Abstract
Reduction of the greenhouse gas N2O to N2 is a trait among denitrifying and non-denitrifying microorganisms having an N2O reductase, encoded by nosZ. The nosZ phylogeny has two major clades, I and II, and physiological differences among organisms within the clades may affect N2O emissions from ecosystems. To increase our understanding of the ecophysiology of N2O reducers, we determined the thermodynamic growth efficiency of N2O reduction and the selection of N2O reducers under N2O- or acetate-limiting conditions in a continuous culture enriched from a natural community with N2O as electron acceptor and acetate as electron donor. The biomass yields were higher during N2O limitation, irrespective of dilution rate and community composition. The former was corroborated in a continuous culture of Pseudomonas stutzeri and was potentially due to cytotoxic effects of surplus N2O. Denitrifiers were favored over non-denitrifying N2O reducers under all conditions and Proteobacteria harboring clade I nosZ dominated. The abundance of nosZ clade II increased when allowing for lower growth rates, but bacteria with nosZ clade I had a higher affinity for N2O, as defined by μmax/Ks. Thus, the specific growth rate is likely a key factor determining the composition of communities living on N2O respiration under growth-limited conditions.
Similar content being viewed by others
Introduction
Nitrous oxide (N2O) is a potent greenhouse gas and the major ozone-depleting substance in the atmosphere [1]. In the era of global warming, it is imperative to find strategies to mitigate N2O emissions, especially from managed ecosystems with high nitrogen loadings like agricultural soils and wastewater treatment plants. For this reason, there is an increasing interest to understand the ecophysiology of N2O-reducing microorganisms [2], which have received much less attention than microbial and chemical processes that can lead to the formation of N2O [3].
The N2O reductase (NosZ), encoded by the gene nosZ, is the only known enzyme converting N2O, by reducing it to N2. There are two major clades in the nosZ phylogeny—clade I and the recently described clade II [4, 5]—and genome comparisons as well as studies focusing on the ecology and physiology of N2O reducers have suggested differences, for example regarding niches and composition of the electron transport chain, between organisms harboring the two clades [2]. Nitrous oxide reduction as part of the denitrification pathway has been extensively studied in model organisms (e.g., Paracoccus denitrificans and Pseudomonas stutzeri) but N2O reduction is not a metabolic feature restricted to denitrifiers. Many non-denitrifying N2O-reducing organisms possess N2O reductases, but lack all or some of the other reductases in the denitrification pathway [6]. Hence, there is an emerging body of literature focusing on non-denitrifying N2O-reducing organisms (which often possess clade II type nosZ; [2] and references therein), since they have the potential to increase the N2O sink capacity of soils and other environments [7, 8].
Nitrous oxide reduction conserves energy and can fully sustain the energetic needs of a cell, as organisms possessing either nosZ clade I or clade II have been successfully grown using N2O as a sole electron acceptor [9,10,11]. Recent work based on a small selection of pure cultures report a lower whole-cell half-saturation constant (Ks) for N2O and up to 1.5 times higher biomass yields among organisms with nosZ clade II compared to those with clade I [5, 12, 13]. A higher energy conservation per electron accepted could potentially be due to differences between the two clades regarding the nos gene cluster, which encodes proteins involved in the electron transport to NosZ [5, 14]. However, the physiological and bioenergetic implications of possessing nosZ clade I or clade II are largely unknown considering the broad taxonomic diversity of N2O reducers detected in the environment (e.g., refs. [7, 15]). To date, there are no reports on long-term growth of bacteria based on their N2O-reducing capacity and the selective effect of N2O as sole electron acceptor in natural communities.
Our aim was to increase the understanding of the ecophysiology of N2O-reducing microorganisms, and more specifically determine (i) the selection of bacteria, and associated nosZ genes, in an environmental community specializing in N2O reduction under N2O-limiting as well as acetate-limiting conditions, and (ii) the thermodynamic growth efficiency of N2O reduction and the efficiency of N2O respiration as reflected in the growth yields of the enrichment cultures under said conditions. We hypothesized that organisms harboring nosZ clade II genes would dominate the enrichment culture during N2O-limiting conditions since these organisms have been suggested to have a lower Ks [13] and thus, likely have a higher overall affinity for N2O (as defined by the ratio of μmax over Ks). For our purpose, we employed chemostat enrichment cultures using activated sludge from a wastewater treatment plant as our seed community and fed with N2O as the sole electron acceptor and acetate as the only energy and carbon source. We worked at two different dilution rates and with either the electron acceptor (N2O-limiting condition) or the donor as a limiting factor (acetate-limiting condition). With the enrichment approach, we could monitor the abundance, diversity, and bioenergetics of naturally occurring N2O reducers potentially relevant in wastewater treatment or other environments instead of narrowing our view to a few model organisms. In contrast to batch cultures, the chemostat set-up also allowed us to effectively address the effects of limiting conditions. We chose acetate as a carbon and energy source mainly to avoid enrichment of non-N2O-reducing microorganisms, as acetate cannot be readily converted in the absence of an external electron acceptor. It also allowed us to compare our results to other studies reporting differences in physiology among nosZ clade I and II organisms grown on acetate [5, 13]. As a control, a pure culture of Pseudomonas stutzeri, closely related to the dominant population in the enrichment culture during high dilution rates, was grown under similar conditions.
Materials and methods
Chemostat operation for enrichment of N2O reducers
An acetate-consuming N2O-reducing culture was enriched from activated sludge and cultivated in a double-jacket glass bioreactor with a working volume of 2 l (Applikon, Delft, the Netherlands) operated as a continuous stirred tank reactor (i.e., a flow-controlled chemostat) during 195 days. Mixing was achieved with a stirrer having two standard geometry six-blade turbines turning at 750 rpm. The dilution rate was controlled by two peristaltic pumps feeding-concentrated medium and water to the system and an effluent pump controlled by a level sensor. The reactor temperature was maintained at 20 ± 1 °C using a cryostat bath (Lauda, Lauda-Köningshofen, Germany). The pH was monitored with a pH electrode (Mettler Toledo) and maintained at 7.0 ± 0.05 by titration of 1 M HCl controlled by an ADI 1030 biocontroller (Applikon).
During the experiment, the chemostat system was operated under four different operational conditions, referred to as periods I–IV (Table 1 and Figure S1), with a minimum of 30 volume changes each. The shifts between N2O and acetate limitation were achieved by changing the N2O supply rate (Fig. S1). A variable flow of N2 and N2O (summing up to a total of 100–800 ml min−1) was sparged into the reactor, controlled by two separate mass flow controllers (Brooks Instruments, Ede, the Netherlands). While N2O served as the electron acceptor, N2 gas was used as a dilution gas to increase mixing and enhance gas–liquid mass transfer for both N2O and CO2. The N2O concentration was not directly measured in the liquid, but calculations based on the N2O concentration in the off-gas using mass transfer laws show that N2O was growth limiting when acetate was present in excess (Table 1). The medium was supplied in two separate flows of concentrated acetate and mineral medium, respectively. A second peristaltic pump supplied tap water to the system to dilute the medium. The final influent contained 45.3 mmol acetate (NaCH3COO·3H2O), 13.3 mmol NH4Cl, 7.4 mmol KH2PO4, 2.1 mmol MgSO4·7H2O, 0.5 mmol NaOH, 2 mg yeast extract and 2.5 ml trace element solution [16] per liter.
Prior to the experimental periods I–IV, the reactor was inoculated with activated sludge from the wastewater treatment plant of Harnaschpolder, the Netherlands, and run for a period of 118 days with similar conditions as in the experimental periods I and II (Table S1). When changing dilution rate, and one additional time shortly after changing to periods III and IV, the reactor was re-inoculated with 20–100 ml sludge from Dokhaven wastewater treatment plant to minimize legacy effects of the previous periods for selection of bacteria under the actual conditions in each period. The operation of the Harnashpolder and Dokhaven plants is described in Gonzalez-Martinez et al. [17]. The chemostat was operated under non-sterile conditions and cleaned approximately every 3 weeks to remove any biofilm present. The contribution of biofilm growth to the amount of biomass inside the reactor was negligible.
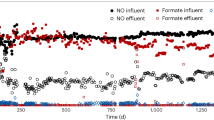
Before terminating the chemostat experiment, a batch experiment was performed on day 192 to compare the maximum acetate conversion rate of the enrichment with either N2O or NO3− as an electron acceptor. During these tests, the influent and effluent pumps of the chemostat were stopped, but flushing with N2 gas was kept constant (at 800 ml min−1). The sparging with N2O was initially increased to obtain maximum conversion rates on N2O and then stopped before adding 1 mM NO3− and monitoring its reduction.
Growth of P. stutzeri JM300 under N2O and acetate-limiting conditions
A pure culture of P. stutzeri JM300, a strain closely related to the dominant population in the enrichment culture during periods I and II, was grown under alternating N2O and acetate-limiting conditions. The chemostat reactor set-up was similar to the one described above, except for operating at 30 ± 1 °C, under sterile conditions, and using 1 M H2SO4 for pH control. The substrate and mineral medium were fed simultaneously, but final concentrations in the influent were the same as in the enrichment culture. The mineral medium was adjusted to a defined medium by removing the yeast extract. The chemostat was inoculated with a pre-culture of P. stutzeri JM300 grown aerobically in a shake flask and harvested at exponential phase. Start-up of the reactor was initially in batch mode with the defined mineral medium supplemented with 30 mM NO3− to induce denitrification. Once the NO3− was depleted, N2O sparging was initiated, and medium was added at a dilution rate of 0.044 h−1. Like in the enrichment, the N2O supply rate varied to achieve N2O-limiting vs. acetate-limiting conditions. The concentration of N2O, diluted in N2 supplied at a rate of 600 ml min−1, was 0.62–0.93%.
Analytical procedures
Samples from the reactor for analysis of acetate and NH4+ were immediately filtered after sampling (0.45-μm pore size poly-vinylidene difluoride membrane, Merck Millipore, Carrigtohill, Ireland). Acetate was measured with a Chrompack CP 9001 gas chromatograph (Chrompack, Middelburg, the Netherlands) equipped with an HP Innowax column (Algilent Technologies, Santa Clara, CA, USA) and a flame ionization detector. Ammonium, as well as NO3− and NO2−, was determined spectrophotometrically using cuvette test kits (Hach Lange, Düsseldorf, Germany). For the estimation of biomass concentration, the volatile suspended solids (VSS) concentration was determined by centrifuging 0.2 l of the enrichment, drying the pellet overnight at 105 °C, and then burning the pellet at 550 °C for 2 h to determine the ash content. Concentrations of N2O, N2, and CO2 in the headspace of the reactor were monitored in dried gas, either by using an infrared gas analyzer (NGA 2000, Rosemount, Chanhassen, MN, USA) or through mass spectrometry (Prima BT, Thermo Scientific).
Elemental and electron balances were set up to determine the conversions taking place within the chemostat. To convert VSS to biomass, a biomass composition of CH1.8 O0.5 N0.2 was assumed [18]. The N2O consumption and N2 and CO2 production rates were computed from the off-gas partial pressure and the gas supply rate. Dissolved N2O as well as dissolved CO2 and ionized species were included in the mass balances. For the electron balance, an average of the N2O consumption and the N2 production was used to estimate the moles of electrons accepted.
To monitor the microbial community structure of the enrichment, the reactor was sampled regularly for microscopy, FISH (fluorescent in situ hybridization), and DNA extraction for quantitative PCR and sequencing as described below.
DNA extraction and quantitative PCR of nirK, nirS, nosZ, and 16S rRNA genes
DNA was extracted from a pellet retrieved from 2 ml of the enrichment culture at each sampling occasion, and from the activated sludge samples used as inoculum using the PowerLyzer PowerSoil DNA Isolation Kit (MoBio Laboratories). DNA was quantified using the Qubit Fluorometer (Life Technologies Corporation).
To determine the abundance of the denitrifier, nitrous oxide reducing, and total bacterial communities in the enrichment, the genes nirS and nirK, nosZ clade I and II, and the 16S rRNA gene were quantified using quantitative real-time PCR (qPCR). Each independent duplicate reaction contained 5 ng template DNA, iQ SYBR Green Supermix (BioRad), 0.1% BSA, and primer concentrations of 0.25 µM for nirK, 0.5 µM for the 16S rRNA gene and nirS, and 0.8 µM for both nosZ clades. Primer sequences and thermal cycling conditions are available in Table S2. Standard curves were obtained using serial dilutions of linearized plasmids containing fragments of the respective genes. We tested for PCR inhibitors in all DNA extracts with a plasmid-specific qPCR assay (pGEM-T; Promega) and no inhibition was detected when comparing to controls with only the plasmid added.
Amplicon sequencing of 16S rRNA genes and nosZ clade I and II
The composition of the bacterial and N2O-reducing communities were determined by amplicon sequencing of 16S rRNA genes and the nosZ genes of both clades. A two-step PCR protocol was used [19]. The 16S rRNA genes were amplified in duplicate using 25 + 8 cycles and the primers pro341F and pro805R [20]. Amplicons were pooled and purified with the AMPure Beads Purification kit (Agilent Technologies) in between and after the final amplification step. The PCR contained 10 ng template DNA, Phusion High-Fidelity PCR Master Mix (Thermo Scientific), 0.75 µg µL−1 BSA and 0.25 µM of each primer, equipped with Nextera adapter sequences in the second PCR. The amplification of nosZ clade I and II were done in duplicate with 20 + 12 cycles for nosZI and 25 + 15 cycles for nosZII as described in Jones et al. [7]. The AMPure Beads Purification kit was used for purification of the final PCR. The reactions contained 20 ng template DNA, DreamTaq Green PCR Master Mix (Fermentas), 0.1% BSA, 1 mM MgCl2, and 0.8 µM of either nosZ clade I or II specific primers. Primer sequences and cycling conditions are listed in Table S2. Sequencing was performed by Microsynth (Microsynth AG) on the MiSeq platform (Illumina) using 2 × 300 bp paired-end chemistry for 16S rRNA genes and on a 454 FLX Genome Sequencer (Roche) using Titanium FLX + chemistry for the nosZ genes. The sequences obtained in this study are available at The Sequence Read Archive under the accession number PRJNA398140.
Bioinformatic analysis
The 16S rRNA gene sequences were trimmed with the FASTX-Toolkit (http://hannonlab.cshl.edu/fastx_toolkit) and paired-end reads were merged with PEAR [21] using a minimum overlap of 30 bp. Further quality filtering was done using VSEARCH [22]. The sequences were clustered at 97% nucleotide similarity into operational taxonomic units (OTU), followed by de novo and reference-based chimera checking (Gold database retrieved from UCHIME; [23]). All OTUs that comprised less than 1% of all sequences within each sample were removed from the data set. This resulted in a total of 2,210,612 sequences clustering into 82 OTUs. Taxonomy assignment was done with the SINA aligner using the SILVA taxonomy [24].
The nosZ sequence data were screened and de-multiplexed using QIIME [25]. Frameshift correction and removal of contaminating sequences was performed using HMMFRAME [26], with HMM profiles based on separate reference alignments of full-length nosZ amino acid sequences obtained from genomes for each clade [27]. Chimera checking (de novo and reference-based using the nosZ database retrieved from FunGene; [28]) and OTU clustering at 97% nucleotide similarity were performed using USEARCH in QIIME. The full set of OTUs was reduced in the same way as for the 16S rRNA gene data set, which resulted in 300,554 sequences for clade I and 143,185 for clade II corresponding to 79 and 102 OTUs, respectively. The representative sequences of all 181 OTUs were aligned to the reference alignment containing 624 full-length nosZ amino acid sequences using HMMER [29] and the alignment was edited manually using ARB [30]. The final nosZ phylogeny was generated from the amino acid alignment with FastTree 2 [31] using the WAG + CAT substitution model [32].
FISH and microscopic analysis of the culture
FISH was performed using the probes listed in Table S3 as described by Johnson et al., [33], using a hybridization buffer containing 35% (v/v) formamide. Slides were observed with an epifluorescence microscope (Axioplan 2, Zeiss, Sliedrecht, the Netherlands) and images were acquired with a Zeiss MRM camera, compiled with the Zeiss microscopy image acquisition software (AxioVision version 4.7, Zeiss)
Results
Conversion rates and biomass yields of the enrichment culture and P. stutzeri JM300
An enrichment culture using acetate as a carbon source and exogenous N2O as the sole electron acceptor was successfully maintained during a period of 195 days under two different dilution rate regimes and subjected to either electron donor (acetate) or electron acceptor (N2O)-limiting conditions (Table 1). The limiting substrates could not be detected in the chemostat, which supports that we had obtained limiting conditions (Table 1). That N2O was growth limiting when acetate was present in excess was further verified by shortly increasing the N2O supply rate and observing an increase in biomass-specific N2O conversion rates during an N2O-limiting period (data not shown). The conversion rates were averaged for each of the four conditions, and checked for consistency by evaluating the carbon and electron balances (>90% of C and electrons recovered; Table 2). The stoichiometry deduced from the conversion rates (Table 2) and the biomass yields (Table 3) shows that the N2O:acetate ratio used by the culture was 2.5–3.1 mol/mol depending on the limitation regime.
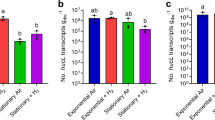
When deriving the biomass yield of the culture from the conversion rates (Table 2), we found higher values during the N2O-limited conditions (periods I and IV) compared to the acetate-limited conditions (periods II and III; Table 3). To validate these results, we determined the biomass growth yields during N2O versus acetate-limiting conditions in a pure culture of Pseudomonas stutzeri JM300, a strain closely related to the dominant OTU in the enrichment culture when the dilution rate was high (periods I and II; see below). Although biomass yields overall were lower for P. stutzeri JM300 compared to the enrichment, the pattern with lower growth per mole of substrate (almost 30% lower) during acetate-limited growth compared to N2O-limited growth was similar (Table 3). Moreover, the NH4+ consumption was higher than expected (Table 3) and this was also the case in the enrichment when considering the measured nitrogen content in the biomass (0.21–0.23 mole N/C-mole biomass; data not shown). At the end of the experiment, on day 192, the enrichment culture was able to reduce NO3− with transient accumulation of NO2− and NO, but no detectable N2O. The maximum acetate oxidation rate during NO3− reduction was roughly 60% of that during N2O reduction (1.05 vs. 1.65 mM h−1 acetate, respectively; Figure S4).
Abundance of bacterial community and functional genes during chemostat operation
Organisms harboring nosZ clade I dominated the system over those with nosZ clade II throughout operation of the chemostat (Fig. 1). At the higher dilution rate (periods I and II), nosZ clade I gene abundance was in the same order of magnitude as that of the 16S rRNA genes and was unaffected by the limitation regime. After the shift to a lower dilution rate (periods III and IV), nosZ clade I remained the dominant clade, although the average gene copy number decreased by one order of magnitude. This co-occurred with an increase by one to two orders of magnitude of the nosZ clade II abundance.
The nirS and nirK genes were abundant in the enrichment culture throughout the entire time of operation (Fig. 1). Overall, nirS dominated over nirK, except for a short period in which both genes were equally abundant (period II). The abundance of nirS was strongly affected by the shift from nitrous oxide to acetate limitation during the low dilution rate, whereas nirK and nosZ genes were mainly affected by the change in dilution rate.
Composition of the enriched N2O-reducing community
A relatively simple community was selected in the N2O-reducing enrichment culture with four to five nosZ OTUs representing the majority of the sequences at any given time period (Fig. 2). At the higher dilution rate, both the nosZ clade I and clade II community structure remained similar regardless of whether the electron acceptor (N2O) or donor (acetate) was the growth-limiting substrate. After decreasing the dilution rate, the community composition changed and then changed again when switching from acetate limitation to N2O limitation. The nosZ clade I OTUs included sequences clustering closely with nosZ from the families Pseudomonadaceae, Rhodocyclaceae, Rhodobacteraceae, and Comamonadaceae, the latter including the genus Acidovorax (Fig. 2 and S2). The nosZII-harboring community was more diverse and most dominant OTUs clustered with Rhodocyclaceae (including the genera Azonexus and Dechloromonas) and Flavobacteriaceae (including Chryseobacterium and Riemerella).
Relative abundance of nosZI (a) and nosZII (b) OTUs with >10% of the sequences at any given date during operation of the N2O-reducing chemostat under four conditions (I–IV). The OTUs are listed on the right-hand side with genus/family indicated in parenthesis (see Figure S2). Closely related OTUs are shown in shades of the same color
Composition of the overall, taxa-based bacterial community
In line with the results for the nosZ communities, the analysis of the 16S rRNA gene reads revealed that a relatively simple community was enriched, dominated by a few OTUs, and the changes in patterns over time were similar to those observed for the nosZ communities (Fig. 3). The main OTU at the high dilution rate belonged to the genus Pseudomonas, and coexisted with OTUs related to Comamonadaceae, Rhizobium, Flavobacteriaceae, and the phylum Gracilibacteria (Fig. 3a and Table S4). With a lower dilution rate and acetate limitation, the enrichment became dominated by Betaproteobacteria sequences, notably Azoarcus-like, but also transiently by sequences affiliated with Rhodocyclaceae. From the relative abundance patterns of 16S rRNA and nosZ gene sequences we could, with some caution, assign some of the main 16S rRNA OTUs to the nosZ OTUs (Fig. 3b). The Pseudomonas sp. (16S rRNA OTU 1), which dominated the community during periods I and II, might have been poorly resolved as two divergent copies of nosZI (nosZI OTUs 0 and 1) co-occurred with this taxon. Alternatively, it possessed two different copies of the gene. It is likely closely related to Pseudomonas stutzeri TS44 (which possesses two closely related copies of nosZI). OTU 2 in the nosZI community, assigned as Rhodocyclaceae, could not be assigned at the genus level, but likely corresponds to the same organism as OTU 3 of the 16S rRNA gene survey, as shown in Fig. 3b. No nosZ OTU matched the relative abundance pattern of the 16S rRNA OTU 2 belonging to the Gracilibacteria phylum.
Changes in the relative abundance of the main 16S rRNA gene OTUs and the related nosZ OTUs during operation of the N2O-reducing chemostat under four conditions (I–IV). a Contribution of the main 16S rRNA gene OTUs with >10% of the sequences at any given date. b Comparison between the major 16S rRNA gene OTUs and the nosZ OTUs with corresponding relative abundance patterns. For taxonomy assignment of the 16S rRNA gene OTUs, see Table S4
When the relative abundances of the main groups present in the enrichment were independently validated using FISH, the results roughly corroborated those obtained by 16S rRNA sequencing (Figure S4). For example, Gammaproteobacteria dominated the enrichment during periods I and II, and were later washed out and replaced by Betaproteobacteria. Microscopy did not reveal the presence of other cells than prokaryotes.
Discussion
An enrichment culture growing by N2O reduction to N2 at the expense of acetate oxidation was maintained for an extended number of generations. The availability of N2O was not a selective driver for non-denitrifying N2O reducers, as mainly bacteria with a denitrification pathway were selected in the enrichment, irrespective of N2O-limiting or acetate-limiting conditions. This was inferred from the abundances of nir and nosZ genes, phylogenetic placement of nosZ genes, classification of 16S rRNA gene sequences and what is known about these organisms. Both the nosZ clade I and II communities were dominated by Proteobacteria and all major OTUs from the 16S rRNA gene survey were affiliated with this phylum. The dominant 16S rRNA gene-based OTUs were closely related to Pseudomonas stuzeri (periods I and II) and Azoarcus (period III). These taxa are well-known denitrifiers that possess both nir and nosZ clade I genes [6]. The most abundant nosZ clade II OTU selected in the enrichment was closely related to nosZ in Dechloromonas—a Betaproteobacteria known to perform full denitrification—even though, in general, nosZ clade II represents a taxonomically highly diverse clade dominated by non-denitrifying N2O reducers [2]. The ratio of nir to nosZ gene abundances based on qPCR also indicates selection for complete denitrifying bacteria. This is further supported by the batch test performed on day 192 to compare the maximum conversion rates of the enrichment with either N2O or NO3− as an electron acceptor. The NO3−-reducing capacity of the culture was in the same order of magnitude as the N2O-reducing capacity. Moreover, these tests showed that the full denitrification pathway was constitutively or rapidly induced even after growth with N2O as a sole electron acceptor for a long period. Apart from the expected N2O reducers, the presence of a relatively abundant OTU closely related to Gracilibacteria is intriguing, since organisms belonging to this phylum are most probably fermentative organisms lacking any type of electron transport chain [34]. However, this phylum is understudied and might include members with other metabolic features than what is currently described. Their presence could also indicate important microbial interactions that we cannot account for (e.g., cross-feeding, secretion of inhibiting substances, etc.) resulting in the co-existence of N2O-reducing denitrifiers and other organisms . We can only speculate about the Gracilibacteria living off the fermentation of byproducts excreted by denitrifiers or products of cell lysis.
In contrast to what was suggested by Yoon et al. [13], our results indicate that organisms with nosZ clade I display a higher overall affinity for N2O under the conditions studied, since organisms harboring nosZ clade I genes dominated the enrichment even when N2O was limiting. This does not necessarily imply that organisms with nosZ clade I have a lower affinity constant (KS) for N2O, as the overall affinity for a substrate in a continuous culture is determined not only by the Ks, but by the ratio of μmax to KS [35]. During the periods with a high dilution rate, an OTU closely related to P. stutzeri, known to be a fast grower with a high maximum growth rate on a variety of substrates [36, 37], dominated the enrichment. Enrichments with N2O as electron acceptor in batch cultures, where maximum growth rate is the selective factor, often select for this species [38, 39]. Although the apparent Ks for N2O is claimed to be relatively high in certain P. stutzeri strains [13], its high maximum specific growth rate may have very well compensated for it when competing for N2O uptake during N2O-limiting conditions in our reactor. The OTU related to P. stutzeri that dominated under both N2O and acetate-limiting conditions is likely two closely related strains harboring two divergent gene copies of nosZ (clade I) according to the nosZ phylogeny. One could speculate that these divergent copies offer different advantages if they are expressed and translated into functional proteins under different conditions. Within a single isolate of a Bacillus sp. having two phylogenetically divergent nosZ clade II genes, N2O reduction could be maintained at different pH values when compared to closely related strains with only one nosZ copy [40].
After the switch to a lower dilution rate, the P. stutzeri-dominated community was washed out and replaced by a more diverse community, but still dominated by nosZ clade I bacteria. The dilution rate imposed in this study was low enough to be able to sustain growth of the nosZ clade II bacterium Dechloromonas aromatica, but too high for the three other nosZ clade II species for which μmax has been determined [13]. The evidence for a lower μmax among organisms with clade II than organisms with nosZ type I is limited, but if this would be the case, nosZ clade II could potentially increase when decreasing the dilution rate even further. In agreement, the qPCR results indicate a significant increase in the abundance of nosZ clade II when the dilution rate was lowered, irrespective of the limiting substrate. In natural and engineered systems, microorganisms typically grow at rates lower than those used in our experiment (e.g., in the order of 0.2–0.05 d−1 in activated sludge WWTP). This could explain why nosZ clade II genes are often detected in equal or higher abundance compared to nosZ clade I in many environments [4]. In contrast, the ratios between nosZ clade I and II observed in root-associated communities are higher compared to those in bulk soil [41, 42]. The rhizosphere is an example of an environment where carbon supply is high, and high growth rate or overall affinity would be a competitive advantage. It would be interesting to address the competition of nosZ clade I vs. clade II under N2O limitation under a larger range of growth rates, especially lower than those used here.
The growth yield per electron accepted of the enrichment was in the same range (or lower) as the reported yields of denitrifying pure cultures using acetate as a carbon source (refs. [5, 13, 43]; Table S6). However, growth yields were significantly different depending on whether acetate or N2O was the limiting substrate, independent of the dilution rate. The biomass yield per electron accepted identified during the N2O-limiting periods was comparable to two denitrifying enrichments grown under either acetate or NO3− limitation with comparable conditions regarding pH, temperature, inoculum, etc. (0.38 ± 0.04 and 0.34 ± 0.05 CmolX/CmolS, respectively; data not shown). This suggests that the yield during acetate limitation was unexpectedly low. The difference in yield cannot be related to the selection of different communities with different electron transport pathways for N2O respiration, since it was observed not only between periods I and II, which had a similar community structure, but also in the pure culture of P. stutzeri JM300. We therefore hypothesize that the stunted growth during acetate limitation was caused by cytotoxic effects of excess N2O. Cytotoxicity of N2O with concentrations as low as 0.1 μM, which is far below the concentration in our system during acetate limitation, has been reported for Paracoccus denitrificans [44]. This effect was attributed to the destruction of vitamin B12, which is necessary for methionine and DNA synthesis. In P. denitrificans, high levels of N2O presumably lead to a switch to vitamin B12-independent pathways, which are energetically costly. An increased energy expense in the presence of excess N2O could explain why we found lower growth yields in the enrichment during the acetate-limited periods. In broader perspective, cytotoxic effects of N2O on certain organisms would be another mechanism of selection effects of N2O in a community, in addition to the organisms’ μmax and KS for N2O. Metatranscriptomics when N2O is present in excess may provide more supporting evidence for this hypothesis. In the environment, N2O reduction could be a detoxification mechanism without being linked to energy conservation. The relative importance of detoxification and anaerobic respiration for N2O reduction in the environment is not known [2], but in our chemostat, N2O reducers would necessarily have to harvest energy during N2O reduction, as it was the sole electron acceptor provided.
Overall, denitrifying bacteria were favored over non-denitrifying N2O reducers when N2O was the sole electron acceptor, even under N2O-limiting conditions. A high affinity for N2O may be advantageous for complete denitrifiers to avoid scavenging, by other microorganisms, of the N2O produced in the first steps of denitrification. This strategy would allow them to harvest the energy available in N2O reduction to N2 in addition to the rest of the denitrification steps, which may be particularly advantageous when there is limited access to electron acceptors in the environment. Denitrifiers with nosZ clade I were favored over microorganisms with nosZ clade II under all conditions and our results suggest that the μmax was important in the selection of different N2O-reducing communities. The conditions are not typical for specific ecosystems, but rather reflect growth-limited conditions determined by substrate supply rates. We cannot exclude that the choice of acetate as an electron donor and carbon source, or the absence of O2 and N oxides other than N2O, which are known to be involved in the regulation of nosZ expression in model denitrifiers (e.g., Paracoccus denitrificans [45, 46]) and in Gemmatimonas aurantiaca (nosZ clade II [47]), influenced the clade I vs. clade II competition, and further studies are required to look into this. Thus, other conditions than evaluated in the present study could be needed to enrich and study the elusive non-denitrifying N2O reducers with clade II nosZ. Recent work suggests that these organisms form biotic interactions with truncated denitrifiers in soils and sediments [27, 48] and further work should address if these interactions could be exploited in engineered and managed systems to mitigate N2O emissions.
References
Ravishankara AR, Daniel JS, Portmann RW. Nitrous oxide (N2O): the dominant ozone-depleting substance emitted in the 21st century. Science. 2009;326:123–5.
Hallin S, Philippot L, Löffler FE, Sanford RA, Jones CM. Genomics and ecology of novel N2O-reducing microorganisms. Trends Microbiol. 2018;26:43–55.
Schreiber F, Wunderlin P, Udert KM, Wells GF. Nitric oxide and nitrous oxide turnover in natural and engineered microbial communities: biological pathways, chemical reactions, and novel technologies. Front Microbiol. 2012;3:372.
Jones CM, Graf DRH, Bru D, Philippot L, Hallin S. The unaccounted yet abundant nitrous oxide−reducing microbial community: a potential nitrous oxide sink. ISME J. 2013;7:417–26.
Sanford RA, Wagner DD, Wu Q, Chee-Sanford JC, Thomas SH, Cruz-García C, et al. Unexpected nondenitrifier nitrous oxide reductase gene diversity and abundance in soils. Proc Natl Acad Sci USA. 2012;109:19709–14.
Graf DRH, Jones CM, Hallin S. Intergenomic comparisons highlight modularity of the denitrification pathway and underpin the importance of community structure for N2O emissions. PLoS ONE. 2014;9:e114118.
Jones CM, Spor A, Brennan FP, Breuil M-C, Bru D, Lemanceau P, et al. Recently identified microbial guild mediates soil N2O sink capacity. Nat Clim Chang. 2014;4:801–5.
Domeignoz-Horta LA, Putz M, Spor A, Bru D, Breuil MC, Hallin S, et al. Non-denitrifying nitrous oxide-reducing bacteria - an effective N2O sink in soil. Soil Biol Biochem. 2016;103:376–9.
Allen MB, Van Niel CB. Experiments on bacterial denitrification. J Bacteriol. 1952;64:397–412.
Sacks LE, Barker HA. Substrate oxidation and nitrous oxide utilization in denitrification. J Bacteriol. 1952;64:247–52.
Yoshinari T. N2O reduction by Vibrio succinogenes. Appl Environ Microbiol. 1980;39:81–84.
Kern M, Simon J. Three transcription regulators of the Nss family mediate the adaptive response induced by nitrate, nitric oxide or nitrous oxide in Wolinella succinogenes. Environ Microbiol. 2016;18:2899–912.
Yoon S, Nissen S, Park D, Sanford RA, Löffler FE. Nitrous oxide reduction kinetics distinguish bacteria harboring clade I NosZ from those harboring clade II NosZ. Appl Environ Microbiol. 2016;82:3793–800.
Simon J, Klotz MG. Diversity and evolution of bioenergetic systems involved in microbial nitrogen compound transformations. Biochim Biophys Acta. 2013;1827:114–35.
Wittorf L, Bonilla-Rosso G, Jones CM, Bäckman O, Hulth S, Hallin S. Habitat partitioning of marine benthic denitrifier communities in response to oxygen availability. Environ Microbiol Rep. 2016;8:486–92.
Vishniac W, Santer M. The Thiobacilli. Microbiol Mol Biol Rev. 1957;21:195–213.
Gonzalez-Martinez A, Rodriguez-Sanchez A, Lotti T, Garcia-Ruiz MJ, Osorio F, Gonzalez-Lopez J, et al. Comparison of bacterial communities of conventional and A-stage activated sludge systems. Sci Rep. 2016;6:18786.
Roels JA. Simple model for the energetics of growth on substrates with different degrees of reduction. Biotechnol Bioeng. 1980;22:33–53.
Berry D, Ben Mahfoudh K, Wagner M, Loy A. Barcoded primers used in multiplex amplicon pyrosequencing bias amplification. Appl Environ Microbiol. 2011;77:7846–9.
Takahashi S, Tomita J, Nishioka K, Hisada T, Nishijima M. Development of a prokaryotic universal primer for simultaneous analysis of Bacteria and Archaea using next-generation sequencing Bourtzis K (ed). PLoS ONE. 2014;9:e105592.
Zhang J, Kobert K, Flouri T, Stamatakis A. PEAR: a fast and accurate Illumina paired-end reAd mergeR. Bioinformatics. 2014;30:614–20.
Rognes T, Flouri T, Nichols B, Quince C, Mahé F. VSEARCH: a versatile open source tool for metagenomics. PeerJ. 2016;4:e2584.
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics. 2011;27:2194–2200.
Pruesse E, Peplies J, Glöckner FO. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics. 2012;28:1823–9.
Caporaso J, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Zhang Y, Sun Y. HMM-FRAME: accurate protein domain classification for metagenomic sequences containing frameshift errors. BMC Bioinforma. 2011;12:198.
Juhanson J, Hallin S, Söderström M, Stenberg M, Jones CM. Spatial and phyloecological analyses of nosZ genes underscore niche differentiation amongst terrestrial N2O reducing communities. Soil Biol Biochem. 2017;115:82–91.
Fish J, Chai B, Wang Q, Sun Y, Brown CT. FunGene: the functional gene pipeline and repository. Front Microbiol. 2013;4:291.
Eddy SR. Profile hidden Markov models. Bioinformatics. 1998;14:755–63.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar A, et al. ARB: a software environment for sequence data. Nucleic Acids Res. 2004;32:1363–71.
Price MN, Dehal PS, Arkin AP. FastTree 2 - approximately maximum-likelihood trees for large alignments Poon AFY (ed). PLoS ONE. 2010;5:e9490.
Whelan S, Goldman N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol. 2001;18:691–9.
Johnson K, Jiang Y, Kleerebezem R, Muyzer G, Van Loosdrecht MCM. Enrichment of a mixed bacterial culture with a high polyhydroxyalkanoate storage capacity. Biomacromolecules. 2009;10:670–6. American Chemical Society
Rinke C, Schwientek P, Sczyrba A, Ivanova NN, Anderson IJ, Cheng J-F, et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature. 2013;499:431–7.
Kuenen JG. Continuous cultures (Chemostats). Reference Module in Biomedical Sciences. Philadelphia: Elsevier; 2015. 2015
Lalucat J, Bennasar A, Bosch R, García-Valdés E, Palleroni NJ. Biology of Pseudomonas stutzeri. Microbiol Mol Biol Rev. 2006;70:510–47.
Sorokin DY, Teske A, Robertson LA, Kuenen JG. Anaerobic oxidation of thiosulfate to tetrathionate by obligately heterotrophic bacteria, belonging to the Pseudomonas stutzeri group. FEMS Microbiol Ecol. 1999;30:113–23.
Pichinoty F, Mandel M, Greenway B, Garcia J-L. Isolation and properties of a denitrifying bacterium related to Pseudomonas lemoignei. Int J Syst Bacteriol. 1977;27:346–8.
Desloover J, Roobroeck D, Heylen K, Puig S, Boeckx P, Verstraete W, et al. Pathway of nitrous oxide consumption in isolated Pseudomonas stutzeri strains under anoxic and oxic conditions. Environ Microbiol. 2014;16:3143–52.
Jones CM, Welsh A, Throbäck IN, Dörsch P, Bakken LR, Hallin S. Phenotypic and genotypic heterogeneity among closely related soil-borne N2- and N2O-producing Bacillus isolates harboring the nosZ gene. FEMS Microbiol Ecol. 2011;76:541–52.
Graf DRH, Zhao M, Jones CM, Hallin S. Soil type overrides plant effect on genetic and enzymatic N2O production potential in arable soils. Soil Biol Biochem. 2016;100:125–8.
Zhao M, Jones CM, Meijer J, Lundquist PO, Fransson P, Carlsson G, et al. Intercropping affects genetic potential for inorganic nitrogen cycling by root-associated microorganisms in Medicago sativa and Dactylis glomerata. Appl Soil Ecol. 2017;119:260–6.
Strohm TO, Griffin B, Zumft WG, Schink B. Growth yields in bacterial denitrification and nitrate ammonification. Appl Environ Microbiol. 2007;73:1420–4.
Sullivan MJ, Gates AJ, Appia-Ayme C, Rowley G, Richardson DJ. Copper control of bacterial nitrous oxide emission and its impact on vitamin B12-dependent metabolism. Proc Natl Acad Sci USA. 2013;110:19926–31.
Bergaust L, van Spanning RJM, Frostegård A, Bakken LR. Expression of nitrous oxide reductase in Paracoccus denitrificans is regulated by oxygen and nitric oxide through FnrP and NNR. Microbiology. 2012;158:826–34.
Qu Z, Bakken LR, Molstad L, Frostegård Å, Bergaust L. Transcriptional and metabolic regulation of denitrification in Paracoccus denitrificans allows low but significant activity of nitrous oxide reductase under oxic conditions. Environ Microbiol. 2015;18:2951–63. https://doi.org/10.1111/1462-2920.13128. e-pub ahead of print
Park D, Kim H, Yoon S. Nitrous oxide reduction by an obligate aerobic bacterium, Gemmatimonas aurantiaca strain T-27. Appl Environ Microbiol. 2017;83:e00502–17.
Baker BJ, Lazar CS, Teske AP, Dick GJ. Genomic resolution of linkages in carbon, nitrogen, and sulfur cycling among widespread estuary sediment bacteria. Microbiome. 2015;3:14.
Acknowledgements
This work was funded by the European Commission (Marie Curie ITN NORA, FP7-316472) and the Swedish Research Council (VR grant 2016-03551 to SH). We like to warmly thank Koen Verhagen for carrying out preliminary work with the N2O chemostat and Gerben Stouten for his help with the off-gas measurements.
Funding
The European Commission (Marie Sklodpowska-Curie ITN NORA, FP7-316472) and the Swedish Research Council (grant 2016-03551)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Conthe, M., Wittorf, L., Kuenen, J.G. et al. Life on N2O: deciphering the ecophysiology of N2O respiring bacterial communities in a continuous culture. ISME J 12, 1142–1153 (2018). https://doi.org/10.1038/s41396-018-0063-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0063-7
This article is cited by
-
Determining how oxygen legacy affects trajectories of soil denitrifier community dynamics and N2O emissions
Nature Communications (2024)
-
Synergistic effect of elevated CO2 and straw amendment on N2O emissions from a rice–wheat cropping system
Biology and Fertility of Soils (2024)
-
Niche Differentiation Among Canonical Nitrifiers and N2O Reducers Is Linked to Varying Effects of Nitrification Inhibitors DCD and DMPP in Two Arable Soils
Microbial Ecology (2023)
-
Higher N2O production in sequencing batch reactors compared to continuous stirred tank reactors: effect of feast-famine cycles
Frontiers of Environmental Science & Engineering (2023)
-
Identification of nosZ-expressing microorganisms consuming trace N2O in microaerobic chemostat consortia dominated by an uncultured Burkholderiales
The ISME Journal (2022)