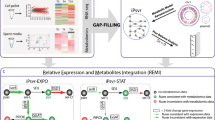
Abstract
Microbial communities play a vital role in biogeochemical cycles, allowing the biodegradation of a wide range of pollutants. The composition of the community and the interactions between its members affect degradation rate and determine the identity of the final products. Here, we demonstrate the application of sequencing technologies and metabolic modeling approaches towards enhancing biodegradation of atrazine—a herbicide causing environmental pollution. Treatment of agriculture soil with atrazine is shown to induce significant changes in community structure and functional performances. Genome-scale metabolic models were constructed for Arthrobacter, the atrazine degrader, and four other non-atrazine degrading species whose relative abundance in soil was changed following exposure to the herbicide. By modeling community function we show that consortia including the direct degrader and non-degrader differentially abundant species perform better than Arthrobacter alone. Simulations predict that growth/degradation enhancement is derived by metabolic exchanges between community members. Based on simulations we designed endogenous consortia optimized for enhanced degradation whose performances were validated in vitro and biostimulation strategies that were tested in pot experiments. Overall, our analysis demonstrates that understanding community function in its wider context, beyond the single direct degrader perspective, promotes the design of biostimulation strategies.
Similar content being viewed by others
Introduction
Microorganisms in nature co-exit as communities. Members in microbial communities may interact bidirectionally or unidirectionally, leading to synergism, commensalism, mutualism, parasitism or competition among them [1, 2]. Such interactions are in many cases derived by metabolism—food chains, substrate competition, syntrophy, and waste product inhibition [3], playing important roles in almost all processes occurring on this planet. Examples include element cycling [4], biodegradation of pollutants [5,6,7,8], food fermentation [9, 10] and many other biogeochemical processes essential for eco-system sustainability and human health [11, 12]. Despite vital impacts of microbial function, design principles that rationally promote specific metabolic activities towards optimized performances have still remained largely obscure.
In soil, the efficiency of community function is critical to processes of environmental interest such as bioremediation—the breakdown of pollutants through microbial metabolism. Key bioremediation strategies include bioaugmentation—the optimization of microbial composition for bioremediation; and biostimulation—the optimization of environmental conditions. Bioaugmentation practices involve the addition of living cells capable of degradation and biostimulation practices involve the addition of limiting nutrients to stimulate microbial growth [13]. To date, both bioaugmentation and biostimulation strategies are mostly driven by intuition and rely on experience derived from trial and error experiments [14]. Typically, enrichment cultures from samples exposed to a specific pollutant lead to the isolation of consortia with pollutant degrading ability. Bioaugmentation practices based on returning such consortia to soil often fail, as myriad of biotic and abiotic factors influence the effectiveness of the treatment and in many cases the consortia are not successful when re-introduced to soil [15, 16]. Understanding the required conditions for natural enhancement of desired endogenous consortia can enhance the rate of success of biostimulation based bioremediation treatments. New sequencing technologies allow now revealing the dynamics of community shifts and together with modeling approaches lay foundations for the educated design of community function [17]. Progress in sequencing technologies promotes the description of the bio-diversity and metabolic activity of microorganisms in ecological niches [18,19,20,21]. Parallel advancement of computational tools such as Genome-scale metabolic models (GSMM) and respective simulation algorithms such as Flux Balance Analysis (FBA) further enable in silico analysis of microbial interactions [19, 20, 22, 23]. By modeling of microbial interactions in a community, the metabolic features could be simulated and optimal solutions can be predicted. Notably, natural communities are not necessarily optimized to provide environmental solutions; hence the educated directing towards pre-defined functions is of potential to induce significant change in efficiency.
Here, we applied sequence-based modeling approach to explore the role of microbial function in the degradation of the atrazine. The s-triazine herbicide atrazine is one of the world’s most heavily applied herbicides, in particular used on crops such as maize, sugarcane and sorghum. Atrazine and its degradation products can be found in the soil for decades after application [24, 25], often leading to widespread contamination of both water and soils [26, 27]. Even at very low concentrations atrazine may act as endocrine-disrupting chemical in frogs [28], among other effects leading to sexual transformation [29, 30]. In human, the main target of atrazine is the endocrine system together with considerable evidence supporting its damaging effects on the central nervous system, reproductive system, immune system and cardiovascular function [31]. Given the widespread dispersal and long persistence of atrazine, its removal from polluted sites is crucial for a safe and chemical free environment.
Is soil, microbial communities take central part in atrazine degradation [32,33,34]. In many cases, the degradation involved a consortium rather than a single species. For example, a consortium containing Klebsiella sp. A1 and Comamonas sp. A2 showed very high atrazine-mineralizing efficiency as strain A2 metabolize N-ethylammelide, a product of atrazine degradation produced by strain A1 whose degrading activity was suppressed by N-ethylammelide [34]. In another consortium, Clavibacter michiganese ATZ1 and Pseudomonas sp. CN1 collectively mineralized atrazine with a much higher degradation rate than did C. michiganese alone [32]. These examples show that, to a large extent, the structure of communities determines their functions, and the composition of communities affects both the rate of the degradation process and the identity of the final products.
Here, we used sequence-based information on the dynamics in soil community from atrazine-treated fields in order to construct a corresponding imputed in silico community, and used simulations for exploring functional significance of community dynamics and predicting possible biostimulation strategies. First we used high-throughput sequencing approaches for describing the structure of soil communities exposed to atrazine treatment, showing that atrazine treatments alter the microbial communities and that different microbial communities are associated with different functional performances. Then, GSMM of five species, chosen based on the analysis of the high-throughput data from the respective soil samples, were constructed and manually curated. Using these GSMM we were able to model community function. Simulations of the performances of alternative community combinations predict variations in atrazine degradation efficiencies and were correlated with the observed activity profile. The simulations provided functional interpretation for observed co-occurrence patterns. The functional interpretation provided a basis for the educated design of optimized consortia and biostimulation strategies that were experimentally tested.
Methods
Field experiment and soil sampling
The field experiment was conducted in Newe Ya’ar Research Center, Israel (32°42’N, 35°11’E). The soil and climate types and properties are described in Supplementary Materials. The plot was irrigated by 300 m3 ha−1, cultivated (by rototiller) to a depth of 12 cm, maize (var. Royalty) was sown, atrazine (at rate of 500 active ingredients per hectare, a.i. ha−1) was applied at 21.6.2015, using a motorized sprayer equipped with Tee Jet 8001E nozzles (Spraying Systems Co., Wheaton, IL, USA) operated at a pressure of 300 kPa and delivering a spray volume of 200 l ha−1. After herbicide application, the field was irrigated by 300 m3 ha−1. Ten sub-plots at size of 20 m2 were used for soil sampling; five atrazine treated and five non-treated used as a control. Beginning at sowing date and germinating irrigation, and at seven days interval up to 77 days from sowing, 500 g soil was sampled from each plot (total of ten samples at a time) and kept in dark under dry conditions at 4 °C until use. Soil samples were classified into two sub-samples; 400 g were taken for bioassay (all sampling dates); tubes were filled soil for enrichment culture (volume of 30 ml) and sequencing analysis (volume of 40 ml), respectively.
Determining atrazine degradation in soil using a bioassay
The design of bioassay experiments followed the experiments designed in Onofri et al. [35]. Briefly, the bioassays monitor the growth performances of atrazine-sensitive plant. Here, wheat (cv. “Jordan”) was used as the reporter plant based on a demonstrated dose dependent sensitivity of shoot development performance (biomass and height) to atrazine concentration in soil (Supplementary Figure 1), growth performances are hence indicative of atrazine level in soil [36]. Two bioassay experiments were carried: (i) estimating atrazine degradation in samples delivered to pots from treated vs. non treated field in seven days intervals (as described above, Supplementary Figure 2); and (ii) estimating atrazine degradation following soil amendment treatments in pots with soil from a non-herbicide treated field. The soil amendments include combinations of atrazine and glucose at different doses. Experiments were carried in replicates of five pots (0.5 l), 10 seeds sown in each. In experiment ii (soil amendments), the soil was mixed with glucose at dose of 5, 10, 15 g/Kg using a cement machine (Shatal, 150 l), delivered into pots and sprayed with atrazine (500 g a.i. ha−1) on soil surface. Herbicides were applied using a motorized laboratory sprayer equipped with a flat fan nozzle (8001E, Degania Sprayers Co. Ltd., Degania Bet, Israel) calibrated to deliver 300 l ha−1 at 245 kPa, as described by Eizenberg et al. [37]. Glucose free pots non-atrazine treated served as control. In both experiments, pots were irrigated as needed by sprinklers. Twenty days after planting, wheat height was measured, shoots were cut and dry biomass (60 °C for 72 h) was measured.
All experiments were arranged in a completely randomized design. Effect of herbicide phytotoxicity was computed by one-way ANOVA. Means were compared by Tukey–Kramer honestly significant difference test (a, 0.05) using JMP software (vers. 7, SAS). Non-linear regressions were computed using SigmaPlot version 11.01 (SPSS Inc., Chicago, IL, USA).
Determining atrazine degradation in enrichment cultures using HPLC and identification of atrazine-degrading isolates
Ten gram of soil from the respective soil was suspended in R medium (based on ATCC 2662 R) containing glucose as carbon source and atrazine (30 mg/l) as nitrogen source (allowing maximal solubility of atrazine in water at room temperature). The soil was shaken (150 rpm) and incubated at 25 °C for 7 weeks. Control (autoclaved soils) was incubated as well. Periodically samples were taken from 0 to 7 weeks after incubation for atrazine residues analysis. Atrazine in filtered (0.20 µm) supernatant of the enrichment cultures was determined by Agilent 1100 HPLC (Waldbronn Germany) equipped with a DAD detector. Samples (20 µl) were separated on a Kinetex C18 column (Phenomenex Torrance, CA) and the mobile phase consisted of 70% methanol and 30% water flowing at a 1 ml/min. Detection and quantitation of atrazine was done at 240 nm by the external standard that was linear between 0.15 to 30 mg/l.
Isolation assays were carried by collecting serial dilutions of the suspension of the enrichment culture using 0.85% NaCl and spreading them on solid R-media plates amended with glucose and atrazine (30 mg/l). Plates were incubated at 30 °C for 48 h and single colonies were picked based on distinct colony morphology and sub cultured on ATZ-R media plates to obtain pure isolates. Resulting isolates were grown in 10 ml R-medium containing atrazine (30 mg/l) as sole nitrogen source and incubated at 30 °C with shaking. Culture suspensions were extracted in methanol to screen for degradation potential (measuring atrazine concentrations by HPLC) leading to the identification of two degrader-isolates. The taxonomic characterization of the isolates is described in Supplementary Table 1.
Amplicon sequencing and data analysis
In total, 30 and 32 sequencing libraries were constructed for soil and enrichment samples, respectively. The details of sequencing library construction are described in Supplementary Materials. Sequences were deposited in SRA (SUB2541893). After filtration (Supplementary Materials), the operational taxonomic units (OTUs) were analyzed using the UPARSE pipeline [38]. Sequences were assigned to OTUs at 97% identity. The RDP classifier was used for picking representative sequences for each OTU and to assign taxonomic data to each representative sequence at the 70% threshold [39]. Rarefaction curves were analyzed using the QIIME pipeline [40]. The Chao1 richness index, Shannon diversity index, Simpson diversity index, and nonmetric multidimensional scaling (NMDS) ordination based on Rho similarities were performed using PAST software [41]. OTUs were sorted according to the ratio of reads between treated and untreated soil samples at different time points. Significance of differential abundance distribution pattern was tested using the group significance function in QIIME (Supplementary Table 2).
Reconstruction of single species metabolic network models
Five species were selected for model construction based on differential abundance pattern and isolation tests (Supplementary Tables 1 & 2). Genome sequences representing the respective species were selected based on blast search screening and respective sequences were retrieved from public resources. In cases where highly-scored hits were retrieved from several species, the closest species for which genome sequence was available were selected based on phylogenetic relatedness as inferred from a 16S rRNA based phylogenetic tree (Supplementary Figure 3). Differentially abundant OTUs and the respective genome sequences used for constructing the respective metabolic networks are listed in Table 1. Model SEED was used for constructing the initial draft metabolic models from the genome sequence data [42]. The RAST annotation algorithm was used for structural and functional annotation [43]. The genome was then imported into Kbase (www.kbase.us) and a draft metabolic model was constructed.
After having a working draft model (that is a biomass flux > 0 when all exchange reactions are open), each of the models were manually curated according to literature and other available resources such as KEGG [44], UniProt [45], and JGI [46] to ensure that it captures the biochemical and physiological knowledge available. Overall, manual curation processes involved (i) the addition of new reactions based on the literature and the additional annotations schemes; (ii) standardization - all reactions’ Ids from the different databases were converted to KBASE rxn conventions and validation of reaction reversibility and balance; and (iii) removal of futile loops. Curation procedures were carried iteratively vs. growth simulation ensuring that the reconstruction version was able to produce all biomass components in minimal mineral media (MMM: K+, Mn2+, CO2, Zn2+, SO42−, Cu2+, Ca2+, HPO42−, Mg2+, Fe2+, Cl−) with alternative C and N sources. The different C and N sources that the species can grow on were determined based on literature reporting performances of the selected species [47,48,49,50]. The final GSMM were consistent with the experimental knowledge on the nutrients required for culturing each species.
The GSMM of each species (k) was represented in a mathematic format in a stoichiometric matrix (Sk). In the assumed pseudo-steady-state, the model can be represented as: S × v = 0, where vector v signifies the reaction flux. Flux balance analysis (FBA) was used for predicting activity. The objective of the optimization was maximizing the flux thorough the biomass objective function. Constrains for the optimization include upper and lower bounds (LBk and UBk) for selected uptake reaction fluxes [51, 52]. FBA was performed using COBRAToolbox-2.0 [53] in MATLAB, with GLPK as the linear programming solver.
Community network model construction
The five single species models were combined into a single dynamic model and analyzed following conventions described for COMETS [54] with some revisions detailed below. Briefly, like COMETS, our algorithm of community dynamic modeling uses dynamic flux balance analysis (dFBA) for simulating the growth of multiple species in a given media across time. The model is updated after each time tick. The amount of biomass of each species is changed after each time tick based on the biomass reaction flux of the given species in that time tick; uptaken metabolites are then removed from the media and secreted metabolites are added to it. Whereas COMETS simulations occur on a spatially structured lattice of interacting metabolic subsystems (“boxes”) [14], our simulations assume a simplistic single-box world with no spatial differentiation and equal accessibility of all organisms to all nutritional sources. This simplification reflects our lack of knowledge of the conditions in soil and corresponds to conditions in in vitro experiments. As in [14], we define the initial conditions of the community dynamic model by setting initial concentrations (at time 0) of each substrate k (Xk(0)) in the media and the setting of the initial biomass of each species (Bkbiomass). Each reaction is limited by lower and upper bounds (LB and UB respectively), set to −1 mmol/(gDW*hr), respectively.
The total time period of each simulation cycle was subdivided into predefined discrete time intervals (Δt). For each time interval, the following three steps were executed:
(1) the substrates were divided equally between the species based on the relative abundance of each species (biomass). The amount of metabolites that can be uptaken by each species in a given time tick follows:
Where:
LBki (t+1) is the amount of metabolite (k) that can be uptaken (uptaken values are negatives) at time (t+1) by species (i)
Xki (t) is the amount of metabolite Xk for species (i) at time tick (t)
Bibiomass (t) is the amount of biomass of species(i) in time-tick (t)
Xtotal biomass defines the sum of Xkbiomass of all the species,
UBki (t) is the amount of metabolite (k) that can be secreted (secreted values are positives) at time (t). This value was set to be 1000.
(2) At each time point we optimized the biomass flux for each species using the standard FBA optimization:
(3) Following each time tick, media uptake bounds and species biomass are updated to reflect secretions and uptakes & biomass fluxes.
Where:
Xk (t+Δt) is the amount of metabolite (k) available in the media at time (t+Δt)
V [ex]ik is the flux of the exchange reaction for metabolite (k) in the metabolic model of species (i),
This value can be negative (uptaken) or positive (secreted)
Where:
Bibiomass (t+Δt) is the amount of biomass of species (i) at time (t+Δt)
Vbiomassi (t) is the flux of the biomass reaction at time (t)
The new concentrations were then used as a starting point for the next iteration. Simulations assumed an equal initial biomass for each species (Xkbiomass = 1), aimed at gaining a qualitative prediction for the nature of interaction (e.g., enhancing vs. repressing degradation), rather the quantitative description of activity in soil.
Co-culture experiments and metabolomics profiling
The isolates Arthrobacter sp. AT5 (MG763151), Halomonas sp. N8 (MG763150), and Halobacillus sp. NY15 (MG763149) from the Agricultural Culture Collection of China (ACCC) were used for testing experimentally the computational predictions. In vitro experiments were designed to correspond to simulation conditions considering growth media and species ratio. Atrazine degradation carried by two- (combination of N8 or NY15 with AT5, 1:1) and three-member consortia (combination of N8, NY15, and AT5, 1:1:1) were measured. In addition to combinations based on species selected according to differential abundance following atrazine application, two arbitrary bacterial species were used for constructing reference consortia: Mycobacterium sp. S8 (MG763148), a soil species whose abundance in soil is not effected by atrazine, and the model microorganism Escherichia coli. The isolate S8 was abundant in soils (>6.5% on average), representing endogenous species that is not effected by atrazine treatments. The two negative controls were combinations of AT5 and S8 (1:1), and AT5 and E. coli (1:1). The similarities between 16s of the OTUs and the used isolates were 97%, 98%, 99%, and 98% for AT5, N8, NY15, and S8 respectively. All strains were grown in Luria-Bertani (LB) medium at 30 °C. To measure the atrazine degradation, the single strain and consortia were separately grown in R media supplemented with atrazine or both atrazine and glucose for 36 h in 50 ml flasks. In addition R media were supplemented with predicted exchanged metabolites (EM): for NY15 R medium was supplemented with atrazine, aminoethanol, ethylamine and hypoxanthine (EM1); for N8 R medium was supplemented with atrazine, aminoethanol, ethylamine, hypoxanthine and urea (EM2); for AT5 R medium was supplemented with atrazine, leucine and NH4 (EM3). As in simulations, all compounds were added in equal concentration (30 mg/l). The concentration of atrazine in each culture was measured every 6 h by HPLC as described above. All treatments were carried out in triplicates.
To test secretion of EM by the three species, consortia (combination of AT5, NY15, and N8) were initially grown in R media supplemented with atrazine as a sole carbon and nitrogen sources for 30 h, and then we screened for the hypothesized EM by liquid chromatography–mass spectrometry (LC–MS). Pure compounds, including aminnethanol, ethylamine, glycerol, urea, hypoxanthine, leucine, and mannitol were used as reference standards. To facilitate the separation of the small molecules (including aminnethanol, ethylamine, glycerol, and urea) by liquid chromatography, a derivatization step was performed before analysis. A C18 reverse phase column (2.1 × 100 mm, 1.7 μm particles, ACQUITY UPLC BEH, WATERS) was used for liquid chromatography. Mass spectrometry was performed using electrospray ionization in positive or negative ion mode with MSe acquisition mode, with a selected mass range of 50–1200 m/z. The details of derivatization and LC-MS analysis are described in Supplementary Materials.
Results
Functional assays for biodegradation activity following introduction of atrazine to agriculture soil
To study the role of bacterial communities in atrazine biodegradation in agricultural land, corn fields were treated with atrazine. Based on a bioassay, we detected a 90%, 50%, and 0 decrease in phytotoxicity following 35, 49, and 77 days from treatment, respectively (Supplementary Figure 2). To further determine biodegradation activity and community structure, soil samples from 0, 49, and 77 days from atrazine application were taken for direct degradation assay and bacterial community analysis.
For degradation assays, soil samples were incubated in an atrazine containing R medium. Rapid atrazine degradation was detected only in samples taken from the atrazine-treated soil (T49 & T77 in Fig. 1). In comparison, no degradation or minor degradation was observed in samples from time 0 (T0), samples from untreated soil (U0, U49, U77), or autoclaved samples (U and T control), precluding the possible contribution of abiotic reaction to the degradation of atrazine.
Atrazine degradation in enrichment cultured samples taken from agricultural soil. T, soil treated with atrazine; U, untreated soil (control); T control, autoclaved treated soil of T0; U control, autoclaved untreated soil of U0; 0/49/77, soil samples collected 0/49/77 days after atrazine applied in the field experiment. X axis show time of inoculation in minimal R medium. The results are average of duplicates. ANOVA of the regression, coefficients and significance of log logistic regression for T0, T49, and T77 are provided in Supplementary Table 5. All three regressions are significantly different according to F test (P value < 0.01). For all other treatments, regression coefficients and ANOVA were not significant and were omitted from the table
The degradation assay together with the bioassay point at enhanced biodegradation activity in atrazine-treated soil. The application of atrazine to soil followed the common agricultural practices. Despite the relatively low amounts and the complexity of agricultural soil, both analyses suggest that the application of atrazine triggered functional modification in the soil community associated with increased degradation rate. We next examined whether a corresponding change in community structure can be detected.
Bacterial community dynamics and structure
Community structure in the atrazine-treated and untreated soil was determined using Illumina MiSeq (Supplementary Materials). Multidimensional scaling of community similarity points at a clear separation between treated and untreated soil samples in both soil and enrichment samples (Fig. 2). Unlike samples from untreated soils, samples from treated soil diverge with time and show a growing distance from untreated soil, or treated soil at day 0. The analysis points at a shift in community structure that corresponds with the shift in community function, induced by the application of atrazine.
To identify species that are effected by the application of atrazine, we screened for OTUs whose abundance differentiated following treatments. The three OTUs with the highest fold-change following atrazine application were classified as Halobacillus_unclutrued, Bacillus decolorationis, and Cesiribacter sp. JJ021 (Supplementary Table 2). These three species were barely detected in T0 and in any of the untreated soil samples (Supplementary Table 3). Surprisingly, none of the ten OTUs with highest fold-change was assigned to a genus that includes species with atrazine degradation activity. In parallel to the genomic survey (16S rRNA analysis), we carried independent isolation screens for atrazine degraders from the respective atrazine-treated field samples resulting in the identification of two isolates classified as Sinorhizobium and Arthrobacter that were confirmed as atrazine degraders (Supplementary Table 1). OTU that can potentially represent the Sinorhizobium isolate (identity in 16S rRNA sequences > 97%) was detected in relatively low and stable abundances across all time points and do not show a quantitative response to the application of atrazine (Supplementary Table 1). The OTU with the highest sequence similarity to the 16S rRNA of the Arthrobacter degrader-isolate (99% identity, Supplementary Table 1) is OTU2327. OTU2327 was detected in a relatively high abundance (~6-15 times more abundant in atrazine-treated samples in comparison to OTU representing the other isolate with degradation activity, Supplementary Table 1), supporting a potential role in the degradation of atrazine in the soil samples. OTU2327 showed an approximately two fold increase in abundance following exposure to atrazine (P value 0.065, below significance threshold, Supplementary Table 2).
In order to explore functional modification in microbial activity in the respective soil samples following atrazine application we chose to focus on five species including the potential atrazine degrader Arthrobacter_unclassified (OTU2327), three species whose respective OTUs show significant increase in abundance following atrazine application— Halobacillus_unclutrued, Bacillus decolorationis, and Cesiribacter sp. JJ021 and Halomonas stevensii—a species whose respective OTU (OTU1826) demonstrates a reverse pattern—a significant decrease following atrazine treatment.
Predicting performances of different bacterial combinations in atrazine only vs. glucose supplemented media
GSMM were constructed for the five species whose partial abundance was affected following atrazine application (Table 1). In order to compare atrazine degradation efficiencies of different consortia, we constructed 16 multi-species models representing all possible combinations that contain Arthrobacter, the only atrazine degrader (Supplementary Table 4). Performances were simulated using constraint-based modeling (CBM). A single species of Arthrobacter was used as a reference point, allowing predicting positive or negative contribution of species interactions.
For each combination we initially measured two parameters of community performances: total biomass and atrazine degradation efficiency (Fig. 3). Simulations were carried in a medium containing atrazine as a sole nitrogen and carbon source and in a medium containing an equal amount of atrazine (acting as a sole nitrogen source) supplemented by glucose as the major carbon source. As expected, supplementing the medium with glucose, in comparison to atrazine only medium, improved performances—in the rich-nutritional medium (glucose containing) total biomass increased in 2–5 fold and atrazine degradation was about twofold faster. Based on simulation results we repeated pot bioassays while supplementing the soil in glucose in different concentrations. In support of the simulations, we find that in high glucose concentrations, atrazine degradation in soil was expedited in comparison to non-treated control (Fig. 3d).
Simulations and experimental validations of atrazine degradation and bacterial growth performances. a–c Performances of bacterial combinations. Line colors are indicative of the combinations as indexed on the right grid (grey/white cells indicate species included/not included in the combination, respectively. I – 16 combinations formed by species represented by GSMM. All combinations include Arthrobacter aurescens TC1 (atrazine degrader) together with all possible combinations of the species modeled (chosen based on differential abundance). Red, A. aurescens TC1 only; Blue, combinations with Halobacillus sp. BAB-2008 but without Halomonas stevensii S18214; Orange, combinations with both H. sp. BAB-2008 and H. stevensii S18214; Green, combinations with H. stevensii S18214 but without H. sp. BAB-2008, Dark grey, combinations without H. sp. BAB-2008 or H. stevensii S18214. II, combinations used as control in the experimental validation. Purple, A. aurescens TC1 (atrazine degrader) with an exogenous strain (Escherichia coli) or an endogenous strains whose abundance was not affected by the application of atrazine (Mycobacterium sp. S8). Light grey—Halobacillus sp. NY15 and Halomonas sp. N8 without A. aurescens AT5 (atrazine degrader); Black—autoclaved strains. In both simulations (a, b) and in vitro experiments (c) performances were tested in two media: medium that contains atrazine as the sole carbon and nitrogen source (left) and medium with atrazine and glucose as carbon and nitrogen sources (right). a Predicted relative increase in biomass (1/h). The relative increase values indicate biomass(t)/biomass(0) -biomass(0). b Predicted atrazine degradation (mmol/gDW) due to degradation activity. c In vitro decrease in atrazine (mg/l). d Experimental validation of simulations by pot experiments following supplementing the soil with glucose. Fraction of atrazine left was estimated according to a bioassay (Supplementary Figure 1). T1, no atrazine; T2, atrazine only; T3, atrazine and 0.5% glucose; T4, atrazine and 1.0% glucose; T5, atrazine and 1.5% glucose. Different letters indicated statistically significant differences (P < 0.05) according to the Tukey test. e In vitro atrazine degradation in media with atrazine and supplemented/not supplemented with glucose (+glucose/-glucose, respectively). C1, AT5 only; C2, combinations of AT5 and S8; C3, combinations of AT5 and E. coli; C4, combinations of AT5 and NY15; C5, combinations of AT5 and N8; C6, combinations of AT5, NY15 and S8
Based on simulations results, we compared the performances of different species combinations (Fig. 3). In both media types, variations between combinations were recorded for both performance parameters. The reference community— containing only the atrazine degrader Arthrobacter (red, Fig. 3a, b), had the worst performances considering both growth (community biomass) and atrazine degradation. Multi-species combinations improve performances almost in all cases. All top performing combinations contain Halobacillus (blue and orange, Fig. 3a, b), followed by four combinations containing Halomonas (green). To test predicted performances in the lab we created artificial consortia of Arthrobacter, Halomonas and Halobacillus isolates. Experimental results provided validation to simulation outputs. First, no atrazine degradation was detected by Halomonas or Halobacillus (Fig. 3c). Second, performances of all consortia were better than the Arthrobacter alone on both glucose amended and non-amended media (Fig. 3c). Halobacillus best supported the atrazine degradation followed by the combination Halobacillus and Halomonas, and then Halomonas. In comparison, control consortia composed of Arthrobacter and a random bacterium, not associated with the original community (E. coli), or an endogenous soil bacterium, non-differentially abundant after atrazine application in the original community (S8), did not show enhanced performances in comparison to the performances of Arthrobacter alone (Fig. 3c). Third, atrazine degradation efficiencies of all combinations were higher on rich-nutritional medium compared to poor-nutritional medium, showing the degradation enhancement by glucose (Fig. 3e).
Characterization of interactions and exchange fluxes
Within each combination, we looked at the individual growth of each member according to biomass mass production in the corresponding compartment. Plots in Fig. 4 indicate the individual growth of Arthrobacter, Halobacillus and Halomonas in different combinations (Fig. 4 top, middle and bottom, respectively). Growth pattern of Arthrobacter differ between the two media types: in the relatively rich, glucose-supplemented medium, Arthrobacter grows best as a single species; in the poor medium (atrazine only), Arthrobacter growth as a single species is the worst in comparison to all combinations. Reliance of species on community members for improved growth is typical of minimal media and is likely to reflect dependency in exchange fluxe,s which are masked in rich media [22, 55]. Unlike the pattern observed for Arthrobacter, improved growth in all combinations in the poor media, an antagonistic growth pattern is predicted between Halobacillus and Halomonas, both species performs better in mutual exclusive combinations in comparison to combinations containing both species. This antagonistic pattern corresponds with the dynamics in community structure in soil where an increase in the level of Halobacillus was associated in reduction in the abundance of Halomonas (Supplementary Table 3).
Simulation of biomass of each species within different combinations. Relative increase in biomass of specific species in different combinations on defined media with atrazine, either as the sole carbon and nitrogen source (left) or supplemented by glucose (right). The relative increase shows biomass(t)/biomass(0) -biomass(0). Line colors are indicative of the combinations as indexed in the right grid and are identical to these in Fig. 3
In order to suggest patterns of metabolic interactions that might explain co-occurrence patterns in both simulations and soil we predicted the mutual exchange fluxes in the three-species in silico consortia. Simulations predicted that Arthrobacter secretes aminoethanol, ethylamine and hypoxanthine during atrazine degradation, that are consumed by both Halobacillus and Halomonas (Fig. 5a). The antagonistic association in their relative abundance can hence be related to co-dependency on these secretion products. In return, Halobacillus and Halomonas secret ammonium (NH4+) and leucine (Halobacillus only) that are consumed by Arthrobacter. In glucose amended medium, though secretion pattern of Arthrobacrer remains the same (Fig. 5a), mutual exchanges are conserved only with Halobacillus and not with Halomonas. To provide experimental support for the predicted exchange fluxes we examined growth and degradation of atrazine in monocultures of the three species (Halobacillus. Halomonas and Arthrobacter) grown on minimal media, each supplemented by the relevant exchange metabolites (EM1, EM2 and EM3, respectively, Fig. 5a). In agreement with predictions, Halobacillus and Halomonas growth was recovered in the supplemented media in comparison to no growth in an atrazine only media; Arthrobacter growth and atrazine degradation was enhanced in the supplemented medium (Fig. 5b). We further screened for the presence of the eight predicted exchange metabolites in the atrazine only media (Fig. 5a, left). Reassuringly, six metabolites—aminnethanol, ethylamine, glycerol, urea, hypoxanthine and ammonium—were detected in a co-culture of the three species (Supplementary Figure 4). Two exchange metabolites, leucine and mannitol were not detected, possibly due to a rapid turnover. Finally, quantitative screens of ammonium in mono and co-cultures are consistent with model predictions: When grown in atrazine-only medium, Arthrobacter is predicted to consume ammonium secreted by Halobacillus and Halomonas (Fig. 5a, left). In accordance with predictions, ammonium concentrations in co-cultures (Arthrobacter- Halobacillus and Arthrobacter- Halomonas) are much higher than concentrations detected in the mono-cultures of Halobacillus and Halomonas that are un-capable of growing alone on atrazine as a sole carbon and nitrogen source or in the mono-culture of Arthrobacter that does not secret ammonium in atrazine only medium according to model predictions (Fig. 5c). Supplementing the medium with glucose, is predicted to induce ammonium secretion of Arthrobacter (Fig. 5a, right). In accordance with predictions, ammonium concentrations in a mono-culture of Arthrobacter are much higher in glucose supplemented vs. non-supplemented medium (Fig. 5c).
Prediction and validation of potential exchange fluxes in combinations. a Predicted exchange fluxes in media with atrazine as the only carbon and nitrogen sources (left) and atrazine and glucose nutrition (right). EM1, EM2 and EM3 represent directional exchanges in a specific combination. b In vitro measurements of growth (right) and fraction of atrazine left (left) in medium containing atrazine as a sole nitrogen and carbon source vs. the same medium supplemented by exchange metabolites. Experiments were carried in monocultures of Halobacillus sp. (top), Halomonas sp. (middle) and Arthrobacter sp. (bottom). c Levels of NH4+ concentrations in media. Species in (b, c) were represented by the same isolates as in Fig. 3 (NY15, N8 and AT5, respectively). Bars represent the standard errors of the three replicates
Discussion
Manipulating bacterial communities is a key emerging challenge in microbial ecology with potential applications for medical, agricultural and environmental practices [56,57,58]. Especially, the “in situ microbiome engineering” could be a new paradigm of community-scale microbial engineering [57]. Microbiome’s activities and capacities are to a large extent determined by complex networks of metabolic interactions and exchanges [59,60,61,62]. Traditionally, the study of bacterial interactions required the use of laboratory experiments such as growth and co-culture assays [59, 63]. Major obstacles are difficulties in isolation and culturing of all community members and the complexity of the microbiome’s interactions. Furthermore, the composition of taxa and their interactions in the microbiome can vary substantially over short time scales and nutrient environment modification. In order to detect metabolic interactions, methods able of capturing species identity, dependencies and the nature of exchanged metabolites are needed, and thus multiple combinations of diverse techniques, such as metagenomics, mass spectrometry, and isotope labeling are required [64]. Although multi-technique strategies have been successfully applied for some model systems such as enrichment cultures and synthetic communities [3, 65], their application for the study of natural communities is far from trivial [63]. Mathematical models of bacterial community expand the toolbox for detecting metabolic dependencies in natural bacterial consortia [14, 17, 62].
Here we aimed at applying mathematical modeling approaches for an “in situ microbiome engineering” targeted for accelerating pollutant degradation in soil contaminated with the herbicide atrazine. Our results first demonstrate that atrazine applications in soil triggers a compositional shift of bacterial communities (Fig. 2), and that the shift is associated with functional modification, atrazine-degrading ability of the corresponding communities (Fig. 1). Despite the functional shift, the predicted atrazine degrader, Arthrobacter, was detected in both herbicide treated and untreated soil and significant abundance modifications were detected for non-degrading species. These non-intuitive observations, where abundance shift cannot provide a straightforward justification for the functional modification, can be related to community interactions. In order to reach a system-level view of the activity in soil, we first made use of the information of the compositional shifts associated with the degrading ability. Then, we applied dynamic modeling methods and explored the performances and exchanges in a range of environmental microbial consortia. The modeling aimed at charactering performances (atrazine degradation efficiencies) and exchanges in different consortia in a range of simulated media. Simulation results demonstrated the importance of the non-degraders species to the degradation process, providing functional interpretation to the observed compositional shifts. Simulations predicted an exchange of metabolites between the atrazine degrader (Arthrobacter) and non-degrader (Halomonas and Halobacillus). Exchange metabolites include aminoethanol and ethylamine—products of atrazine degradation by Arthrobacter [65], providing carbon and nitrogen sources for the non-degrader. The improved atrazine-degrading efficiency and increase in the biomass of community suggested mutual benefits associated with cross-feeding. Based on simulation we designed bacterial communities and validated in vitro improved performances of selected vs. control combinations. Relevant exchange fluxes, predicted by simulation were also supported by experimental results. Simulations supported the amendment of the medium in additional carbon source, here glucose, as previously reported [66], and also validated in our experimental system. Moreover, Halobacillus was not detected in the enrichment samples, possibly because the culture condition do not support and maintain the composition of the source sample community, and pointing at the importance of genomic approaches and genomic-driven analysis for studying soil microbial function, circumventing the need to isolate. Our analyses provide a comprehensive mechanistic description that explains community dynamics leading to enhanced degradation, not only through affecting Arthrobacter performances but through affecting community function. Hence the analysis paves the way for the educated design of biostimulation strategies. Glucose, the carbon source used here is not common in soil, and its application to agricultural practice is questionable. Further research will focus on the simulation-based examination of alternative carbon, sources that are present in soil [66], and can act as specified biostimulation agents of selected bacterial combinations.
Limitations of the analysis should be acknowledged, to a large extent reflecting current state of technology. First, genome sequences for model construction and simulations were retrieved from available public resources based on similarity in 16S rRNA gene, rather than assembled directly from the soil sample. Similarly, isolates for in vitro validation were chosen from public resources based on 16S rRNA. It is important to acknowledge that the genome of bacteria with close phylogenetic proximity (inferred by similarity in the 16S rRNA) can vary substantially in their genomes, and hence simulations and validation represent an approximation of the in situ combinations. Such an approximation, despite its obvious limitations, is applied here as in many other recent explorations of microbial function and interactions in complex environments, relying on a demonstrated overall correlation between phylogeny and function in tightly associated species [67,68,69,70,71]. Such approximations allow a relatively straightforward approach to circumvent the difficulty in isolating and supporting growth in culture for most endogenous soil species. Sequencing technologies advance rapidly, and pioneering projects demonstrate that the full assembly of abundant species based on direct sequencing from the sample is possible even in complex communities [72]. Whereas future projects are expected to make use of the genomes that were directly sequenced in the relevant environment, phylogeny-based approximations are currently significantly less costly and more feasible and can hence provide a realistic bridge between microbial ecology and system biology and allow to study ecosystem-level function and dynamics [14, 17, 62]. Here, all approximations require at least 97% identity in the 16S rRNA between OTUs and representative species, following the common practice in the large majority of ecological surveys of co-clustering sequences above this degree of similarity into a common taxonomic unit. An alternative approach to looking for the most closely related sequence species (above a threshold) is to define a set of core genes that are conserved between all species of a wider corresponding phylogenetic group. However, such pan-genome approaches have their own cones where the set of shared genes is heavily dependent on the number of species considered and the variability in lifestyle and genomes sizes [73,74,75]. In particular, soil samples were demonstrated to be poorly represented in genome depositories in comparison to the relatively high coverage of sequenced species in the human microbiome and mammalian gut datasets, creating a bias against the inclusion of soil-associated functions [71].
Finally, due to the complexity of model construction process, we limited the analysis to five soil species. Advances in sequencing technologies together with improvement in platforms for genome annotation and model construction [42, 76,77,78,79] are expected to lead to in silico representation of complex microbial communities [70]. In parallel to the advent of sequencing technologies, metabolomics technologies are now rapidly emerging and in the very near future a growing number of ecosystems will be subject to an extensive profiling [78, 80, 81]. Simulations accuracy is expected to improve when based on a detailed description of the metabolic environmental conditions. Though the current analysis is based on species and environment approximation, representing only a sub-set of the original community, it demonstrates the strength of metabolic-modelling in producing testable hypotheses that were supported by in vitro and pot experiments. In the near future, where new technologies will reduce the need in approximations and will facilitate model construction and analyses, reliability and success rate of genomic driven predictions are likely to increase and make them an integral part of microbial community engineering.
Conclusion
Modeling of microbial communities from atrazine-treated soils allows predicting community performances considering growth, atrazine degradation, and specific exchange fluxes. Both simulations and experimental results indicated that adding Halobacillus, and/or Halomonas to the community remarkably improve atrazine-degrading efficiency. The efficiency of each community could also be clearly enhanced by adding to the media additional carbon sources such as glucose. The outcome of this study can assist to reduce atrazine contamination in soil and water. This methodology will be addressed to reduce the soil contamination of other herbicide families.
References
Hansen SK, Rainey PB, Haagensen JAJ, Molin S. Evolution of species interactions in a biofilm community. Nature. 2007;445:533–6.
Fuhrman JA. Microbial community structure and its functional implications. Nature. 2009;459:193–9.
Großkopf T, Soyer OS. Synthetic microbial communities. Curr Opin Microbiol. 2014;18:72–7.
Falkowski PG, Fenchel T, Delong EF. The microbial engines that drive Earth’s biogeochemical cycles. Science. 2008;320:1034–9.
Li X, Li P, Lin X, Zhang C, Li Q, Gong Z. Biodegradation of aged polycyclic aromatic hydrocarbons (PAHs) by microbial consortia in soil and slurry phases. J Hazard Mater. 2008;150:21–26.
Bacosa HP, Suto K, Inoue C. Bacterial community dynamics during the preferential degradation of aromatic hydrocarbons by a microbial consortium. Int Biodeterior Biodegrad. 2012;74:109–15.
Keller AH, Kleinsteuber S, Vogt C. Anaerobic benzene mineralization by nitrate-reducing and sulfate-reducing microbial consortia enriched from the same site: comparison of community composition and degradation characteristics. Microb Ecol. 2018;75:941–53.
Fu H, Zhang JJ, Xu Y, Chao HJ, Zhou NY. Simultaneous biodegradation of three mononitrophenol isomers by a tailor-made microbial consortium immobilized in sequential batch reactors. Lett Appl Microbiol. 2017;64:203–9.
Roh SW, Kim K-H, Nam Y-D, Chang H-W, Park E-J, Bae J-W. Investigation of archaeal and bacterial diversity in fermented seafood using barcoded pyrosequencing. ISME J. 2010;4:1–16.
Ercolini D. High-throughput sequencing and metagenomics: moving forward in the culture-independent analysis of food microbial ecology. Appl Environ Microbiol. 2013;79:3148–55.
Walter J, Ley R. The human gut microbiome: ecology and recent evolutionary changes. Annu Rev Microbiol. 2011;65:411–29.
Huttenhower C, Gevers D, Knight R, Abubucker S, Badger JH, Chinwalla AT, et al. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207–14.
Adams GO, Fufeyin PT, Okoro SE, Ehinomen I. Bioremediation, biostimulation and bioaugmention: a review. Int J Environ Bioremediation Biodegrad. 2015;3:28–39.
Zomorrodi AR, Segrè D. Synthetic ecology of microbes: mathematical models and applications. J Mol Biol. 2016;428:837–61.
Bento FM, Camargo FAO, Okeke BC, Frankenberger WT. Comparative bioremediation of soils contaminated with diesel oil by natural attenuation, biostimulation and bioaugmentation. Bioresour Technol. 2005;96:1049–55.
Mrozik A, Piotrowska-Seget Z. Bioaugmentation as a strategy for cleaning up of soils contaminated with aromatic compounds. Microbiol Res. 2010;165:363–75.
Widder S, Allen RJ, Pfeiffer T, Curtis TP, Wiuf C, Sloan WT, et al. Challenges in microbial ecology: building predictive understanding of community function and dynamics. ISME J. 2016;10:2557–68.
Ofaim S, Ofek-Lalzar M, Sela N, Jinag J, Kashi Y, Minz D, et al. Analysis of microbial functions in the rhizosphere using a metabolic-network based framework for metagenomics interpretation. Front Microbiol. 2017;8:1606.
Nagarajan H, Embree M, Rotaru A-E, Shrestha PM, Feist AM, Palsson BØ, et al. Characterization and modelling of interspecies electron transfer mechanisms and microbial community dynamics of a syntrophic association. Nat Commun. 2013;4:ncomms 3809.
Zelezniak A, Andrejev S, Ponomarova O, Mende DR, Bork P, Patil KR. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc Natl Acad Sci USA. 2015;112:6449–54.
Opatovsky I, Santos-Garcia D, Ruan Z, Lahav T, Ofaim S, Mouton L, et al. Modeling trophic dependencies and exchanges among insects’ bacterial symbionts in a host-simulated environment. BMC Genom. 2018;19:402.
Freilich S, Zarecki R, Eilam O, Segal ES, Henry CS, Kupiec M, et al. Competitive and cooperative metabolic interactions in bacterial communities. Nat Commun. 2011;2:589.
Zhuang K, Izallalen M, Mouser P, Richter H, Risso C, Mahadevan R, et al. Genome-scale dynamic modeling of the competition between Rhodoferax and Geobacter in anoxic subsurface environments. ISME J. 2011;5:305–16.
Jablonowski ND, Köppchen S, Hofmann D, Schäffer A, Burauel P. Persistence of 14C-labeled atrazine and its residues in a field lysimeter soil after 22 years. Environ Pollut. 2009;157:2126–31.
Chiaia-Hernandez AC, Keller A, Wächter D, Steinlin C, Camenzuli L, Hollender J, et al. Long-term persistence of pesticides and TPs in archived agricultural soil samples and comparison with pesticide application. Environ Sci Technol. 2017;51:10642–51.
Tappe W, Groeneweg J, Jantsch B. Diffuse atrazine pollution in German aquifers. Biodegradation. 2002;13:3–10.
Mahía J, Martín A, Díaz-Raviña M. Extractable atrazine and its metabolites in agricultural soils from the temperate humid zone. Environ Geochem Health. 2008;30:147–52.
Murphy MB, Hecker M, Coady KK, Tompsett AR, Jones PD, Du Preez LH, et al. Atrazine concentrations, gonadal gross morphology and histology in ranid frogs collected in Michigan agricultural areas. Aquat Toxicol. 2006;76:230–45.
Dalton R. Frogs put in the gender blender by America’s favourite herbicide. Nature. 2002;416:665–6.
Hayes TB, Khoury V, Narayan A, Nazir M, Park A, Brown T, et al. Atrazine induces complete feminization and chemical castration in male African clawed frogs (Xenopus laevis). Proc Natl Acad Sci USA. 2010;107:4612–7.
Hénault-Ethier L. Backgrounder: atrazine: banned in Europe, common in Canada. Canada: equiterre; 2016.
de Souza ML, Newcombe D, Alvey S, Crowley DE, Hay A, Sadowsky MJ, et al. Molecular basis of a bacterial consortium: interspecies catabolism of atrazine. Appl Environ Microbiol. 1998;64:178–84.
Smith D, Alvey S, Crowley DE. Cooperative catabolic pathways within an atrazine-degrading enrichment culture isolated from soil. FEMS Microbiol Ecol. 2005;53:265–75.
Yang C, Li Y, Zhang K, Wang X, Ma C, Tang H, et al. Atrazine degradation by a simple consortium of Klebsiella sp. A1 and Comamonas sp. A2 in nitrogen enriched medium. Biodegradation. 2010;21:97–105.
Onofri A, Carbonell EA, Piepho H-P, Mortimer AM, Cousens RD. Current statistical issues in weed research. Weed Res. 2010;50:5–24.
Jettner RJ, Walker SR, Churchett JD, Blamey FPC, Adkins SW, Bell K. Plant sensitivity to atrazine and chlorsulfuron residues in a soil-free system. Weed Res. 1999;39:287–95.
Eizenberg H, Goldwasser Y, Achdary G, Hershenhorn J. The potential of sulfosulfuron to control troublesome weeds in tomato. Weed Technol. 2003;17:133–7.
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics. 2011;27:2194–2200.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA. 2011;108(Suppl 1):4516–22.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Hammer Ø, Harper DAT, Ryan PD. PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron. 2001;4:9.
Henry CS, DeJongh M, Best AA, Frybarger PM, Linsay B, Stevens RL. High-throughput generation, optimization and analysis of genome-scale metabolic models. Nat Biotechnol. 2010;28:977–82.
Meyer F, Paarmann D, D’Souza M, Olson R, Glass E, Kubal M, et al. The metagenomics RAST server – a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinforma. 2008;9:386.
Kanehisa M, Goto S, Sato Y, Kawashima M, Furumichi M, Tanabe M. Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res. 2014;42:D199–D205.
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S, et al. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 2004;32:D115–9.
Nordberg H, Cantor M, Dusheyko S, Hua S, Poliakov A, Shabalov I, et al. The genome portal of the Department of Energy Joint Genome Institute: 2014 updates. Nucleic Acids Res. 2014;42:D26–D31.
Jiang J, Pan Y, Hu S, Zhang X, Hu B, Huang H, et al. Halomonas songnenensis sp. nov., a moderately halophilic bacterium isolated from saline and alkaline soils. Int J Syst Evol Microbiol. 2014;64:1662–9.
Liu WY, Zeng J, Wang L, Dou YT, Yang SS. Halobacillus dabanensis sp. nov. and Halobacillus aidingensis sp. nov., isolated from salt lakes in Xinjiang, China. Int J Syst Evol Microbiol. 2005;55:1991–6.
Ma C, Zhuang L, Zhou SG, Yang GQ, Yuan Y, Xu RX. Alkaline extracellular reduction: isolation and characterization of an alkaliphilic and halotolerant bacterium, Bacillus pseudofirmus MC02. J Appl Microbiol. 2012;112:883–91.
Srinivas TNR, Anil Kumar P, Madhu S, Sunil B, Sharma TVRS, Shivaji S. Cesiribacter andamanensis gen. nov., sp. nov., isolated from a soil sample from a mud volcano. Int J Syst Evol Microbiol. 2011;61:1521–7.
Orth JD, Thiele I, Palsson BØ. What is flux balance analysis? Nat Biotechnol. 2010;28:245–8.
Thiele I, Palsson BØ. A protocol for generating a high-quality genome-scale metabolic reconstruction. Nat Protoc. 2010;5:93–121.
Schellenberger J, Que R, Fleming RMT, Thiele I, Orth JD, Feist AM, et al. Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox v2.0. Nat Protoc. 2011;6:1290–307.
Harcombe WR, Riehl WJ, Dukovski I, Granger BR, Betts A, Lang AH, et al. Metabolic resource allocation in individual microbes determines ecosystem interactions and spatial dynamics. Cell Rep. 2014;7:1104–15.
Wintermute EH, Silver PA. Emergent cooperation in microbial metabolism. Mol Syst Biol. 2010;6:407.
Johns NI, Blazejewski T, Gomes AL, Wang HH. Principles for designing synthetic microbial communities. Curr Opin Microbiol. 2016;31:146–53.
Sheth RU, Cabral V, Chen SP, Wang HH. Manipulating bacterial communities by in situ microbiome engineering. Trends Genet. 2016;32:189–200.
Huang X, He J, Yan X, Hong Q, Chen K, He Q, et al. Microbial catabolism of chemical herbicides: microbial resources, metabolic pathways and catabolic genes. Pestic Biochem Physiol. 2017;143:272–97.
Kato S, Haruta S, Cui ZJ, Ishii M, Igarashi Y. Stable coexistence of five bacterial strains as a cellulose-degrading community. Appl Environ Microbiol. 2005;71:7099–106.
Lawrence D, Fiegna F, Behrends V, Bundy JG, Phillimore AB, Bell T, et al. Species interactions alter evolutionary responses to a novel environment Ellner SP (ed). PLoS Biol. 2012;10:e1001330.
Chen K, Huang L, Xu C, Liu X, He J, Zinder SH, et al. Molecular characterization of the enzymes involved in the degradation of a brominated aromatic herbicide. Mol Microbiol. 2013;89:1121–39.
Muller EEL, Faust K, Widder S, Herold M, Martínez Arbas S, Wilmes P. Using metabolic networks to resolve ecological properties of microbiomes. Curr Opin Syst Biol. 2018;8:73–80.
Zeidan AA, Rådström P, van Niel EW. Stable coexistence of two Caldicellulosiruptor species in a de novo constructed hydrogen-producing co-culture. Microb Cell Fact. 2010;9:102.
Ponomarova O, Patil KR. Metabolic interactions in microbial communities: untangling the Gordian knot. Curr Opin Microbiol. 2015;27:37–44.
Embree M, Nagarajan H, Movahedi N, Chitsaz H, Zengler K. Single-cell genome and metatranscriptome sequencing reveal metabolic interactions of an alkane-degrading methanogenic community. ISME J. 2014;8:757–67.
Strong LC, Rosendahl C, Johnson G, Sadowsky MJ, Wackett LP. Arthrobacter aurescens TC1 metabolizes diverse s-triazine ring compounds. Appl Environ Microbiol. 2002;68:5973–80.
Abdelhafid R, Houot S, Barriuso E. Dependence of atrazine degradation on C and N availability in adapted and non-adapted soils. Soil Biol Biochem. 2000;32:389–401.
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol. 2013;31:814–21.
Friedman N, Shriker E, Gold B, Durman T, Zarecki R, Ruppin E, et al. Diet-induced changes of redox potential underlie compositional shifts in the rumen archaeal community. Environ Microbiol. 2017;19:174–84.
Magnúsdóttir S, Heinken A, Kutt L, Ravcheev DA, Bauer E, Noronha A, et al. Generation of genome-scale metabolic reconstructions for 773 members of the human gut microbiota. Nat Biotechnol. 2017;35:81–89.
Aßhauer KP, Wemheuer B, Daniel R, Meinicke P. Tax4Fun: predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics. 2015;31:2882–4.
Anantharaman K, Brown CT, Hug LA, Sharon I, Castelle CJ, Probst AJ, et al. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat Commun. 2016;7:13219.
Udaondo Z, Molina L, Segura A, Duque E, Ramos JL. Analysis of the core genome and pangenome of P seudomonas putida. Environ Microbiol. 2016;18:3268–83.
Garcia-Garcera M, Touchon M, Brisse S, Rocha EPC. Metagenomic assessment of the interplay between the environment and the genetic diversification of Acinetobacter. Environ Microbiol. 2017;19:5010–24.
Herbold CW, Lehtovirta-Morley LE, Jung M-Y, Jehmlich N, Hausmann B, Han P, et al. Ammonia-oxidising archaea living at low pH: Insights from comparative genomics. Environ Microbiol. 2017;19:4939–52.
Latendresse M, Krummenacker M, Trupp M, Karp PD. Construction and completion of flux balance models from pathway databases. Bioinformatics. 2012;28:388–96.
Agren R, Liu L, Shoaie S, Vongsangnak W, Nookaew I, Nielsen J. The RAVEN toolbox and its use for generating a genome-scale metabolic model for Penicillium chrysogenum. PLoS Comput Biol. 2013;9:e1002980.
Machado D, Andrejev S, Tramontano M, Patil KR. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 2018;46:7542–53. https://doi.org/10.1093/nar/gky537.
El Amrani A, Dumas AS, Wick LY, Yergeau E, Berthomé R. “Omics” insights into PAH degradation toward improved green remediation biotechnologies. Environ Sci Technol. 2015;49:11281–91.
Daliri EB, Wei S, Oh DH, Lee BH. The human microbiome and metabolomics: current concepts and applications. Crit Rev Food Sci Nutr. 2017;57:3565–76.
Parmar KM, Gaikwad SL, Dhakephalkar PK, Kothari R, Singh RP. Intriguing interaction of bacteriophage-host association: an understanding in the era of omics. Front Microbiol. 2017;8:559.
Acknowledgements
This work was supported by grants from National Key Research and Development Program (2016YFD0800203), the NSFC-ISF joint program (31461143009), Israel Science Foundation Grant no. 1416/14, and the Science Foundation of Jiangsu Province, China (BK20150670).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Xu, X., Zarecki, R., Medina, S. et al. Modeling microbial communities from atrazine contaminated soils promotes the development of biostimulation solutions. ISME J 13, 494–508 (2019). https://doi.org/10.1038/s41396-018-0288-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0288-5
This article is cited by
-
Designing a synthetic microbial community through genome metabolic modeling to enhance plant–microbe interaction
Environmental Microbiome (2023)
-
Establishment of microbial model communities capable of removing trace organic chemicals for biotransformation mechanisms research
Microbial Cell Factories (2023)
-
A framework for the targeted recruitment of crop-beneficial soil taxa based on network analysis of metagenomics data
Microbiome (2023)
-
Nitrogen transfer and cross-feeding between Azotobacter chroococcum and Paracoccus aminovorans promotes pyrene degradation
The ISME Journal (2023)
-
Coupling multifactor dominated the biochemical response and the alterations of intestinal microflora of earthworm Pheretima guillelmi due to typical herbicides
Environmental Science and Pollution Research (2023)