxidizing Thaumarchaeota and NOB [73]. While metabolic handoffs via reciprocal feeding is an obvious explanation for the co-occurrence of Thaumarchaeota and Nitrospina OTUs in our networks, it could also result from shared environmental preferences, as suggested by the clustering patterns in the CCA plot (Fig. 6b).
Concluding remarks
Co-occurrence of taxa primarily results from two related factors: (i) niche-overlap due to shared environmental preferences; and (ii) metabolic interactions involving exchange of nutrients. Many such relationships may reflect niche overlap more than realized ecological interactions between taxa [74], and parsing out the specific effect of each is challenging given they often reinforce each other. The patterns we observe here for thaumarchaeal ecotype populations are compelling because these are non-random associations that potentially suggest clade-specific associations between taxa. Of particular importance is the co-occurrence of specific phylotypes of Thaumarchaeota and Nitrospina, largely incompatible with the association patterns expected based on their relative abundances. Recent single-cell genomics and metagenomics studies point to potential ecotype segregation within marine Nitrospinae populations [73, 75], although it remains unclear whether water column depth plays a role in this diversification. In light of observations that Thaumarchaeota and Nitrospinae are major contributors to both nitrogen cycling and carbon fixation within and below the mesopelagic [73], and consistently co-occur (e.g., ref. 39), we believe further investigation into this important partnership in ocean waters is clearly warranted. Finally, the association patterns uncovered here can be used to design co-culture experiments and mine metagenomes to examine the relative importance of biology and environmental factors in defining ecotype niches for the globally important ammonia-oxidizing Thaumarchaeota.
Data availability
Sequence data have been deposited in the NCBI Sequence Read Archive (SRA) database under BioProject PRJNA488312 (SRA accession: SRP159037).
References
Karner MB, DeLong EF, Karl DM. Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature. 2001;409:507–10.
Teira E, van Aken H, Veth C, Herndl GJ. Archaeal uptake of enantiomeric amino acids in the meso‐ and bathypelagic waters of the North Atlantic. Limnol Oceanogr. 2006;51:60–69.
Könneke M, Bernhard AE, la Torre de JR, Walker CB, Waterbury JB, Stahl DA. Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature. 2005;437:543–6.
Hatzenpichler R, Lebedeva EV, Spieck E, Stoecker K, Richter A, Daims H, et al. A moderately thermophilic ammonia-oxidizing crenarchaeote from a hot spring. PNAS. 2008;105:2134–9.
Santoro AE, Casciotti KL. Enrichment and characterization of ammonia-oxidizing archaea from the open ocean: phylogeny, physiology and stable isotope fractionation. ISME J. 2011;5:1796–808.
Stieglmeier M, Klingl A, Alves RJE, Rittmann SKMR, Melcher M, Leisch N, et al. Nitrososphaera viennensis gen. nov., sp. nov., an aerobic and mesophilic, ammonia-oxidizing archaeon from soil and a member of the archaeal phylum Thaumarchaeota. Int J Syst Evol Micro. 2014;64:2738–52.
Qin W, Amin SA, Martens-Habbena W, Walker CB, Urakawa H, Devol AH, et al. Marine ammonia-oxidizing archaeal isolates display obligate mixotrophy and wide ecotypic variation. PNAS. 2014;111:12504–9.
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB. Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. PNAS. 2005;102:14683–8.
Mincer TJ, Church MJ, Taylor LT, Preston C, Karl DM, DeLong EF. Quantitative distribution of presumptive archaeal and bacterial nitrifiers in Monterey Bay and the North Pacific Subtropical Gyre. Environ Micro. 2007;9:1162–75.
Beman JM, Popp BN, Francis CA. Molecular and biogeochemical evidence for ammonia oxidation by marine Crenarchaeota in the Gulf of California. ISME J. 2008;2:453–63.
Yakimov MM, Cono VL, Denaro R. A first insight into the occurrence and expression of functional amoA and accA genes of autotrophic and ammonia-oxidizing bathypelagic Crenarchaeota of Tyrrhenian Sea. Deep Sea Res Part II. 2009;56:748–54.
Santoro AE, Casciotti KL, Francis CA. Activity, abundance and diversity of nitrifying archaea and bacteria in the central California Current. Environ Microbiol. 2010;12:1989–2006.
Tolar BB, King GM, Hollibaugh JT. An analysis of Thaumarchaeota populations from the Northern Gulf of Mexico. Front Microbiol. 2013;4:72. https://doi.org/10.3389/fmicb.2013.00072.
Tolar BB, Ross MJ, Wallsgrove NJ, Liu Q, Aluwihare LI, Popp BN, et al. Contribution of ammonia oxidation to chemoautotrophy in Antarctic coastal waters. ISME J. 2016;10:2605–19.
Hallam SJ, Konstantinidis KT, Putnam N, Schleper C, Watanabe Y-I, Sugahara J, et al. Genomic analysis of the uncultivated marine crenarchaeote Cenarchaeum symbiosum. PNAS. 2006;103:18296–301.
Church MJ, Wai B, Karl DM, DeLong EF. Abundances of crenarchaeal amoA genes and transcripts in the Pacific Ocean. Environ Microbiol. 2010;12:679–88.
Hu A, Jiao N, Zhang CL. Community structure and function of planktonic Crenarchaeota: changes with depth in the South China sea. Microb Ecol. 2011;62:549–63.
Mosier AC, Francis CA. Determining the distribution of marine and coastal ammonia-oxidizing archaea and bacteria using a quantitative approach. In: Martin G. Klotz (ed.) Research on Nitrification and Related Processes, Part A, vol. 486. Elsevier: Methods in Enzymology; 2011, pp. 205–21.
Smith JM, Casciotti KL, Chavez FP, Francis CA. Differential contributions of archaeal ammonia oxidizer ecotypes to nitrification in coastal surface waters. ISME J. 2014;8:1704–14.
Smith JM, Damashek J, Chavez FP, Francis CA. Factors influencing nitrification rates and the abundance and transcriptional activity of ammonia‐oxidizing microorganisms in the dark northeast Pacific Ocean. Limnol Oceanogr. 2016;61:596–609.
Luo H, Sun Y, Hollibaugh JT, Moran MA. Low genome content diversity of marine planktonic Thaumarchaeota. Environ Microbiol Rep. 2016;8:501–7.
Santoro AE, Saito MA, Goepfert TJ, Lamborg CH, Dupont CL, DiTullio GR. Thaumarchaeal ecotype distributions across the equatorial Pacific Ocean and their potential roles in nitrification and sinking flux attenuation. Limnol Oceanogr. 2017;62:1984–2003.
Müller O, Wilson B, Paulsen ML, Rumińska A, Armo HR, Bratbak G, et al. Spatiotemporal dynamics of ammonia-oxidizing Thaumarchaeota in distinct Arctic water masses. Front Microbiol. 2018;9:24.
Luo H, Tolar BB, Swan BK, Zhang CL, Stepanauskas R, Moran MA, et al. Single-cell genomics shedding light on marine Thaumarchaeota diversification. ISME J. 2014;8:732–6.
Merbt SN, Stahl DA, Casamayor EO, Martí E, Nicol GW, Prosser JI. Differential photoinhibition of bacterial and archaeal ammonia oxidation. FEMS Microbiol Lett. 2012;327:41–46.
Tolar BB, Powers LC, Miller WL, Wallsgrove NJ, Popp BN, Hollibaugh JT. Ammonia oxidation in the ocean can be inhibited by nanomolar concentrations of hydrogen peroxide. Front Mar Sci. 2016;3:499.
Sintes E, Bergauer K, De Corte D, Yokokawa T, Herndl GJ. Archaeal amoA gene diversity points to distinct biogeography of ammonia-oxidizing Crenarchaeota in the ocean. Enviro Microbiol. 2013;15:1647–58.
Faust K, Sathirapongsasuti JF, Izard J, Segata N, Gevers D, Raes J, et al. Microbial co-occurrence relationships in the human microbiome. PLoS Comp Biol. 2012;8:e1002606.
Layeghifard M, Hwang DM, Guttman DS. Disentangling interactions in the microbiome: a network perspective. Trends Microbiol. 2017;25:217–28.
He S, Guo L, Niu M, Miao F, Jiao S, Hu T, et al. Ecological diversity and co-occurrence patterns of bacterial community through soil profile in response to long-term switchgrass cultivation. Sci Rep. 2017;7:3608.
Fuhrman JA, Steele JA. Community structure of marine bacterioplankton: patterns, networks, and relationships to function. Aquat Microb Ecol. 2008;53:69–81.
Steele JA, Countway PD, Xia L, Vigil PD, Beman JM, Kim DY, et al. Marine bacterial, archaeal and protistan association networks reveal ecological linkages. ISME J. 2011;5:1414–25.
Eiler A, Heinrich F, Bertilsson S. Coherent dynamics and association networks among lake bacterioplankton taxa. ISME J. 2012;6:330–42.
Chow C-ET, Sachdeva R, Cram JA, Steele JA, Needham DM, Patel A, et al. Temporal variability and coherence of euphotic zone bacterial communities over a decade in the Southern California Bight. ISME J. 2013;7:2259–73.
Vick-Majors TJ, Priscu JC, Amaral-Zettler LA. Modular community structure suggests metabolic plasticity during the transition to polar night in ice-covered Antarctic lakes. ISME J. 2013;8:778–89.
Cram JA, Xia LC, Needham DM, Sachdeva R, Sun F, Fuhrman JA. Cross-depth analysis of marine bacterial networks suggests downward propagation of temporal changes. ISME J. 2015;9:2573–86.
Baldassano SN, Bassett DS. Topological distortion and reorganized modular structure of gut microbial co-occurrence networks in inflammatory bowel disease. Sci Rep. 2016;6:26087.
Tapio I, Fischer D, Blasco L, Tapio M, Wallace RJ, Bayat AR, et al. Taxon abundance, diversity, co-occurrence and network analysis of the ruminal microbiota in response to dietary changes in dairy cows. Smidt H, editor. PLoS ONE. 2017;12:e0180260.
Parada AE, Fuhrman JA. Marine archaeal dynamics and interactions with the microbial community over 5 years from surface to seafloor. ISME J. 2017;11:2510–25.
Daebeler A, Bodelier PL, Yan Z, Hefting MM, Jia Z, Laanbroek HJ. Interactions between Thaumarchaea, Nitrospira and methanotrophs modulate autotrophic nitrification in volcanic grassland soil. ISME J. 2014;8:2397–410.
Xiao Y, Angulo MT, Friedman J, Waldor MK, Weiss ST, Liu Y-Y. Mapping the ecological networks of microbial communities. Nat Comm. 2017;8:2042.
Timothy Pennington J, Chavez FP. Seasonal fluctuations of temperature, salinity, nitrate, chlorophyll and primary production at station H3/M1 over 1989-1996 in Monterey Bay, California. Deep Sea Res Part 2. 2000;47:947–73.
Smith JM, Chavez FP, Francis CA. Ammonium uptake by phytoplankton regulates nitrification in the sunlit ocean. PLoS ONE. 2014;9:e108173.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2012;41:D590–6. https://doi.org/10.1093/nar/gks1219
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics. 2011;27:2194–200.
R CT. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2017.
McMurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE. 2013;8:e61217.
Oksanen J, Kindt R, Legendre P, O'Hara B, Simpson GL, Solymos P et al. (2008), vegan: Community Ecology Package. R package version 2.5-2.
Kurtz ZD, Müller CL, Miraldi ER, Littman DR, Blaser MJ, Bonneau RA. Sparse and compositionally robust inference of microbial ecological networks. PLoS Comp Biol. 2015;11:e1004226.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard N-U, et al. Community genomics among stratified microbial assemblages in the ocean’s interior. Science. 2006;311:496–503.
Brown MV, Philip GK, Bunge JA, Smith MC, Bissett A, Lauro FM, et al. Microbial community structure in the North Pacific ocean. ISME J. 2009;3:1374–86.
Treusch AH, Vergin KL, Finlay LA, Donatz MG, Burton RM, Carlson CA, et al. Seasonality and vertical structure of microbial communities in an ocean gyre. ISME J. 2009;3:1148–63.
Gilbert JA, Steele JA, Caporaso JG, Steinbrück L, Reeder J, Ben T, et al. Defining seasonal marine microbial community dynamics. ISME J. 2012;6:298–308.
Ghiglione J-F, Galand PE, Pommier T, Pedrós-Alió C, Maas EW, Bakker K, et al. Pole-to-pole biogeography of surface and deep marine bacterial communities. PNAS. 2012;109:17633–88.
Hatosy SM, Martiny JBH, Sachdeva R, Steele J, Fuhrman JA, Martiny AC. Beta diversity of marine bacteria depends on temporal scale. Ecology. 2013;94:1898–904.
Walsh EA, Kirkpatrick JB, Rutherford SD, Smith DC, Sogin M, D’Hondt S. Bacterial diversity and community composition from seasurface to subseafloor. ISME J. 2015;10:978–89.
Yaveroğlu ÖN, Malod-Dognin N, Davis D, Levnajic Z, Janjic V, Karapandza R, et al. Revealing the hidden language of complex networks. Sci Rep. 2014;4:04547.
Peura S, Bertilsson S, Jones RI, Eiler A. Resistant microbial cooccurrence patterns inferred by network topology. Appl Environ Microbiol. 2015;81:2090–97.
Williams RJ, Howe A, Hofmockel KS. Demonstrating microbial co-occurrence pattern analyses within and between ecosystems. Front Microbiol. 2014;5:358.
Fillol M, Auguet J-C, Casamayor EO, Borrego CM. Insights in the ecology and evolutionary history of the Miscellaneous Crenarchaeotic Group lineage. ISME J. 2015;10:665–77.
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE. Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem. 2016;97:188–98.
van der Heijden MGA, Hartmann M. Networking in the plant microbiome. PLoS Biol. 2016;14:e1002378.
Hawley AK, Nobu MK, Wright JJ, Durno WE, Morgan-Lang C, Sage B, et al. Diverse marinimicrobia bacteria may mediate coupled biogeochemical cycles along eco-thermodynamic gradients. Nat Comm. 2017;8:1507.
Santoro AE, Buchwald C, McIlvin MR, Casciotti KL. Isotopic signature of N2O produced by marine ammonia-oxidizing archaea. Science. 2011;333:1282–5.
Martens-Habbena W, Qin W, Horak REA, Urakawa H, Schauer AJ, Moffett JW, et al. The production of nitric oxide by marine ammonia‐oxidizing archaea and inhibition of archaeal ammonia oxidation by a nitric oxide scavenger. Environ Microbiol. 2015;17:2261–74.
Kozlowski JA, Kits KD, Stein LY. Comparison of nitrogen oxide metabolism among diverse ammonia-oxidizing bacteria. Front Microbiol. 2016;7:1703.
Fernández-Gómez B, Richter M, Schüler M, Pinhassi J, Acinas SG, González JM, et al. Ecology of marine bacteroidetes: a comparative genomics approach. ISME J. 2013;7:1026–37.
Bergauer K, Fernandez-Guerra A, Garcia JAL, Sprenger RR, Stepanauskas R, Pachiadaki MG, et al. Organic matter processing by microbial communities throughout the Atlantic water column as revealed by metaproteomics. PNAS. 2017;57:1708779115.
Beman JM, Popp BN, Francis CA. Molecular and biogeochemical evidence for ammonia oxidation by marine Crenarchaeota in the Gulf of California. ISME J. 2008;2:429–41.
Fuhrman JA. Microbial community structure and its functional implications. Nature. 2009;459:193–9.
Pachiadaki MG, Sintes E, Bergauer K, Brown JM, Record NR, Swan BK, et al. Major role of nitrite-oxidizing bacteria in dark ocean carbon fixation. Science. 2017;358:1046–51.
Barberán A, Bates ST, Casamayor EO, Fierer N. Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 2012;6:343–51.
Ngugi DK, Blom J, Stepanauskas R, Stingl U. Diversification and niche adaptations of Nitrospina-like bacteria in the polyextreme interfaces of Red Sea brines. ISME J. 2016;10:1383–99.
Acknowledgments
We thank Tim Pennington, Marguerite Blum, and other members of the Biological Ocean Group at MBARI for intellectual and logistical support. We also thank the captain & crew of the RV Western Flyer and RV Rachel Carson (MBARI) for support in sample acquisition efforts at sea. Jessica Lee, Emily Cardarelli and Madison Jackson assisted with sample collection; additional assistance with sample processing was done by Tynan Challenor. Sequencing support came from a DOE JGI CSP project to CAF. JGI project manager Tijana Glavina del Rio facilitated sample submission and data acquisition. Field research was supported (in part) by the grant OCE-1357024 from NSF Biological Oceanography (to CAF), an ARCS Foundation Graduate Research Fellowship and an MBARI Postdoctoral Fellowship (to JMS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Reji, L., Tolar, B.B., Smith, J.M. et al. Differential co-occurrence relationships shaping ecotype diversification within Thaumarchaeota populations in the coastal ocean water column. ISME J 13, 1144–1158 (2019). https://doi.org/10.1038/s41396-018-0311-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0311-x
This article is cited by
-
Elevated hydrostatic pressure enhances the potential for microbially mediated carbon sequestration at the sediment–water interface in a deep-water reservoir by modulating functional genes and metabolic pathways
Carbon Research (2024)
-
Distinct patterns of distribution, community assembly and cross-domain co-occurrence of planktonic archaea in four major estuaries of China
Environmental Microbiome (2023)
-
Network analysis of 16S rRNA sequences suggests microbial keystone taxa contribute to marine N2O cycling
Communications Biology (2023)
-
Phylotype resolved spatial variation and association patterns of planktonic Thaumarchaeota in eastern Chinese marginal seas
Marine Life Science & Technology (2023)
-
Dynamics of actively dividing prokaryotes in the western Mediterranean Sea
Scientific Reports (2022)