Abstract
A critical step in the biogeochemical cycle of sulfur on Earth is microbial sulfate reduction, yet organisms from relatively few lineages have been implicated in this process. Previous studies using functional marker genes have detected abundant, novel dissimilatory sulfite reductases (DsrAB) that could confer the capacity for microbial sulfite/sulfate reduction but were not affiliated with known organisms. Thus, the identity of a significant fraction of sulfate/sulfite-reducing microbes has remained elusive. Here we report the discovery of the capacity for sulfate/sulfite reduction in the genomes of organisms from 13 bacterial and archaeal phyla, thereby more than doubling the number of microbial phyla associated with this process. Eight of the 13 newly identified groups are candidate phyla that lack isolated representatives, a finding only possible given genomes from metagenomes. Organisms from Verrucomicrobia and two candidate phyla, Candidatus Rokubacteria and Candidatus Hydrothermarchaeota, contain some of the earliest evolved dsrAB genes. The capacity for sulfite reduction has been laterally transferred in multiple events within some phyla, and a key gene potentially capable of modulating sulfur metabolism in associated cells has been acquired by putatively symbiotic bacteria. We conclude that current functional predictions based on phylogeny significantly underestimate the extent of sulfate/sulfite reduction across Earth’s ecosystems. Understanding the prevalence of this capacity is integral to interpreting the carbon cycle because sulfate reduction is often coupled to turnover of buried organic carbon. Our findings expand the diversity of microbial groups associated with sulfur transformations in the environment and motivate revision of biogeochemical process models based on microbial community composition.
Similar content being viewed by others
Introduction
The cycling of sulfur is one of Earth’s major biogeochemical processes. Sulfate reduction in conjunction with sulfur disproportionation may be an early evolved microbial metabolism, given evidence for biological fractionation of sulfur isotopes around 3.5 billion years ago [1, 2], and it remains an important energy metabolism for anaerobic life [3]. In natural ecosystems, human microbiomes, and engineered systems, this process is important because the product hydrogen sulfide (H2S) can be toxic [4], can corrode steel [5], and sour oil reservoirs [6]. Overall, sulfate reduction is a primary driver in the carbon cycle, and is responsible for a large part of the organic carbon flux to CO2 in marine sedimentary environments [7, 8] and in wetlands [9]. Importantly, the coupling of sulfate/sulfite reduction to oxidation of H2, small chain fatty acids, or other carbon compounds limits the availability of these substrates to other organisms like methanogens and alters the energetics via syntrophic interactions [10, 11]. All of these processes also impact methane production. Given the many reasons why the biological conversion of sulfate/sulfite to sulfide is important, it is vital that we understand which organisms can carry out the reactions and the pathways involved.
The canonical microbial pathway for dissimilatory sulfate reduction involves the initial reduction of sulfate to sulfite by a combination of sulfate adenylyltransferase (Sat) and adenylyl-sulfate reductase (AprBA) followed by reduction of sulfite by sulfite reductases. Sulfite reductase genes catalyze the rate-limiting steps in the global sulfur cycle [12, 13] and confer bacteria and archaea the ability to grow via reduction of sulfite, and can function in reverse in some organisms that disproportionate or oxidize elemental sulfur [14,15,16]. Four different groups of sulfite reductases function in dissimilatory sulfur metabolism. Of these, siroheme-dependent dissimilatory sulfite reductase (dsr), siroheme-dependent anaerobic sulfite reductase (asr) genes, and octaheme cytochrome c sulfite reductase (mccA) catalyze the reduction of sulfite to sulfide, while reverse dissimilatory sulfite reductase genes (rdsr) are involved in sulfur oxidation. All of these sulfite reductases except for mccA constitute an ancient lineage of enzymes that may predate the separation of Bacteria and Archaea [17].
The taxonomic distribution of dissimilatory sulfite reductases has been considered to be restricted to organisms from selected bacterial and archaeal phyla [18]. Only organisms from nine microbial phylum-level lineages, namely Deltaproteobacteria, Firmicutes, Thermodesulfobacteria, Actinobacteria, Nitrospirae, Caldiserica, Euryarchaeota, Crenarcheota, and Aigarchaeota are known to possess the genetic capacity to reduce sulfite to sulfide using the dsr system. The asr enzymes have a far more limited distribution and are known to be present only in organisms from four phylum-level lineages, Gammaproteobacteria, Firmicutes, Spirochaetes, and Fusobacteria. The distribution of MccA enzymes is restricted to organisms from Epsilonproteobacteria [19] and Gammaproteobacteria [20]. Finally, the rdsr enzyme complex for sulfur oxidation is associated with organisms from five phylum-level lineages including Alphaproteobacteria, Betaproteobacteria, Gammaproteobacteria, Deltaproteobacteria, and Chlorobi. This diversification of sulfite reductases was likely driven by speciation and functional divergence, and to a lesser extent, lateral gene transfer (LGT) [21].
The recent availability of thousands of genomes from organisms belonging to many newly sampled phyla has provided the opportunity to test for the presence of sulfite reductase genes in bacteria and archaea that have not previously been associated with dissimilatory sulfur metabolism [22]. Here we use shotgun metagenomic sequencing and recovery of metagenome-assembled genomes (MAGs) from a diverse set of marine and terrestrial environments to show that organisms from novel lineages contain sulfite reductases that implicate them in the dissimilatory cycling of sulfur. In total, we more than doubled the number of microbial lineages that can catalyze dissimilatory sulfate/sulfite reduction or sulfur oxidation in the environment. We shed light on the complicated evolutionary history of dissimilatory sulfite reductases and show that LGT of catalytic sulfite reductase genes is much more common than previously thought.
Materials and methods
Sample collection and data processing
Details of sample collection (Sampling, DNA Extraction), individual sample geochemical measurements, and data processing (DNA sequencing, assembly, annotation, binning, genome completion estimates) are described in detail elsewhere [23,24,25,26,27].
Identification of selected sulfate/sulfite reduction genes
Genome-specific metabolic potential for sulfate/sulfite reduction was determined in an iterative manner by (A) searching all predicted ORFs in a genome with HMM profiles for dsrA and dsrB from TIGRFAM [28], and dsrD from Pfam [29] using hmmscan v3.1b2 [30], and (B) searching against custom hmm profiles for dsrA, dsrB, and dsrD using hits generated from step (A) and searching all predicted ORFs again for the above genes. (C) Identification of anaerobic sulfite reductase genes was conducted by searching all predicted ORFs against asrA, asrB, and asrC hmm databases from TIGRfam [28]. (D) Identification of genes for the reduction of sulfate to sulfite was conducted by searching all predicted ORFs against aprA, aprB, and sat hmm databases from TIGRfam [28].
Taxonomic confirmation
The taxonomy of organisms represented by the 123 identified genomes was determined by using a concatenated RP tree. Briefly 16 RPs (L2, L3, L4, L5, L6, L14, L15, L16, 18, L22, L24, S3, S8, S10, S17, S19) were each aligned separately using MUSCLE v3.8.31 [31]. All alignments were end trimmed manually and all columns with >95% gaps were trimmed. The individual alignments were concatenated and the phylogenetic tree was inferred by RAxML v8.0.26 [32] implemented by the CIPRES Science Gateway [33]. This analysis sampled a total of 204 bootstrap replicates before being stopped by the autoMRE algorithm. The complete RP tree is presented as Data File S1.
RAxML was called as follows:
raxmlHPC-HYBRID -s input -N autoMRE -n result -f a -p 12345 -x 12345 -m PROTCATLG
Other dsr genes
To study the dsr operon structure in the newly identified organisms, we identified other dsr genes, namely C, E, F, H, M, K, J, O, P, N, L, R, and S. These genes were identified in the vicinity of dsrAB using domain similarity identified by hits to TIGRfam [28], COG [34], and Pfam databases [29], as well certain key traits and conserved residues in specific genes.
Sequence alignment and phylogeny
Phylogenetic analyses were performed as follows:
Each individual gene (dsrA, dsrB, dsrC, dsrD, aprA, aprB, dsrT, qmoA, qmoB, sat) was aligned along with reference sequences using MUSCLE [31] with default parameters. All alignments were manually refined by trimming the start and ends and removing all columns with >95% gaps. For generation of concatenated alignments (dsrAB, qmoAB, and aprBA), individual alignments were concatenated in Geneious version 7 [35]. In construction of the concatenated qmo tree, only subunits A and B were used since subunit C is not universally present in sulfate/sulfite-reducing organisms. All phylogenetic analyses were inferred by RAxML v8.0.26 [32] implemented by the CIPRES Science Gateway [33]. RAxML was called as follows:
For AsrABC, DsrD, Sat, DsrT, QmoAB, DsrEFH trees:
raxmlHPC-PTHREADS -s input -N 1000 -n result -f a -p 12345 -x 12345 -m PROTGAMMAGTR.
For DsrAB, AprBA trees:
raxmlHPC-HYBRID -s input -N autoMRE -n result -f a -p 12345 -x 12345 -m PROTGAMMAGTR.
The complete DsrAB tree is presented as Data File S2.
Conserved residues and motifs
Conserved residues and motifs in DsrA, DsrB, DsrC, AsrC, DsrD, and DsrE proteins were identified by aligning the identified genes from all 123 genomes in this study with reference proteins [36,37,38]. All conserved residues identified by us were also compared with the model sulfate-reducing organism, Desulfovibrio vulgaris [39].
Rooting of dsrA/dsrB tree
A reference alignment was calculated de novo using MAFFT based on a non-redundant (90% identity clustering with uclust [40]) set of length-filtered (300–500 nt) sulfite reductase superfamily sequences from this study [18], and additional sequences collected from UniProt and UniParc [41] using the eggNOG COG2221 and TIGRFAM DsrA/DsrB HMM models. The reference alignment was end trimmed and filtered with noisy [42]. Selected full-length DsrA, DsrB, and outgroup sequences were reference-aligned and used for phylogenetic inference using PhyloBayes with the CAT-GTR model [43].
Trees for inferring LGT by comparison of 16S rRNA and reductive-type DsrAB
16S rRNA genes were aligned with the SINA aligner [44] with default parameters. Alignments were then manually refined by trimming the start and ends and removing all columns with gaps >95%. DsrA and DsrB sequences were aligned as described above. 16S rRNA and DsrAB were sourced from the same organisms where possible. In the case of MAGs not containing 16S genes, the closest related organisms with 16S rRNA genes were chosen for this analysis. Organisms with oxidative-type DsrAB were excluded from this analysis. 16S rRNA trees were calculated using maximum likelihood (RAxML) and neighbor-joining (Geneious) methods. The neighbor joining tree was constructed using the Jukes-Cantor Genetic distance model with 1000 bootstrap replicates.
RAxML was run as follows:
raxmlHPC-PTHREADS -f a -s input -n result -m GTRGAMMA -x 12345 -# autoMRE -p 12345 -T 4.
The 16S rRNA consensus tree was constructed using the majority (extended) consensus rule setting using CONSENSE and branch lengths were adjusted using DNAML implemented in the PHYLIP package [45]. Similarly, DsrAB trees were combined into a consensus tree by using the majority (extended) consensus rule in CONSENSE and branch lengths of the consensus tree were inferred by using PROML (JTT model).
Structural models
We selected the DsrT proteins identified in Desulfovibrio vulgaris (WP_012611240) and Candidatus Rokubacteria CSP1-6 (KRT71371) for structural modeling. Protein models were predicted using the I-TASSER suite [46]. Only the top predicted models out of the top five I-TASSER simulations are shown. Both DsrT proteins used the identical top threading template from the sporulation inhibitor protein pXO1-118 from Bacillus anthracis [47].
Analyses of electron donors for sulfate/sulfite reduction
Analyses of putative electron donors and other metabolic potential were centered around identification of genes for hydrogen oxidation, complex carbon compounds (carbohydrates), fatty acid metabolism, and carbon fixation. For identification of the potential for hydrogen oxidation, hmm searches were conducted by searching all predicted ORFs against individual HMM profiles for nickel–iron hydrogenases from Groups I, IIa, IIb, IIIa, IIIb, IIIc, and IIId, and Fe–Fe hydrohenases. All hits above the noise cutoffs were inspected manually. The details are described in detail previously [24, 26, 48].
For identification of carbohydrate substrates for sulfate/sulfite reduction, all predicted ORFs were searched against the CAZy HMM database [49]. Pre-filtering of hits was conducted using the following cutoffs: coverage: 0.40; e-value: 1e–18. To determine the specificity of enzymes, we established a set of 84 distinct reactions involving 189 enzyme families that allowed us to track specific substrates and products. All hits to glycosyltransferases (GT) and Carbohydrate Binding Modules (CBM) were excluded from this analysis due to high incidence of false positives and/or difficulty in determining substrate specificity.
Generation of custom hmm models
For generation of custom HMM profiles, reference sequences and identified genes from the 123 genomes in this study were aligned using MUSCLE [31] with default parameters followed by manually trimming the start and ends of the alignment. The alignment was converted into Stockholm format and databases were built using hmmscan [30]. Individual noise and trusted cutoffs for all HMMs were determined by manual inspection and are built into the custom HMM profiles.
Results and discussion
To investigate the diversity of microorganisms that contain sulfite reductases involved in dissimilatory sulfate/sulfite reduction or sulfur oxidation in the environment, we analyzed genomes reconstructed from metagenomic sequence datasets recovered from six distinct terrestrial and marine subsurface environments where geochemical conditions have suggested active microbial sulfur cycling. Our sampling sites included an aquifer adjacent to the Colorado River, USA [24, 26], a deep subsurface CO2 geyser in Utah, USA [27], a deep borehole in Japan [23], an acidic sulfide mine waste rock site in Canada, deep subseafloor basaltic crustal fluids of the hydrothermally active Juan de Fuca ridge flank in the Pacific Ocean [25], and an acidic peatland in Germany [50]. In total, we searched in excess of 4000 near-complete MAGs for the presence of sulfite reductase genes.
Identification of dissimilatory sulfur cycling organisms from MAGs
We identified sulfite reductase genes in 123 near-complete microbial genomes (Supplementary Table 1). Phylogenetic analyses using a set of 16 concatenated ribosomal proteins (RP) and the small subunit ribosomal (SSU) RNA gene show that these genomes belong to organisms from 20 distinct phylum-level lineages (Table 1), 13 of which were not known to have dsr genes [18]. In addition, we identified anaerobic sulfite reductase (asr) genes required for sulfite reduction in three bacterial groups not previously reported to have this capacity [51]. All of the identified catalytic protein subunits (DsrA, DsrB, and AsrC) contained all conserved sulfite reductase residues and secondary structure elements for the formation of α helices and β sheets [36] (Supplementary Figs. 1– 3).
Dissimilatory sulfite reductase containing organisms
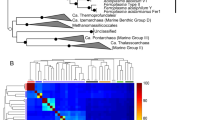
Given our interest in identifying organisms with the capacity to produce sulfide, we initially searched the genomes for operons that contained genes encoding DsrD [52]. This gene was considered a marker for sulfite reduction because it is absent in bacteria that use the rdsr pathway for sulfur oxidation [53]. It is however important to note that the dsrD gene is present and highly expressed in sulfur disproportionating organisms like Desulfurivibrio alkaliphilus that cannot be distinguished from canonical sulfate-reducing bacteria using gene synteny or other genomic features [16]. Although the exact function of the DsrD protein is unclear, the presence of winged-helix domains in its structure and its association with other core proteins of the dsr complex (dsrABC) suggest a regulatory role in bacterial sulfite reduction [37]. We identified 78 genomes that encode at least dsrABCD (Supplementary Fig. 4). A multiple alignment of DsrD sequences confirmed highly conserved residues, indicating that the proteins are likely active (Supplementary Fig. 5). These putative sulfate/sulfite-reducing microorganisms affiliate with eight distinct phyla not previously reported to be capable of these processes. Four are phyla with isolated representatives (Acidobacteria, Armatimonadetes, Ignavibacteria, Planctomycetes) and four are uncultivated candidate phyla (Candidatus Zixibacteria, Candidatus Schekmanbacteria, Candidatus Desantisbacteria, Candidatus Lambdaproteobacteria) (Fig. 1a).
A. Concatenated DsrAB protein tree showing the diversity of organisms involved in dissimilatory sulfur cycling using the dsr system. Lineages in blue contain genomes reported in this study. Phylum-level lineages with first report of evidence for sulfur cycling are indicated by blue letters. Only bootstrap values >50 are shown. The complete tree is available with full bootstrap support values as Additional Data File S2. b Concatenated AsrABC protein tree showing the diversity of organisms that possess the anaerobic sulfite reductase system. Lineages in colors were identified in this study. Only bootstrap values >50 are shown
Importantly, organisms from Verrucomicrobia and two candidate phyla, Candidatus Rokubacteria and Candidatus Hydrothermarchaeota, lack dsrD genes and their dsrAB sequences form completely novel lineages outside the four known main phylogenetic DsrAB clusters, namely the reductive bacterial-type, the oxidative bacterial-type, the reductive archaeal-type, and the second dsrAB copies of Moorella species (Fig. 1a). To determine the earliest evolved and most basal lineages in the DsrAB tree, we performed paralogous rooting analysis on a representative subset of sequences. In accordance with previous reports, our results show that the second copies of dsrAB in Moorella spp. likely represent the most basal DsrAB branch [18]. This was followed by the newly identified sequences from Candidatus Rokubacteria, Verrucomicrobia, and Candidatus Hydrothermarchaeota (Fig. 2). Interestingly, Candidatus Hydrothermarchaeota sequences were not monophyletic with one sequence (JdFR-18 JGI24020J35080_1000005) clustering with Verrucomicrobia and Candidatus Rokubacteria, while the remaining were affiliated with bacterial-type DsrAB. Other organisms lacking dsrD genes cluster together with organisms known to be sulfur oxidizers in the dsrAB tree. Based on this clustering, the group implicated in sulfur oxidation using the rdsr pathway now includes bacteria from three additional phylum-level lineages: Nitrospirae, Nitrospinae,and Candidatus Muproteobacteria (Fig. 1a).
Paralogous rooting analysis of DsrAB. Bayesian inference tree showing the phylogenetic relationship between DsrA and DsrB (50 sequences, 377 alignment positions). Arrow indicates outgroup of other sulfite, non-DsrAB reductase superfamily (COG2221) sequences. Branch supports (posterior probability) higher than 0.9 are indicated by black circles. DsrA/DsrB sequences from this study are marked in bold. Assignment of oxidative/reductive, bacterial/archaeal-type DsrAB is according to Müller et al. [18]
Anaerobic sulfite reductase-encoding organisms
The asr pathway for sulfite reduction was found in three bacterial phyla not previously known to possess this pathway, namely Planctomycetes and members of two candidate phyla, Candidatus Omnitrophica and Candidatus Riflebacteria. Concatenated protein trees of all three subunits AsrA, AsrB, and AsrC showed that sequences from these phyla clustered with those from Firmicutes, suggesting that they were acquired by LGT (Fig. 1b). Investigations into the operon structure of the asr complex revealed that while organisms from Planctomycetes and Candidatus Riflebacteria had a canonical gene organization in the order asrA, asrB, and asrC, Candidatus Omnitrophica had a fourth gene (asrD) as an insertion between asrB and asrC subunits. Analyses of conserved domains show that AsrD is related to the family of formate and nitrite transporters (pfam01226, COG2116, TIGRfam00790). We hypothesize that this may in fact serve as a bisulfide channel associated with dissimilatory sulfite reduction using the asr enzyme complex as observed in Clostridium difficile [54].
dsrD genes in candidate phyla radiation organisms
Surprisingly, we identified dsrD genes in eight genomes of organisms affiliating with Candidatus Falkowbacteria, putatively symbiotic bacteria within the Parcubacteria superphylum of the candidate phyla radiation (CPR) [55]. There is no indication of the presence of other dsr genes in these genomes. Given the predicted close physical and metabolic interactions between CPR bacteria and their hosts, we suggest that this small protein could modulate host metabolism, as sometimes occurs with viruses/phage and their hosts [56]. CPR organisms are common in aquifers where conditions oscillate between oxic and anoxic [24, 26]. The predicted Falkowbacteria DsrD protein sequences cluster with sequences from well-characterized Deltaproteobacteria capable of sulfate reduction, suggesting that deltaproteobacterial sulfate reducers served as dsrD-donors during LGT to these CPR bacteria (Supplementary Fig. 6). Considering the presence of dsrD genes in CPR organisms and putative sulfur-oxidizing/sulfur disproportionating bacteria [16], we propose that dsrD is not a good marker for sulfite reduction. Therefore, we suggest an alternate set of rules for utilizing a combination of dsr genes to distinguish DsrAB-based sulfite reduction from sulfur oxidation on the basis of genomic features (Table 2).
Lateral gene transfer of DsrAB sulfite reductases
Prior analyses have suggested that LGT has influenced the evolution of dsrAB among extant microorganisms but only by comparably few events among major taxonomic lineages [14, 18, 57]. We used a comparison of 16S ribosomal RNA and concatenated DsrAB protein trees to reevaluate the extent to which LGT has influenced the organismal distribution of dsrAB genes (Fig. 3). Mismatching branching pattern between the two trees indicates that dsrAB has been introduced into most of the candidate phyla members by multiple independent LGT events. Our analyses show that organisms from five bacterial and archaeal phyla, Deltaproteobacteria, Nitrospirae, Candidatus Hydrothermarchaeota, Actinobacteria, Chloroflexi likely acquired dsrAB genes in multiple events. Amongst these, Nitrospirae and Deltaproteobacteria displayed the highest number of LGT involving five independent events spanning across both reductive and oxidative branches of the DsrAB tree. These findings provide evidence for the complex evolutionary history of dsr genes. Currently, it may not be possible to identify the specific lineage in which sulfite reduction first appeared; however, our extensive dataset sets the stage for future studies to investigate the evolution of dissimilatory sulfur metabolism.
Comparison of 16S rRNA and concatenated DsrAB trees for sulfate/sulfite-reducing microorganisms. Sequences are grouped at the phylum level. Trees were constructed using a consensus of neighbor-joining and maximum-likelihood phylogenies with 1000 bootstrap re-samplings each. Each phylum is colored differently to identify LGT based on inconsistent branching patterns. Phylum names with an asterisk represent sulfate/sulfite-reducing lineages that were discovered in this study. Numbers indicate number of independent LGT events associated with the specific phylum. Complete phylogenetic trees with bootstrap values are available in Data Files S3–S6. LGT events involving oxidative-type DsrAB for Nitrospirae (2 events) and Deltaproteobacteria (1 event) are not shown
Sulfate vs. sulfite reduction in organisms
To determine whether these newly identified organisms reduce sulfate vs. sulfite to sulfide we looked for the genes involved in the reduction of sulfate to sulfite, specifically adenylyl-sulfate reductase reductase subunits A and B (aprBA), sulfate adenylyl transferase (sat), and quinone-interacting membrane-bound oxidoreductase subunits A, B, and C (qmoABC) [58,59,60]. Organisms from three phyla, the dsr-containing Candidatus Lambdaproteobacteria, and asr-containing Candidatus Riflebacteria, and Candidatus Omnitrophica, lacked genes for the reduction of sulfate to sulfite suggesting that they were sulfite reducers. Sulfite utilized by these organisms may derive from the environment or is produced inside the cell as part of other sulfur metabolism pathways such as tetrathionate or thiosulfate reduction, sulfur disproportionation, or by organosulfonate respiration. This suggests that recent genome-based observations supporting potential “metabolic handoffs” between organisms (transfer of metabolites associated with energy metabolism) in the oxidative cycle of sulfur [24, 26, 61] likely extend to the reductive cycle as well [62]. Interestingly, Candidatus Rokubacteria whose DsrAB sequences represent a novel deep-branching lineage in the DsrAB tree also have apr, sat, and qmo genes that are required for sulfate reduction or sulfite oxidation. Phylogenetic analyses of the individual dsr proteins shows that the sulfate reduction system of Candidatus Rokubacteria is of mosaic evolutionary origin (Fig. 4) (Supplementary Figs. 7– 9).
Dsr operon structure and enzymatic roles of proteins involved in sulfate reduction in Candidatus Rokubacteria. Purple: genes involved in sulfate reduction to sulfite. Orange: putative enzymatic roles of genes; blue: microbial lineages with closest homologs as determined by phylogeny/BLAST against NCBI GenBank. APS refers to adenosine-5′-phosphosulfate. Green: genes involved in sulfite reduction to sulfide. This is the first case in which dsrE, dsrF, and dsrH genes are present in organisms other than sulfur-oxidizing bacteria
Prevalence of dsrT in dsrAB-containing microorganisms
In addition to dsrD, we sought evidence for hypothetical genes in proximity to known dsr genes that may help in distinguishing between DsrAB-based sulfate/sulfite reduction and sulfur oxidation pathways. We identified a hypothetical gene that encodes for the N-terminal domain of an anti-sigma factor antagonist protein [63] that almost always occurs within the operon encoding dsr genes (Fig. 5; Supplementary Fig. 4). This hypothetical protein is part of a protein family that includes the Bacillus subtilis RsbT co-antagonist protein rsbRD, which are important components of the stressosome and function as negative regulators of the general stress transcription factor sigma-B [64]. This gene is unique to DsrAB-based sulfite-reducing organisms and is mostly absent in recognized sulfur-oxidizing organisms, except for those within the phylum Chlorobi (Supplementary Fig. 10). We refer to this gene as ‘dsrT’ in accordance with homologous genes in phototrophic green sulfur bacteria from the phylum Chlorobi [65]. This gene always precedes the electron transport components encoded by dsrMKJOP genes [66] and is fused with dsrM in some organisms (Supplementary Fig. 11). Fused dsrT-dsrM genes are oriented with dsrT in the N-terminal and dsrM in the C-terminal, thereby maintaining the gene order observed in canonical dsr operons: dsrT, dsrM, dsrK. From structural predictions and conserved motifs, we hypothesize that it likely performs a regulatory function (Supplementary Fig. 12).
Dsr operon structure in previously reported (black names) and newly reported groups (blue names). Interestingly, and in contrast to the previously studied organisms for which the operon is interrupted (=SS=), the entire dsr pathway (including electron transport chain and ancillary proteins) is often encoded in a single genomic region
dsrEFH in the newly discovered dsrAB-containing microorganisms
Recent studies looking into the distribution of genes associated with the dsr operon have suggested that dsrE, dsrF, and dsrH are unique to sulfur oxidizing microorganisms and are absent in sulfate/sulfite-reducing and putative sulfur disproportionating microorganisms [16, 38, 67]. In sulfur-oxidizing microorganisms, DsrEFH can serve as an effective sulfur donor for DsrC [68]. On the other hand, co-located dsrE, dsrF, and dsrH genes are present in ~24% of the newly identified DsrAB-encoding microorganisms (30 out of 123 genomes) (Supplementary Fig. 4). These dsrEFH genes were identified in organisms from six phylum-level lineages, Actinobacteria, Candidatus Rokubacteria, Candidatus Lambdaproteobacteria, Candidatus Muproteobacteria, Nitrospirae, and Nitrospinae. Phylogenetic analysis of all identified DsrEFH shows that they cluster with well-characterized sulfur-oxidizing organisms (Fig. 6). The presence of dsrEFH genes (with well-known roles in sulfur oxidation) in organisms from Actinobacteria and Candidatus Lambdaproteobacteria is perplexing since these organisms also possess the dsrD gene that is unique to DsrAB-containing sulfite-reducing organisms. Further, the presence of dsrEFH genes in organisms within the phylum-level lineage Candidatus Rokubacteria (with a novel deep-branching clade of DsrAB) suggests that these organisms are likely involved in sulfur oxidation rather than sulfite reduction. Finally, sequences from Candidatus Muproteobacteria formed two distinct clades (Group 1, Group 2) and clustered with two separate groups, Alphaproteobacteria and Gammaproteobacteria respectively. All Candidatus Muproteobacteria with dsrEFH sequences in Group 1 also possessed a second copy of these genes that clustered with sequences from Magnetofaba australis and Candidatus Rokubacteria.
Concatenated DsrEFH protein tree inferred by maximum likelihood. Phylum-level lineages with first report of the presence of dsrEFH genes are shown in blue (from organisms with unknown-type DsrAB) and orange (from organisms with oxidative type DsrAB). Homologous TusBCD from E. coli and S. enterica were used to root the tree. Only bootstrap values >50 are shown
Electron donors and other metabolic potential associated with sulfur cycling
In order to better understand the energy metabolism and ecology of these newly identified organisms, we investigated potential electron donors for putative sulfate/sulfite reduction. Specifically, we targeted genes involved in the oxidation of hydrogen [69] (Ni–Fe hydrogenase groups I, IIa, IIb, IIIa, IIIb, IIIc, IIId, Fe–Fe hydrogenase groups A, B1/B2) and transformation of organic carbon compounds (genes involved in breakdown of cellulose, hemicellulose, chitin, pectin, starch, amino sugars, other monosaccharides, and polysaccharides) [49] and short chain fatty acids. Our analyses show that organisms from 12 of the 13 newly identified putative sulfate/sulfite-reducing DsrAB-containing lineages identified in this study possess the ability to utilize hydrogen as an electron donor (Supplementary Table 2). On the other hand, organisms from all 13 lineages possessed the ability to breakdown complex carbon compounds although the diversity of genes encoding for specific carbohydrate-active enzymes varied greatly across phyla (Supplementary Table 3). Organisms from 8 of the 13 putative sulfate/sulfite-reducing lineages possessed the ability to oxidize short chain fatty acids by beta-oxidation (Supplementary Table 4). In order to understand if these newly identified organisms were heterotrophs or autotrophs, we looked at the carbon fixation potential encoded in the genomes. We identified three different carbon fixation mechanisms, the Calvin–Benson cycle (CBB), the reverse (reductive) tricarboxylic acid (rTCA) cycle, and the Wood–Ljungdahl pathway in ~50% of all organisms (Supplementary Table 5). In total, 11 organisms contained genes encoding for the CBB cycle with 10 possessing the Form I RuBisCO and 1 organism possessed the Form II RuBisCO. Genes for the Wood–Ljungdahl pathway were encoded in 42 genomes while genes for the rTCA cycle were encoded in 7 genomes. We propose that sulfate/sulfite reduction by organisms from these newly identified lineages likely serves an important control on carbon cycling in the terrestrial and marine subsurface.
Conclusions
By the Proterozoic Eon, sulfate reduction had become a significant biological process in the oceans [70, 71]. Based on phylogenomic arguments and isotopic records, it was suggested that the capacity to reduce sulfite to sulfide emerged in thermophilic archaea around 3.5 billion years ago, and that mesophilic sulfate reducers evolved only after the rise in atmospheric oxygen level [2, 72]. Our findings indicate a complex evolutionary history of this capacity involving extensive LGT of dsr genes. Consequently, it may be impossible to constrain the specific lineage in which this metabolism first appeared. The ability for a DsrAB-based dissimilatory sulfur metabolism is now predicted in a much wider diversity of mesophilic bacterial and archaeal groups than was recognized previously. We conclude that many groups of microorganisms now known to have genes involved in dissimilatory sulfur metabolism impact biogeochemical processes in marine and terrestrial sediments, aquifers, wetlands, methane seeps, coastal marshes and estuaries, as well as agricultural and human microbiomes. Many are organisms from well-studied phyla, but still novel at the genus to class levels, but others are organisms from candidate phyla known only based on their genomes. The results underline the value of genomic analyses for prediction of key ecosystem capacities that cannot be made based on rRNA gene surveys and motivate targeted cultivation strategies for organisms currently lacking laboratory tractable representatives. Finally, these findings will better inform future microbial trait-based ecosystem models that can predict the outcomes of global change on biogeochemical processes and planetary elemental cycles [73].
Data availability
NCBI Genbank, BioProject, BioSample, and Taxonomy ID (TaxID) accession numbers for individual genomes are listed in Supplementary Table 1. Genomes are also available through ggKbase: http://ggkbase.berkeley.edu/novel_sulfate_reducers (ggKbase is a ‘live’ site, genomes may be updated after publication). The JdFR-17, JdFR-18, and JdFR-19 genomes are also available through the Integrated Microbial Genomes and Microbiomes database (IMG) through Genome IDs: 2728369317, 2728369320, 2728369322. Hmm databases used in this study are available from https://github.com/banfieldlab/metabolic-hmms. The authors declare that all other data supporting the findings of this study are available within the article and its supplementary information files, or from the corresponding author on request.
References
Finster K. Microbiological disproportionation of inorganic sulfur compounds. J Sulfur Chem. 2008;29:281–92.
Shen Y, Buick R, Canfield DE. Isotopic evidence for microbial sulphate reduction in the early Archaean era. Nature. 2001;410:77–81.
Grein F, Ramos AR, Venceslau SS, Pereira IAC. Unifying concepts in anaerobic respiration: insights from dissimilatory sulfur metabolism. Biochim Biophys Acta. 2013;1827:145–60.
Huycke MM, Gaskins HR. Commensal Bacteria, Redox Stress, and Colorectal Cancer: Mechanisms and Models. Exp Biol Med . 2004;229:586–597.
King RA, Miller JDA. Corrosion by the sulphate-reducing bacteria. Nature. 1971;233:491–2.
Voordouw G, Armstrong SM, Reimer MF, Fouts B, Telang AJ, Shen Y, et al. Characterization of 16S rRNA genes from oil field microbial communities indicates the presence of a variety of sulfate-reducing, fermentative, and sulfide-oxidizing bacteria. Appl Environ Microbiol. 1996;62:1623–9.
Jørgensen BB. Mineralization of organic matter in the sea bed—the role of sulphate reduction. Nature. 1982;296:643–5.
Bowles MW, Mogollón JM, Kasten S, Zabel M, Hinrichs K-U. Global rates of marine sulfate reduction and implications for sub-sea-floor metabolic activities. Science. 2014;344:889-891.
Pester M, Knorr K-H, Friedrich MW, Wagner M, Loy A. Sulfate-reducing microorganisms in wetlands—fameless actors in carbon cycling and climate change. Front Microbiol. 2012;3:72.
Pereira IAC, Ramos AR, Grein F, Marques MC, da Silva SM, Venceslau SS. A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front Microbiol. 2011;2:69.
Schink B, Stams AJM. Syntrophism among prokaryotes. In: Eugene Rosenberg, Edward F. DeLong, Stephen Lory, Erko Stackebrandt, Fabiano Thompson (eds) The prokaryotes. Berlin, Heidelberg: Springer; 2013. p. 471–93.
Leavitt WD, Bradley AS, Santos AA, Pereira IAC, Johnston DT. Sulfur isotope effects of dissimilatory sulfite reductase. Front Microbiol. 2015;6:1392.
Bradley AS, Leavitt WD, Johnston DT. Revisiting the dissimilatory sulfate reduction pathway. Geobiology. 2011;9:446–57.
Loy A, Duller S, Baranyi C, Mußmann M, Ott J, Sharon I, et al. Reverse dissimilatory sulfite reductase as phylogenetic marker for a subgroup of sulfur-oxidizing prokaryotes. Environ Microbiol. 2009;11:289–99.
Simon J, Kroneck PMH. Microbial sulfite respiration. Adv Microb Physiol. 2013;62:45–117.
Thorup C, Schramm A, Findlay AJ, Finster KW, Schreiber L. Disguised as a sulfate reducer: growth of the deltaproteobacterium desulfurivibrio alkaliphilus by sulfide oxidation with nitrate. MBio. 2017;8:e00671–17.
Wagner M, Roger AJ, Flax JL, Brusseau GA, Stahl DA. Phylogeny of dissimilatory sulfite reductases supports an early origin of sulfate respiration. J Bacteriol. 1998;180:2975–82.
Müller AL, Kjeldsen KU, Rattei T, Pester M, Loy A. Phylogenetic and environmental diversity of DsrAB-type dissimilatory (bi)sulfite reductases. ISME J. 2015;9:1152–65.
Hermann B, Kern M, La Pietra L, Simon J, Einsle O. The octahaem MccA is a haem c–copper sulfite reductase. Nature. 2015;520:706–9.
Shirodkar S, Reed S, Romine M, Saffarini D. The octahaem SirA catalyses dissimilatory sulfite reduction in Shewanella oneidensis MR-1. Environ Microbiol. 2011;13:108–15.
Loy A, Duller S, Wagner M. Evolution and ecology of microbes dissimilating sulfur compounds: insights from siroheme sulfite reductases. In: Christiane Dahl, Cornelius G. Friedrich (eds) Microbial sulfur metabolism. Berlin, Heidelberg: Springer; 2008. p. 46–59.
Hug LA, Baker BJ, Anantharaman K, Brown CT, Probst AJ, Castelle CJ, et al. A new view of the tree of life. Nat Microbiol. 2016;1:16048.
Hernsdorf AW, Amano Y, Miyakawa K, Ise K, Suzuki Y, Anantharaman K, et al. Potential for microbial H2 and metal transformations associated with novel bacteria and archaea in deep terrestrial subsurface sediments. ISME J. 2017;11:1915–1929.
Anantharaman K, Brown CT, Burstein D, Castelle CJ, Probst AJ, Thomas BC, et al. Analysis of five complete genome sequences for members of the class Peribacteria in the recently recognized Peregrinibacteria bacterial phylum. PeerJ. 2016;4:e1607.
Jungbluth SP, Amend JP, Rappé MS. Metagenome sequencing and 98 microbial genomes from Juan de Fuca Ridge flank subsurface fluids. Sci Data. 2017;4:170037.
Anantharaman K, Brown CT, Hug LA, Sharon I, Castelle CJ, Probst AJ, et al. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat Commun. 2016.13219.
Probst AJ, Castelle CJ, Singh A, Brown CT, Anantharaman K, Sharon I, et al. Genomic resolution of a cold subsurface aquifer community provides metabolic insights for novel microbes adapted to high CO 2 concentrations. Environ Microbiol. 2017;19:459–74.
Haft DH, Selengut JD, White O. The TIGRFAMs database of protein families. Nucleic Acids Res. 2003;31:371–3.
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, et al. Pfam: the protein families database. Nucleic Acids Res. 2014;42:D222–30.
Eddy SR. Accelerated Profile HMM Searches. PLoS Comput Biol. 2011;7:e1002195.
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7.
Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3.
Miller MA, Pfeiffer W, Schwartz T. The CIPRES science gateway. Proceedings of the 2011 TeraGrid conference on extreme digital discovery—TG ’11. New York, NY, USA: ACM Press; 2011. p 1.
Tatusov RL, Koonin EV, Lipman DJ. A genomic perspective on protein families. Science. 1997;278:631–7.
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647–9.
Dhillon A, Goswami S, Riley M, Teske A, Sogin M. Domain evolution and functional diversification of sulfite reductases. Astrobiology. 2005;5:18–29.
Mizuno N, Voordouw G, Miki K, Sarai A, Higuchi Y. Crystal structure of dissimilatory sulfite reductase D (DsrD) protein—possible interaction with B- and Z-DNA by its winged-helix motif. Structure. 2003;11:1133–40.
Venceslau SS, Stockdreher Y, Dahl C, Pereira IAC. The ‘bacterial heterodisulfide’ DsrC is a key protein in dissimilatory sulfur metabolism. Biochim Biophys Acta. 2014;1837:1148–64.
Heidelberg JF, Seshadri R, Haveman SA, Hemme CL, Paulsen IT, Kolonay JF, et al. The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. Nat Biotechnol. 2004;22:554–9.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
The UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 2015;43:D204–D212.
Dress AW, Flamm C, Fritzsch G, Grünewald S, Kruspe M, Prohaska SJ, et al. Noisy: Identification of problematic columns in multiple sequence alignments. Algorithms Mol Biol. 2008;3:7.
Lartillot N, Lepage T, Blanquart S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics. 2009;25:2286–8.
Pruesse E, Peplies J, Glöckner FO. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics. 2012;28:1823–9.
Felsenstein J. PHYLIP—Phylogeny inference package—v3.2. Cladistics. 1989;5:164–6.
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The I-TASSER Suite: protein structure and function prediction. Nat Methods. 2014;12:7–8.
Stranzl GR, Santelli E, Bankston LA, La Clair C, Bobkov A, Schwarzenbacher R, et al. Structural insights into inhibition of Bacillus anthracis sporulation by a novel class of non-heme globin sensor domains. J Biol Chem. 2011;286:8448–58.
Anantharaman K, Breier JA, Dick GJ. Metagenomic resolution of microbial functions in deep-sea hydrothermal plumes across the Eastern Lau Spreading Center. ISME J. 2015;10:225–39.
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014;42:D490–D495.
Hausmann B, Knorr K-H, Schreck K, Tringe SG, Glavina del Rio T, Loy A, et al. Consortia of low-abundance bacteria drive sulfate reduction-dependent degradation of fermentation products in peat soil microcosms. ISME J. 2016;10:2365–75.
Hallenbeck PC, Clark MA, Barrett EL. Characterization of anaerobic sulfite reduction by Salmonella typhimurium and purification of the anaerobically induced sulfite reductase. J Bacteriol. 1989;171:3008–15.
Mussmann M, Richter M, Lombardot T, Meyerdierks A, Kuever J, Kube M, et al. Clustered genes related to sulfate respiration in uncultured prokaryotes support the theory of their concomitant horizontal transfer. J Bacteriol. 2005;187:7126–37.
Rabus R, Venceslau SS, Wöhlbrand L, Voordouw G, Wall JD, Pereira IAC. Chapter Two—A post-genomic view of the ecophysiology, catabolism and biotechnological relevance of sulphate-reducing prokaryotes. Adv Microb Physiol. 2015;66:55–321.
Czyzewski BK, Wang D-N. Identification and characterization of a bacterial hydrosulphide ion channel. Nature. 2012;483:494–7.
Brown CT, Hug LA, Thomas BC, Sharon I, Castelle CJ, Singh A, et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature. 2015;523:208–11.
Anantharaman K, Duhaime MB, Breier JA, Wendt K, Toner BM, Dick GJ. Sulfur oxidation genes in diverse deep-sea viruses. Science. 2014;344:757–60.
Klein M, Friedrich M, Roger AJ, Hugenholtz P, Fishbain S, Abicht H, et al. Multiple lateral transfers of dissimilatory sulfite reductase genes between major lineages of sulfate-reducing prokaryotes. J Bacteriol. 2001;183:6028–35.
Meyer B, Kuever J. Phylogeny of the alpha and beta subunits of the dissimilatory adenosine-5’-phosphosulfate (APS) reductase from sulfate-reducing prokaryotes—origin and evolution of the dissimilatory sulfate-reduction pathway. Microbiology. 2007;153:2026–44.
Pires RH, Lourenço AI, Morais F, Teixeira M, Xavier AV, Saraiva LM, et al. A novel membrane-bound respiratory complex from Desulfovibrio desulfuricans ATCC 27774. Biochim Biophys Acta. 2003;1605:67–82.
Sperling D, Kappler U, Wynen A, Dahl C, Trüper HG. Dissimilatory ATP sulfurylase from the hyperthermophilic sulfate reducer Archaeoglobus fulgidus belongs to the group of homo-oligomeric ATP sulfurylases. FEMS Microbiol Lett. 1998;162:257–64.
Hug LA, Thomas BC, Sharon I, Brown CT, Sharma R, Hettich RL, et al. Critical biogeochemical functions in the subsurface are associated with bacteria from new phyla and little studied lineages. Environ Microbiol. 2016;18:159-73.
Wasmund K, Mußmann M, Loy A. The life sulfuric: microbial ecology of sulfur cycling in marine sediments. Environ Microbiol Rep. 2017;9:323–44.
Kim T-J, Gaidenko TA, Price CW. A multicomponent protein complex mediates environmental stress signaling in Bacillus subtilis. J Mol Biol. 2004;341:135–50.
Akbar S, Gaidenko TA, Kang CM, O’Reilly M, Devine KM, Price CW. New family of regulators in the environmental signaling pathway which activates the general stress transcription factor sigma(B) of Bacillus subtilis. J Bacteriol. 2001;183:1329–38.
Holkenbrink C, Barbas SO, Mellerup A, Otaki H, Frigaard N-U. Sulfur globule oxidation in green sulfur bacteria is dependent on the dissimilatory sulfite reductase system. Microbiology. 2011;157:1229–39.
Sander J, Engels-Schwarzlose S, Dahl C. Importance of the DsrMKJOP complex for sulfur oxidation in Allochromatium vinosum and phylogenetic analysis of related complexes in other prokaryotes. Arch Microbiol. 2006;186:357–66.
Grimm F, Franz B, Dahl C. Regulation of dissimilatory sulfur oxidation in the purple sulfur bacterium Allochromatium Vinosum. Front Microbiol. 2011;2:51.
Stockdreher Y, Venceslau SS, Josten M, Sahl H-G, Pereira IAC, Dahl C. Cytoplasmic sulfurtransferases in the purple sulfur bacterium Allochromatium vinosum: evidence for sulfur transfer from DsrEFH to DsrC Battista, JR (ed). PLoS ONE. 2012;7:e40785.
Vignais PM, Billoud B. Occurrence, classification, and biological function of hydrogenases: an overview. Chem Rev. 2007;107:4206–72.
Canfield DE, Habicht KS, Thamdrup B. The Archean sulfur cycle and the early history of atmospheric oxygen. Science. 2000;288:658–61.
Anbar AD, Knoll AH. Proterozoic ocean chemistry and evolution: a bioinorganic bridge? Science. 2002; 297:1137–1142.
Blank CE. Evolutionary timing of the origins of mesophilic sulphate reduction and oxygenic photosynthesis: a phylogenomic dating approach. Geobiology. 2004;2:1–20.
Coles VJ, Stukel MR, Brooks MT, Burd A, Crump BC, Moran MA, et al. Ocean biogeochemistry modeled with emergent trait-based genomics. Science. 2017;358:1149–54.
Acknowledgements
This work was supported by Lawrence Berkeley National Laboratory’s Sustainable Systems Scientific Focus Area funded by the U.S. Department of Energy, Office of Science, Office of Biological and Environmental Research under contract DE-AC02-05CH11231 and by the Austrian Science Fund (FWF, P25111-B22 and I2320-B22).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
:The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Anantharaman, K., Hausmann, B., Jungbluth, S.P. et al. Expanded diversity of microbial groups that shape the dissimilatory sulfur cycle. ISME J 12, 1715–1728 (2018). https://doi.org/10.1038/s41396-018-0078-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0078-0
This article is cited by
-
Microbial assimilatory sulfate reduction-mediated H2S: an overlooked role in Crohn’s disease development
Microbiome (2024)
-
Serial cultures in invert emulsion and monophase systems for microbial community shaping and propagation
Microbial Cell Factories (2024)
-
Microbial life in preferential flow paths in subsurface clayey till revealed by metataxonomy and metagenomics
BMC Microbiology (2024)
-
Autotrophic biofilms sustained by deeply sourced groundwater host diverse bacteria implicated in sulfur and hydrogen metabolism
Microbiome (2024)
-
Globally distributed marine Gemmatimonadota have unique genomic potentials
Microbiome (2024)