Abstract
Metabolic abnormalities lead to the dysfunction of metabolic pathways and metabolite accumulation or deficiency which is well-recognized hallmarks of diseases. Metabolite signatures that have close proximity to subject’s phenotypic informative dimension, are useful for predicting diagnosis and prognosis of diseases as well as monitoring treatments. The lack of early biomarkers could lead to poor diagnosis and serious outcomes. Therefore, noninvasive diagnosis and monitoring methods with high specificity and selectivity are desperately needed. Small molecule metabolites-based metabolomics has become a specialized tool for metabolic biomarker and pathway analysis, for revealing possible mechanisms of human various diseases and deciphering therapeutic potentials. It could help identify functional biomarkers related to phenotypic variation and delineate biochemical pathways changes as early indicators of pathological dysfunction and damage prior to disease development. Recently, scientists have established a large number of metabolic profiles to reveal the underlying mechanisms and metabolic networks for therapeutic target exploration in biomedicine. This review summarized the metabolic analysis on the potential value of small-molecule candidate metabolites as biomarkers with clinical events, which may lead to better diagnosis, prognosis, drug screening and treatment. We also discuss challenges that need to be addressed to fuel the next wave of breakthroughs.
Similar content being viewed by others
Introduction
Metabolite biosignatures from human biofluids providing a link between genotype, environment and phenotype, are attractive biomarkers for the clinical diagnosis, prognosis, and diseases classification.1,2,3,4,5,6,7,8 It can provide a unique metabolic readout and snapshot of the health/disease status of key information about the downstream products related to various metabolic processes.9,10,11,12 Differential metabolites can improve the specificity and accuracy as biomarkers for patient diagnosis, patient monitoring, risk prediction and prognosis.13,14,15,16 Discovery and identification of small molecule metabolites or metabolic pathway alterations is useful for understanding the pathophysiology of diseases, and help identify therapeutic targets.17,18,19,20,21,22,23,24,25,26,27 Metabolome represent the upstream input from environment and downstream output of genome, the collection of bioactive small molecule metabolites including nucleotides, carbohydrates, amino acid, and fatty acid, has used for discovery of early prediction and diagnosis biomarkers of diseases that insight into the best use of interventions.28,29,30,31,32,33,34,35 Endogenous metabolites could provide unique metabolic insights into the mechanistic basis and therapeutic targets of disease and also leads to personalized metabolic phenotype.36
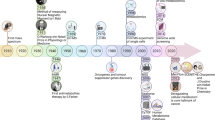
Bioactive functions and detail molecular mechanisms of small metabolites have gradually raised attention of scientists and researchers.37,38,39,40,41,42,43 Fortunately, advancements in metabolomics technologies hold promise as non-invasive and high-throughput tool that conventionally divided into untargeted and untargeted analysis has demonstrated high value in investigation of metabolite signatures, and allowed researcher to establish mass spectrometry-based comprehensive profiling of small molecule metabolites to provide insight into metabolic function.44,45,46,47,48,49,50,51,52,53,54,55. Metabolomics, the science of characterizing known and unknown small molecule metabolites, appears to be an ideally tool for disease characterization and monitoring as well as the investigation of disease pathophysiology and biochemical characteristics in body systems.56,57,58,59,60 Major approaches include metabolic phenotyping, metabolic fingerprinting, metabolic profiling and targeted metabolite analysis.61,62,63,64,65,66,67,68,69,70,71,72,73,74,75 Fig. 1 shows general workflow for biomarker discovery from small-molecule metabolites through metabolomics approach. Metabolic phenotypes could reflect the metabolic response feature variation to pathophysiological stimuli at a certain time point.76,77,78 According to specific profiles, metabolic regulation associated with therapeutic responses is new therapeutic strategy for diseases.79,80,81 Since metabolomics aims to identify small metabolites from biological system, insights into metabolism and its regulation mechanisms that symptom generation and therapeutic response, provides an innovative approach to answer phenotype-related questions distinctly altered in diseases, elucidate the biochemical functions and delineate the associated mechanisms implicated in the dysregulated metabolism from patients within clinical settings.82,83,84,85,86,87,88,89,90,91,92
Analytical workflow of small molecule metabolites-based metabolomics. The first stage involves experimental design, followed by election of biological subjects, sample collection, preparation, and metabolite extraction. Next is acquisition and processing of data, then data analysis, and finally, making sense of the data through biomarker discovery, and functional interpretation. The images were obtained using the example data provided by the MetaboAnalyst 5.0 and figures created by BioRender
Identification of the small metabolites and molecular mechanisms using high-throughput metabolomics may allow for the rapid development of biomarker and improving disease diagnosis, prognosis, treatment response, and for revealing mechanisms and disease etiology, therapeutic target for ameliorating the quality of life in patients.93,94,95,96,97,98,99 Using small molecule metabolites-based metabolomics for discovery of metabolic biomarkers to diagnosis and then providing key information for biomarker validation and elucidation of the molecular mechanism of disease, has attracted broad interest.100,101 This review focused on functional features of small molecule metabolites, utility of them as biomarkers and therapeutic targets for disease via the function relationship and associated molecular mechanisms, and also discussed its progress in the early diagnosis, prognosis, and pathogenesis of disease at the level of metabolism in vivo, which is expected to translate the milestone findings into clinical trials to enhance the efficacy and provide new sights for human clinical use in the future.
Advanced technology platform
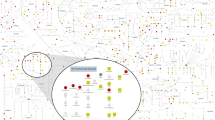
Metabolites are the final downstream products of protein translation and gene transcription or cellular perturbations to the proteome, genome or transcriptome, have potentially crucial linkage between genotype and environment, and provide a closer image of the final phenotype.102,103,104,105 A human metabolome mainly contains the detailed information of 41,993 small-molecule metabolites, has been implemented for public.106,107 Metabolites act as signaling molecules, serve as cofactors, energy production and storage, and can trigger regulation processes.108,109,110,111,112,113,114,115,116,117,118,119 Small molecule metabolites-based metabolomics have several advantages over the other omics approaches. Genomics may have little impact on expression outcome in the function of a protein, but metabolomics can directly detect the biochemical response to a stimulus.120,121,122,123 Unlike metabolomics, genomics, transcriptomics and proteomics is unable to dynamically analyze the detailed information of metabolic function in living systems.124 Considering time sensitive and accurate phenotypic analysis of live organisms, their individual diagnostic ability is lower than that of metabolomics.125,126,127,128,129 As a downstream product of transcriptome, genome and proteome, metabolome includes small molecule metabolites correlate to specific metabolic phenotype and insights into the mechanistic basis and therapeutic targets of diseases (illustrated in Fig. 2). Over the past few years, it has demonstrated significant benefits for discovering biomarkers, disease diagnosis and treatment, and delineating metabolic regulation mechanism.130,131,132,133,134,135,136 Metabolic signatures from complete system can infer the possible mechanism of diseases and identify therapeutic targets.137,138,139,140,141,142,143,144,145,146
Schematic representation of the most commonly used omic platforms for multi-omics studies. Metabolites are the downstream products of the genome, transcriptome, proteome, and enzymatic reactions, which are also affected by environmental exposures. The metabolome provides a functional readout of these upstream changes. Multi-omics (including genome, transcriptome, proteome, metabolome, and microbiome data) are collected from patients and integrated to identify personalized functional signatures using complex and comprehensive network analysis. The figures created by BioRender
Metabolome covers a suite of small metabolites with a molecular mass less than 1500 Da, including but not limited to amino acids, lipids, organic acids, and some exogenous chemicals.40,42,147,148,149,150,151 All metabolite repertoire is influenced by the physiological activity or exogenous environmental factors.152,153,154,155,156 It makes metabolome information data more difficult to interpret.157,158 Small metabolites classified as endogenous and exogenous analytes could participate in various metabolic pathways, such as urea cycle, tricarboxylic acid cycle, or fat oxidation.159,160,161 The former includes amino acids such as glycosylation products, histidine and cystine, organic acids such as succinate and citrate, lipids such as glycerolipids and sphingolipids, and other endogenous molecules.162,163,164,165,166 A wide variety of biological media has been used from all available body fluids and tissues, including serum, plasma, cerebrospinal fluid, saliva, feces, sweat, tears, urine, breast milk, cervicovaginal secretions.127,167,168,169,170,171
Molecular profiling of minor molecules offers invaluable insights into the metabolic function and targets. A disruption of metabolic pathways indicates that metabolomics might be used as a more precise tool for patients when compared with the conventional biomarkers.126,172,173 It is vital to understand the biological role of metabolites in regulating biological functions. Numerous strategies have been showed to expand small-molecule metabolites coverage.174,175,176 Mass spectrometry (MS) has been applied to the detection of small molecule metabolites, and allowing interpretation of metabolic changes at the systems-level in health and disease, from whole organisms to single cells.177,178,179,180,181,182,183,184,185,186,187,188,189,190 Metabolomics mass spectrometry-based can rapidly discover small molecule metabolites and improve the understanding metabolic mechanism of numerous diseases, and improve the ability for monitoring various metabolic changes in clinical settings.191,192,193,194 Mass spectrometry coupled with liquid chromatography platforms enhances versatility and sensitivity of identification and quantification of metabolites, precisely facilitates exploration of a large number of small-molecule metabolites from bio-samples, and describes a main picture of general metabolic changes that related to disease alteration.49,136,195 Emerging mass spectrometry imaging is a powerful analytical approach for spatial detection, quantification and imaging of endogenous and exogenous molecules.196,197,198 A cross-platform approach by integration of systems biology and small molecule data could discover the regulators of human metabolism into clinical insights.199,200,201 High-throughput metabolic profiling can reveal credible information on the underlying functional metabolic mechanisms.202,203,204,205
Technique breakthroughs have provided new opportunities to explore metabolic dimensions of diseases. Major analytic techniques for endogenous molecules include nuclear magnetic resonance (NMR) and mass spectrometry. MS can identify the low-abundance metabolites and metabolic alteration along key pathways is identifies by NMR. Recent efforts are directed towards revealing globally spatial distribution of small molecule metabolites and identifying active metabolites beyond their trend analysis and metabolites characterization.64,206,207,208 High-throughput MS imaging (MSI) technology allows for simultaneous visualization of spatial distribution of small metabolite molecules, providing attractive platforms for spatial visualization of metabolic processes to understand the complex communication networks.209,210,211 It is noteworthy that MSI technology has been successfully applied to imaging various human and animal tissues, such as liver, kidney, brain, heart, skin, breast and lens.212,213,214,215,216 NMR profiles has been largely used for characterizing biomarker and classified numerous diseases, including kidney diseases, cancer, cardiovascular diseases, Alzheimer’s disease and etc.217,218,219,220,221,222,223,224,225 At present, no single analytical method or instrument can fulfill the mission of identification of entire metabolome.226 Many reviews have recognized about the combination platform to maximize metabolomics data.227,228,229 Multiple technologies have greatly broadened the level of metabolite coverage, and several reviews have also been widely discussed regarding how different MS and NMR platforms works and their own advantages and disadvantages.135,230,231,232,233,234,235,236,237,238,239,240,241
Small molecule metabolites-based metabolomics can be categorized into targeted and untargeted approaches.65,75,134,242,243 Untargeted metabolomics reveals previously unknown metabolic information, and conversely, targeted approach highlight analyzing a set of metabolites, tend to be more sensitive and higher reproducibility relative to untargeted approach.133,244,245,246,247,248,249 Targeted metabolomics tends to analyze a specific known metabolic pathway for the metabolite quantification.250,251 However, untargeted metabolomics often focuses on a large number of unknown metabolites without bias and metabolite identification.252,253,254,255,256,257,258 Untargeted (discovery-based) approach enables global detection of all metabolites that linked phenotype information. Targeted (validated-based) metabolomics focused on the metabolites related to a metabolic pathway of interest. Due to the complex of metabolome, robust data analysis requires the preprocessing raw data followed by multivariate statistical analysis, omics data mining and bioinformatics integration.259,260,261,262,263,264,265,266,267,268,269 The larger data sets require the specialized tools for rapid analysis.270,271,272,273 The progressions such as automatic annotation, in-silico fragmentation and databases construction have advanced to solving these problems.274,275,276 Multivariate statistical techniques are widely applied in mechanistic understanding of metabolic processes, beyond phenotyping and biomarker discovery of various diseases.277,278,279,280,281,282,283,284,285,286,287,288 Data pre-processing software and numerous pattern recognition analysis packages have been reviewed elsewhere.289,290,291,292,293,294,295 Human Metabolome Database and Kyoto Encyclopedia of Genes and Genomes are the frequently used databases currently in small molecule applications field.296,297,298,299,300,301 Metabolome data can be processed automatically by bioinformatic tools.302,303,304 For instance, MetaboAnalyst tools can generalize network interaction and visualization map derive meaningful biological inferences, which includes numerous modules for pathway analysis and metabolite enrichment analysis with network topology approaches.305,306,307,308,309,310 It can provide a ranked list of potential metabolite biomarkers and fundamental metabolic pathways by allocating small metabolites to relevant biological pathways with pathophysiological basis of disease.
Identification of bioactive metabolites
Endogenous metabolites are biosynthesized by the host organism or microflora. In 1971, Linus Pauling et al. had used endogenous metabolites to reveal physiological status in biological system. Small molecule metabolites can be produced by catabolism or anabolism, such as peptides, sugars, amino acids, nucleic acids, organic acids, lipids, and fatty acids. Metabolites are the closest link between the genotype and phenotypes, and that reflects the genome, proteome, transcriptome, epigenome, and the interactions with environment.311 They play critical roles in biological pathways and serve as valuable bioindicators during cellular processes.91,312,313 Metabolic profile could provide a snapshot of complex interplay between environment and intermediary processes.314,315,316 Once specific metabolites to disease pathophysiology are identified and then gain interest in understanding biological biomarkers within mechanistic pathways, using an invasive approach to monitor disease progression and distinguish diseased subjects. To date, metabolic signatures have already been discovered from investigations to uncover biomarkers and gain insight into the ongoing metabolism and treatment targets for numerous diseases.317,318,319,320,321,322,323,324
Metabolome are comprehensively characterizing small metabolites in cells, biofluids, organs, or other biological systems. Due to the chemical complexity and dynamic range of the metabolome, the simultaneous identification and reliable quantification of metabolite features are greatly complicated. Biomarker identification can facilitate the diagnosis and prognosis of diseases or individualized treatment, better understanding and exploring potential molecular pathways and mechanisms within disease progression or modulated by drugs. Identification of active metabolite is part of the most important processes in the discovery stage.325,326,327,328,329 The biological matrices are complex with thousands of small metabolites in them, the use of analytical profiling techniques identify (global, untargeted, and top-down approach) and quantify (specific, targeted, and bottom-up approach) metabolites contribute to understanding the pathology mechanisms. Given the metabolic profile alterations, the qualitative and quantitative study technique of small metabolite molecules, provides an opportunity for identifying promising biomarkers and predictive model330,331,332,333,334,335,336,337,338,339,340,341,342.
The identification of the selected minor metabolites can be carried out by a range of analytical technology.342,343,344 Major analytical platforms for small molecules are nuclear magnetic resonance (NMR), liquid chromatography-mass spectrometry (LC-MS), and gas chromatography-mass spectrometry (GC-MS) (Fig. 3). Each technique offers unique advantages in the sensitivity, accuracy, resolution, dynamic range, reproducibility and throughput. A mass spectrometry-based strategy for identifying a list of biological activity metabolites. Moreover, it detects entire metabolites rather than a single metabolite. MS scan, high-precision MS/MS analysis combined with database (e.g., HMDB and METLIN) can provide a large number of the relatively abundant ions and acquire more reliable identifications.345,346,347 It also needs analysis software for highly complex data to complete the metabolite identification and metabolic pathways analysis.348 XCMS Online, Open-MS, MZmine and MS-DIAL software are available for peak detection and alignment.349,350,351 Untargeted approach deals with a vast number of unknown molecules and reveals functional changes, and then targeted approach focus on accurate identification and quantitation is subsequent validation including sample preparation, data acquisition and analysis. NMR data processing has been accessible via NMRbox for metabolite identification. Software tools and substantial spectral databases facilitate the identification of small metabolites by both 1D-NMR (e.g., B.I. QUANT, Chenomx NMR Suite, Bayesil, MagMet) and 2D-NMR (e.g., COLMAR).352,353,354,355,356,357,358 However, combining NMR and MS data greatly improves the metabolome coverage and enhances the accuracy of small metabolite identification, greatly benefit the quality of data.359
Overview of advanced technology platform for metabolite quantification in biomedicine. Step 1: Sample preparation through deproteinization and/or centrifugation of biofluids. Step 2: Detection of analyte signal through NMR or MS spectroscopy. Step 3: Small metabolites is filtered and quantified for significant biomarkers of interest. The images were obtained using the example data provided by the MetaboAnalyst 5.0 and figures created by BioRender
Numerous groups seek to provide the available online tools for statistical and bioinformatic analysis, e.g., Metlin, MetaboAnalyst, KEGG.58,360,361,362,363,364 Small metabolite abundance is quantified depending on peak intensity. The biological activity metabolites are selected by specific statistical cutoff (e.g. a fold changeå 2 or a p value < 0.01). Correlations calculated the association between metabolites and clinical features and further evaluated the underlying metabolism differences. To find a panel of metabolites as possible biomarkers for the specific condition, each metabolite needs to be independently analyzed to illustrate the diagnostic ability. The area under the ROC curve (AUC) measured accuracy to see how the metabolites contribute to group separation and ROC analysis could check of the performance of particular metabolites for a diagnostic test.172,365,366,367,368 To evaluate the overall performance of small metabolites for diagnosis, the sensitivities, AUC values, specificities were evaluated. A total of 24368 metabolites has been published according to recent HMDB 5.0 database.106 Number of small metabolites was identified in urine and blood are 5661 and 38,036, respectively. In recent decades, many small metabolites have been discovered in diseases progression, and these studies need emphasize metabolite bioactivity and provide their relevant biological significance.
Exploring phenotype signatures
Metabolites are end-products or intermediates of the metabolism processes and closely linked to the phenotype of a biological system, which govern the modulating the phenotype function. Level changes of small metabolites could be used as diagnostic and prognostic biomarkers as well as therapeutic targets.320,369,370,371,372 Metabolome is in constant change, and thus a more reflection of body phenotype than the other “-omics”, such as transcriptomics, proteomics or genomics. Metabolomics tool in clinic measuring variations of metabolites will play a key role for biomarker research, the identification of biochemical pathways involved in the treatment follow-up.373,374,375,376,377 Metabolomics obtaining global metabolic profile in biological systems can measure low-molecular-weight metabolites in the biological systems associated with various pathological conditions, could fill gaps between end-phenotypes and genotype.378,379,380
Small metabolites are correlation with the functional status in a biological system. Exploring metabolites and the related metabolic pathways allow a better understanding of how the abnormal metabolism could lead to disease’s onset, and progression.381,382,383,384,385 They enter body circulation and then is transferred to target organ and tissues, and then exert a series of biological effects that modulate cell function.386,387,388,389,390 Small metabolites could hint proteins acting as modulators of various biological phenotypes and could develop targets for early intervention.391,392,393,394,395 Metabolic signatures associated with human phenotype can be identified by various ways including by exploring associations between small metabolites and phenotypes.396,397 In addition, research of metabolic signatures has help discovery of potential biomarkers for the diseases. Toward developing effective approaches to evaluate disease progression and therapy responses, a robust and reproducible method is necessary to accurately depict their phenotype. A challenge is to identify the key “signals” of interest in metabolomic data that real influence on phenotype. Identifying molecular signatures that modulate phenotype could be achieved by an appropriate screening way. Mass spectrometry (MS) can detect all the ionizable metabolites without labeling or preselection.57,255,351,398,399,400,401,402,403 High-throughput screening of metabolic signatures that are closest to phenotypes advances to quantify and identify small-molecule metabolites at microenvironments.404,405,406,407,408,409 Single-cell metabolomic methods provide a direct understanding of the phenotypes of cellular activity and environmental changes.410,411,412,413,414 High-throughput molecular fingerprints in a wide range of pathological conditions were generated from metabolic profiling of biofluids and have been evaluated neurodegenerative conditions, cardiovascular diseases, metabolic disorders, and various types of cancers.415,416,417,418,419,420,421,422,423
Understanding manifestations of each patient’s metabolic map will allow for precision therapies rather than for the “average patient”. Metabolic profile, a collection of distinct metabolites, could describe human phenotype using small chemical metabolites as index for biochemical traits.68,424,425,426,427 Because it could reflect a patient’s phenotype, metabolic profile offers a comprehensive, precise and dynamic picture of the phenotype, allows the discovery of small metabolites related with various human phenotypes that link to health, disease or drug monitoring.100,428,429,430,431,432,433,434,435,436,437,438,439,440,441,442 Discrimination between the metabolite profile of diseases could result in potential benefits of identification of early diagnostic or prognostic biomarkers response to predictions.443,444,445,446,447,448,449 The elucidation of specific metabolic phenotype is essential for identifying potential biomarkers and drug targets, better understanding the underlying pathogenesis during disease progression (Fig. 4). Metabolic phenotyping from biological samples based on the fundamental paradigm of the homeostasis could reflect the substantial changes in the whole metabolism.450 The metabolic profiles of patients are dynamic and can be influenced by lifestyle, disease, external or internal stimuli and physiological and pathological condition changes.451,452,453,454,455,456,457,458 The biological processes that be related with gender, age, obesity, disease, medication, etc., could change the metabolic profile of an individual.459,460,461,462,463,464 Metabolic profile of biofluid media can directly reflect the particular metabolic status of different tissues or organs, also determine metabolic signatures for identifying the distinct patient subgroups according to disease characteristics.168,465,466,467,468,469,470,471,472 Since changes in various pathological conditions can be revealed by metabolic profiles, exploring metabolome could help towards enhancing the disease diagnosis, prognosis, surveillance, and personalized treatments. Metabolic changes serving as biomarkers for early diagnosis and potential therapeutic target, play significant pathological effects on regulated metabolism.
Representative metabolite biomarkers associated with human diseases in clinical studies for disease phenotype, diagnosis, classification, prognosis, and treatment (the detailed information showed in Table 1)
Several small metabolites analyses have been carried out to identify the specific metabolic phenotype profile relevant with disease progression and characterize alterations of metabolic signatures which may be used as potential biomarkers in clinic. The study by Liu et al. was aimed at characterizing the distinctive inflammatory phenotypes, and then identify the metabolic signatures and pathways. It demonstrated adenosine 5'-monophosphate, allantoin and nicotinamide correlate with metabolic changes to predict asthma inflammatory phenotypes.473 Another study has reported that the differential metabolites including glycerophosphocholine, rosterone sulfate, and elaidic carnitine as potential indicators can predict abortion rate of polycystic ovary syndrome, with an AUC of 0.933, 0.941, 0.933 for high predictive performance, respectively.474 LC-tandem MS was performed to characterize the serum metabolic signatures of hepatolithiasis, and identified 277 metabolites, AUC values for metabolites including18-β-glycyrrhetinic acid and PC (4:0/16:2) were up to 0.90, may have clinical value for hepatolithiasis.475 A study showed that polyunsaturated fatty acids and bile acids as potent markers were closely related to the severity and chronicity of drug-induced liver injury patients, respectively.476 In addition, the distinct metabolic signatures at the acute phase of COVID-19 patients compared to the recovery period, suggesting arginine and tryptophan metabolism as main pathways with a probable link to disease severity.477 In a quantitative profiling study that focused on urinary metabolic signatures including homovanillate, L-methionine, and thymine as indicators of traumatic brain injury.478 Concerning metabolic signatures of tumor growth stage, such as colorectal cancer, sporadic colorectal adenoma, and the potential metabolites such as D-mannose, sarcosine, 4,5-trimethoxybenzoic acid are found by serum metabolic screening.479 By metabolic features of PCa analyses, Yu et al. discovered a series of altered metabolites that were related to TCA, glycine cleavage system, fatty acid metabolism. Importantly, Glu/Gln had high predictive power when detecting PCa patients (AUC = 0.984), with a higher sensitivity (96.6%) than PSA (94.4%).480 In summary, these studies show the small molecule phenotype signatures offers new avenues for better understanding biological metabolic processes of diseases, and for developing new biomarkers to improve patient management in clinic.
Promising biomarkers
According to NIH Biomarkers Definitions Group, biomarkers were defined as features which are measured as an index or sign for physiological, biological, pathological, or pharmacological processes. Biomarker have characteristics that can be quantified, analyzed and associated with human phenotype and could be used for early disease detection, improve outcomes of treatments and selection of therapeutic strategy, and reduce disease-related mortalities, and lead to the identification of the therapeutic targets. Although the extensive efforts, currently used biomarkers in clinic are lacking adequate sensitivity and specificity for disease early detection and treatment monitoring. A growing number of biomarkers in urine, blood, plasma or saliva, have been considered to identify intermediate phenotypes with a clearer picture for predicting the response to therapy. Altered metabolisms have recognized as biomarkers. Metabolic profile can describe the underlying molecular picture of disease disorder or phenotype. Therefore, to improve the patient management, more precise biomarkers in biofluids are needed. Discovery of metabolic biomarkers will improve patient pretreatment and response to therapy.
Recently, a variety of biomarkers were discovered and employed to detect early-stage disease and predict disease progression, clinical outcome or drug response (Table 1). It can be a group of metabolites, a metabolite, or a molecular feature. The presence of a disease suggests the metabolite concentration has abnormal change (lower or higher concentration) is a sign of a perturbed or dysfunctional metabolic pathways of systemic homeostasis. There are huge advantages to consider and apply metabolic information during discovery phase that focusing on the understanding of the biological system associated with the metabolic pathways and can provide novel biomarkers and targets. Thus, unlike proteins and genes, metabolites as signatures of biochemical activity are closely correlate with human phenotype, since they play a key role in cellular signaling regulation and physiological function control. Therefore, discovery of altered metabolic features related to phenotypic variation produced insights into pathophysiology, mechanistic basis and therapeutic targets of metabolic diseases.
Given metabolism plays fundamental roles in characteristic metabolic alterations to gain deep insights into disease pathogenesis, small metabolites could emerge as potential targets for developing predictive biomarkers, and therapeutic targets. The precision treatment of metabolic disorders remains a huge challenge due to the imprecise diagnosis and involved incomplete understanding of pathophysiological process. To practice precision treatment, it is necessary to investigate small biomarkers that carefully consider phenotype determination. To establishing quantitative fingerprint and detection of endogenous metabolite biomarkers in easily obtainable and less intrusive biofluid may help to establish the close relationship between disease process and metabolic changes that contribute to body dysfunction of mechanistic basis of metabolic diseases. Currently, it is a challenge to rapidly detect disease using specific metabolite signatures at initial stages. Despite many biomarkers have been discovered in clinic, other biomarkers have not undergone their clinical validity and usefulness, preventing them advanced into clinical treatment. Advanced technology has greatly facilitated the discovery of biomarkers insights into metabolic regulatory and signaling activities that are strongly associated with human phenotype. Furthermore, biomarkers for the prediction, prognosis, and monitoring therapy, after the biomarker discovery phase, need GC or LC-MS, and NMR spectroscopy analytical techniques. Advanced analytical techniques could insight into the concentration detection of potential metabolite biomarkers within its early stages. Advanced platforms, especially using LC/MS/MS, facilitate detection, quantification, and characterization of small metabolic molecules (e.g., peptides, carbohydrates, amino acids, and fatty acids) involved in metabolic and catabolic processes, and greatly enhanced their translational capability.
Some representative potential metabolite biomarkers are currently screened (Fig. 4). A six-metabolite panel (beta-alanine, homoserine, 3-hydroxykynurenine, aspartate, tyrosine and ornithine) was quantified as potential blood-based biomarkers, and considers as a potential diagnostic or prognostic assay for Parkinson’s disease.481 Eva et al. had profiled serum metabolite signatures in early breast cancer participants and found that circulating metabolites: glutamine, tyrosine, proline, histidine, alanine and citrate can significantly correlate with tumor proliferation.482 Interestingly, a panel of two potential predictive metabolites (palmitic amide and deoxycholic acid) in serum was reported as potential biomarker of Crohn’s disease patients, and its metabolic disturbance involved the fatty acids, bile acid biosynthesis, and energy metabolism.483 Additionally with the use of correlation analysis and ROC curve analysis, the characteristic metabolites including alanine, glucose, lactate, glycine and threonine were identified in pulmonary arterial hypertension patients, and threonine and lactate were markedly correlated with pulmonary vascular resistance and arterial pressure.484
In a study that focused on biomarkers associated with gouty arthritis progression in patients, serum metabolic profiles were screened N1-Methyl-2-pyridone-5-carboxamide, kynurenic acid, 5-and hydroxyindole acetic acid.485 A multi-omics model with machine learning approaches was developed for discovering metabolite biomarkers predicting COVID-19 patients.486 Interestingly, 5-oxoproline can be used as a key biomarker for acute ischemic stroke.487 Metabolic profiling model based on seven metabolite candidates in plasma samples can provide powerful early survival prediction capabilities for ST-segment elevation myocardial infarction patients.488 Potential small metabolites included LysoPC(15:0), docosapentaenoic acid, propionyl carnitine, LysoPC(14:0), and phenylalanine were constructed a risk score for dose-response relationship with metabolism abnormalities and metabolic syndrome.381 Plasma metabolic profiling revealed four circulating metabolites (glutamate, pseudouridine, N-acetyltryptophan and leucylleucine) were identified in diabetic retinopathy patients.489 It has been reported that candidate biomarkers arachidonic acid and 13(S)-HODE associated with Akt pathway were potential biomarkers of non-small-cell lung cancer.490
Diagnostic biomarkers
Early diagnosis and effective prevention are of great importance and has attracted great attention for improving treatment and new therapeutic targets. For ideal biomarkers, molecular compound should be readily measurable in invasive biological media. Given metabolites are downstream expression of genome, closely indicate phenotypic fingerprints at a particular physiological period.491,492,493,494 One of the major advantages of metabolome over genome is that it can reflect environmental impact and provide global photograph of individual pathological conditions at any time point. Timely diagnosis is crucial, and the screening of small metabolites could play pivotal role in disease diagnosis. Therefore, the need for prompt diagnosis indicates the huge potential of advanced methods that reflect phenotype and therefore function changes. Since small metabolites indicate end-products of physiological processes, exploring whole metabolome can better understand disease pathology and mechanisms of intervention.495,496,497,498
Advanced analytical technology for small metabolites profiling features in distinguishing or determining disease pathophysiology associated with disease subtypes, progression, and treatment. Disease detection techniques over traditional methods are necessary for initial diagnosis, and also provide an effective approach to screen the right populations, assess drug efficacy, guide the choice of treatment or track disease progression, provide better patient care. Rapid progress in omics by high-throughput technology including LC-MS, GC-MS, and NMR, focused on characterization of metabolic phenotype, has allowed for simultaneous determination of a large number of small metabolic products in biological specimens.499,500,501,502,503 Omics approaches for biomarker discovery of early disease diagnosis could be achieved by analytical tools together with pattern recognition analysis (Fig. 5). Typical examples of these approaches consist of metabolic profiling, metabolic footprinting, metabolic fingerprinting, flux and target analysis, each of which has played a significant effect in clarifying the related metabolic pathways, understanding disease mechanisms and pathological processes.504,505,506,507,508 It can accurately detect the changes in distinguished features of metabolism, remains indispensable for disease detection.
Potential roles and applications of small-molecule candidate metabolites for biomarker discovery, diseases diagnosis, prognosis, and monitoring treatments in biomedicine. Compound detection, metabolites are detected by using specific detection techniques; data pre-processing, raw signals are then pre-processed to produce data in a suitable format for subsequent statistical analysis; then, data normalization is used to reduce the system and technical bias; data processing, for untargeted studies, metabolites are identified from spectral information in some given database; statistical analyses, univariate and multivariate statistical analyses are used to identify significantly expressed metabolites; biomarker discovery from multicenter, the discriminant metabolites originated from metabolomics approaches may become promising candidate molecules to aid disease diagnosis, and risk stratification; function analyses, next, the significantly expressed metabolites are subsequently linked to the biological context by using enrichment and pathway analysis. The images were obtained using the example data provided by the MetaboAnalyst 5.0 and figures created by BioRender
The metabolites linking between genotype and phenotype will result in biomarker identification for the early diagnosis, detection, and response to treatment, better understanding the complex disease pathophysiology that dramatic functional changes. Metabolic signature of disease could assess the risk or earlier diagnosis, detection, treatment monitoring, specific disease subtypes, and help selection of targeted treatment to match metabolic alterations of diseases related to phenotypic variation.509,510,511,512,513,514 To identify metabolite profile changes in early diagnosis stage of diseases is important for improving the prognosis, treatment and management. A larger number of single metabolite or a panel of the dysregulated metabolites can build diagnostic models that hold diagnostic power and are capable of differentiating patients.515,516,517,518 Small molecule metabolites reflecting dynamic pathological information is gradually moving towards clinical practice, and has been proven accurate enough for satisfactory diagnostic performance to predict diseases or early diagnosis or discriminate patients. Figure 6 shows how small metabolites could build metabolic blueprint of predictors in identifying biomarkers for early complex disease detection. Pathway analysis could expound altered metabolic alteration and show disease treatment options. Its application in all aspects of diagnostic potential has been proved in the research of metabolic disorders involved in disease progression, such as diabetes, metabolic syndrome and obesity.
Schematic diagram of an integrated pharmacology framework for discovery of bioactivity-correlated constituents, target identification and action mechanism of herbal medicine and natural products. The first stage discovers active compounds of treatment-related herbs followed by construction of correlation analysis network of treatment-related herb-compound and small molecule metabolite (Correlations based on the abundance scored value). Next is that highlight the main active constituents from identification of new candidates from natural products, and then elucidate the underlying mechanisms by target virtual screening and identification, until the final step of in vitro and in vivo tests. The images were obtained using the example data provided by the MetaboAnalyst 5.0 and figures created by BioRender
Regarding establishing early clinical diagnostic tool, a study was performed to identify differential and functional metabolites of early NAFLD. New candidates were discovered, including the upregulated theophylline and 1-naphthylmethanol, downregulated lysophosphatidylcholine (24:1(15Z)) and 2-hydroxyphenylacetic acid. It can achieve a high diagnostic power in the discovery phase (80.99%) and validation phase (75.23%).519 Study carried out by Xin et al. highlighted how metabolite biomarkers and metabolic profile can serve as biomarkers for precision diagnosis of various types of tuberculosis.520 Further, it also demonstrated that potential of machine learning method combining metabolome in screening out diagnostic biomarkers from big data set. Parallel study has been carried out in plasma samples focusing on the metabolic characterization of breast cancer patient, and revealed specific metabolic profiles, identified a panel of glutamate, sphingomyelins, and cysteine that showed high predictability that can be used as diagnostic biomarkers.521 Previous study reported LPC (18:2/0:0) level correlate with diastolic dysfunction and glycyl tyrosine correlate with reduced lower left ventricular ejection fraction, indicating they can detect cardiovascular risk.522 An integrated multi-platform analyses were used to screen biologically significant metabolites linked to Esophageal squamous cell carcinoma patients.523 It found the close link lipid, amino acid metabolism, and a diagnosis panel of citrulline, l-carnitine, acetyl-carnitine, tryptophan and lysine selected as potential biomarkers in distinguishing patients. Metabolic pathway analysis obtained biomarkers associated with oral squamous cell carcinoma that closely related to amino acid and cholic acid metabolism. Further, a diagnostic panel was established and constituted of cysteine, cholic acid, decanoylcarnitine, and had high early diagnosis power (AUC = 0.998).524 According to Lunyera et al., urine tricarboxylic acid cycle signatures are potential indicators at early-stage diabetic kidney disease progression.525 It has proved that glycerolipid metabolism and galactose metabolism are the main metabolic pathways, and serum metabolite glycerol-3-galactoside can be used as an independent indicator to predict diabetic kidney disease.526 Here, these instances of clinical trials based on endogenous small molecule metabolites expand the coverage of metabolic biomarkers for disease diagnosis.
Disease classification and stratification
Clinicians need rapidly assess disease stratification risk and with adequate accuracy. Recently, omics combination approach has employed as a promising strategy for generating information on detecting early metabolic alterations which could contribute to the disease classification, stratification and progression for diseases that are immediately associated to biologically meaningful metabolism, such as cardiovascular diseases, cancer, diabetes, and obesity.527,528,529,530 The right choice of small molecule metabolites that correlate with pathological states can help making decision and lower-costs from the pilot testing into the clinic. Metabolic profiles of diseases are able to characterize disease signatures for discovering and identifying diagnostic biomarkers and many unexpected mechanistic pathways that involved in disease pathogenesis. Classically, endogenous small molecules metabolite screening combined with the traditional risk assessments enable characterization of metabolic phenotypes even before manifestation of symptoms and have the potential to improve non-invasive diagnostics and disease classification with great potential to translate them into clinical settings.531,532,533,534,535 According to small molecule metabolite profiles or fingerprints shown ability to predict disease risks, the big data being collected on artificial intelligence or big data mining will contribute to disease stratification analysis as an integrative tool that assists clinicians in making decisions.536,537,538 Over the past few years, one of the most striking aspects of screening of endogenous small molecules metabolite of systemic metabolome alterations particularly has evolved to gain a much broader dimension, also showed great potential for differentiating disease subtypes.
Importantly, a large number of cohort studies have been carried out to help establish a more effective and reliable risk performance model for disease stratified risk events. In pancreatic ductal adenocarcinoma, a panel of three small metabolites including creatine, proline, and palmitic acid can exhibit a beneficial performance for distinguishing pancreatic ductal adenocarcinoma from benign pancreatic neoplasms or healthy controls.539 A study has focused on characterizing the metabolic subtypes of pancreatic ductal adenocarcinoma and analyzing the relationship between long-term prognosis and metabolic subtype.540 It did not reveal the metabolic differences at the clinical stages and choline-like type showed better prognosis among metabolic subtypes. Interestingly, a metabolites biomarker panel can precisely predict the overall survival of pancreatic cancer and distinguish tumors from normal pancreatic tissues in a clinical setting.541 A recent study investigating potential biomarkers for screening and diagnosis of lung metastases, some low-molecular metabolites such as indoleacrylic acid, L-tyrosine, retinol, L-octanoylcarnitine and decanoylcarnitine were selected and found they had high AUCs values and showed a strong ability to differentiate between pulmonary metastatic carcinoma and other subtypes.542 An integrated metabolome and lipidome platform discovered four differential metabolites including D-glyceric acid, cortisol, 2-(methylthio) ethanol, N-acetylhistamine and then established a differentiation model for precise pathological classification of squamous carcinoma and non-small cell lung adenocarcinoma.543
Serum metabolic profiles in salivary gland tumors patients were investigated to gain a better understanding of the disease risk stratification. A total of 32 small metabolites were identified, and a risk predicting model based on the gradually upregulated serine and lactic acid was developed in benign and malignant stages.544 In medulloblastoma, a panel of two urine metabolites including cortolone and tetrahydrocortisone showed a high accuracy for diagnosis and monitoring.545 A machine learning-derived nomogram models using thiamine triphosphate, diabetes duration, and systolic blood pressure were established for early diagnosis and accurate prediction of diabetic retinopathy.546 Metabolic alterations in amino acid, energy, lipid, and metabolism could distinguish the different stable and unstable types of coronary atherosclerotic heart disease.547 Moreau et al. analyzed salivary metabolome in primary burning mouth syndrome and found tyrosine pathway (L-tyrosine, tyramine, L-dopa) can differ patients according to the levels of pain.548 A study performed by tissue-based spatial metabolomics with mass resolution imaging had developed classification system of gastric cancer subtypes and insight into their distinct metabolic pathways and molecular characteristics.549
A metabolic biomarker panel was discovered in discovery cohort to discriminate papillary thyroid cancer from benign thyroid nodule with 91.89% sensitivity, AUC of 97.03%, and 92.63% specificity, and in validation cohort displayed 86.57% sensitivity, AUC of 92.72%, and 92.50% specificity, and can improve stratification of thyroid microcarcinoma.550 In recent work, several metabolites such as carnitines, fatty acids, ketone bodies, bile acids, purines and tryptophan, were obtained as early biomarkers to distinguish from the early-stage and end-stage coronavirus disease 2019.551 Using a ROC curve and logistic regression analysis, a biomarker panel of PA(37:4), 6-keto-PGF1α, PS(36:0), and LysoPC(20:1) demonstrated good classification and diagnostic ability in distinguishing endometrial polyps from endometrial hyperplasia or endometrial cancer in the validation set.552 Notably, targeted metabolomics analyses identified gamma-aminobutyric acid markedly reduced in COVID-19 patients and its change levels with high sensitivity that allowed for COVID-19 stratification.553
Urinary metabolic features of prostate cancer, bladder cancer, and renal cell carcinoma have been carried out to determine and reveal that N-methylhydantoin, 4-hydroxybenzoate, creatinine, acetate and glutamine had significantly discriminatory accuracy among groups.554 When the level of a specific antigen is located in the range of 4–10 ng/ml, differential metabolites are screened to effectively distinguish between benign prostatic hyperplasia and prostate cancer.555 As for the metabolic perturbations in vivo, Alotaibi and colleagues followed the bioactive lipid molecules screening approach and reported the 5 metabolites as biomarkers of disease severity differed between pulmonary artery hypertension with systemic sclerosis and idiopathic pulmonary arterial hypertension, and provide important underlying mechanistic basis in subgroups of pulmonary artery hypertension.556 Furthermore, Luo et al. performed a comprehensive analysis of metabolome data, and relevant metabolite dehydrophytosphingosine and 9-cis-retinoic acid had proved to be the most discriminative biomarkers for ventricular fibrillation phenotype, had the high predictive probability based on their combination model.557 Albillos et al. conducted metabolome with multivariate analysis to examine potential biomarkers such as acyl-carnitines, bilirubin, tyramine, for differentiation between Parkinson’s disease and essential tremor.558 These studies show a great potential for screening biomarkers for better disease stratification to advance the understanding pathophysiology, allowing therapeutic options.
Prognosis biomarkers
The lack of symptoms in the prognosis stages makes early disease diagnosis difficult. Prognosis biomarkers are important in order to reduce complex disease mortality. It is essential to identify prognostic biomarkers that could facilitate decision making by clinicians and promote individual therapy. Identification of useful prognosis biomarkers remains a huge challenge in clinic. However, regular tests offer low specificity and sensitivity, leading to inadequate early-stage diagnosis or risk assessment. To improve the risk stratification and prevention of disease, benefit from therapy, it need insight into multiple prognosis biomarkers and simultaneously quantify in a high-throughput way. Interestingly, a great advantage of small metabolites as biomarkers likely occurs a panel of multiple metabolites with markedly concentration changes correlated with disease status.559,560,561,562 Interestingly, one of the significant advantages of metabolite biomarkers may be that they are composed of multiple metabolites, and their concentration changes are significantly related to disease status.563,564,565,566,567 The response to drug therapy can accurately monitor the changes of small metabolites in biological media (e.g., urine and blood).
Large amounts of studies have analyzed the metabolic profiles of patients to identify potential small biomarkers with prognosis utility in the clinic (Table 1). These researches include huge efforts to develop simple, inexpensive, and novel diagnostic applications, to enhance knowledge on the predictive or prognosis biomarkers of the diseases and its complications. Metabolic deregulation could affect various molecular biological processes (e.g., cell apoptosis or invasion) that contribute to disease progression and impact patient survival. Of interest to physician is the great potential of small molecule metabolites an invaluable tool from a prognostic point of view. Instead of a single biomarker, multiple small metabolites corresponding to particular phenotypes are anticipated to yield a higher selectivity and sensitivity.568,569,570 Both un-targeted and targeted approach have also been conducted identifying specific metabolites or predictor biomarkers which were linked to metabolic alterations.571,572,573,574
Analyses of the relative level of tricarboxylic acid by semi-targeted serum metabolomics shows that circulating pyruvate is an effective prognostic biomarker of COVID-19, which means that the quantification of pyruvate is a clinical support for prognosis prediction.575 Metabolic profiling of plasma reveals COVID-19 affected porphyrin and glycerophospholipid metabolism, respectively.576 Small metabolites in porphyrin and purine pathways were markedly elevated in severe group, indicating that they can be used for prognostic biomarkers. Prognostic tests based on intermediary metabolites such as ureidopropionate, deoxycytidine and kynurenine could improve COVID-19 patient treatment outcome and severity.577 One example comes from Barco et al., who used a targeted metabolite profiling approach to discover the high expression of 3-O-methyldopa was associated with worse prognosis in neuroblastoma patients.578 A study performed untargeted metabolomics had revealed eight metabolic biomarkers were identified as prognostic biomarkers of acute ischemic stroke.579 As an example according to Brunmair et al., metabolic phenotyping of tear fluid has been successfully established and revealed taurine and nicotinic acid represent new biomarkers were elevated in the diabetic cohort, and supports prediction of disease development.580 Furthermore, metabolite profiles also showed that asparagine synthesis was increased and associated to poor prognosis for female colorectal cancer patients.581
Metabolic pathways
Aberrant metabolism is a necessary pillar as a hallmark of disease, e.g. lactate, pyruvate metabolites can assist in cellular proliferation. Comprehensive understanding and investigating mechanistic pathways can provide powerful evidence for precise diagnosis, phenotypic classification, prognosis and treatment of patients. Metabolic pathway analysis can be performed with benefiting mechanistic explanations of therapeutic targets for metabolism-related diseases.582,583,584 The altered metabolites are significantly correlated with metabolic pathways and biological processes involved in the disease progression. In addition, the differential metabolites are likely to be one of the most important information to explain the pathogenesis mechanism. From a metabolic perspective, small molecule metabolites whose altered concentrations could reflect phenotypes and elucidate pathophysiological changes of complex diseases, provides clues regarding alteration of metabolism in dysfunction, helps functional interpretation of metabolic perturbations in vivo related to phenotypic variation.585,586,587,588 In this context, small molecule metabolites are associated with diagnosis or prognosis in metabolic processes and alteration in treatment of systemic homeostasis. Targeting metabolic pathways can regulate the abnormal metabolisms and finally alleviate disease syndromes.
Small molecule metabolites associated with specific metabolic phenotype can be used to screen early disease symptoms and monitoring its progression, through measuring endogenous metabolite alterations in biofluids or tissues.589,590,591 Discovery of small metabolite by high-throughput, non-invasive, and cost-effective metabolomics are quite useful to compute metabolic pathways that link complex chemical reactions involved in the biological process. Advanced metabolomics technology could amplify the small changes of differential metabolite expression to achieve a wide coverage and then reflect functional changes, deeply reveal action mechanism.592,593 This approach enables providing the key information for further exploration of metabolic signatures and potential biomarkers, mechanistic in-depth understanding, and therapeutic targets for treatment. Metabolite can be used as an early indicator of pathological changes prior to development of disease symptom. Several available software platforms have been designed to facilitate metabolic pathway analysis for small molecule metabolites.594,595,596 Particularly, Ingenuity Pathway Analysis and MetaboAnalyst could be used to clarify the relevant metabolic pathway network change associated with small molecule metabolites found in omics data, enable integration for biological interpretations.58,360,597,598,599 The online databases, such as KEGG, provide huge information about a large number of biological pathways and can be easily used to determine and visualize the metabolic pathways and metabolite interaction network involved in fundamental biological processes. These comprehensive tools to the biological interpretation help the identification of differentially altered analytes and dysregulations of pathways.
Previous report had shown that metabolic alterations in clinical hypothyroidism and subclinical hypothyroidism linked to various potential metabolite biomarkers suggesting that impacting onsteroid hormone biosynthesis, primary bile acid biosynthesis, lysine degradation, purine metabolism and tryptophan metabolism.600 Recently, Marino et al. performed multivariate network analysis to identify the core pathways in the advanced stage of Amyotrophic lateral sclerosis, and suggested the metabolic alteration of lysophosphatidylcholine, sphingomyelin, and phosphocholine metabolism, consistent with repairing inflammation and neuronal degeneration.601 Metabolic dysfunction in glycerophospholipid metabolism, arginine and proline metabolism, and tryptophan biosynthesis of invasive ductal carcinoma patients was also observed by pathway enrichment analysis.602 Alterations to metabolic pathways included glycerophospholipid metabolism, D-glutamine and D-glutamate metabolism associated with atrial fibrillation have been broadly explored at small metabolites level. A study has focused on characterizing the specific and precise metabolic features of atrial fibrillation subtypes, indicated that small-molecule metabolites may facilitate effective treatment.603 Additionally with the use of untargeted metabolomics, a study demonstrated serum biomarkers of progression of diabetic retinopathy in Asians, and there were 171 metabolic features including glutamine, N-acetyl-l-glutamate, glutamate, aspartate, N-acetyl-l-aspartate, docosahexaenoic, icosapentaenoic, and dihomo-gamma-linolenate distinguished proliferative diabetic retinopathy patients from T2DM patients.604 Enrichment pathway analyses for major metabolite biomarkers indicated arginine biosynthesis metabolism, d-glutamine and d-glutamate metabolism were dysregulated in advanced stages of diabetic retinopathy.
Metabolic snapshot of COVID-19 revealed some additional interconnection pathways implicated in disease pathogenesis, including citrulline, phenylalanine and histidine, 2-aminobutyric acid, asymmetric dimethylarginine.605 The disordered metabolic pathways of primary Sjögren’s syndrome patients are associated with tyrosine metabolism, tryptophan metabolism, aspartate and asparagine metabolism, carbon fixation and affect neurological cognitive impairment, inflammatory injury, and the immune response.606 Pathway analysis by urinary metabolomic study demonstrated that aberrant metabolisms involved in aspartate metabolism, glycine metabolism, glycolysis, glyoxylate metabolism, and TCA cycle.607 Metabolic signatures enriched metabolic pathways of multiple myeloma patients were linked to amino acid metabolism and biosynthesis, and insight into elucidating disease pathogenesis.608 Characteristic biomarkers succinic acid semialdehyde, uracil, uridine or metabolic pathways enriched in lipid metabolism, amino acid metabolism, nucleotide metabolism and glycometabolism were identified and related to specific multiple trauma complicated with sepsis.609 Through untargeted analysis, a total of 120 candidate differential metabolites were detected in patients with ischemic stroke and markedly altered metabolic pathways were purine metabolism, steroid hormone biosynthesis, or CoA biosynthesis.610 Metabolic profiling using high-resolution mass spectrometry of cystic renal disease patients was collected and impact several pathways involved in purine and pyrimidine, aminoacyl-tRNA biosynthesis, glutathione, TCA cycle, etc.611
Enabling precision treatment
There is not any specific therapy for satisfying all the patients. Thus, to predict the therapeutic response with matching the right patients at the right treatment is necessary in clinic. Additional techniques are critical to discover effective and potential biomarkers to guide patient management matching the proper treatment. Metabolite profiling as cost-effective and productive way enables holistic and systematic analyses of metabolites and can be utilized to predict and monitor the response to drug treatment, uncover therapeutic target for drug discovery, personalized management to reduce disease burden. Application of small metabolites to predict specific response to drug therapy is closely related to patient’s pharmacological phenotype and could generate more information than other omics data for interpretation of the metabolome data.612,613,614 Furthermore, it enables exploring promising models to predict therapeutic response.171,615,616 Small molecule metabolites can be used for diagnosis and prognosis of patients, predicting pharmacological responses to the peculiar treatment. Furthermore, metabolic signatures can provide the huge information from targeted metabolic pathways or precision drug therapy. Distinctive metabolite signatures that are useful for identifying different therapies responses are summarized (Table 1).
Irajizad et al. conducted plasma metabolomics profiles and artificial intelligence using a deep learning model to identify biomarkers for predicting response to neoadjuvant chemotherapy in triple-negative breast cancer.617 According to metabolic profiles, taurine, glutamine, glycine and hypoxanthine were potential biomarkers of ladder cancer patients treated with neoadjuvant chemotherapy and pathway enrichment analysis characterized significant alterations were related to amino acid metabolism.618 Amino acid metabolism seems to be a predominant pathway altered in ladder cancer patients and has potential value in enhancing the efficacy of chemotherapy. Notably, hypotaurine and taurine metabolism, pentose and glucuronate interconversions were the most altered pathway for subcutaneous immunotherapy.619 In this study, the authors found taurine, l-alanine, and hypotaurine, considered to be predictive biomarkers relevant with effective subcutaneous immunotherapy.
A recent study investigating the relationship between anti-VEGF therapy and serum metabolome and described differential metabolite LPC 18:0 may be a potential biomarker for guiding treatment options for macular degeneration and choroidal vasculopathy.620 It has been reported that decreased kynurenic acid in cerebrospinal fluid and kynurenic acid/kynurenine ratio represent a biomarker of epileptic spasms and further therapeutics method should be explored to increase the kynurenic acid level.621 Previous report has shown that after 4 weeks of olanzapine monotherapy in schizophrenic patients, methyl n-methylaminobenzoate as response biomarkers in the kynurenine pathway is associated with treatment outcomes.622 Metabolic profile alteration to molecular phenotype of psoriasis vulgaris patients showed that SM (d16:1/16:1) and Cer (d18:1/18:0) correlated with the biochemical indicators and could contribute to precision treatment.623 Another study used metabolomic profiling and small metabolites (N-methylisoleucine, nornicotine, 2,3-dihydroxybutanoic acid) were able to discriminate rheumatoid arthritis patients with early response to methotrexate therapy.624 These clinical applications of small metabolites provide excellent examples to illustrate new channels for targeted therapies and enabling precision treatment.
Modulating metabolism
Modulating metabolisms with small molecules have been known for decades. Metabolic therapies are imperative and bring new opportunities for patients. Metabolic disorders are caused by various mechanisms. Recently, metabolism has acquired interest regarding the relationship with environmental factors, host genes and diseases.625,626,627,628,629,630 How does small molecule metabolites drive phenotype modulation? The common regulating mechanism of active pathway is metabolites bind allosteric sites on enzymes. Discovery of the relationship network or pathway of metabolite interaction can uncover the action modes of regulation. Numerous works have revealed differences changes in small molecular metabolites associated metabolic pathways are closely related to therapy efficacy and potential drug targets.631,632,633 Perhaps the application of these metabolic pathways involves small metabolites could better clarify the development of complex diseases in the future.
To exploiting the unique features of modulating metabolism with small molecules for treatment and monitoring is a very promising direction. The endogenous metabolite profiling provides the best view of disease phenotypes. Advanced screening approaches by analyzing the metabolic profiles have become increasingly application in metabolism study.634,635,636,637 Simultaneously, continuous development in high-throughput metabolomics technology has allowed considerable progress to be made in determining disease pathogenesis, understanding the various relationship between metabolic regulation and disease. Single-cell metabolomics technologies will reveal new insights into modulating metabolism with small molecules.198,214,413,638,639 It can provide meaningful cell phenotype, enabling us to analyze cell status and obtain the overall biological information.
Metabolic perturbations in vivo contribute to early discovery and mechanisms of phenotype modulation. Decoding the molecular mechanism of metabolic alterations will provide a promising way for novel therapeutic interventions. Metabolic alterations can modulate the cell signaling pathways to maintain the systemic homeostasis. The most diseases (e.g., obesity, diabetes, hypertension, or depression) have strong metabolic disorders, many chronic diseases (e.g., Alzheimer disease, or cancer) have unexpected metabolic basis of associations.640,641,642 It still needs a significant treatment window for effectively optimize therapies by precisely inhibiting the metabolic targets. To blocking metabolic pathways or inhibiting metabolic enzymes are almost impossible to generate an effective treatment. Metabolic disorders have become a feature of several cancers. Interestingly, several studies showed that targeting metabolic enzymes could significantly inhibit tumors to promote an effective therapeutic intervention.370,643,644,645,646
Major findings of previous studies in small molecule metabolites drive metabolism were summarized in Table 1. Targeted metabolomics identified energy metabolites of lung adenocarcinoma cells and found that KCNK3 can inhibit proliferation and glucose metabolism through activation AMPK/TXNIP pathways, indicating KCNK3 may be a potential therapeutic target.647 The authors examined a total of 202 relationship features between various cancers and metabolites, and showed gamma-glutamylisoleucine, 7-alpha-hydroxy-3-oxo-4-cholestenoate, gamma-glutamylleucine, and 1-oleoylglycerophosphocholine were the most dangerous metabolites for ovarian cancer, lung cancer, glioma and breast cancer, respectively. Analyses in these causal links demonstrated these small metabolites play a key role in phenotypic regulation to distinguish cancer patients in clinic.648 Pathway enrichment analyses indicated that the imbalance of purine and amino acid metabolism could affect the prognosis of patients with oral squamous cell carcinoma.649
A recent study conducting an inquiry into the relationship of small molecule metabolite hydroxyasparagine in blood samples associated with the progression of chronic kidney disease patients.650 Another study used serum metabolomic analysis and differentially expressed metabolites, such as triethanolamine, chavicol and alpha-methylstyrene, that involved in platelet degranulation and immune responses, and metabolism process were firstly identified as biomarkers in COVID-19 progression.651 A study suggested that the mechanism of lipid metabolism plays a critical role in pathological process of osteoarticular tuberculosis.652 Multivariate statistical analysis based on open database, metabolic differences of altered small metabolites were identified in superior limbic keratoconjunctivitis patients, and fundamental processes mainly involved in the inoleic acid metabolism, butyrate metabolism, ketone body metabolism, carnitine synthesis, and etc.653 Glutamate metabolism and urea cycle are related to psychiatric symptoms and accounted for the highest proportion in the altered metabolic pathway, and decreased in the schizophrenia group.654 A study has demonstrated that glycerophospholipid metabolism and arginine and proline metabolism pathways are related to inflammatory states and β-pseudouridine, may participate in inflammation regulation.655
Functional target
Metabolite has a wide range of biochemical function, a growing area of researches is the usage of small molecule metabolites to discover the metabolic targets with optimal therapeutic response for precision medicine. A change of metabolite levels as results of the modified enzyme activities indicates a phenotype alteration because of metabolite concentrations provide a close association with biochemical activity. Endogenous metabolites as therapeutic molecules targeting regulators prone to modulate metabolism activity with key metabolic pathways such as regulating multiple enzymatic reactions. Recent advances in high-throughput metabolic flux analysis technologies using stable isotope tracer methods make characterization of a large scale of endogenous metabolite for characterizing and tracking the metabolic activities.656,657,658,659,660 It could provide potential therapeutic targets depending on the improved and detailed understanding of the interaction between metabolism in vivo and functional status. A clear understanding of molecular mechanisms about targeting central metabolic pathway always plays a key role in the discovery of drug targets for optimal therapies.661,662,663 Metabolomics directly contributed to uncover novel targets can elucidate disease mechanisms in various diseases. From a point of view of metabolism, such knowledge will uncover new therapeutic targets related to phenotypic variation. Understanding the metabolic dysregulation can facilitate drug development and provide therapeutic targets for disease therapy. Numerous active compounds as modulators of metabolism and could target metabolic regulation mechanism.664,665,666,667,668,669
The disordered metabolic pathways associated with COVID-19 patients performed by quasi-targeted metabolomics with pathway enrichment, and showed glutamine/glutamate ratio markedly related with severe disease.670 It is therefore proposed that elevated glutamate level is associated with the increased risk of disease infection. Nevertheless, elevated glutamine was associated with a reduced risk of severe. This study has provided the probable targets for COVID-19 patients. In another study, concerning plasma metabolites, Ozaki et al. found that small metabolites can predict the progression of cognitive impairment in Alzheimer’s disease.671 Metabolic profiling of cerebrospinal fluid revealed pentose phosphate pathway is an important target for sedatives to change brain metabolism.672 The study carried out by Thomas et al. highlighted pathophysiological mechanisms from serum metabolome, and demonstrated metabolic disruption of choline phospholipids as among the strongest predictors were associated with severity of traumatic brain injury patients.673 Furthermore, a study proven palmitic acid is a key metabolite as promising therapeutic target, which accelerates cellular senescence by producing living oxygen in kawasaki disease.674
According to Scarale et al., tryptophan, kynurenine and hexanoylcarnitine are associated to improve the mortality prediction of type 2 diabetes.675 Metabolomics and mass spectrometry analysis have identified succinate as a therapeutic target for aortic aneurysm and dissection.164 Metabolic alterations of colorectal cancer patients were assayed by functional metabolome profiling and glycodeoxycholic acid positively showed high specificity and sensitivity correlated with CRC.676 Further findings showed GDCA can promote cell proliferation and migration, and PARP-1 was identified as a key target. A study profiled metabolome and identified subtype-specific N-acetyl-aspartyl-glutamate as a key tumor-promoting metabolite and therapeutic targets for advance precision treatment for triple-negative breast cancer.22
Targeting tryptophan metabolism in cancer has curative potential. For instance, tryptophan metabolism is a major metabolic pathway which restricts anti-tumor immunity and promotes intrinsic malignant properties of tumor cells, and considers as a target for cancer immunotherapy.677,678,679 Tryptophan metabolism changes could result in a series of alterations in tumor microenvironment and tumor cells, and then promote tumor progression. Via hindering DNA repair, small molecule metabolites had accumulated in tumors involved in abnormal metabolism.680 Mechanistically, targeting isocitrate dehydrogenase 1 and 2 (IDH1 and IDH2) enzymes result in elevated levels of 2-hydroxyglutarate in cells and its accumulation boosts rapid development of tumors. Functionally, this better define relationships between small molecule metabolite disrupting DNA repair and biochemical function that benefit efficient treatment. In summary, these metabolic targets can enhance precise treatment in the upcoming era of precision medicine.
Metabolic networks exploration
Metabolic perturbations could be modified by drug or natural products treatment as a crucial mechanism for its effects. Most natural products influence on multiple rather than single targets to exert the bioactivity. Both targeted and untargeted small molecule metabolites-based metabolomics have been used to characterize unexpected metabolic changes in biological samples, understand the metabolic processes and explore network targets and mechanism in various organisms.681,682,683 This analytical process always generates increasingly complex datasets at a large scale (thousands of metabolites), cause processing, analyzing, and interpreting relationships of small molecule metabolites are major challenges. Biological interpretation of the connected informative relationships of small molecule metabolites could be formalized as metabolic networks based on the prior knowledge, where the feature metabolites as nodes and the related metabolites are connected as edges.
Integrating multi-layer networks to use prior metabolic knowledge would help to improve the identification of metabolites and derive new interpretation of biological contexts.684,685,686 Once the metabolic networks of co-regulated metabolites are established, and then metabolism information will be mined using advanced algorithms. Importantly, the recent use of multiple network constructions and graph-based methods to perform topological analysis focused on analyzing the metabolic processes or metabolites data associated with a phenotype of interest.687,688,689 Metabolic networks or graph are generated depend on the prior biological knowledge. Network metabolites are co-regulated or connected within metabolic pathways. Instead, it represents the interconnections of metabolism network connected metabolites via distinct pathways. If correlation value of metabolites in a metabolic network are reaches a given threshold. Based on similarity or correlation of the identified metabolites, the graph analysis (e.g., metabolite graph and compound reaction graph), advanced statistical methods, and data analysis can be used to explore the inter-connected data to reveal metabolite relationship in biological samples.
During the past few years, network-based approaches towards multi-targeted compounds represent an important tool owing to its potential for ascertaining and investigating new drug targets and complex relationships.690,691,692,693 The ‘network pharmacology’ created by Hopkins and focus on a therapeutic concept from ‘one target-one drug’ to ‘target-network-component’ to combat the complex diseases.694,695,696,697 It used bioinformatics and high-throughput screening method to facilitate the prediction of various drug targets network based on the establishment of biological models, and is becoming more important in revealing the underlying mechanisms of drug actions. By analyzing the highly connected nodes in metabolic networks may open new avenues for discovery of mechanistically relevant signals for specific multi-target natural compounds. Thus, systems analysis of diverse metabolic pathways to identify novel targets may overcome pitfalls and facilitate change concepts of current drug design and develop new diagnostic as well as targeted therapeutic tool via exhibiting multiple targets and action modes.
Multi-omics interaction networks were constructed and showed that multiple biomarkers included pyridoxamine phosphate, folic acid, pyridoxal phosphate, and vitamin metabolism disorder was pathological characteristics of pulmonary tuberculosis patients.698 Based on 127 metabolic signatures from the Alzheimer’s Disease, specific metabolic networks modeling for diagnosis were constructed and provided key insights for personalized late-onset.699 Recently, Guo et al. performed a metabolic network-based identification modeling for mapping the differential metabolites, a panel of eight candidate metabolites (i.e., palmitic acid, pyruvate, tryptophan) were further indicated a high discrimination for non-small cell lung cancer (accuracy > 97.7%).700
Efficacy evaluation
Metabolic profile change of complex diseases suggests distinctive aberrations of the metabolism can be due to drug’s treatment efficacy on patients’ genotype. Small molecule metabolite-based metabolomics plays an important role in discovering biomarkers to evaluate the efficacy of therapies and have become critical tools for investigating modes of drug action, identifying novel drug targets701,702,703,704 Particularly, by generating metabolic signature, it is increasingly being implemented to diagnose disease, monitor treatment and uncover the underlying mechanisms of complex diseases, seek to understand drug efficacy. Moreover, in-depth research on small molecule metabolite may guide drug efficacy, development, and safety. Selecting the most effective treatment drugs is an extremely important event. Identifying small molecule metabolites as biomarkers associated metabolic alteration by drug response before administration could greatly reduce costs of treatment. This is compatible with the notion that we need screening strategies of small molecule metabolite to determine each stage of treatment efficacy, and to develop more effective therapies. Clinical models combining small molecule metabolites have shed light on the search for biomarkers and therapeutic targets, could improve the accuracy of identifying patients.
The decrease of plasma kynurenine level may indicate the therapeutic response of escitalopram, suggesting that it may participate in the pathophysiological response of severe depression caused by escitalopram treatment.705 Metabolic signatures of cholangiocarcinoma patients showed that the TCA cycle was reversed, which was obviously manifested by the increase in the level of amino acid and citric acid as intermediate products of TCA cycle and have the ability to predict patients' response to chemotherapy.706 Stratification of methotrexate efficacy identified significant alterations to various metabolites such as phosphatidylcholines, glucosylceramides, sphingomyelins, hypoxanthine, etc, involved in nucleotide, energy, fatty acid/lipid metabolism.707 Serum-based metabolites involving L-arginine and arachidonic acid can serve as diagnostic biomarkers for breast cancer predicting therapeutic effects of trastuzumab.708
Medicinal plants usually depend on complex components are a great resource for treatment of metabolic disorders. However, due to complex components and multiple molecular targets, molecular action mechanisms of herbs and formulations are still not very largely clear. Usual methods are not enough sensitive to evaluate drug efficacy, even small effects of drugs can be sensitively detected by small molecule metabolites as disease-related biomarkers in clinical trials, by monitoring differences of metabolite profiles. Knowledge about metabolic regulation mechanism by herbal medicines can help to predict and understand the efficacy and toxicity. Recent years some studies utilizing metabolomics to elucidate the biological basis and mechanism of the effect.709,710,711 It focuses on fluctuations of small molecule metabolites and insights into drug efficacy assessment and investigates molecular mechanism of herbal plants as adjuvant therapy for aberrant metabolism-related diseases. Research has shown that small molecule metabolite-based metabolomics can be further used to identify the active compounds and targets, which develop new therapies.712,713,714,715
Active ingredients discovery
Over the past decades, more than half of new drugs and drug leads have been developed from natural products that possess immense chemical structure with various biological properties. Recently, natural products in medicinal plants, such as alkaloids, flavonoids, terpenoids, carotenoids, and glycosides, possess therapeutic effects and are used as new therapeutic drugs.716,717,718 In clinic, herbal products are combined with conventional drugs to improve pharmacological effects. Active ingredients or drug leads from natural products have been a key source but their identification is always a challenge due to their complexity. Understanding effective mechanism of natural products or their derivatives or synthetic mimic requires elucidation of pharmacological response to complex phytoconstituents. Although a great advance achieved, one major challenge in discovering new active ingredients is unclear pharmacological mechanisms. The action mechanisms and efficacy profiles of herbal medicines for their potential use should provide in-depth information on elucidating the underlying mechanisms for active ingredients discovery.
Medicinal plants such as herbal extracts, formulae, and different compounds showed the pharmacological effects through regulating metabolic disorder and mechanism pathways due to multi-compound interactions and diverse chemical structures. High-throughput metabolomics agrees with holistic view and insight into a comprehensive mechanistic efficacy of herbal medicines, including medicinal plants, preparation, active compounds, aqueous extracts and formulas or patent medicines.719,720,721,722,723 Target small molecule metabolite based-screens offer numerous advantages for functional ingredients discovery from natural products as a treasure trove for drug development, and allows in-depth understanding of the possible targets and action mechanisms. Advanced metabolomics techniques consist of LC-MS, NMR, and GC-MS in combination with pattern recognition analysis or multivariate statistical analyses could identify a large number of metabolites and impact on diagnosing disease, discovery of biomarkers, investigation of phenotypes, classify physiological status and response to treatment, unravel efficacy of metabolic-targeting drug, cover the full pipeline of lead compound discovery and development from medicinal plants.719,724,725,726,727 Based on multiple metabolic alterations involved in disease pathogenesis, it is particularly pivotal to explore herb-derived bioactive ingredients for these mechanistic basis, damaged metabolic pathways and therapeutic targets of metabolic diseases. Moreover, herb-derived bioactive ingredients have been screened and validated in vitro and in vivo, to investigate the underlying changes of small metabolites and metabolic pathways, and to find potential targets (Fig. 6). They can provide a functional relationship between chemical diversity and metabolite changes.
Natural compounds as potential therapeutic agents have gained increasing interest due to ability to target metabolism and their diverse structures. Different metabolites involved in metabolic alterations could be targeted by the active components due to their efficacy in the clinic. Various herb-derived bioactive compounds could target the metabolic regulation mechanism of diseases and exhibit therapeutic potential. Cell culture and animal model experiments had been to analyze the potential effect and metabolic activity, the additional clinical studies are necessary to fully elucidate therapeutic efficacy and mechanisms of action. High-resolution prediction technology or visualization approaches enhances screen and validates the lead compounds from natural products.728 Eighty-nine compounds were identified and calceolarioside B, isoacteoside, and 2'-acetylacteoside being validated to treat renal fibrosis. The functional mechanisms modulate the metabolic pathways or whole metabolism of natural bioactive compounds needs to be elucidated for use as therapeutic agents. Bioactive compounds could target the small molecule metabolites associated metabolic process of a specific phenotype and modulate metabolic activity of distinct pathways, hold great potential as therapeutic preparations for highly complex diseases.
The potentially vasodilative compounds from Uncaria were screened as isocorynoxeine, corynoxeine, rhynchophylline, isorhynchophylline, by correlation analysis of small metabolites.729 In vitro and in vivo constituents of American ginseng were in-depth investigated using mass spectrometry, and then natural bioactive compounds associated with therapeutic effects were explored using correlation analysis between in vivo constituents and marker metabolites, and revealed ginsenoside Rd, and pseudoginsenoside F11 may be potential active markers of American ginseng.730 By correlation analysis between anti-inflammatory activity of Scutellariae radix and small metabolites, a total of ten potential components were screened out with high correlation coefficients. An in vivo study revealed oroxylin A had the potential effect of antisepsis by inhibiting TLR4/NF-κB signaling pathway.731
Metabolic process of active components
Since the beginning of the 19th century, natural products such as morphine isolated from opium plant have been explored in drug development. With the concept of returning to nature, natural products have attracted great attention and are useful agents for lead compound discovery and new effective drugs. The herb-derived phytochemicals known as natural products have health benefits and their activities and the underlying mechanisms have remained elusive, due to lack a method about how to characterize active components to the whole effect that visualizes dynamic changes in vivo and these components possess diverse effects. To reveal the pharmacological effects of herb-derived phytochemicals with multi-components and multi-targets, the following issues should be solved with emphasis: how to detect active components with low content, elucidate metabolic pathway, reveal overall in vivo metabolic process and effective mechanism. Considering complexity, meanwhile it has numerous metabolic reactions in vivo, producing diverse metabolites. Metabolic processes study in vivo is important to determine multi-component characteristics of efficacy and guide the new drug development, or speed up drug discovery from natural products.732,733,734
Drug discovery process from natural products exhibits some obstacles that presented by the extraction, purification and separating active components. Several of the obstacles have been addressed by employing small molecule metabolites-based screening. Metabolic processes undergo biotransformation mediated by phase I and II reactions or gut microbiota direct impact on the efficacy and are important for determination of pharmacokinetic parameters on the concentrations of active components to the organs and tissues over time.735,736,737 To reveal in vivo processes (absorption, distribution, metabolism, excretion) of multi-active components, it can elaborate the efficacy material basis. Metabolic parameters in vivo, Cmax, Tmax, t1/2, and AUC0–t were the most calculated. Active substances are from the prototypes and the metabolites entering into human circulation, which is directly related to the metabolic process. Endogenous small molecule metabolites can be linked to specific metabolic phenotype, activities, or functions, are closely linked to therapeutic efficacy to screen out the key components in the whole in vivo process.
Understanding metabolic fate of active components is a key factor for elaboration of new therapeutic agents. However, the identified herb-derived bioactive metabolites suggesting the curative potential by modulating multiple targets of disease-associated networks. Owing to the high sensitivity and stability, modern mass spectrometry coupled with all kinds of hyphenated chromatography separation techniques has a pivotal role in the exploration of in vivo metabolism of active components.738,739,740,741,742,743,744,745,746,747,748,749,750,751,752,753,754,755,756,757 Cheminformatics utilizing computer-aided, high-throughput virtual screening, network-based and machine learning techniques have opened up a new avenue in exploiting naturally-inspired products for lead compound and active components discovery.758,759,760,761,762,763,764,765,766,767 A non-targeted metabolomics screening strategy is carried out focusing on the in vivo metabolites that exist in the administrated samples and do not exist in blank biological samples, by exploring the dosage-effect relationships. To selecting in vivo metabolites being associated with therapeutic effect as pharmacological index reflecting overall efficacy is of great significance. Simultaneous determination in vivo of herbal components is technically challenging due to complex interactions with co-existing components. Indeed, LC-MS with selective reaction monitoring mode or background deduction method and pattern recognition analyses were adopted to eliminate possible interference as an effective technique for the identification and quantitative analysis of in vivo components in biological media. Due to active components are complex and their contribution weight to effect is different, it is necessary to explore in vivo metabolic processes of combination multiple active components based on AUC-weighting approach, so that can guide practice administration in the clinic.
Metabolic whole-process in vivo could be realized by efficiently constructing the relationship among endogenous metabolites and compounds. Based on different scores of relationships building in vivo metabolic network, active components markers including metabolites or prototypes that are highly related to small molecule metabolites can be effectively screened out by molecular network technology.768,769,770,771,772 To screen potential active candidates for revealing overall effects, compounds with highly relevant and large VIP values (>1) rankings were selected and identified by the correlation analysis model. The appropriate mix of active components could be optimized via metabolic phenotypic screening and their targets and molecular mechanisms can be revealed by network pharmacology, artificial intelligence, or computer docking.
Metabolic homeostasis and gut microbiota
Increasing evidence shows that homeostasis balance of the human body depends on reciprocal interaction with gut microbiome. Microbial dysbiosis is a contributing factor for onset and progression of diseases. Gut microbiota contribution to control homeostasis, modulating immunity environment, and maintaining systemic health is an emerging.773,774,775,776 It produces a large number of small metabolites that regulates host metabolism responses and metabolic disorders. To date, there is growing evidence that most of small molecular metabolites have beneficial impacts on the host.777,778,779,780,781,782 Gut microbiota can produce various metabolites as messengers between host and microorganisms. Major communication between host and gut microbiota takes place through the metabolites, such as acetate, butyrate, and propionate, and via the microbiota composition modulating metabolism processes.783,784,785,786 Gut microbiota acts as an “invisible organ” and produces active metabolites via their receptor signal to regulate the host metabolism that affects systemic health, and plays a variety of effects on the host from shaping gut structure and function to the modulation of the host status. The increasing evidence showing that gut environment disorder could affect various organs lead to metabolic diseases.787,788,789,790 Imbalance of gut microbiota is closely associated with disease mechanisms, and implying a new therapeutic avenue.
Probiotics can improve the intestinal microecological balance of the host, and play a positive role in enhancing the immunity of the body and helping the absorption of nutrients. The host also could produce a variety of important metabolites that also affect the balance of microbiota. Gut microbiota by producing various bioactive compounds is linked to pharmacological effects and plays an important role in drug absorption, metabolism and efficacy.791,792,793,794 The gut microbiota as an indispensable “organ” to regulation of drug metabolism affects the inherent bioavailability of drugs to reduce toxicity and increase the target efficacy. It can activate or inactivate the pharmacological effect of natural products. Natural products can alter the microbiota compositions or their metabolites via modulating the host metabolism, which could enhance therapeutic effects and attenuate adverse reaction in pharmaceutical development. For instance, some metabolites as ligand metabolic signaling can activate cell-surface GPCRs and may provide potential targets.795 Herbal ingredients could regulate the metabolic disruptions by altering gut microbiota, particularly to reveal the dysregulated metabolites in metabolic pathways interacting with gut microbiota. Since natural products are metabolized and mainly absorbed in the intestinal tract, the secondary metabolites could regulate metabolic perturbations. Microbial-derived products receive considerable attention in disease treatment based on their efficacy by modulating expression of metabolic regulators.
Human metabolism is also influenced by regulating gut microbiota that could regulate metabolic products of the host metabolism enter into circulation system of host and participate in metabolic regulation mechanism in vivo. It continues the awareness of gut microbiota influences on metabolism, due to the complexity and interplay between the host gut and microbiota. Trimethylamine as a toxic molecule cross the gut-blood barrier for circulatory system homeostasis of cardiovascular diseases.796 Gut microbiota as signal distant organs via the systemic circulation affects the pathological processes on controlling host vascular and energy homeostasis. Due to gut microbial dysbiosis, disorder of energy homeostasis balance has a key role in the disease progression. Some evidence points to the important role of gut-derived metabolites in regulating energy metabolism.797,798,799,800,801,802 For instance, SCFA as ligands activating cellular signaling cascades have a variety of beneficial impacts in regulating energy metabolism, especially in obesity-related diseases.803 Small metabolites of gut microbiota, such as bile acids (BAs), and amino acids could decrease the insulin sensitivity and regulate metabolic dysfunction and immune homeostasis, which play a crucial role of the glucose regulation. Interactivity relationship of gut microbiota and BAs as signaling regulators have a profound impact on disease progression through modulating metabolic homeostasis (Fig. 7). In addition, many works have shown that relationship between gut microbiota and BAs plays a critical role in the systemic homeostasis.
Schematic summary of interactions of bile acids and gut microbe participate in the host metabolism. Note: BAs, bile acids; BSEP, bile salt export protein; FGF, fibroblast growth factor; FGFR, FGF receptor; RXR, retinoid X receptor; NTCP, sodium taurocholate cotransporting polypeptide; OATP, organic anion-transporting polypeptide; SHP, small heterodimer partner; JNK, c-Jun N-terminal kinase; ERK, extracellular signal-regulated kinase; T3, thyroid hormone; T4, thyroxine; DIO2, type 2 iodothyronine deiodinase; ASBT, apical sodium-dependent bile acid transporter; OST, organic solute transporter. Primary bile acids are synthesized and then conjugated with taurine or glycine in hepatocytes. Conjugated bile acids are transported into the bile duct by BSEP. Most conjugated bile acids are reabsorbed via ASBT and circulate to the liver by OATP, OSTa/b, and NTCP. Bile acids acts as the endogenous ligands for FXR and TGR5 to generate distinct effects on metabolism regulation. The figures created by BioRender
Reciprocal relationship between host and gut microbiota to maintain homeostasis of biological processes is in dynamic balance and that influenced by endogenous metabolites or biological effects of their precursor. Gut microbiota could produce various small molecules and metabolites, plays a physiological role in maintaining hot homeostasis and internal stability through its metabolites serve as messenger affecting disease states. In addition, some metabolites are only released from gut microbiota, such as bacteriocins, short-chain fatty acids, etc. It showed that tryptophan could also be metabolized by gut microbiota, leading to synthesis of bioactive indoles.804 Additionally, gut microbiota metabolizes primary bile acids, showing a protective role in gastrointestinal diseases. Gut microbiota and their metabolites, as a whole, have been considered as versatile “organ” maintaining the body homeostasis. As gut microbial metabolites, such as secondary bile acids and short-chain fatty acids, could alter the metabolic fluxes by interacting with host receptors, thereby significantly affect metabolic homeostasis leading diseases.805,806,807 These metabolites participate in diverse metabolic processes, such as cell communication, energy metabolism, and host immunity, could influence human physiology, and future studies should define the functional relevance involved in disease pathogenesis.
Gut microbiota plays an important role in affecting the metabolic homeostasis. It has been increasingly appreciated small molecule metabolites-based metabolomics integrating with gut microbiota to explore systemic interaction between metabolites, gut microbiome and disease subtype. Small metabolic biomarkers have used to analyze the interaction between gut microbial composition and metabolism via omics analysis. Recently, Brial et al. discovered that hippurate was a key co-metabolite of host-microbial, could mediate the metabolic improvements associated with high-richness microbiota.808 This work has provided a beneficial biomarker hippurate as a mediator of metabolic health that contributes to metabolic improvements in terms of metabolic phenotype control for the host. In a recent study in Nature, Wu et al. had discovered host-microbe interaction that microbiota-derived inositol phosphate metabolism contributed intestinal homeostasis through mediating the HDAC3 activity in the intestine.809
Analyses in airway serum composition and microbiome datasets demonstrated the gut microbiota can influence on metabolic activity.810 Through depicting the overall landscape of metabolome and microbiome in rheumatoid arthritis patients, 26 genera and 41 metabolites were remarkably altered and function prediction model observed the depleted dysregulation pathways of amino acids biosynthesis.811 Integrated metagenomic and metabolomic analysis had characterized the interactions between metabolites and gut microbiome in early-onset colorectal cancer patients and helps explain the disease pathogenesis. Microbiome-derived metabolites such as bile acid, tryptophan, and choline could be used for the accurate and rapid detection of disease.812 Changes in the levels of several amino acid derived metabolites in a cohort of obese patients, such as the decrease of phenylacetylglutamine and the increase of L-histidine, linked to changes in the gut microbiota composition and function.813 Particularly, P. pentosaceus and L. lactis can ameliorate NAFLD progression by regulating tryptophan metabolism of the gut-liver axis and also closed associated with metabolic dysregulations of bile acid and indole.814 A study analyzed the correlation between serum metabolites and gut microbiota in elderly patients with chronic heart failure insights into metabolic phenotypes.815 Further, biocytin was negatively correlated with Escherichia Shigella; lactose and sucrose were negatively correlated with Haemophilus; bilirubin was positively correlated with Klebsiella; inosine and riboflavin were negatively correlated with Klebsiella. From the plasma metabolome analysis, a positive correlation between metabolite levels and the amount was observed and provide a guide for modulating gut microbiome may help shape a healthy metabolome.816
Influence factors
Duo to the influence of metabolism is multifaceted factors including exogenous or endogenous. Small metabolites could be widely varied by sex, age, weight, nutrition, medications, lifestyle, and circadian rhythm. If they changed, accordingly, the metabolism will also change and then different small metabolites will produce. Certainly, these factors are impossible or even difficult to monitor and may represent the most challenging in metabolic biomarkers studies. These factors can lead to metabolic dysregulation of body, which can cause metabolic pathway alterations in disease-associated patients. When some influencing factors were intervened by drugs, early prevention, timely and effective disease treatment can achieve.
Small metabolites could participate in the entire “metabolic chain” in body, and has important implications for identifying the metabolic features related to disease phenotypic variation. Although the potential metabolite markers for diagnosis and treatment of diseases have screened and made great progress, and still needed to be validated the feasibility in a large number of cohort studies. Differences in some marker metabolites between the humans and animal experiment need to consider it. Compared with animal experiments, some elements, e.g. gender, age, psychological status, disease history, diet, exercise pattern, habits and customs, and other lifestyle differences have been considered to be significant variation factors for the altered metabolism within the body.
Metabolome is affected by genetic factors, intestinal flora and environmental exposures.817,818,819,820 These changes might originate from the environment, physiological or pathological status, diseases, drugs or other external stimuli, which can significantly contribute to the metabolome composition. Moreover, small molecule metabolites-based metabolomics can identify as many marker metabolites as possible and then reveal the metabolic network associations of bioactive substances and effective targets, needs to validate these associations, and further mechanistic research on dysregulated metabolites should be implemented and reproduced. Metabolomics contrary to genomics, proteomics, or transcriptomics, can rapidly and accurately reflect a more clearly phenotypic state of organisms, however it influenced by outside confounding factors. Previous studies have shown these factors could influence the metabolic phenotypes of metabolism-related diseases.
Limitations and challenges
As we have shown, small molecule metabolites-based metabolomics has enormous potentials and many applications, however, several challenges and limitations need to be further addressed. The complexity of a large number of metabolic signatures that present in “dark metabolome” associated with manifestations of transient phenotypic state is one of the huge challenges. Advantage of omics as tools for biomarker discovery simultaneously quantifying a large number of small molecules in biological media. Since metabolome associated metabolic alteration is highly complex, and no uniform strategy and distinct analytical platform can analyze all the small metabolites of the whole phenotype. Others are the challenges in small metabolite analysis associated metabolic alteration include the genetic factors, environmental factors, or gut microbiota. Numerous small metabolites from different biological media were considered to be candidate biomarker of predicting the of disease onset, and therapeutic response. Determination of small metabolites as excellent candidate are measured from no-invasive samples such as urine and blood. Potential role of the target metabolites needs further verify and identify as biomarkers in the disease diagnosis, management and prognosis of diseases. Predictive ability, disease prognosis and diagnostic value of the established metabolic profiles need further validate with larger sample sizes in real-world medicine. In addition, method standardization of large randomized clinical trials is needed.
Previous studies have shown that quality of metabolic data lead to significant variability that influenced by sampling techniques and analytic approaches.821,822,823 Technological limitations or insufficient use of metabolomics are partly caused by structurally diverse metabolites, standardization and uniformity of instrumentation, temperature variation, sample preparation and handling, proficiency and availability of trained staff. It is important to emphasize the standardization of laboratory procedures, such as extraction, sample processing, and other analytical protocols, is a fundamental step to obtain biologically meaningful metabolic results. It is necessary to establish the commonly accepted standardized criteria or protocols for sample extraction, data mining and data reporting. This could resolve the major challenges with metabolite identification by used various spectroscopy, chromatographic methodology to influence specific constituent, result evaluation by employed different statistical methods and interpretation of clinical significance, all of which could affect experimental or clinical outcome and thus limit the application of small molecule metabolites-based metabolomics into clinical aspects. In the past decade, the significant progress and improvements in technical aspects have been made for small metabolite analysis from metabolic perturbations in tissues and biofluids to further promote understanding of molecular mechanisms to advance meaningful interpretation of metabolic features related to phenotypic variation. Analyzing and revealing metabolic changes in disease response to drugs could provide opportunities to discover the potential targets and biologically meaningful metabolic pathways for metabolism-related diseases therapy. Fortunately, targeted metabolic profiling of some metabolites has been endorsed to be applied in clinical practice for disease markers and potential targets identification for monitoring, diagnosis, and drug efficacy.
The accurate masses, fragment mass spectra and retention time should be provided to identify metabolites via database-based search methods. However, considering that significant amounts of datasets, special statistical software, complexity of computational processing, bioinformatics tool, lead to detecting specific molecules, validate the pathways and associations, analyze data even more difficult. Databases for metabolome analysis with extensive metabolite coverage with help of multivariate analysis have been significantly developed for data identification and visualization. Small metabolites are downstream of transcriptome-proteome and their metabolism were affected by various microbiota in vivo, so that multi-omics can create approach to explore the interactions of proteins, metabolites, genes, and microbiota, and then reveal the pathophysiological mechanisms in both diseased and non-diseased states. Fortunately, integration with other omics could insight into the characteristic metabolic alterations.824,825 Integrative analysis of omics data by multi-omics technology could provide the mechanistic insights into diseases and bring precision treatment.
One of the biggest challenges is mainly in the realm of data integration still in early stages and needs additional consideration. To achieve this goal, high-throughput integration multi-omics with help improvement of computational and bioinformatics techniques has greatly contributed to accurately identify the relevant small-metabolites and their biological processes involved in metabolic perturbations in vivo. Such integration analysis approach can effectively use it to understand abnormal biological mechanisms in the underlying metabolic network of interest by visualizing metabolic pathways. The integration importance of metabolomics with other omics is seen in some very recent research.826,827,828 The high complexity of metabolome poses another challenge for identifying metabolic features related to phenotypic variation. Fortunately, computational approaches and artificial intelligence-based algorithms representing promising tools for biomarker discovery is provided to overcome the above problems. Advanced analytical techniques, such as artificial intelligence, computational algorithms, metabolite imaging, statistical, and big data mining are urgently needed to improve coverage of the low-abundance metabolites for clinical validity and utility of small molecule metabolites involved in disease pathogenesis.
The full validation stage of the small molecule metabolites-based metabolomics workflow is missing and prohibits the biomarker discovery to clinical translation over time. However, there are several limitations should be addressed in future research. Due to lack of standardization in process research and external validation and some works need to be done before ascertaining biomarkers for clinical practice. Utilization of standardization process should be embodied in each stage, such as patient enrollment, sample collection and processing, storage, preparation treatment, data acquisition and in-depth analysis. In addition, the standardization of sample preparation and processing, data analysis and variation factors will help to promote the research of small molecule metabolites to beyond the discovery stage of biomarkers and towards the development and validation of clinical trials.
Indeed, small molecule signatures provide crucial information for diagnostic and prognostic biomarkers, and therapeutic response. However, it remains a subsequent challenge of translation of small molecule signatures from laboratory results to clinical and industrial application. An open issue is most researches still have done with smaller cohorts, and it requires future studies with a larger cohort dataset from multicenter studies in clinic should include many metabolite biomarkers and metabolic pathways for accurate diagnosis of diseases and better understanding metabolic alterations, and should be addressed in the future.
Future outlook
Altered metabolism leads to characteristic metabolic phenotypes as the hallmark of disease that drive identification of new targets related to metabolic regulation mechanism could be applied for developing effective screening strategies for predicting early disease, or evaluation treatment monitoring responses. Metabolomics, as mentioned above, is a relatively young discipline relative to other omics, has identified small molecule metabolite biomarkers for the disease diagnosis, prediction, screening, and monitoring treatment. Compared to genome or transcriptome, coverage of metabolome remains limited and lead inadequate interpretation of the final results. Due to no single technology can offer an entire metabolic spectrum, thereby different advanced analytical chemistry platforms are recommended to integrate metabolomics with upstream omics and network target analysis. Integration of multi-omics datasets can represent a powerful method to reveal metabolic signatures related to phenotypic variation of patients.829 Moreover, multi-omics integrative analysis can uncover disease biomarkers and new pathological pathways, deepen understanding of mechanistic basis and therapeutic targets of metabolic diseases, and accelerate new drug development for better therapy, significantly enhanced translational capability.830,831,832,833 The integration of multi-omics analysis may present precise metabolic biomarkers, a global metabolic snapshot and metabolic networks, which can deepen exploration of underlying mechanisms towards improving clinical management of disease. However, due to the complexity of metabolic pathway data and the interaction between metabolic network and other factors, the integration of multi omics data is a huge challenge. It needs to establish an international network or a different platform with modern instruments for the integrative multi-omics data by biologists, statistician and chemists.
Future work should strive to solve many of the following clinical problems, e.g. limited in sample size and control groups, validation of candidate biomarkers in clinical settings. It is still difficult to rapidly separate the small molecule metabolite while keeping their metabolic states, especially under metabolic disturbance, because metabolite biomarker is susceptible to change of environmental factors. Future studies should validate the selected small metabolites in larger sample datasets to increase the analysis credibility and statistical power that help to our understanding of the aberrant metabolism. In this scenario, it needs to improve the detection precision and accuracy of small metabolites from the entire metabolome in real time analysis. Particularly, break the technical bottlenecks by substantial efforts needed to acquire high sensitivity and accuracy for extensive coverage detection of metabolome. Recently, high-resolution mass spectrometry improves detection capability and enable us to identify metabolic biomarkers towards clinical validation.834,835,836
Forthcoming research of diseases will address questions about the integrated metabolic phenotypes, and then how to transform them into clinical applications for better therapies. Large, prospective clinical practice can validate the discovered small molecule metabolites with high translational chances for diagnostics, prevention and therapeutics. Nevertheless, upcoming challenges will include harmonization and normalization of disparate datasets, protocols standardizations and new algorithmic analysis to better explore the underlying mechanisms and insights. Likewise, software for data processing and interpretation is becoming standardized and widespread. Standardization of procedures for meaningful and accurate management of metabolite biomarker research that modulates biological processes should be further refined for untargeted or targeted analysis towards ensuring laboratory results become clinical translational. Therefore, targeted and functional analysis approach to overcome limitations of conventional metabolomics is a new strategy for exploring the small-molecule metabolism associated mechanisms of complex alterations of systemic homeostasis. A large number of small metabolites have been identified as disease biomarkers or predictors.466,837,838,839,840,841,842 Therefore, future research should reduce the number of diagnostic or prognostic biomarkers to the most appropriate number.
With help of the enhanced AI or big data analytics, machine learning or developing algorithms, a clear understanding of underlying pathological pathways and metabolism-related diseases can offer a tangible route or evidence to support clinical decisions in addressing the formalized utilization of small molecule metabolites or metabolic phenotyping into routine clinic. On localizing metabolic alterations is key to driving our understanding of disease, advances in metabolic imaging in an unprecedented approach actively realized better resolution of metabolic property to quantify metabolites and illuminating metabolic pathways. In the near future, the significant improvement in metabolic imaging tool combined with innovative algorithms enables to differentiate tissue states for various diseases. Current applications of multi-omics via modern imaging techniques and computational improvements translate to clinical diagnosis, prevention and precision treatment, and facilitate selective treatment in future medical care.
Future work should explore the non-invasive, specific biomarkers with high diagnostic value, simplify the evaluation process assessment of candidate biomarkers into clinical application, develop new algorithms or bioinformatics software for exploring molecular interactions associated metabolic alteration, and establish effective association integrating laboratory and clinical results. Additionally, the further characterization of small molecule metabolites associated gut microbiome could shed light on metabolic features related to phenotypic variation of disease pathogenesis, enable to assist diagnosis, prognosis, treatment strategy selection, and realize the benefits that of small molecule metabolites bring to precision treatment. Small molecule metabolites-based metabolomics has greatly changed biomedical research. In the future, metabolite biomarkers will need to be effectively validated and transferred to clinical applications, thereby researchers should work closely with clinicians. For increasing the accessibility of metabolite biomarkers, its analytical instruments should become far cheaper and simpler, insights into specific metabolic phenotypes. With the joint efforts of the government and industrial community, all this will be achieved.
Conclusion
Abnormal metabolites could serve as potential biomarkers for evaluating diagnosis and monitoring treatment response and prognosis, will provide abundant evidence for future precise medicine. A variety of metabolic pathways are altered in human diseases, including fatty acid oxidation, amino acid metabolism, lipid and energy metabolism, glycolysis, phospholipid metabolism, and tricarboxylic acid cycle that maybe considered as potential targets for further clinical trials. Metabolome has provided a comprehensive, dynamic, and precise picture of metabolic phenotype that could confer the personalized clinical practice. Metabolomics has changed the world of metabolite biomarkers, for its utilization of identification of pre-disease states, diagnosis, subtyping, prognosticating and monitoring treatment response, provide evidence for the early diagnosis, prevention, and mechanistic exploration. Current milestone findings encourage further investigations to urgent application in the clinic. Numerous studies are also going on metabolomics-based discovery of identifying the unique metabolic signatures, opens up the complex metabolite networks in physiological or pathophysiological processes; its biomedical applications can already be foreseen to monitor health and disease, assess disease severity or drug development, predict the time-course, monitor progression of diseases, and predict potential treatments, and elucidation of disease mechanisms.
The present review summarized the outcomes of most significant researches to extend the knowledge of small-molecule metabolite biomarkers. A major limitation is the absence of validation of metabolic phenotype-related mechanistic targets of diseases, which leads to lack the focused therapy and become increasingly prevalent. It is vital that we should dig out the most sensitive and accurate, specific metabolite signatures and conduct more studies to corroborate and validate these findings. However, standardization studies of metabolite applications methods and validation with a greater degree of certainty in large-scale clinical samples are needed before these tests could be a wide range of applications in clinical settings. In addition, multicenter exploration with large-scale populations in validation of small-molecule metabolite biomarkers is still needed for clinical translation and utilization. Integration multi-omics data combined with clinical measures has potential to facilitate delineating of disease progress and treatment. The combined analysis of multi-omics data focused on the precise metabolic phenotype characterization offer opportunities to facilitate deciphering the molecular changes underlying metabolic mechanisms in human diseases. Upcoming research should improve the diagnostic ability of potential biomarkers to easily predict diagnosis and prognosis. Additionally, future direction to address clinical application relates to the establishment of the relationships between metabolic profiles and clinical parameters. Future endeavors should increase the confidence in metabolite identification by systematic large-scale profiling analysis and emphasize determining the applicability of metabolomic-derived biomarkers and their clinical utility in large-scale clinical settings. Further targeted metabolic profile is needed to better explore the suitability of small-molecule metabolite as initial indicators of diseases, better understanding of pathophysiologic mechanisms, mitigating the risk and benefit from the best treatment, may open novel avenues for future precise medicine.
References
Goodman, R. P. et al. Hepatic NADH reductive stress underlies common variation in metabolic traits. Nature 583, 122–126 (2020).
White, P. J. & Newgard, C. B. Branched-chain amino acids in disease. Science 363, 582–583 (2019).
Sovio, U. et al. A maternal serum metabolite ratio predicts fetal growth restriction at term. Nat. Med. 26, 348–353 (2020).
Qiu, S. et al. Decoding functional significance of small molecule metabolites. Biomed. Pharmacother. 158, 114188 (2022).
Imperlini, E. et al. Mass spectrometry-based metabolomic and proteomic strategies in organic acidemias. Biomed. Res. Int. 2016, 9210408 (2016).
Bergers, G. & Fendt, S. M. The metabolism of cancer cells during metastasis. Nat. Rev. Cancer 21, 162–180 (2021).
Girdhar, K. et al. Dynamics of the gut microbiome, IgA response, and plasma metabolome in the development of pediatric celiac disease. Microbiome 11, 9 (2023).
DeBerardinis, R. J. & Keshari, K. R. Metabolic analysis as a driver for discovery, diagnosis, and therapy. Cell 185, 2678–2689 (2022).
Su, Y. et al. Multi-omics resolves a sharp disease-state shift between mild and moderate COVID-19. Cell 183, 1479–1495.e1420 (2020).
Llufrio, E. M., Cho, K. & Patti, G. J. Systems-level analysis of isotopic labeling in untargeted metabolomic data by X(13)CMS. Nat. Protoc. 14, 1970–1990 (2019).
Chen, Y. et al. Genomic atlas of the plasma metabolome prioritizes metabolites implicated in human diseases. Nat. Genet. 55, 44–53 (2023).
Qiu, S. et al. Innovation in identifying metabolites from complex metabolome—Highlights of recent analytical platforms and protocols. Front. Chem. 11, 1129717 (2023).
Alexander, J. L. et al. The gut microbiota and metabolome are associated with diminished COVID-19 vaccine-induced antibody responses in immunosuppressed inflammatory bowel disease patients. EBioMedicine 88, 104430 (2023).
Wilmanski, T. et al. Blood metabolome predicts gut microbiome α-diversity in humans. Nat. Biotechnol. 37, 1217–1228 (2019).
Liu, Q. et al. Altered faecal microbiome and metabolome in IgG4-related sclerosing cholangitis and primary sclerosing cholangitis. Gut 71, 899–909 (2022).
Coyle, S. et al. Predicting dying from lung cancer: Urine metabolites predict the last weeks and days of life. J. Clin. Oncol. 39, 12030–12030 (2021).
Wang, B. et al. Metabolism pathways of arachidonic acids: mechanisms and potential therapeutic targets. Signal Transduct. Target Ther. 6, 94 (2021).
Di'Narzo, A. F. et al. Integrative analysis of the inflammatory bowel disease serum metabolome improves our understanding of genetic etiology and points to novel putative therapeutic targets. Gastroenterology 162, 828–843.e811 (2022).
Perea-Gil, I. et al. Serine biosynthesis as a novel therapeutic target for dilated cardiomyopathy. Eur. Heart J. 43, 3477–3489 (2022).
Genchi A. et al. Neural stem cell transplantation in patients with progressive multiple sclerosis: an open-label, phase 1 study. Nat. Med. https://doi.org/10.1038/s41591-022-02097-3 (2023).
Gong, Y. et al. Metabolic-pathway-based subtyping of triple-negative breast cancer reveals potential therapeutic targets. Cell Metab. 33, 51–64.e59 (2021).
Xiao, Y. et al. Comprehensive metabolomics expands precision medicine for triple-negative breast cancer. Cell Res. 32, 477–490 (2022).
Liu, P. et al. Critical roles of functional molecule metabolites. Front. Mol. Biosci. 10, 1119588 (2023).
Wrzosek, L. et al. Microbiota tryptophan metabolism induces aryl hydrocarbon receptor activation and improves alcohol-induced liver injury. Gut 70, 1299–1308 (2021).
Ginsberg, H. N. et al. Triglyceride-rich lipoproteins and their remnants: metabolic insights, role in atherosclerotic cardiovascular disease, and emerging therapeutic strategies-a consensus statement from the European Atherosclerosis Society. Eur. Heart J. 42, 4791–4806 (2021).
Xie, N. et al. NAD(+) metabolism: pathophysiologic mechanisms and therapeutic potential. Signal Transduct. Target Ther. 5, 227 (2020).
Huang, L. et al. Rapid, label-free histopathological diagnosis of liver cancer based on Raman spectroscopy and deep learning. Nat. Commun. 14, 48 (2023).
Dong, R. et al. Principal components from untargeted CSF metabolomics associated with tau. Alzheimer’s Dement. 16, e046065 (2020).
Bar, N. et al. A reference map of potential determinants for the human serum metabolome. Nature 588, 135–140 (2020).
Patti, G. J., Tautenhahn, R. & Siuzdak, G. Meta-analysis of untargeted metabolomic data from multiple profiling experiments. Nat. Protoc. 7, 508–516 (2012).
Ogawa, T. et al. Novel regulation of cardiac branched-chain amino acid metabolism through AMP deaminase: a possible therapeutic target for diabetic cardiomyopathy. Eur. Heart J. 41, ehaa946.3619 (2020).
Chen, F. et al. Integrated analysis of the faecal metagenome and serum metabolome reveals the role of gut microbiome-associated metabolites in the detection of colorectal cancer and adenoma. Gut 71, 1315–1325 (2022).
Li, Z. B. et al. Novel potential metabolic biomarker panel for early detection of severe COVID-19 using full-spectrum metabolome and whole-transcriptome analyses. Signal Transduct. Target Ther. 7, 129 (2022).
Adam, M. G. et al. Identification and validation of a multivariable prediction model based on blood plasma and serum metabolomics for the distinction of chronic pancreatitis subjects from non-pancreas disease control subjects. Gut 70, 2150–2158 (2021).
Esther, C. R. Jr et al. Metabolomic biomarkers predictive of early structural lung disease in cystic fibrosis. Eur. Respir. J. 48, 1612–1621 (2016).
Zhang, S. J. et al. Ketone body 3-hydroxybutyrate ameliorates atherosclerosis via receptor Gpr109a-mediated calcium influx. Adv. Sci. (Weinh.). 8, 2003410 (2021).
Rinschen, M. M., Ivanisevic, J., Giera, M. & Siuzdak, G. Identification of bioactive metabolites using activity metabolomics. Nat. Rev. Mol. Cell Biol. 20, 353–367 (2019).
Hu, L. et al. Functional metabolomics decipher biochemical functions and associated mechanisms underlie small-molecule metabolism. Mass Spectrom. Rev. 39, 417–433 (2020).
Odom, J. D. & Sutton, V. R. Metabolomics in clinical practice: improving diagnosis and informing management. Clin. Chem. 67, 1606–1617 (2021).
Liang, Y., Zhang, H. & Cai, Z. New insights into the cellular mechanism of triclosan-induced dermal toxicity from a combined metabolomic and lipidomic approach. Sci. Total Environ. 757, 143976 (2021).
Handakas, E. et al. A systematic review of metabolomic studies of childhood obesity: State of the evidence for metabolic determinants and consequences. Obes. Rev. 23(Suppl 1), e13384 (2022).
Borges, R. M. et al. Quantum chemistry calculations for metabolomics. Chem. Rev. 121, 5633–5670 (2021).
Badhwar, A. et al. A multiomics approach to heterogeneity in Alzheimer’s disease: focused review and roadmap. Brain 143, 1315–1331 (2020).
Chen, L. et al. Metabolite discovery through global annotation of untargeted metabolomics data. Nat. Methods 18, 1377–1385 (2021).
Perakakis, N., Stefanakis, K. & Mantzoros, C. S. The role of omics in the pathophysiology, diagnosis and treatment of non-alcoholic fatty liver disease. Metabolism 111s, 154320 (2020).
Guo, Q., He, Z., Liu, X., Liu, B. & Zhang, Y. High-throughput non-targeted metabolomics study of the effects of perfluorooctane sulfonate (PFOS) on the metabolic characteristics of A. thaliana leaves. Sci. Total Environ. 710, 135542 (2020).
Dührkop, K. et al. Systematic classification of unknown metabolites using high-resolution fragmentation mass spectra. Nat. Biotechnol. 39, 462–471 (2021).
Han, S. et al. A metabolomics pipeline for the mechanistic interrogation of the gut microbiome. Nature 595, 415–420 (2021).
Harju K. et al. Simultaneous metabolomics analysis of atrial tissue, pericardial fluid and blood reveal novel metabolite signatures of the pathophysiology and biomarkers related to permanent atrial fibrillation. Eur. Heart J. 43, ehac544.503 (2022).
Carter, B. Z. et al. Inhibition of anti-apoptotic Mcl-1 exerts anti-leukemia activity through modulation of leukemia-stromal interactions and metabolic functions in AML. Blood 134, 3727–3727 (2019).
Yackoubov, D. et al. Transcriptional and metabolic profiling of nicotinamide-enhanced natural killer (NAM-NK) cells (GDA-201). Blood 138, 4791–4791 (2021).
Niu, L. et al. Noninvasive proteomic biomarkers for alcohol-related liver disease. Nat. Med. 28, 1277–1287 (2022).
Hinshaw, D. C. et al. Hedgehog signaling regulates metabolism and polarization of mammary tumor-associated macrophages. Cancer Res. 81, 5425–5437 (2021).
Forte, D. et al. Circulating extracellular vesicles from acute myeloid leukemia patients drive distinct metabolic profile of leukemic cells and reveal crucial lipidomic biomarkers. Blood 138, 3471–3471 (2021).
Mallick, H. et al. Predictive metabolomic profiling of microbial communities using amplicon or metagenomic sequences. Nat. Commun. 10, 3136 (2019).
Zhou, Z. et al. Ion mobility collision cross-section atlas for known and unknown metabolite annotation in untargeted metabolomics. Nat. Commun. 11, 4334 (2020).
Sarvin, B. et al. Fast and sensitive flow-injection mass spectrometry metabolomics by analyzing sample-specific ion distributions. Nat. Commun. 11, 3186 (2020).
Pang, Z. et al. MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 49, W388–w396 (2021).
Shen, X. et al. Metabolic reaction network-based recursive metabolite annotation for untargeted metabolomics. Nat. Commun. 10, 1516 (2019).
Li, Y. et al. Sub-nanoliter metabolomics via mass spectrometry to characterize volume-limited samples. Nat. Commun. 11, 5625 (2020).
Voss, K. et al. A guide to interrogating immunometabolism. Nat. Rev. Immunol. 21, 637–652 (2021).
Faubert, B., Solmonson, A. & DeBerardinis, R. J. Metabolic reprogramming and cancer progression. Science 368, eaaw5473 (2020).
Brewer, M. K. et al. Targeting pathogenic lafora bodies in lafora disease using an antibody-enzyme fusion. Cell Metab. 30, 689–705.e686 (2019).
Yuan, Z. et al. SEAM is a spatial single nuclear metabolomics method for dissecting tissue microenvironment. Nat. Methods 18, 1223–1232 (2021).
Van Dooijeweert, B. et al. Untargeted metabolomic fingerprinting as a potential tool in the diagnostic evaluation of diamond blackfan anemia. Blood 136, 7–8 (2020).
Caocci, G. et al. Metabolomics profile of patients with chronic myeloid leukemia and cardiovascular adverse events after treatment with tyrosine kinase inhibitors. Blood 134, 4144–4144 (2019).
Jin, H. et al. Novel oncogenic non-coding RNA:circRIC8B regulates lipid metabolism Via Mir-199b-5p /LPL axis in chronic lymphocytic leukemia. Blood 138, 3712–3712 (2021).
Vitko, D. et al. Urinary tract infections in children with vesicoureteral reflux are accompanied by alterations in urinary microbiota and metabolome profiles. Eur. Urol. 81, 151–154 (2022).
van Zyl, C. W., Loots, D. T., Solomons, R., van Reenen, M. & Mason, S. Metabolic characterization of tuberculous meningitis in a South African paediatric population using (1)H NMR metabolomics. J. Infect. 81, 743–752 (2020).
Mukhopadhyay, M. Metabolic profiling of CD8(+) T cells at the single-cell level. Nat. Methods 17, 1071 (2020).
Li, T. W., Huang, Y., Zhong, Z. & Huang, Q. THU0405 serum metabolic profiling analysis of gout patients based on Uhplc-Q-Tof/Ms. Ann. Rheum. Dis. 79, 440 (2020).
Aragon Herrera, A. et al. Empaglifozin induces changes in the liver metabolome of diabetic rats. Eur. Heart J. 41, ehaa946.3825 (2020).
Shen, B. et al. Proteomic and metabolomic characterization of COVID-19 patient sera. Cell 182, 59–72.e15 (2020).
Bonnay, F. et al. Oxidative metabolism drives immortalization of neural stem cells during tumorigenesis. Cell 182, 1490–1507.e1419 (2020).
Moreau, R. et al. Blood metabolomics uncovers inflammation-associated mitochondrial dysfunction as a potential mechanism underlying ACLF. J. Hepatol. 72, 688–701 (2020).
Xie, X. et al. Activation of anxiogenic circuits instigates resistance to diet-induced obesity via increased energy expenditure. Cell Metab. 29, 917–931.e914 (2019).
McMillan, A. & Hazen, S. L. Gut microbiota involvement in ventricular remodeling post-myocardial infarction. Circulation 139, 660–662 (2019).
Koundouros, N. et al. Metabolic fingerprinting links oncogenic PIK3CA with enhanced arachidonic acid-derived eicosanoids. Cell 181, 1596–1611.e1527 (2020).
Watson, M. J. et al. Metabolic support of tumour-infiltrating regulatory T cells by lactic acid. Nature 591, 645–651 (2021).
He, Y. et al. Gut microbial metabolites facilitate anticancer therapy efficacy by modulating cytotoxic CD8(+) T cell immunity. Cell Metab. 33, 988–1000.e1007 (2021).
Decano, J. L. et al. Systems approach to discovery of therapeutic targets for vein graft disease: PPARα pivotally regulates metabolism, activation, and heterogeneity of macrophages and lesion development. Circulation 143, 2454–2470 (2021).
Mars, R. A. T. et al. Longitudinal multi-omics reveals subset-specific mechanisms underlying irritable bowel syndrome. Cell 182, 1460–1473.e1417 (2020).
Liang, L. et al. Metabolic dynamics and prediction of gestational age and time to delivery in pregnant women. Cell 181, 1680–1692.e1615 (2020).
Garcia-Bermudez, J. et al. Squalene accumulation in cholesterol auxotrophic lymphomas prevents oxidative cell death. Nature 567, 118–122 (2019).
Chen, L. et al. The long-term genetic stability and individual specificity of the human gut microbiome. Cell 184, 2302–2315.e2312 (2021).
Augustijn, H. E. & Medema, M. H. Freedom of expression: A synthetic route to metabolites. Cell 185, 1449–1451 (2022).
Parker, B. L. et al. An integrative systems genetic analysis of mammalian lipid metabolism. Nature 567, 187–193 (2019).
Hang, S. et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nature 576, 143–148 (2019).
Choi, W. S. et al. The CH25H-CYP7B1-RORalpha axis of cholesterol metabolism regulates osteoarthritis. Nature 566, 254–258 (2019).
Gao, X. et al. Dietary methionine influences therapy in mouse cancer models and alters human metabolism. Nature 572, 397–401 (2019).
Ogawa, T. et al. Intracellular localization of AMP deaminase and its novel role in BCAA and lipid metabolism in diabetic cardiomyopathy. Eur. Heart J. 42, ehab724.3228 (2021).
Johnson, C. H., Ivanisevic, J. & Siuzdak, G. Metabolomics: beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 17, 451–459 (2016).
Allesøe, R. L. et al. Discovery of drug-omics associations in type 2 diabetes with generative deep-learning models. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01520-x (2023).
Vatanen, T. et al. Mobile genetic elements from the maternal microbiome shape infant gut microbial assembly and metabolism. Cell 185, 4921–4936 (2023).
Campos, A. I. & Zampieri, M. Metabolomics-driven exploration of the chemical drug space to predict combination antimicrobial therapies. Mol. Cell. 74, 1291–1303.e1296 (2019).
Chung, K. P. et al. Mitofusins regulate lipid metabolism to mediate the development of lung fibrosis. Nat. Commun. 10, 3390 (2019).
Xie, J. et al. Akkermansia muciniphila protects mice against an emerging tick-borne viral pathogen. Nat. Microbiol. 8, 91–106 (2023).
Jacobs, J. P. et al. Multi-omics profiles of the intestinal microbiome in irritable bowel syndrome and its bowel habit subtypes. Microbiome 11, 5 (2023).
Storbeck, K. H. et al. Steroid metabolome analysis in disorders of adrenal steroid biosynthesis and metabolism. Endocr. Rev. 40, 1605–1625 (2019).
Yachida, S. et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat. Med. 25, 968–976 (2019).
Masoodi, M. et al. Metabolomics and lipidomics in NAFLD: biomarkers and non-invasive diagnostic tests. Nat. Rev. Gastroenterol. Hepatol. 18, 835–856 (2021).
Macedo, A. N. et al. The sweat metabolome of screen-positive cystic fibrosis infants: revealing mechanisms beyond impaired chloride transport. ACS Cent. Sci. 3, 904–913 (2017).
Cheng, Y. et al. Rare genetic variants affecting urine metabolite levels link population variation to inborn errors of metabolism. Nat. Commun. 12, 964 (2021).
Huang, Y. H. et al. Cancer-associated fibroblast-derived interleukin-1β activates protumor C-C motif chemokine ligand 22 signaling in head and neck cancer. Cancer Sci. 110, 2783–2793 (2019).
Dey, S. K. et al. Repurposing an adenine riboswitch into a fluorogenic imaging and sensing tag. Nat. Chem. Biol. 18, 180–190 (2022).
Wishart, D. S. et al. HMDB 5.0: the human metabolome database for 2022. Nucleic Acids Res. 50, D622–d631 (2022).
Gallois, A. et al. A comprehensive study of metabolite genetics reveals strong pleiotropy and heterogeneity across time and context. Nat. Commun. 10, 4788 (2019).
Zhou, Y., Hu, G. & Wang, M. C. Host and microbiota metabolic signals in aging and longevity. Nat. Chem. Biol. 17, 1027–1036 (2021).
Zhang, B. et al. B cell-derived GABA elicits IL-10(+) macrophages to limit anti-tumour immunity. Nature 599, 471–476 (2021).
Xia, Y. et al. Mesenchymal stromal cells overexpressing farnesoid X receptor exert cardioprotective effects against acute ischemic heart injury by binding endogenous bile acids. Adv. Sci. (Weinh.). 9, e2200431 (2022).
Shigeta, K. et al. IDH2 stabilizes HIF-1α-induced metabolic reprogramming and promotes chemoresistance in urothelial cancer. EMBO J. 42, 110620 (2023).
Perino, A. & Schoonjans, K. Metabolic messengers: bile acids. Nat. Metab. 4, 416–423 (2022).
Li, W. et al. A bacterial bile acid metabolite modulates Treg activity through the nuclear hormone receptor NR4A1. Cell Host Microbe 29, 1366–1377.e1369 (2021).
Hoogerland, J. A. et al. Glucose-6-phosphate regulates hepatic bile acid synthesis in mice. Hepatology 70, 2171–2184 (2019).
Galmozzi, A. et al. PGRMC2 is an intracellular haem chaperone critical for adipocyte function. Nature 576, 138–142 (2019).
Venegas-Molina, J., Molina-Hidalgo, F. J., Clicque, E. & Goossens, A. Why and how to dig into plant metabolite-protein interactions. Trends Plant Sci. 26, 472–483 (2021).
Shimizu, K. & Matsuoka, Y. Redox rebalance against genetic perturbations and modulation of central carbon metabolism by the oxidative stress regulation. Biotechnol. Adv. 37, 107441 (2019).
Julius, C., Salgado, P. S. & Yuzenkova, Y. Metabolic cofactors NADH and FAD act as non-canonical initiating substrates for a primase and affect replication primer processing in vitro. Nucleic Acids Res. 48, 7298–7306 (2020).
Chen, D. et al. Lysine acetylation restricts mutant IDH2 activity to optimize transformation in AML cells. Mol. Cell. 81, 3833–3847.e3811 (2021).
Li, Z. et al. Single-cell lipidomics with high structural specificity by mass spectrometry. Nat. Commun. 12, 2869 (2021).
da Silveira, W. A. et al. Comprehensive multi-omics analysis reveals mitochondrial stress as a central biological hub for spaceflight impact. Cell 183, 1185–1201.e1120 (2020).
Chan, K. R. et al. Metabolic perturbations and cellular stress underpin susceptibility to symptomatic live-attenuated yellow fever infection. Nat. Med. 25, 1218–1224 (2019).
Wozniak, J. M. et al. Mortality risk profiling of staphylococcus aureus bacteremia by multi-omic serum analysis reveals early predictive and pathogenic signatures. Cell 182, 1311–1327.e1314 (2020).
Keshavan, M. S. Characterizing transdiagnostic premorbid biotypes can help progress in selective prevention in psychiatry. World Psychiatry 20, 231–232 (2021).
Özer, Ö. et al. Detection of brain metastasis by metabolomics methods in metastatic breast cancer patients. J. Clin. Oncol. 37, e12572–e12572 (2019).
Thyparambil, S. P. et al. Deviation from the precisely timed age-associated patterns revealed by blood metabolomics to find CRC patients at risk of relapse at the CRC diagnosis. J. Clin. Oncol. 40, 206–206 (2022).
Murata, T. et al. Machine learning methods with salivary metabolomics for breast cancer detection. J. Clin. Oncol. 37, 3135–3135 (2019).
Wang, L. B. et al. Proteogenomic and metabolomic characterization of human glioblastoma. Cancer Cell. 39, 509–528.e520 (2021).
Bancos, I. et al. Urine steroid metabolomics for the differential diagnosis of adrenal incidentalomas in the EURINE-ACT study: a prospective test validation study. Lancet Diabetes Endocrinol. 8, 773–781 (2020).
Han, W. et al. OP0304 metabolomics profiling of human serum for discovering biomarkers to diagnose psoriatic arthritis and ankylosing spondylitis with high specificity. Ann. Rheum. Dis. 79, 188–189 (2020).
El Zarif, T. et al. Comprehensive metabolomic profiling of plasma from patients (pts) with metastatic urothelial carcinoma (mUC) receiving immune checkpoint inhibitors (ICI) or platinum-based chemotherapy (PBC). J. Clin. Oncol. 40, 565–565 (2022).
Cocco, D. et al. Defining the metabolomic profile associated with early cardiotoxicity in patients with breast cancer treated with anthracyclines. Eur. Heart J. 41, ehaa946.3289 (2020).
Frias, M. et al. Evaluation of antiretroviral therapy on metabolomics and atherogenic markers in HIV patients. Eur. Heart J. 41, ehaa946.3307 (2020).
Frias, M. et al. HIV-infected patients display increased proatherogenic anti-apolipoprotein A1 autoantibodies, inflammatory and metabolomic markers. Eur. Heart J. 42, ehab724.2906 (2021).
Muranaka, H. et al. Plasma metabolomics to predict chemotherapy (CTX) response in advanced pancreatic cancer (PC) patients (pts) on enteral feeding for cachexia. J. Clin. Oncol. 40, 600–600 (2022).
Tanigawara, Y., Sugimoto, S. & Muro, K. Pretreatment metabolomic markers associated with therapeutic responses to FOLFOX with bevacizumab in chemotherapy-naive patients with colorectal cancer. J. Clin. Oncol. 37, 540–540 (2019).
Ranjbarvaziri, S. et al. Altered cardiac energetics and mitochondrial dysfunction in hypertrophic cardiomyopathy. Circulation 144, 1714–1731 (2021).
Wang, X. et al. ATF4 protects the heart from failure by antagonizing oxidative stress. Circ. Res. 131, 91–105 (2022).
van Beek, S. M. M. et al. Effect of β2-agonist treatment on insulin-stimulated peripheral glucose disposal in healthy men in a randomised placebo-controlled trial. Nat. Commun. 14, 173 (2023).
Li, L. et al. Hypoxia-induced GBE1 expression promotes tumor progression through metabolic reprogramming in lung adenocarcinoma. Signal Transduct. Target Ther. 5, 54 (2020).
Wei, Z., Oh, J., Flavell, R. A. & Crawford, J. M. LACC1 bridges NOS2 and polyamine metabolism in inflammatory macrophages. Nature 609, 348–353 (2022).
Lim, S. A. et al. Lipid signalling enforces functional specialization of Treg cells in tumours. Nature 591, 306–311 (2021).
Verma, S. et al. NRF2 mediates melanoma addiction to GCDH by modulating apoptotic signalling. Nat. Cell Biol. 24, 1422–1432 (2022).
Liu, Z. & Xiao, T. S. Partners with a killer: Metabolic signaling promotes inflammatory cell death. Cell 184, 4374–4376 (2021).
Qiu, S. et al. Functional metabolomics using UPLC-Q/TOF-MS combined with ingenuity pathway analysis as a promising strategy for evaluating the efficacy and discovering amino acid metabolism as a potential therapeutic mechanism-related target for geniposide against alcoholic liver disease. RSC Adv. 10, 2677–2690 (2020).
Awan, S. et al. Wnt5a promotes lysosomal cholesterol egress and protects against atherosclerosis. Circ. Res. 130, 184–199 (2022).
Zhang, A. H. et al. High-throughput lipidomics analysis to discover lipid biomarkers and profiles as potential targets for evaluating efficacy of Kai-Xin-San against APP/PS1 transgenic mice based on UPLC-Q/TOF-MS. Biomed. Chromatogr. 34, e4724 (2020).
Anglada-Girotto, M. et al. Combining CRISPRi and metabolomics for functional annotation of compound libraries. Nat. Chem. Biol. 18, 482–491 (2022).
Zhang, A. H. et al. High-throughput metabolomics evaluate the efficacy of total lignans from acanthophanax senticosus stem against ovariectomized osteoporosis rat. Front. Pharmacol. 10, 553 (2019).
Schmidt, D. R. et al. Metabolomics in cancer research and emerging applications in clinical oncology. CA Cancer J. Clin. 71, 333–358 (2021).
Zhang, A. H. et al. Chinmedomics: a powerful approach integrating metabolomics with serum pharmacochemistry to evaluate the efficacy of traditional Chinese medicine. Engineering 5, 60–68 (2019).
Baixauli, F. et al. An LKB1-mitochondria axis controls TH17 effector function. Nature 610, 555–561 (2022).
Blacher, E. et al. Potential roles of gut microbiome and metabolites in modulating ALS in mice. Nature 572, 474–480 (2019).
Chen, Y., McAndrews, K. M. & Kalluri, R. Clinical and therapeutic relevance of cancer-associated fibroblasts. Nat. Rev. Clin. Oncol. 18, 792–804 (2021).
Serger, E. et al. The gut metabolite indole-3 propionate promotes nerve regeneration and repair. Nature 607, 585–592 (2022).
Song, X. et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nature 577, 410–415 (2020).
Bauermeister, A., Mannochio-Russo, H., Costa-Lotufo, L. V., Jarmusch, A. K. & Dorrestein, P. C. Mass spectrometry-based metabolomics in microbiome investigations. Nat. Rev. Microbiol. 20, 143–160 (2022).
Hoffmann, M. A. et al. High-confidence structural annotation of metabolites absent from spectral libraries. Nat. Biotechnol. 40, 411–421 (2022).
Jalota, A. et al. Unbiased metabolomic screening reveals pre-existing plasma signatures in large B-cell lymphoma patients treated with anti-CD19 chimeric antigen receptor (CAR) T-cells: association with cytokine release syndrome (CRS) and neurotoxicity (ICANS). Blood 136, 42–43 (2020).
Sato, S. et al. Time of exercise specifies the impact on muscle metabolic pathways and systemic energy homeostasis. Cell Metab. 30, 92–110.e114 (2019).
Yang, H., Lei, T., Li, C., Yu, H. & Chen, Z. Potential metabolites with diagnostic value in plasma for angioimmunoblastic T-cell lymphoma By LC-MS based untargeted metabonomics study. Blood 134, 5234–5234 (2019).
Li, H. et al. Alterations of gut microbiota contribute to the progression of unruptured intracranial aneurysms. Nat. Commun. 11, 3218 (2020).
Baptista, E. et al. P3483 Diet governs metabolic and electrical properties of the atrial myocardium in mice. Eur. Heart J. 40, ehz745.0352 (2019).
Cui, H. et al. Untargeted metabolomics identifies succinate as a biomarker and therapeutic target in aortic aneurysm and dissection. Eur. Heart J. 42, 4373–4385 (2021).
Fu, A. et al. Glucose-dependent partitioning of arginine to the urea cycle protects beta-cells from inflammation. Nat. Metab. 2, 432–446 (2020).
Souto-Carneiro, M. et al. Differences in the serum metabolome and lipidome identify potential biomarkers for seronegative rheumatoid arthritis versus psoriatic arthritis. Ann. Rheum. Dis. 79, 499–506 (2020).
Zhang, A. H. et al. Identifying quality-markers from Shengmai San protects against transgenic mouse model of Alzheimer’s disease using chinmedomics approach. Phytomedicine 45, 84–92 (2018).
Olshan, K. et al. 248 Metagenomic and metabolomic breast milk analysis reflects similar composition in subjects with celiac disease on a gluten-free diet and healthy controls. Gastroenterology 160, S-55 (2021).
Qiu, S. et al. Dissect new mechanistic insights for geniposide efficacy on the hepatoprotection using multiomics approach. Oncotarget 8, 108760–108770 (2017).
Zhang, A. et al. Mass spectrometry-driven drug discovery for development of herbal medicine. Mass Spectrom. Rev. 37, 307–320 (2018).
Sinclair, E. et al. Metabolomics of sebum reveals lipid dysregulation in Parkinson’s disease. Nat. Commun. 12, 1592 (2021).
Coorey, C., Tang, O., Yang, J. Y. H. & Figtree, G. Machine learning analysis of metabolomic biomarkers for diagnosis of heart failure. Eur. Heart J. 42, ehab724.0864 (2021).
Lin, L.-I. et al. Metabolic profiling reveals cellular reprogramming of acute myeloid leukemia by omipalisib through serine synthesis pathway. Blood 138, 3296–3296 (2021).
Perez de Souza, L., Alseekh, S., Scossa, F. & Fernie, A. R. Ultra-high-performance liquid chromatography high-resolution mass spectrometry variants for metabolomics research. Nat. Methods 18, 733–746 (2021).
Shao, Y. & Le, W. Recent advances and perspectives of metabolomics-based investigations in Parkinson’s disease. Mol. Neurodegeneration. 14, 3 (2019).
Li, D. & Gaquerel, E. Next-generation mass spectrometry metabolomics revives the functional analysis of plant metabolic diversity. Annu Rev. Plant Biol. 72, 867–891 (2021).
Dibble, C. C. et al. PI3K drives the de novo synthesis of coenzyme A from vitamin B5. Nature 608, 192–198 (2022).
Pareek, V., Tian, H., Winograd, N. & Benkovic, S. J. Metabolomics and mass spectrometry imaging reveal channeled de novo purine synthesis in cells. Science 368, 283–290 (2020).
Ali, A. et al. Single-cell metabolomics by mass spectrometry: Advances, challenges, and future applications. TrAC Trends Anal. Chem. 120, 115436 (2019).
Li, Y. F. et al. Metabolomic estimation of the diagnosis of hepatocellular carcinoma based on ultrahigh performance liquid chromatography coupled with time-of-flight mass spectrometry. RSC Adv. 8, 9375–9382 (2018).
Li, Y. et al. High-throughput metabolomics to identify metabolites to serve as diagnostic biomarkers of prostate cancer. Anal. Methods 8, 3284–3290 (2016).
Lacalle-Bergeron, L. et al. Chromatography hyphenated to high resolution mass spectrometry in untargeted metabolomics for investigation of food (bio)markers. TrAC Trends Anal. Chem. 135, 116161 (2021).
Zhang, Y. et al. Exploration of metabolite signatures using high-throughput mass spectrometry coupled with multivariate data analysis. RSC Adv. 7, 6780–6787 (2017).
Wang, X. et al. Urine metabolic phenotypes analysis of extrahepatic cholangiocarcinoma disease using ultra-high performance liquid chromatography-mass spectrometry. RSC Adv. 6, 63049–63057 (2016).
Liang, Q. et al. High-throughput metabolomics analysis discovers salivary biomarkers for predicting mild cognitive impairment and Alzheimer’s disease. RSC Adv. 6, 75499–75504 (2016).
Zheng, F. et al. Development of a plasma pseudotargeted metabolomics method based on ultra-high-performance liquid chromatography-mass spectrometry. Nat. Protoc. 15, 2519–2537 (2020).
Liang, Q. et al. Discovery of serum metabolites for diagnosis of progression of mild cognitive impairment to Alzheimer’s disease using an optimized metabolomics method. RSC Adv. 6, 3586–3591 (2016).
Ramautar, R., Somsen, G. W. & de Jong, G. J. CE-MS for metabolomics: Developments and applications in the period 2016-2018. Electrophoresis 40, 165–179 (2019).
Jang, C., Chen, L. & Rabinowitz, J. D. Metabolomics and isotope tracing. Cell 173, 822–837 (2018).
Cui, L., Lu, H. & Lee, Y. H. Challenges and emergent solutions for LC-MS/MS based untargeted metabolomics in diseases. Mass Spectrom. Rev. 37, 772–792 (2018).
Alseekh, S. et al. Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices. Nat. Methods 18, 747–756 (2021).
Wawrzyniak, R. et al. Untargeted metabolomics towards understanding molecular mechanisms of pulmonary arterial hypertension. Eur. Heart J. 42, ehab724.3421 (2021).
Stevens, B. M. et al. Unique metabolic vulnerabilities of myelodysplastic syndrome stem cells. Blood 138, 1511–1511 (2021).
Chen, Z. et al. Effects of ibrutinib on metabolic alterations and micro-environmental signalling in chronic lymphocytic leukaemia. Blood 136, 36–37 (2020).
Tian, H. et al. Precise metabolomics reveals a diversity of aging-associated metabolic features. Small Methods 6, e2200130 (2022).
Martin, J. K. 2nd et al. A dual-mechanism antibiotic kills gram-negative bacteria and avoids drug resistance. Cell 181, 1518–1532.e1514 (2020).
Funk, M. & Funk, M. A. Signs of a metabolon in action. Science 368, 278.210-280 (2020).
Capolupo, L. et al. Sphingolipids control dermal fibroblast heterogeneity. Science 376, eabh1623 (2022).
Forsberg, E. M. et al. Data processing, multi-omic pathway mapping, and metabolite activity analysis using XCMS Online. Nat. Protoc. 13, 633–651 (2018).
Eveque, M. et al. OP0240 A multimodal mass spectrometry approach reveals specific cartilage molecular profiles associated to type 2 diabetic patients. Ann. Rheum. Dis. 79, 151–152 (2020).
Prag, H. A. et al. Ischemia-selective cardioprotection by malonate for ischemia/reperfusion injury. Circ. Res. 131, 528–541 (2022).
Zhang, A. et al. Modern analytical techniques in metabolomics analysis. Analyst 137, 293–300 (2012).
Goossens, P. et al. Integrating multiplex immunofluorescent and mass spectrometry imaging to map myeloid heterogeneity in its metabolic and cellular context. Cell Metab. 34, 1214–1225.e1216 (2022).
Garg, H. et al. Role of matrix assisted laser desorption/ionization (MALDI)- mass spectrometry imaging (MSI): A novel tool to study bioenergetic signature in kidney cancer. Eur. Urol. 81, S1654–S1655 (2022).
Liang, Q. et al. Metabolomics-based screening of salivary biomarkers for early diagnosis of Alzheimer’s disease. RSC Adv. 5, 96074–96079 (2015).
Fan, Z. et al. Exercise-induced angiogenesis is dependent on metabolically primed ATF3/4(+) endothelial cells. Cell Metab. 33, 1793–1807.e1799 (2021).
Huang, C. et al. Spatial-temporal lipidomics profile of acute myocardial injury. Eur. Heart J. 43, ehac544.2919 (2022).
Alexandrov, T. Probing metabolism in time and space. Science 368, 241–242 (2020).
Gouw, A. M. et al. The MYC oncogene cooperates with sterol-regulated element-binding protein to regulate lipogenesis essential for neoplastic growth. Cell Metab. 30, 556–572.e555 (2019).
Paizs, P. et al. Mo1076 spatially resolved analysis of faecal metabolites in gastrointestinal health and disease for biomarker identification using optimised laser assisted - rapid evaporative ionization - mass spectrometry imaging(LA-REI-MSI). Gastroenterology 158, S-780-S-781 (2020).
Borodinov, N., Lorenz, M., King, S. T., Ievlev, A. V. & Ovchinnikova, O. S. Toward nanoscale molecular mass spectrometry imaging via physically constrained machine learning on co-registered multimodal data. npj Comput. Mater. 6, 83 (2020).
Manifold, B., Men, S., Hu, R. & Fu, D. A versatile deep learning architecture for classification and label-free prediction of hyperspectral images. Nat. Mach. Intell. 3, 306–315 (2021).
Ghallab, A. et al. Interruption of bile acid uptake by hepatocytes after acetaminophen overdose ameliorates hepatotoxicity. J. Hepatol. 77, 71–83 (2022).
Wang, G. et al. Analyzing cell-type-specific dynamics of metabolism in kidney repair. Nat. Metab. 4, 1109–1118 (2022).
Philipsen, M. H., Ranjbari, E., Gu, C. & Ewing, A. G. Mass spectrometry imaging shows modafinil, a student study drug, changes the lipid composition of the fly brain. Angew. Chem. Int Ed. Engl. 60, 17378–17382 (2021).
Gregoire, S. et al. Imaging and quantifying drug delivery in skin - Part 1: Autoradiography and mass spectrometry imaging. Adv. Drug Deliv. Rev. 153, 137–146 (2020).
Randall, D. W. et al. Batch effect exerts a bigger influence on the rat urinary metabolome and gut microbiota than uraemia: a cautionary tale. Microbiome 7, 127 (2019).
Gisewhite, S., Stewart, I. J., Beilman, G. & Lusczek, E. Urinary metabolites predict mortality or need for renal replacement therapy after combat injury. Crit. Care. 25, 119 (2021).
Boguszewicz, L. et al. 885P Molecular response to induction chemotherapy and its correlation with treatment outcome in head and neck cancer patients by means of NMR-based metabolomics. Ann. Oncol. 32, S796 (2021).
Tzoulaki, I. et al. Serum metabolic signatures of coronary and carotid atherosclerosis and subsequent cardiovascular disease. Eur. Heart J. 40, 2883–2896 (2019).
Hong, C. et al. Application of machine learning to identify top determinants of fibrofatty plaque burden by CCTA in humans with psoriasis. Eur. Heart J. 43, ehac544.213 (2022).
Cediel Calderon, G. et al. Clinical and prognostic significance of the inflammatory markers GlycA and GlycB in chronic heart failure of both ischemic and non-ischemic etiologies. Eur. Heart J. 42, ehab724.0870 (2021).
Trushin, S., Stojakovic, A., Chang, S. Y. & Trushina, E. Partial mitochondrial complex I inhibitors as disease‐modifying therapeutics for Alzheimer’s disease. Alzheimer’s. Dement. 16, e045529 (2020).
Trujillo-Estrada, L. et al. P4-522: Type 2 diabetes mellitus induces tau-independent cognitive and synaptic deficits in a mouse model. Alzheimer’s Dement. 15, P1514–P1514 (2019).
Duan, P. et al. Binding sites of a positron emission tomography imaging agent in Alzheimer’s beta-amyloid fibrils studied using (19)F solid-state NMR. J. Am. Chem. Soc. 144, 1416–1430 (2022).
Yang, T. L. et al. A road map for understanding molecular and genetic determinants of osteoporosis. Nat. Rev. Endocrinol. 16, 91–103 (2020).
Xuan, Q. et al. Multiplatform metabolomics reveals novel serum metabolite biomarkers in diabetic retinopathy subjects. Adv. Sci. (Weinh.). 7, 2001714 (2020).
Garcia-Perez, I. et al. Identifying unknown metabolites using NMR-based metabolic profiling techniques. Nat. Protoc. 15, 2538–2567 (2020).
Tillmann, T. Atherosclerotic metabolites: basic science is progressing, so we need to think about clinical implications. Eur. Heart J. 40, 2897–2898 (2019).
Galan-Diez, M. et al. A targetable bone marrow-niche axis for the treatment of acute myeloid leukemia. Blood 138, 4456–4456 (2021).
Yokota, A. et al. Myelodysplastic syndromes-associated gene mutations lead to pseudohypoxia condition and epigenome hyper-methylation in mouse genetic models. Blood 134, 1696–1696 (2019).
Quintero, M., Montalvão, S. A. D. L., Tasic, L., Huber, S. C. & Annichino- Bizzacchi, J. M. Comparison of the serum metabolic signatures based on 1 H NMR between thrombotic antiphospholipid syndrome (APS) patients and healthy individuals. Blood 134, 5769–5769 (2019).
Marx, D. et al. POS0472 Comparative metabolomic analysis of serum samples from patients with coincidental rheumatological and malignant diseases. Ann. Rheum. Dis. 80, 467–468 (2021).
Manolakou, T. et al. POS0421 combined analysis of metabolic and transcriptomic kidney profiles of NZW/B-F1 murine lupus uncovers biological mechanisms preceding the onset of nephritis. Ann. Rheum. Dis. 80, 439–440 (2021).
Donato, S. D. et al. 574P A metabolomic recurrence score for risk-stratification of elderly patients (pts) with early colorectal cancer (eCRC). Ann. Oncol. 30, v217 (2019).
Bruzzone, C. et al. Unravelling the molecular determinants of metabolic syndrome thanks to NMR-metabolomics of urine and serum samples. J. Hepatol. 73, S288–S289 (2020).
Lin, W., Conway, L. P., Vujasinovic, M., Löhr, J. M. & Globisch, D. Chemoselective andhighly sensitive quantification of gut microbiome and human metabolites. Angew. Chem. Int Ed. Engl. 60, 23232–23240 (2021).
Zhang, D. et al. Integrated metabolomics revealed the fibromyalgia-alleviation effect of Mo(2)C nanozyme through regulated homeostasis of oxidative stress and energy metabolism. Biomaterials 287, 121678 (2022).
Zhang, Q. et al. Integrated multiomic analysis reveals comprehensive tumour heterogeneity and novel immunophenotypic classification in hepatocellular carcinomas. Gut 68, 2019–2031 (2019).
Zang, X., Monge, M. E. & Fernández, F. M. Mass spectrometry-based non-targeted metabolic profiling for disease detection: recent developments. Trends Anal. Chem. 118, 158–169 (2019).
Mulder, F. A. A., Tenori, L. & Luchinat, C. Fast and quantitative NMR metabolite analysis afforded by a paramagnetic co-solute. Angew. Chem. Int Ed. Engl. 58, 15283–15286 (2019).
Yuan, H. et al. Development of a widely targeted volatilomics method for profiling volatilomes in plants. Mol. Plant. 15, 189–202 (2022).
Liu, J. et al. Integrative metabolomic characterisation identifies altered portal vein serum metabolome contributing to human hepatocellular carcinoma. Gut 71, 1203–1213 (2022).
Santos-Gallego, C. G., Mayr, M. & Badimon, J. SGLT2 inhibitors in heart failure: targeted metabolomics and energetic metabolism. Circulation 146, 819–821 (2022).
Madapoosi, S. S. et al. Lung microbiota and metabolites collectively associate with clinical outcomes in milder stage chronic obstructive pulmonary disease. Am. J. Respir. Crit. Care Med. 206, 427–439 (2022).
Leiherer, A. et al. Targeted metabolomics identifies elevated serotonin levels in carriers of a TCF7L2 diabetes-risk allele. J. Am. Coll. Cardiol. 73, 2119 (2019).
Colaco, K. et al. Targeted metabolomic profiling and prediction of cardiovascular events: a prospective study of patients with psoriatic arthritis and psoriasis. Ann. Rheum. Dis. 80, 1429–1435 (2021).
Belghasem, M. et al. Metabolites in a mouse cancer model enhance venous thrombogenicity through the aryl hydrocarbon receptor-tissue factor axis. Blood 134, 2399–2413 (2019).
Rosenberger, G. et al. Statistical control of peptide and protein error rates in large-scale targeted data-independent acquisition analyses. Nat. Methods 14, 921–927 (2017).
Colaco, K. et al. OP0221 Targeted metabolomic profiling and prediction of cardiovascular events: A prospective study of patients with psoriatic arthritis and psoriasis. Ann. Rheum. Dis. 80, 132–133 (2021).
Hourmozdi, J. N. et al. Plasma metabolite profiles are associated with right ventricular dysfunction and prognosis in pulmonary arterial hypertension. J. Am. Coll. Cardiol. 75, 2082 (2020).
Ewald, J. C., Kuehne, A., Zamboni, N. & Skotheim, J. M. The yeast cyclin-dependent kinase routes carbon fluxes to fuel cell cycle progression. Mol. Cell. 62, 532–545 (2016).
Dührkop, K. et al. Systematic classification of unknown metabolites using high-resolution fragmentation mass spectra. Nat. Biotechnol. 39, 462–471 (2020).
Mitchell, S. J. et al. Nicotinamide improves aspects of healthspan, but not lifespan, in mice. Cell Metab. 27, 667–676.e664 (2018).
Zhao, H. et al. Paraben exposure related to purine metabolism and other pathways revealed by mass spectrometry-based metabolomics. Environ. Sci. Technol. 54, 3447–3454 (2020).
Hoki, J. S. et al. Deep interrogation of metabolism using a pathway-targeted click-chemistry approach. J. Am. Chem. Soc. 142, 18449–18459 (2020).
Chen, D. Q. et al. Identification of serum metabolites associating with chronic kidney disease progression and anti-fibrotic effect of 5-methoxytryptophan. Nat. Commun. 10, 1476 (2019).
Chen, C. et al. Metabolomic profiling reveals amino acid and carnitine alterations as metabolic signatures in psoriasis. Theranostics 11, 754–767 (2021).
Taverna, F. et al. BIOMEX: an interactive workflow for (single cell) omics data interpretation and visualization. Nucleic Acids Res. 48, W385–W394 (2020).
Marx, V. Boost that metabolomic confidence. Nat. Methods 17, 33–36 (2020).
Hendrickx, J. O., van Gastel, J., Leysen, H., Martin, B. & Maudsley, S. High-dimensionality data analysis of pharmacological systems associated with complex diseases. Pharm. Rev. 72, 191–217 (2020).
Sun, F. et al. An integrated data-dependent and data-independent acquisition method for hazardous compounds screening in foods using a single UHPLC-Q-Orbitrap run. J. Hazard Mater. 401, 123266 (2021).
Schorn, M. A. et al. A community resource for paired genomic and metabolomic data mining. Nat. Chem. Biol. 17, 363–368 (2021).
Zhang, D. et al. Exploring the biological effect of biosynthesized Au-Pd core-shell nanoparticles through an untargeted metabolomics approach. ACS Appl Mater. Interfaces 13, 59633–59648 (2021).
Azad, R. K. & Shulaev, V. Metabolomics technology and bioinformatics for precision medicine. Brief. Bioinform. 20, 1957–1971 (2019).
Ma, X. et al. Bioinformatics-assisted, integrated omics studies on medicinal plants. Brief. Bioinform. 21, 1857–1874 (2020).
Li, S. et al. Recent advances in the application of metabolomics for food safety control and food quality analyses. Crit. Rev. Food Sci. Nutr. 61, 1448–1469 (2021).
Keane, T. M., O'Donovan, C. & Vizcaino, J. A. The growing need for controlled data access models in clinical proteomics and metabolomics. Nat. Commun. 12, 5787 (2021).
Li, L. et al. An alignment algorithm for LC-MS-based metabolomics dataset assisted by MS/MS information. Anal. Chim. Acta 990, 96–102 (2017).
Mathema, V. B. et al. CRISP: a deep learning architecture for GC × GC–TOFMS contour ROI identification, simulation and analysis in imaging metabolomics. Brief. Bioinforma. 23, bbab550 (2021).
Notararigo, S., Martin-Pastor, M., Dominguez Munoz, JE. & Barreiro-de Acosta, M.P052 Nuclear magnetic resonance metabolomic profiling of IBD patients under anti-TNF treatment. Are the pathways network deregulated?. J. Crohn’s. Colitis 14, S160–S160 (2020).
Krivitsky, V. et al. Cellular metabolomics by a universal redox-reactive nanosensors array: from the cell level to tumor-on-a-chip analysis. Nano Lett. 19, 2478–2488 (2019).
Olivera, P., Danese, S., Jay, N., Natoli, G. & Peyrin-Biroulet, L. Big data in IBD: a look into the future. Nat. Rev. Gastroenterol. Hepatol. 16, 312–321 (2019).
Stancliffe, E., Schwaiger-Haber, M., Sindelar, M. & Patti, G. J. DecoID improves identification rates in metabolomics through database-assisted MS/MS deconvolution. Nat. Methods 18, 779–787 (2021).
Arif, M. et al. iNetModels 2.0: an interactive visualization and database of multi-omics data. Nucleic Acids Res. 49, W271–W276 (2021).
Wishart, D. S. et al. PathBank: a comprehensive pathway database for model organisms. Nucleic Acids Res. 48, D470–D478 (2020).
Zhang, R. Z. et al. Metabolomics-based comparative analysis of the effects of host and environment on Viscum coloratum metabolites and antioxidative activities. J. Pharm. Anal. 12, 243–252 (2022).
Subburaj, D. et al. Metabolomic identification of alpha-ketoglutaric acid elevation in pediatric chronic graft-versus-host disease. Blood 139, 287–299 (2022).
Jun, G., Aguilar, D., Evans, C., Burant, C. F. & Hanis, C. L. Metabolomic profiles associated with subtypes of prediabetes among Mexican Americans in Starr County, Texas, USA. Diabetologia 63, 287–295 (2020).
Eiden, M. et al. Discovery and validation of temporal patterns involved in human brain ketometabolism in cerebral microdialysis fluids of traumatic brain injury patients. EBioMedicine 44, 607–617 (2019).
Duan, Y., Sun, H., Yao, Y., Han, L. & Chen, L. Perturbation of serum metabolome in relation to type 2 diabetes mellitus and urinary levels of phthalate metabolites and bisphenols. Environ. Int. 155, 106609 (2021).
Hsu, J. F. et al. Using a high-resolution mass spectrometry-based metabolomics strategy for comprehensively screening and identifying biomarkers of phthalate exposure: Method development and application. Environ. Int. 128, 261–270 (2019).
Perez-Riverol, Y. et al. Quantifying the impact of public omics data. Nat. Commun. 10, 3512 (2019).
Rattray, N. J. W. et al. Metabolic dysregulation in vitamin E and carnitine shuttle energy mechanisms associate with human frailty. Nat. Commun. 10, 5027 (2019).
Xu, K. et al. Toxicity of three carbon-based nanomaterials to earthworms: Effect of morphology on biomarkers, cytotoxicity, and metabolomics. Sci. Total Environ. 777, 146224 (2021).
Lee, E., Kim, D. J., Cho, J.-Y., & Jung, K.-h.Abstract WMP114: putrescine and kynurenine are associated with large artery atherosclerosis stroke: targeted metabolomics study. Stroke 53, AWMP114 (2022).
Feizi, N., Hashemi-Nasab, F. S., Golpelichi, F., Saburouh, N. & Parastar, H. Recent trends in application of chemometric methods for GC-MS and GC×GC-MS-based metabolomic studies. TrAC Trends Anal. Chem. 138, 116239 (2021).
De Oliveira, M., Alabarse, P. V., Farinon, M., Cavalheiro Do Espírito Santo, R. & Xavier, R. AB0185 Prospective profile of urine metabolome in rheumatoid arthritis. Ann. Rheum. Dis. 79, 1392 (2020).
Wilinski, D. et al. Rapid metabolic shifts occur during the transition between hunger and satiety in Drosophila melanogaster. Nat. Commun. 10, 4052 (2019).
Li, M. et al. Core functional nodes and sex-specific pathways in human ischaemic and dilated cardiomyopathy. Nat. Commun. 11, 2843 (2020).
Hollenberg, A. M., Smith, C. O., Shum, L. C., Awad, H. & Eliseev, R. A. Lactate dehydrogenase inhibition with oxamate exerts bone anabolic effect. J. Bone Min. Res. 35, 2432–2443 (2020).
Zhang, L. et al. Inhibition of UBA6 by inosine augments tumour immunogenicity and responses. Nat. Commun. 13, 5413 (2022).
Sun, Y. et al. Noninvasive urinary protein signatures associated with colorectal cancer diagnosis and metastasis. Nat. Commun. 13, 2757 (2022).
Huang, X. et al. LINC00842 inactivates transcription co-regulator PGC-1α to promote pancreatic cancer malignancy through metabolic remodelling. Nat. Commun. 12, 3830 (2021).
Tripathi, A. et al. Chemically informed analyses of metabolomics mass spectrometry data with Qemistree. Nat. Chem. Biol. 17, 146–151 (2021).
Schuijs, M. J. et al. ILC2-driven innate immune checkpoint mechanism antagonizes NK cell antimetastatic function in the lung. Nat. Immunol. 21, 998–1009 (2020).
Hagenbeek, F. A. et al. Heritability estimates for 361 blood metabolites across 40 genome-wide association studies. Nat. Commun. 11, 39 (2020).
Barylyuk, K. et al. A comprehensive subcellular atlas of the toxoplasma proteome via hyperLOPIT provides spatial context for protein functions. Cell Host Microbe 28, 752–766.e759 (2020).
Bensard, C. L. et al. Regulation of tumor initiation by the mitochondrial pyruvate carrier. Cell Metab. 31, 284–300.e287 (2020).
Xiao, N. et al. Integrated cytokine and metabolite analysis reveals immunometabolic reprogramming in COVID-19 patients with therapeutic implications. Nat. Commun. 12, 1618 (2021).
Ung, C. Y. et al. Regulostat Inferelator: a novel network biology platform to uncover molecular devices that predetermine cellular response phenotypes. Nucleic Acids Res. 47, e82 (2019).
Woodcock, D. J. et al. Prostate cancer evolution from multilineage primary to single lineage metastases with implications for liquid biopsy. Nat. Commun. 11, 5070 (2020).
Habtetsion, T. et al. Alteration of tumor metabolism by CD4+ T cells leads to TNF-α-dependent intensification of oxidative stress and tumor cell death. Cell Metab. 28, 228–242.e226 (2018).
Gordin, D. et al. Characterization of glycolytic enzymes and pyruvate kinase M2 in type 1 and 2 diabetic nephropathy. Diabetes Care. 42, 1263–1273 (2019).
Villa, E. et al. mTORC1 stimulates cell growth through SAM synthesis and m(6)A mRNA-dependent control of protein synthesis. Mol. Cell. 81, 2076–2093.e2079 (2021).
Whitehead, A. et al. Brown and beige adipose tissue regulate systemic metabolism through a metabolite interorgan signaling axis. Nat. Commun. 12, 1905 (2021).
Huang, X., Gan, G., Wang, X., Xu, T. & Xie, W. The HGF-MET axis coordinates liver cancer metabolism and autophagy for chemotherapeutic resistance. Autophagy 15, 1258–1279 (2019).
Polyzos, A. A. et al. Metabolic reprogramming in astrocytes distinguishes region-specific neuronal susceptibility in Huntington Mice. Cell Metab. 29, 1258–1273.e1211 (2019).
Kidiyoor, G. R. et al. ATR is essential for preservation of cell mechanics and nuclear integrity during interstitial migration. Nat. Commun. 11, 4828 (2020).
Huang, L. et al. Machine learning of serum metabolic patterns encodes early-stage lung adenocarcinoma. Nat. Commun. 11, 3556 (2020).
Yadav, A. K., Carroll, A. J., Estavillo, G. M., Rebetzke, G. J. & Pogson, B. J. Wheat drought tolerance in the field is predicted by amino acid responses to glasshouse-imposed drought. J. Exp. Bot. 70, 4931–4948 (2019).
Li, L. Y. et al. Alterations of gut microbiota diversity, composition and metabonomics in testosterone-induced benign prostatic hyperplasia rats. Mil. Med Res. 9, 12 (2022).
Bengel, P. et al. Metabolic modulation as a common adaptive mechanism in patients with different subtypes of aortic valve stenosis. Eur. Heart J. 43, ehac544.2940 (2022).
Zhu, Y. et al. Integrative proteomics and metabolomics approach to elucidate metabolic dysfunction induced by silica nanoparticles in hepatocytes. J. Hazard Mater. 434, 128820 (2022).
Shahid, N., Rolle-Kampczyk, U., Siddique, A., von Bergen, M. & Liess, M. Pesticide-induced metabolic changes are amplified by food stress. Sci. Total Environ. 792, 148350 (2021).
Gao, P. et al. Peroxisome proliferator-activated receptor gamma (PPARγ) activation and metabolism disturbance induced by bisphenol A and its replacement analog bisphenol S using in vitro macrophages and in vivo mouse models. Environ. Int. 134, 105328 (2020).
Tan, A. H. et al. Gut microbial ecosystem in parkinson disease: new clinicobiological insights from multi-omics. Ann. Neurol. 89, 546–559 (2021).
Shao, Y. et al. Comprehensive metabolic profiling of Parkinson’s disease by liquid chromatography-mass spectrometry. Mol. Neurodegener. 16, 4 (2021).
Radford-Smith, D. et al. P198 An inflammatory serum metabolomic signature predicts response to vedolizumab treatment in people with Crohn’s Disease. J. Crohn’s. Colitis 16, i257–i259 (2022).
Li, J.-X. et al. Untargeted metabolomic profiling identifies disease-specific and outcome-related signatures in chronic rhinosinusitis. J. Allergy Clin. Immunol. 150, 727–735.e6 (2022).
Lewis, J. E. & Kemp, M. L. Integration of machine learning and genome-scale metabolic modeling identifies multi-omics biomarkers for radiation resistance. Nat. Commun. 12, 2700 (2021).
Qiu, S. et al. Current status of technical challenges in mass spectrometry-driven metabolomics. In Mass spectrometry-based metabolomics in clinical and herbal medicines (eds A. Zhang and W. Wang). https://doi.org/10.1002/9783527835751.ch7 (2021).
Ke, M. et al. CAR-T therapy alters synthesis of platelet-activating factor in multiple myeloma patients. J. Hematol. Oncol. 14, 90 (2021).
Bell, J. A. et al. Early metabolic features of genetic liability to type 2 diabetes: cohort study with repeated metabolomics across early life. Diabetes Care. 43, 1537–1545 (2020).
Zhao, Q., Wu, Z. E., Li, B. & Li, F. Recent advances in metabolism and toxicity of tyrosine kinase inhibitors. Pharm. Ther. 237, 108256 (2022).
Wang, X. et al. Metabolic tuning of inhibition regulates hippocampal neurogenesis in the adult brain. Proc. Natl. Acad. Sci. USA 117, 25818–25829 (2020).
Schonberger, K. et al. Multilayer omics analysis reveals a non-classical retinoic acid signaling axis that regulates hematopoietic stem cell identity. Cell. Stem Cell. 29, 131–148.e110 (2022).
Mortazavi, A. et al. IDH-mutated gliomas promote epileptogenesis through d-2-hydroxyglutarate-dependent mTOR hyperactivation. Neuro Oncol. 24, 1423–1435 (2022).
Mohamed Amin Mostafa, A., Mostafa, H., Sk Abdul Kader, M. A. & Kah Hay, Y. P258 Pharmacometabolomics analysis of plasma and urine to identify clopidogrel exposure metabolic biomarkers. Eur. Heart J. 41, ehz872.084 (2020).
Luo, P. et al. Metabolic characteristics of large and small extracellular vesicles from pleural effusion reveal biomarker candidates for the diagnosis of tuberculosis and malignancy. J. Extracell. Vesicles. 9, 1790158 (2020).
Khan, A. et al. High-resolution metabolomics study revealing l-homocysteine sulfinic acid, cysteic acid, and carnitine as novel biomarkers for high acute myocardial infarction risk. Metabolism 104, 154051 (2020).
Zhang, A. et al. Metabolomics toward precision medicine. in mass spectrometry-based metabolomics in clinical and herbal medicines (eds A. Zhang and W. Wang). https://doi.org/10.1002/9783527835751.ch11 (2021).
Brierley, D. I. et al. Chemotherapy-induced cachexia dysregulates hypothalamic and systemic lipoamines and is attenuated by cannabigerol. J. Cachexia Sarcopenia Muscle 10, 844–859 (2019).
Annunziato, M. et al. An integrated systems-level model of the toxicity of brevetoxin based on high-resolution magic-angle spinning nuclear magnetic resonance (HRMAS NMR) metabolic profiling of zebrafish embryos. Sci. Total Environ. 803, 149858 (2022).
Qiu S. et al. Mass spectrometry-based metabolomics toward biological function analysis. in mass spectrometry-based metabolomics in clinical and herbal medicines (eds A. Zhang and W. Wang). https://doi.org/10.1002/9783527835751.ch12 (2021).
Zhang, X. et al. Plasma metabolomic profiles of dementia: a prospective study of 110,655 participants in the UK Biobank. BMC Med. 20, 252 (2022).
Sindelar, M. et al. Longitudinal metabolomics of human plasma reveals prognostic markers of COVID-19 disease severity. Cell Rep. Med. 2, 100369 (2021).
Schult, T. A. et al. Screening human lung cancer with predictive models of serum magnetic resonance spectroscopy metabolomics. Proc. Natl. Acad. Sci. USA 118, e2110633118 (2021).
Salvador-Coloma, C. et al. Immunosuppressive profiles in liquid biopsy at diagnosis predict response to neoadjuvant chemotherapy in triple-negative breast cancer. Eur. J. Cancer 139, 119–134 (2020).
McBride, N. et al. Do nuclear magnetic resonance (NMR)-based metabolomics improve the prediction of pregnancy-related disorders? Findings from a UK birth cohort with independent validation. BMC Med. 18, 366 (2020).
Cui, G. Y. et al. Characterization of oral and gut microbiome and plasma metabolomics in COVID-19 patients after 1-year follow-up. Mil. Med Res. 9, 32 (2022).
Adegbola, S. et al. P077 Metabonomic profiling distinguishes Crohn’s perianal fistulas and idiopathic idiopathic (cryptoglandular) perianal fistulas: possible clues to underlying pathogenesis? J. Crohn’s. Colitis 14, S174–S174 (2020).
Sen, P. et al. Deep learning meets metabolomics: a methodological perspective. Brief. Bioinform. 22, 1531–1542 (2021).
Ozcelikay, G. et al. Sensor-based MIP technologies for targeted metabolomics analysis. TrAC Trends Anal. Chem. 146, 116487 (2022).
Pruski, P. et al. Direct on-swab metabolic profiling of vaginal microbiome host interactions during pregnancy and preterm birth. Nat. Commun. 12, 5967 (2021).
Hansen, N. L. et al. Tripterygium wilfordii cytochrome P450s catalyze the methyl shift and epoxidations in the biosynthesis of triptonide. Nat. Commun. 13, 5011 (2022).
Riva, A. et al. A fiber-deprived diet disturbs the fine-scale spatial architecture of the murine colon microbiome. Nat. Commun. 10, 4366 (2019).
Helf, M. J., Fox, B. W., Artyukhin, A. B., Zhang, Y. K. & Schroeder, F. C. Comparative metabolomics with Metaboseek reveals functions of a conserved fat metabolism pathway in C. elegans. Nat. Commun. 13, 782 (2022).
Nothias, L. F. et al. Feature-based molecular networking in the GNPS analysis environment. Nat. Methods 17, 905–908 (2020).
Lai, Z. et al. Identifying metabolites by integrating metabolome databases with mass spectrometry cheminformatics. Nat. Methods 15, 53–56 (2018).
Schmid, R. et al. Ion identity molecular networking for mass spectrometry-based metabolomics in the GNPS environment. Nat. Commun. 12, 3832 (2021).
Darnaud, M. et al. A standardized gnotobiotic mouse model harboring a minimal 15-member mouse gut microbiota recapitulates SOPF/SPF phenotypes. Nat. Commun. 12, 6686 (2021).
Feist, M. et al. Cooperative STAT/NF-κB signaling regulates lymphoma metabolic reprogramming and aberrant GOT2 expression. Nat. Commun. 9, 1514 (2018).
Traube, F. R. et al. Redirected nuclear glutamate dehydrogenase supplies Tet3 with α-ketoglutarate in neurons. Nat. Commun. 12, 4100 (2021).
Yakulov, T. A. et al. CXCL12 and MYC control energy metabolism to support adaptive responses after kidney injury. Nat. Commun. 9, 3660 (2018).
Tadaka, S. et al. jMorp updates in 2020: large enhancement of multi-omics data resources on the general Japanese population. Nucleic Acids Res. 49, D536–d544 (2021).
Giacomelli, E. et al. Human-iPSC-derived cardiac stromal cells enhance maturation in 3D cardiac microtissues and reveal non-cardiomyocyte contributions to heart disease. Cell. Stem Cell. 26, 862–879.e811 (2020).
Wolf, A. R. et al. Bioremediation of a common product of food processing by a human gut bacterium. Cell Host Microbe 26, 463–477.e468 (2019).
Guasch-Ferré, M. et al. Metabolomics in prediabetes and diabetes: a systematic review and meta-analysis. Diabetes Care. 39, 833–846 (2016).
Pang, Z. et al. Using MetaboAnalyst 5.0 for LC-HRMS spectra processing, multi-omics integration and covariate adjustment of global metabolomics data. Nat. Protoc. 17, 1735–1761 (2022).
Dogan, H. O. et al. Understanding the pathophysiological changes via untargeted metabolomics in COVID-19 patients. J. Med Virol. 93, 2340–2349 (2021).
Hu, J. et al. Metabonomic and transcriptomic modulations of HepG2 cells induced by the CuO-catalyzed formation of disinfection byproducts from biofilm extracellular polymeric substances in copper pipes. Water Res. 216, 118318 (2022).
Diab, J. et al. DOP15 Metabolomics coupled with pathway analysis characterise metabolic changes in treatment-naive ulcerative colitis patients. J. Crohn’s Colitis 13, S035–S035 (2019).
Alferink, L. J. M. et al. Microbiomics, metabolomics, predicted metagenomics, and hepatic steatosis in a population-based study of 1,355 adults. Hepatology 73, 968–982 (2021).
Swietlik, E. M. et al. Plasma metabolomics exhibit response to therapy in chronic thromboembolic pulmonary hypertension. Eur. Respir. J. 57, 2003201 (2021).
Mora-Ortiz, M. et al. Metabolomics analysis of type 2 diabetes remission identifies 12 metabolites with predictive capacity: a CORDIOPREV clinical trial study. BMC Med. 20, 373 (2022).
Liu, Z. et al. Nuclear magnetic resonance-based serum metabolomic analysis reveals different disease evolution profiles between septic shock survivors and non-survivors. Crit. Care. 23, 169 (2019).
Larkin, J. R. et al. Metabolomic biomarkers in blood samples identify cancers in a mixed population of patients with nonspecific symptoms. Clin. Cancer Res. 28, 1651–1661 (2022).
McDonald, V. M. & Gibson, P. G. Treatable traits in asthma: moving beyond diagnostic labels. Med. J. Aust. 216, 331–333 (2022).
Massey, V. et al. Integrated multiomics reveals glucose use reprogramming and identifies a novel hexokinase in alcoholic hepatitis. Gastroenterology 160, 1725–1740.e1722 (2021).
Li, W. et al. Multi-omics research strategies in ischemic stroke: A multidimensional perspective. Ageing Res. Rev. 81, 101730 (2022).
Denburg, M. R. et al. Metabolite biomarkers of CKD progression in children. Clin. J. Am. Soc. Nephrol. 16, 1178–1189 (2021).
Yan, Y. et al. Plasma metabolomics in perioperative period of defect repair in patients with pulmonary arterial hypertension associated with congenital heart disease. Eur. Heart J. 42, ehab724.1868 (2021).
Tomita, Y. et al. Vitreous metabolomics profiling of proliferative diabetic retinopathy. Diabetologia 64, 70–82 (2021).
Tateishi, H. et al. Changes in the metabolites of cerebrospinal fluid induced by rTMS in treatment-resistant depression: A pilot study. Psychiatry Res. 313, 114636 (2022).
Ng, S. S. W. et al. Plasma metabolomic profiles in liver cancer patients following stereotactic body radiotherapy. EBioMedicine 59, 102973 (2020).
Fraunhoffer, N. A. et al. Multi-omics data integration and modeling unravels new mechanisms for pancreatic cancer and improves prognostic prediction. NPJ Precis Oncol. 6, 57 (2022).
Wang, Y. et al. An FGF15/19-TFEB regulatory loop controls hepatic cholesterol and bile acid homeostasis. Nat. Commun. 11, 3612 (2020).
Creswell, R. et al. High-resolution temporal profiling of the human gut microbiome reveals consistent and cascading alterations in response to dietary glycans. Genome Med. 12, 59 (2020).
Barupal, D. K. et al. CCDB: A database for exploring inter-chemical correlations in metabolomics and exposomics datasets. Environ. Int. 164, 107240 (2022).
Wu, Q. et al. Multi-stage metabolomics and genetic analyses identified metabolite biomarkers of metabolic syndrome and their genetic determinants. EBioMedicine 74, 103707 (2021).
Talmor-Barkan, Y. et al. Metabolomic and microbiome profiling reveals personalized risk factors for coronary artery disease. Nat. Med. 28, 295–302 (2022).
Lee, A. M. et al. Using machine learning to identify metabolomic signatures of pediatric chronic kidney disease etiology. J. Am. Soc. Nephrol. 33, 375–386 (2022).
Horgusluoglu‐Moloch, E. et al. Integrative metabolomics‐genomics approach reveals that pathways related to the metabolism of acylcarnitines and amines are new potential targets of Alzheimer’s disease. Alzheimer’s. Dement. 16, e045610 (2020).
Choi, S. C. et al. Gut microbiota dysbiosis and altered tryptophan catabolism contribute to autoimmunity in lupus-susceptible mice. Sci. Transl. Med. 12, eaax2220 (2020).
Park, J. & Kim, C. H. Regulation of common neurological disorders by gut microbial metabolites. Exp. Mol. Med. 53, 1821–1833 (2021).
Paik, D. et al. Human gut bacteria produce TH17-modulating bile acid metabolites. Nature 603, 907–912 (2022).
Ke, X. et al. Gut bacterial metabolites modulate endoplasmic reticulum stress. Genome Biol. 22, 292 (2021).
Chouchani, E. T. Logic and mechanisms of metabolite signalling. Nat. Rev. Endocrinol. 18, 71–72 (2022).
Andrade, J. et al. Control of endothelial quiescence by FOXO-regulated metabolites. Nat. Cell Biol. 23, 413–423 (2021).
Xu, Y. et al. The miR-1185-2-3p-GOLPH3L pathway promotes glucose metabolism in breast cancer by stabilizing p53-induced SERPINE1. J. Exp. Clin. Cancer Res. 40, 47 (2021).
Waman, V. P. et al. Mycobacterial genomics and structural bioinformatics: opportunities and challenges in drug discovery. Emerg. Microbes Infect. 8, 109–118 (2019).
Reilly, M. P. & Bornfeldt, K. E. Integrative multiomics approaches for discovery of new drug targets for cardiovascular disease. Circulation 143, 2471–2474 (2021).
Lai, Q. et al. Oxoeicosanoid receptor inhibition alleviates acute myocardial infarction through activation of BCAT1. Basic Res. Cardiol. 116, 3 (2021).
Garana, B. B. & Graham, N. A. Metabolomics paves the way for improved drug target identification. Mol. Syst. Biol. 18, e10914 (2022).
Kaoutari, A. E. et al. Metabolomic profiling of pancreatic adenocarcinoma reveals key features driving clinical outcome and drug resistance. EBioMedicine 66, 103332 (2021).
Dauvilliers, Y., Barateau, L., Middleton, B., van der Veen, D. R. & Skene, D. J. Metabolomics signature of patients with narcolepsy. Neurology 98, e493–e505 (2022).
Geier, B. et al. Spatial metabolomics of in situ host–microbe interactions at the micrometre scale. Nat. Microbiol. 5, 498–510 (2020).
Bao, X. H. et al. Metabolic characterization of the badagan constitution in mongolian medicine by ultrahigh-performance liquid chromatography/quadrupole time-of-flight mass spectrometry/MS. World J. Tradit. Chin. Med. 8, 539–547 (2022).
Liu, R. et al. Quantitative evaluation of the compatibility effects of aidi injection on the treatment of hepatocellular carcinoma using targeted metabolomics: A new strategy on the mechanism study of an anticancer compound in traditional chinese medicine. World J. Tradit. Chin. Med. 7, 111–119 (2021).
Paraskevaidi, M. et al. Laser-assisted rapid evaporative ionisation mass spectrometry (LA-REIMS) as a metabolomics platform in cervical cancer screening. EBioMedicine 60, 103017 (2020).
Hegazi, N. M., Radwan, R. A., Bakry, S. M. & Saad, H. H. Molecular networking aided metabolomic profiling of beet leaves using three extraction solvents and in relation to its anti-obesity effects. J. Adv. Res. 24, 545–555 (2020).
Aron, A. T. et al. Native mass spectrometry-based metabolomics identifies metal-binding compounds. Nat. Chem. 14, 100–109 (2022).
Wang, X. et al. Microfluidic chip and its application in autophagy detection. TrAC Trends Anal. Chem. 117, 300–315 (2019).
Qin, L., Liu, X., Xu, K. & Li, C. Mining and design of biosensors for engineering microbial cell factory. Curr. Opin. Biotechnol. 75, 102694 (2022).
Marella, T. K. et al. Deciphering functional biomolecule potential of marine diatoms through complex network approach. Bioresour. Technol. 342, 125927 (2021).
Guo, Y. et al. An overview of organophosphate esters and their metabolites in humans: Analytical methods, occurrence, and biomonitoring. Sci. Total Environ. 848, 157669 (2022).
Grimm, F. A. et al. Cardiovascular effects of polychlorinated biphenyls and their major metabolites. Environ. Health Perspect. 128, 77008 (2020).
Das, N. K. et al. Microbial metabolite signaling is required for systemic iron homeostasis. Cell Metab. 31, 115–130.e116 (2020).
Yamamoto, K. et al. The complexity of intercellular localisation of alkaloids revealed by single-cell metabolomics. N. Phytol. 224, 848–859 (2019).
Seydel, C. Single-cell metabolomics hits its stride. Nat. Methods 18, 1452–1456 (2021).
Rappez, L. et al. SpaceM reveals metabolic states of single cells. Nat. Methods 18, 799–805 (2021).
Lanekoff, I., Sharma, V. V. & Marques, C. Single-cell metabolomics: where are we and where are we going? Curr. Opin. Biotechnol. 75, 102693 (2022).
Du, J. et al. Raman-guided subcellular pharmaco-metabolomics for metastatic melanoma cells. Nat. Commun. 11, 4830 (2020).
Sade Yazdi, D. et al. Homocysteine fibrillar assemblies display cross-talk with Alzheimer’s disease beta-amyloid polypeptide. Proc. Natl Acad. Sci. U. S. A. 118, e2017575118 (2021).
Parker, A. et al. Fecal microbiota transfer between young and aged mice reverses hallmarks of the aging gut, eye, and brain. Microbiome 10, 68 (2022).
Zhang, Y. W. et al. L-tyrosine metabolic pathway in microorganisms and its application in the biosynthesis of plant-derived natural products. World J. Tradit. Chin. Med. 8, 386–394 (2022).
Mor, D. E. et al. Metformin rescues Parkinson’s disease phenotypes caused by hyperactive mitochondria. Proc. Natl. Acad. Sci. USA 117, 26438–26447 (2020).
Hao, M. et al. Ultra-performance liquid chromatography-quadrupole/time-of-flight mass spectrometry based bile and urine metabonomics study on the ameliorative effects of Curcuma wenyujin rhizoma on acute blood stasis in rats. World J. Tradit. Chin. Med. 8, 141–152 (2022).
Joshi, A., Rienks, M., Theofilatos, K. & Mayr, M. Systems biology in cardiovascular disease: a multiomics approach. Nat. Rev. Cardiol. 18, 313–330 (2021).
Qian, Y. X. et al. Ultra-high performance liquid chromatography/ion mobility-quadrupole time-of-flight massspectrometry and database-driven automatic peak annotation for the rapid profiling and characterization of the multicomponents from Stephaniae Tetrandrae radix (Fang-Ji). World J. Tradit. Chin. Med. 7, 120–12 (2021).
Husain, A. et al. Ephrin-A3/EphA2 axis regulates cellular metabolic plasticity to enhance cancer stemness in hypoxic hepatocellular carcinoma. J. Hepatol. 77, 383–396 (2022).
Dong, R. et al. CSF metabolites associated with CSF NeuroToolKit biomarkers. Alzheimer’s. Dement. 17, e056300 (2021).
Shouval, R. et al. Oral mucositis is associated with distinctive patterns of oral microbiota injury in patients undergoing allogeneic hematopoietic stem cell transplantation. Blood 134, 3265–3265 (2019).
Prabhu, A. H. et al. Integrative cross-platform analyses identify enhanced heterotrophy as a metabolic hallmark in glioblastoma. Neuro Oncol. 21, 337–347 (2019).
Noerman, S., Kolehmainen, M. & Hanhineva, K. Profiling of endogenous and gut microbial metabolites to indicate metabotype-specific dietary responses: a systematic review. Adv. Nutr. 11, 1237–1254 (2020).
Hou, D. et al. Immu-36. B cell-vaccine elicits long term immunity against glioblastoma via activation and differentiation of tumor-specific Cd8+ memory T cells. Neuro-Oncol. 23, vi100-vi100 (2021).
Anyfanti, P., Nikolaidou, B. & Gkaliagkousi, E. Urine metabolomic phenotyping for detection of adrenocortical carcinoma: still a long way to go. Lancet Diabetes Endocrinol. 8, 876–877 (2020).
Schwarzler, J. et al. PUFA-induced metabolic enteritis as a fuel for Crohn’s disease. Gastroenterology 162, 1690–1704 (2022).
Petrus, P. et al. Glutamine links obesity to inflammation in human white adipose tissue. Cell Metab. 31, 375–390.e311 (2020).
Nemet, I. et al. A cardiovascular disease-linked gut microbial metabolite acts via adrenergic receptors. Cell 180, 862–877.e822 (2020).
Liang, C. et al. Localisation of PGK1 determines metabolic phenotype to balance metastasis and proliferation in patients with SMAD4-negative pancreatic cancer. Gut 69, 888–900 (2020).
Murashige, D. et al. Comprehensive quantification of fuel use by the failing and nonfailing human heart. Science 370, 364–368 (2020).
Li, J. et al. The Mediterranean diet, plasma metabolome, and cardiovascular disease risk. Eur. Heart J. 41, 2645–2656 (2020).
Joshi, S. K. et al. The AML microenvironment catalyzes a stepwise evolution to gilteritinib resistance. Cancer Cell. 39, 999–1014.e1018 (2021).
Eddy, S., Mariani, L. H. & Kretzler, M. Integrated multi-omics approaches to improve classification of chronic kidney disease. Nat. Rev. Nephrol. 16, 657–668 (2020).
Zhang, C. et al. A specific gut microbiota and metabolomic profiles shifts related to antidiabetic action: The similar and complementary antidiabetic properties of type 3 resistant starch from Canna edulis and metformin. Pharm. Res. 159, 104985 (2020).
Li, H., Boulougoura, A., Endo, Y. & Tsokos, G. C. Abnormalities of T cells in systemic lupus erythematosus: new insights in pathogenesis and therapeutic strategies. J. Autoimmun. 132, 102870 (2022).
Hor, J. H. et al. ALS motor neurons exhibit hallmark metabolic defects that are rescued by SIRT3 activation. Cell Death Differ. 28, 1379–1397 (2021).
Dong, T. et al. Mitochondrial metabolism mediated macrophage polarization in chronic lung diseases. Pharm. Ther. 239, 108208 (2022).
Bjerrum, J. T., Wang, Y. L., Seidelin, J. B. & Nielsen, O. H. IBD metabonomics predicts phenotype, disease course, and treatment response. EBioMedicine 71, 103551 (2021).
Bekhite, M. M. et al. Longitudinal metabolic profiling of cardiomyocytes derived from human-induced pluripotent stem cells. Basic Res Cardiol. 115, 37 (2020).
Kwan, B. et al. Metabolomic markers of kidney function decline in patients with diabetes: Evidence from the chronic renal insufficiency cohort (CRIC) study. Am. J. Kidney Dis. 76, 511–520 (2020).
Chen, Z., Huang, X., Gao, Y., Zeng, S. & Mao, W. Plasma-metabolite-based machine learning is a promising diagnostic approach for esophageal squamous cell carcinoma investigation. J. Pharm. Anal. 11, 505–514 (2021).
Onesti, C. E. et al. Tryptophan catabolism differentiates breast cancer patients from healthy controls but does not predict outcome. Ann. Oncol. 30, iii18 (2019).
Leaf, D. E. & Ginde, A. A. Vitamin D3 to treat COVID-19: Different disease, same answer. JAMA 325, 1047–1048 (2021).
Lally, P. J. et al. Magnetic resonance spectroscopy assessment of brain injury after moderate hypothermia in neonatal encephalopathy: a prospective multicentre cohort study. Lancet Neurol. 18, 35–45 (2019).
Eross, B., Szentesi, A. & Hegyi, P. Metabolic signature might be an option to identify patients with early CP. Gut 70, 2023–2024 (2021).
Davies, A. et al. Short and medium chain acylcarnitines as markers of outcome in diabetic and non-diabetic subjects with acute coronary syndromes. Eur. Heart J. 41, ehaa946.1561 (2020).
Gallagher, K., Catesson, A., Griffin, J. L., Holmes, E. & Williams, H. R. T. Metabolomic Analysis in Inflammatory Bowel Disease: A Systematic Review. J. Crohns Colitis 15, 813–826 (2021).
Zhang, D. et al. Investigating the effect of Ti3C2 (MXene) nanosheet on human umbilical vein endothelial cells via a combined untargeted and targeted metabolomics approach. Carbon 178, 810–821 (2021).
Wan, X. et al. Metabolomics strategy comprehensively unveils the effect of catechins intervention on the biomarkers of exposure to acrylamide and biomarkers of cardiometabolic risk. Environ. Int. 169, 107517 (2022).
Sun, J. et al. A review of environmental metabolism disrupting chemicals and effect biomarkers associating disease risks: Where exposomics meets metabolomics. Environ. Int. 158, 106941 (2022).
Fu, J., Gong, Z. & Bae, S. Assessment of the effect of methyl-triclosan and its mixture with triclosan on developing zebrafish (Danio rerio) embryos using mass spectrometry-based metabolomics. J. Hazard Mater. 368, 186–196 (2019).
Farag, M. A. et al. Metabolomics reveals impact of seven functional foods on metabolic pathways in a gut microbiota model. J. Adv. Res. 23, 47–59 (2020).
Yuliana, N. D., Hunaefi, D., Goto, M., Ishikawa, Y. T. & Verpoorte, R. Measuring the health effects of food by metabolomics. Crit. Rev. Food Sci. Nutr. 62, 6359–6373 (2022).
Mika, A. et al. The impact of the interplay of the intestinal microbiome and diet on the metabolomic and health outcomes of bariatric surgery. Obes. Rev. 23, e13455 (2022).
Liu, Y., Tang, W., Ao, J., Zhang, J. & Feng, L. Transcriptomics integrated with metabolomics reveals the effect of Bisphenol F (BPF) exposure on intestinal inflammation. Sci. Total Environ. 816, 151644 (2022).
Zhang, H. et al. A metabolomic study on the gender-dependent effects of maternal exposure to fenvalerate on neurodevelopment in offspring mice. Sci. Total Environ. 707, 136130 (2020).
Pu, J. et al. Sex-specific plasma metabolome signatures in major depressive disorder. Psychiatry Clin. Neurosci. 73, 713–714 (2019).
O'Keeffe, L. M. et al. Sex-specific associations of adiposity with cardiometabolic traits in the UK: A multi-life stage cohort study with repeat metabolomics. PLoS Med. 19, e1003636 (2022).
Bell, J. A. et al. Sex differences in systemic metabolites at four life stages: cohort study with repeated metabolomics. BMC Med. 19, 58 (2021).
Lefèvre-Arbogast, S. et al. P1-011: Untargeted metabolomics in a prospective cohort to identify diet‐related metabolites associated with age-related cognitive decline. Alzheimer’s Dement. 15, P234–P234 (2019).
Acar, I. E. et al. Integrating Metabolomics, Genomics, and Disease Pathways in Age-Related Macular Degeneration: The EYE-RISK Consortium. Ophthalmology 127, 1693–1709 (2020).
Teruya, T., Chen, Y. J., Kondoh, H., Fukuji, Y. & Yanagida, M. Whole-blood metabolomics of dementia patients reveal classes of disease-linked metabolites. Proc. Natl. Acad. Sci. USA 118, e2022857118 (2021).
Shen, X. et al. Serum metabolomics identifies dysregulated pathways and potential metabolic biomarkers for hyperuricemia and gout. Arthritis Rheumatol. 73, 1738–1748 (2021).
Yamakawa, P. E. et al. Metabolomic profile in patients with paroxysmal nocturnal hemoglobinuria. Blood 134, 2229–2229 (2019).
Luo, S. et al. Serum metabolomic alterations associated with proteinuria in CKD. Clin. J. Am. Soc. Nephrol. 14, 342–353 (2019).
Hong, H. et al. Cadmium perturbed metabolomic signature in pancreatic beta cells correlates with disturbed metabolite profile in human urine. Environ. Int. 161, 107139 (2022).
Hauser, J. et al. Sialylated human milk oligosaccharides program cognitive development through a non-genomic transmission mode. Mol. Psychiatry 26, 2854–2871 (2021).
Garwolińska, D., Namieśnik, J., Kot-Wasik, A. & Hewelt-Belka, W. State of the art in sample preparation for human breast milk metabolomics—merits and limitations. TrAC Trends Anal. Chem. 114, 1–10 (2019).
Shao, Y. & Le, W. Recent advances and perspectives of metabolomics-based investigations in Parkinson’s disease. Mol. Neurodegener. 14, 3 (2019).
Liu, Y. et al. Sputum metabolomic profiling reveals metabolic pathways and signatures associated with inflammatory phenotypes in patients with asthma. Allergy Asthma Immunol. Res. 14, 393–411 (2022).
Guan, S. Y. et al. Potential biomarkers for clinical outcomes of IVF cycles in women with/without PCOS: Searching with metabolomics. Front Endocrinol. 13, 982200 (2022).
Wang, C. et al. Metabolic signatures of hepatolithiasis using ultra-high performance liquid chromatography-tandem mass spectrometry. Metabolomics 18, 69 (2022).
Zhao, S. et al. Alteration of bile acids and omega-6 PUFAs are correlated with the progression and prognosis of drug-induced liver injury. Front Immunol. 13, 772368 (2022).
Ansone, L. et al. Amino acid metabolism is significantly altered at the time of admission in hospital for severe COVID-19 patients: findings from longitudinal targeted metabolomics analysis. Microbiol Spectr. 9, e0033821 (2021).
Bykowski, E. A. et al. Urinary metabolomic signatures as indicators of injury severity following traumatic brain injury: A pilot study. IBRO Neurosci. Rep. 11, 200–206 (2021).
Guo, J. et al. Serum metabolite signatures in normal individuals and patients with colorectal adenoma or colorectal cancer using UPLC-MS/MS method. J. Proteom. 270, 104741 (2022).
Yu, C. et al. Identification of the metabolic signatures of prostate cancer by mass spectrometry-based plasma and urine metabolomics analysis. Prostate 81, 1320–1328 (2021).
Klatt, S. et al. A six-metabolite panel as potential blood-based biomarkers for Parkinson’s disease. NPJ Parkinsons Dis. 7, 94 (2021).
Baranovicova, E. et al. Circulating metabolites in the early stage of breast cancer were not related to cancer stage or subtypes but associated with ki67 level. Promising statistical discrimination from controls. Mol. Cell Probes. 66, 101862 (2022).
Liu, H. et al. Untargeted serum metabolomics reveals specific metabolite abnormalities in patients with Crohn’s disease. Front Med (Lausanne). 9, 814839 (2022).
Xu, B. et al. NMR-based metabolomic analysis of plasma in patients with adult congenital heart disease and associated pulmonary arterial hypertension: A pilot study. Metabolites 12, 845 (2022).
Lyu, S. et al. Metabolomics analysis reveals four biomarkers associated with the gouty arthritis progression in patients with sequential stages. Semin Arthritis Rheum. 55, 152022 (2022).
Byeon, S. K. et al. Development of a multiomics model for identification of predictive biomarkers for COVID-19 severity: a retrospective cohort study. Lancet Digit Health 4, 632–e645 (2022).
Lan, X. Y. et al. Bone marrow mesenchymal stem cells exert protective effects after ischemic stroke through upregulation of glutathione. Stem Cell Rev. Rep. 18, 585–594 (2022).
Liu, J. et al. Clinical parameters and metabolomic biomarkers that predict inhospital outcomes in patients with ST-segment elevated myocardial infarctions. Front Physiol. 12, 820240 (2022).
Sun, Y. et al. Plasma metabolomics reveals metabolic profiling for diabetic retinopathy and disease progression. Front Endocrinol. 12, 757088 (2021).
Zheng, J. et al. Combined metabolomics with transcriptomics reveals potential plasma biomarkers correlated with non-small-cell lung cancer proliferation through the Akt pathway. Clin. Chim. Acta 530, 66–73 (2022).
Hackshaw, K. V. et al. Vibrational spectroscopy for identification of metabolites in biologic samples. Molecules 25, 4725 (2020).
García-Figueiras, R. et al. Proton magnetic resonance spectroscopy in oncology: the fingerprints of cancer? Diagn Interv. Radiol. 22, 75–89 (2016).
Lin, L. et al. Study on quality markers and action mechanisms of inulae flos on anti-hepatitis through network pharmacology and high-performance liquid chromatography fingerprints. World J. Tradit. Chin. Med. 8, 426–435 (2022).
Meoni, G. et al. The metabolic fingerprints of HCV and HBV infections studied by nuclear magnetic resonance spectroscopy. Sci. Rep. 9, 4128 (2019).
Ismail, M. et al. Noninvasive detection of cocaine and heroin use with single fingerprints: determination of an environmental cutoff. Clin. Chem. 64, 909–917 (2018).
Oluwagbemigun, K. et al. An investigation into the temporal reproducibility of tryptophan metabolite networks among healthy adolescents. Int. J. Tryptophan Res. https://doi.org/10.1177/11786469211041376 (2021).
Kuwayama, K. et al. Effectiveness of saliva and fingerprints as alternative specimens to urine and blood in forensic drug testing. Drug Test. Anal. 8, 644–651 (2016).
Oakman, C. et al. Uncovering the metabolomic fingerprint of breast cancer. Int J. Biochem Cell Biol. 43, 1010–1020 (2021).
Vignoli, A. et al. High-throughput metabolomics by 1D NMR. Angew. Chem. 58, 968–994 (2019).
González-Domínguez, R. et al. High-throughput direct mass spectrometry-based metabolomics to characterize metabolite fingerprints associated with Alzheimer’s disease pathogenesis. Metabolites 8, 52 (2018).
Wu, Z. Y. et al. Semiautomated alignment of high-throughput metabolite profiles with chemometric tools. J. Anal. methods Chem. https://doi.org/10.1155/2017/9402045 (2017).
van Outersterp, R. E. et al. Metabolite identification using infrared ion spectroscopy─novel biomarkers for pyridoxine-dependent epilepsy. Anal. Chem. 93, 15340–15348 (2021).
Beckmann, M. et al. High-throughput, nontargeted metabolite fingerprinting using nominal mass flow injection electrospray mass spectrometry. Nat. Protoc. 3, 486–504 (2008).
Rijk, J. C. et al. Screening for modulatory effects on steroidogenesis using the human H295R adrenocortical cell line: a metabolomics approach. Chem. Res. Toxicol. 25, 1720–1731 (2012).
Beckmann, M. et al. Dietary exposure biomarker-lead discovery based on metabolomics analysis of urine samples. Proc. Nutr. Soc. 72, 352–361 (2013).
Ly-Verdú, S. et al. Combining metabolomic non-targeted GC×GC-ToF-MS analysis and chemometric ASCA-based study of variances to assess dietary influence on type 2 diabetes development in a mouse model. Anal. Bioanal. Chem. 407, 343–354 (2015).
Inoue, K. et al. Blood-based diagnosis of Alzheimer’s disease using fingerprinting metabolomics based on hydrophilic interaction liquid chromatography with mass spectrometry and multivariate statistical analysis. J. Chromatogr. B. 974, 24–34 (2015).
Mastrangelo, A. et al. Metabolomics as a tool for drug discovery and personalised medicine. A review. Curr. Top. Med. Chem. 14, 2627–2636 (2014).
Tai, D. et al. Tissue- and cell-type-specific molecular and functional signatures of 16p11.2 reciprocal genomic disorder across mouse brain and human neuronal models. Am. J. Hum. Genet. 109, 1789–1813 (2022).
Zhou, W. et al. Binding and regulation of transcription by yeast Ste12 variants to drive mating and invasion phenotypes. Genetics 214, 397–407 (2020).
Fernandez-Jimenez, N. et al. The methylome of the celiac intestinal epithelium harbours genotype-independent alterations in the HLA region. Sci. Rep. 9, 1298 (2019).
Enright, E. F. et al. Gut microbiota-mediated bile acid transformations alter the cellular response to multidrug resistant transporter substrates in vitro: focus on P-glycoprotein. Mol. Pharm. 15, 5711–5727 (2018).
Urpi-Sarda, M. et al. Non-targeted metabolomic biomarkers and metabotypes of type 2 diabetes: A cross-sectional study of PREDIMED trial participants. Diabetes Metab. 45, 167–174 (2019).
Li, Y. et al. Penicillin-binding protein transpeptidase signatures for tracking and predicting β-Lactam resistance levels in Streptococcus pneumoniae. mBio 7, 00756 (2016).
Hung, C. I. et al. Metabolomics-based discrimination of patients with remitted depression from healthy controls using 1H-NMR spectroscopy. Sci. Rep. 11, 15608 (2021).
Yu, Z. et al. Potential mechanisms of the anti-hypertensive effects of RVPSL on spontaneously hypertensive rats using non-targeted serum metabolomics. Food Funct. 12, 8561–8569 (2021).
Likhitweerawong, N. et al. Profiles of urine and blood metabolomics in autism spectrum disorders. Metab. Brain Dis. 36, 1641–1671 (2021).
Guan, F. et al. Simultaneous metabolomics and proteomics analysis of plasma-derived extracellular vesicles. Anal. methods 13, 1930–1938 (2021).
Wang X. et al. Serum metabolome alterations in patients with early nonalcoholic fatty liver disease. Biosci. Rep. (2022). https://doi.org/10.1042/BSR20220319.
Hu, X. et al. Combining metabolome and clinical indicators with machine learning provides some promising diagnostic markers to precisely detect smear-positive/negative pulmonary tuberculosis. BMC Infect. Dis. 22, 707 (2022).
An, R. et al. Integrative analysis of plasma metabolomics and proteomics reveals the metabolic landscape of breast cancer. Cancer Metab. 10, 13 (2022).
Ismaiel, A. et al. Metabolic biomarkers related to cardiac dysfunction in metabolic-dysfunction-associated fatty liver disease: a cross-sectional analysis. Nutr. Diabetes 12, 4 (2022).
Zhao, J. et al. A multi-platform metabolomics reveals possible biomarkers for the early-stage esophageal squamous cell carcinoma. Anal. Chim. Acta 1220, 340038 (2022).
Li, X. et al. Metabolomics based plasma biomarkers for diagnosis of oral squamous cell carcinoma and oral erosive lichen planus. J. Cancer 13, 76–87 (2022).
Lunyera, J. et al. Urine tricarboxylic acid cycle signatures of early-stage diabetic kidney disease. Metabolomics 18, 5 (2021).
Liu, S. et al. Serum integrative omics reveals the landscape of human diabetic kidney disease. Mol. Metab. 54, 101367 (2021).
Agnihotri, P. et al. Differential metabolome in rheumatoid arthritis: a brief perspective. Curr. Rheumatol. Rep. 23, 42 (2021).
Castro, A. et al. Understanding the relationship between intrinsic cardiorespiratory fitness and serum and skeletal muscle metabolomics profile. J. proteome Res. 20, 2397–2409 (2021).
Suhre, K. & Zaghlool, S. Connecting the epigenome, metabolome and proteome for a deeper understanding of disease. J. Intern. Med. 290, 527–548 (2021).
Di Minno, A. et al. The evolving landscape of untargeted metabolomics. Nutr., Metab. Cardiovascular Dis. 31, 1645–1652 (2021).
Li, Q. et al. Metabolomics analysis reveals deranged energy metabolism and amino acid metabolic reprogramming in dogs with myxomatous mitral valve disease. J. Am. Heart Assoc. 10, e018923 (2021).
Elmsjö, A. et al. Post-mortem metabolomics: a novel approach in clinical biomarker discovery and a potential tool in death investigations. Chem. Res. Toxicol. 34, 1496–1502 (2021).
Moon, S. et al. Circulating short and medium chain fatty acids are associated with normoalbuminuria in type 1 diabetes of long duration. Sci. Rep. 11, 8592 (2021).
Miller, H. A. et al. Evaluation of disease staging and chemotherapeutic response in non-small cell lung cancer from patient tumor-derived metabolomic data. Lung cancer 156, 20–30 (2021).
Vega-Beyhart, A. et al. Endogenous cortisol excess confers a unique lipid signature and metabolic network. J. Mol. Med. 99, 1085–1099 (2021).
Lord, J. et al. Mendelian randomization identifies blood metabolites previously linked to midlife cognition as causal candidates in Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 118, e2009808118, https://doi.org/10.1073/pnas.2009808118 (2021).
Wegermann, K. et al. Serum bile acid, vitamin E, and serotonin metabolites are associated with future liver-related events in nonalcoholic fatty liver disease. Hepatol. Commun. 5, 608–617 (2021).
Hu, C. et al. Metabolic analysis of early nonalcoholic fatty liver disease in humans using liquid chromatography-mass spectrometry. J. Transl. Med. 19, 152 (2021).
Zhao, R. et al. Biomarkers for pancreatic cancer based on tissue and serum metabolomics analysis in a multicenter study. Cancer Med. https://doi.org/10.1002/cam4.5296 (2022).
Guo, P. et al. Metabolomic analyses redefine the biological classification of pancreatic cancer and correlate with clinical outcomes. Int J. Cancer 151, 1835–1846 (2022).
Liu, C. et al. Tissue metabolomics identified new biomarkers for the diagnosis and prognosis prediction of pancreatic cancer. Front Oncol. 12, 991051 (2022).
Liu Z. et al. Plasm metabolomics study in pulmonary metastatic carcinoma. J Oncol. (2022). https://doi.org/10.1155/2022/9460019
Cao, P. et al. Precise pathological classification of non-small cell lung adenocarcinoma and squamous carcinoma based on an integrated platform of targeted metabolome and lipidome. Metabolomics 17, 98 (2021).
Wu, M. et al. Serum metabolomics reveals an innovative diagnostic model for salivary gland tumors. Anal. Biochem. 655, 114853 (2022).
Liu, X. et al. LC-MS-based urine metabolomics analysis for the diagnosis and monitoring of medulloblastoma. Front Oncol. 12, 949513 (2022).
Li, J. et al. Interpretable machine learning-derived nomogram model for early detection of diabetic retinopathy in type 2 diabetes mellitus: a widely targeted metabolomics study. Nutr. Diabetes 12, 36 (2022).
Fan, Y. et al. The metabolomic characterization of different types of coronary atherosclerotic heart disease in male. Cardiol. Res Pract. 2022, 6491129 (2022).
Moreau, C. et al. Salivary metabolome indicates a shift in tyrosine metabolism in patients with Burning Mouth Syndrome: a prospective case-control study. Pain https://doi.org/10.1097/j.pain.0000000000002733 (2022).
Wang, J. et al. Spatial metabolomics identifies distinct tumor-specific subtypes in gastric cancer patients. Clin. Cancer Res. 28, 2865–2877 (2022).
Yu, S. et al. Integrative metabolomic characterization identifies plasma metabolomic signature in the diagnosis of papillary thyroid cancer. Oncogene 41, 2422–2430 (2022).
Valdés, A. et al. Metabolomics study of COVID-19 patients in four different clinical stages. Sci. Rep. 12, 1650 (2022).
Yan, X. et al. A serum lipidomics study for the identification of specific biomarkers for endometrial polyps to distinguish them from endometrial cancer or hyperplasia. Int J. Cancer 150, 1549–1559 (2022).
Masoodi, M. et al. Disturbed lipid and amino acid metabolisms in COVID-19 patients. J. Mol. Med (Berl.). 100, 555–568 (2022).
Lee, S. et al. A unique urinary metabolic feature for the determination of bladder cancer, prostate cancer, and renal cell carcinoma. Metabolites 11, 591 (2022).
Xu, B. et al. Metabolomics profiling discriminates prostate cancer from benign prostatic hyperplasia within the prostate-specific antigen gray zone. Front Oncol. 11, 730638 (2022).
Alotaibi, M. et al. Metabolomic profiles differentiate scleroderma-PAH from idiopathic PAH and correspond with worsened functional capacity. Chest S0012-3692, 03706 (2022).
Luo, J. et al. Human plasma metabolomics identify 9-cis-retinoic acid and dehydrophytosphingosine levels as novel biomarkers for early ventricular fibrillation after ST-elevated myocardial infarction. Bioengineered 13, 3334–3350 (2022).
Albillos, S. M. et al. Plasma acyl-carnitines, bilirubin, tyramine and tetrahydro-21-deoxycortisol in Parkinson’s disease and essential tremor. A case control biomarker study. Parkinsonism Relat. Disord. 91, 167–172 (2022).
Nalbantoglu, S. & Karadag, A. Metabolomics bridging proteomics along metabolites/oncometabolites and protein modifications: Paving the way toward integrative multiomics. J. Pharm. Biomed. Anal. 199, 114031 (2021).
Laiakis, E. C. et al. Small molecule responses to sequential irradiation with neutrons and photons for biodosimetry applications: An initial assessment. Radiat. Res. 196, 468–477 (2021).
Kettwig, M. et al. Targeted metabolomics revealed changes in phospholipids during the development of neuroinflammation in Abcd1tm1Kds mice and X-linked adrenoleukodystrophy patients. J. Inherit. Metab. Dis. 44, 1174–1185 (2021).
Ouyang, Y. et al. Metabolome-genome-wide association study (mGWAS) reveals novel metabolites associated with future type 2 diabetes risk and susceptibility loci in a case-control study in a Chinese prospective cohort. Glob. Chall. 5, 2000088 (2021).
Aung, M. T. et al. Maternal lipidomic signatures in relation to spontaneous preterm birth and large-for-gestational age neonates. Sci. Rep. 11, 8115 (2021).
Bourdon, M. et al. Adenomyosis is associated with specific proton nuclear magnetic resonance (1H-NMR) serum metabolic profiles. Fertil. Steril. 116, 243–254 (2021).
Yuan, Y. et al. Integrative metabolic profile of myelodysplastic syndrome based on UHPLC-MS. Biomed. Chromatogr. 35, e5136 (2021).
Standage, S. W. et al. NMR-based serum and urine metabolomic profile reveals suppression of mitochondrial pathways in experimental sepsis-associated acute kidney injury. Am. J. Physiol. Ren. Physiol. 320, F984–F1000 (2021).
Steinbusch, L. et al. Targeted urine metabolomics with a graphical reporting tool for rapid diagnosis of inborn errors of metabolism. J. Inherit. Metab. Dis. 44, 1113–1123 (2021).
Li, M. Y. et al. Biomarkers and key pathways in atrial fibrillation associated with mitral valve disease identified by multi-omics study. Ann. Transl. Med. 9, 393 (2021).
Steinbrenner, I. et al. Urine metabolite levels, adverse kidney outcomes, and mortality in CKD patients: A metabolome-wide association study. Am. J. kidney Dis. 78, 669–677 (2021).
Wang, W. et al. GC-MS-based metabolomics reveals new biomarkers to assist the differentiation of prostate cancer and benign prostatic hyperplasia. Clin. Chim. Acta 519, 10–17 (2021).
Qin, Y. et al. Association between plasma free fatty acid levels and primary angle-closure glaucoma based on a mass spectrometry metabolomics analysis. Acta Ophthalmologica. 100, e204–e212 (2022).
Sangaraju, D. et al. Robust and comprehensive targeted metabolomics method for quantification of 50 different primary, secondary, and sulfated bile acids in multiple biological species (human, monkey, rabbit, dog, and rat) and matrices (plasma and urine) using liquid chromatography high resolution mass spectrometry (LC-HRMS) analysis. J. Am. Soc. Mass Spectrom. 32, 2033–2049 (2021).
Chantzichristos, D. et al. Identification of human glucocorticoid response markers using integrated multi-omic analysis from a randomized crossover trial. eLife 10, e62236 (2021).
Rousseau, G. et al. Preliminary metabolomic profiling of the vitreous humor from hypothermia fatalities. J. proteome Res. 20, 2390–2396 (2021).
Ceperuelo-Mallafré, V. et al. Circulating pyruvate is a potent prognostic marker for critical COVID-19 outcomes. Front Immunol. 13, 912579 (2022).
Oliveira, L. B. et al. Metabolomic profiling of plasma reveals differential disease severity markers in COVID-19 patients. Front Microbiol. 13, 844283 (2022).
Roberts, I. et al. Untargeted metabolomics of COVID-19 patient serum reveals potential prognostic markers of both severity and outcome. Metabolomics 18, 6 (2021).
Barco, S. et al. Untargeted LC-HRMS based-plasma metabolomics reveals 3-O-methyldopa as a new biomarker of poor prognosis in high-risk neuroblastoma. Front Oncol. 12, 845936 (2022).
Ke, C. et al. Metabolomics on vascular events and death after acute ischemic stroke: A prospective matched nested case-control study. Atherosclerosis 351, 1–8 (2022).
Brunmair, J. et al. Metabolic phenotyping of tear fluid as a prognostic tool for personalised medicine exemplified by T2DM patients. EPMA J. 13, 107–123 (2022).
Shen, X. et al. Asparagine metabolism in tumors is linked to poor survival in females with colorectal cancer: A cohort study. Metabolites 12, 164 (2022).
Pandey, R. et al. Novel strategy for untargeted chiral metabolomics using liquid chromatography-high resolution tandem mass spectrometry. Anal. Chem. 93, 5805–5814 (2022).
Hu, R. et al. NMR-based metabolomics in cancer research. Adv. Exp. Med. Biol. 1280, 201–218 (2022).
Yin, G. et al. Metabolomics of oral/head and neck cancer. Adv. Exp. Med. Biol. 1280, 277–290 (2021).
Shu, X. et al. A prospective investigation of circulating metabolome identifies potential biomarkers for gastric cancer. Risk Cancer Epidemiol., Biomark. Prev. 30, 1634–1642 (2021).
Ishibashi, Y. et al. Reliability of urinary charged metabolite concentrations in a large-scale cohort study using capillary electrophoresis-mass spectrometry. Sci. Rep. 11, 7407 (2021).
Xie, G. et al. A metabolite array technology for precision medicine. Anal. Chem. 93, 5709–5717 (2021).
Răchieriu, C. et al. Lipidomic signatures for colorectal cancer diagnosis and progression using UPLC-QTOF-ESI+MS. Biomolecules 11, 417 (2021).
McCullough, M. L. et al. Pre-diagnostic circulating metabolites and colorectal cancer risk in the cancer prevention study-II nutrition cohort. Metabolites 11, 156 (2021).
Eick, C. et al. Broad metabolome alterations associated with the intake of oral contraceptives are mediated by cortisol in premenopausal women. Metabolites 11, 193 (2022).
van Driel, B. O. et al. Metabolomics in severe aortic stenosis reveals intermediates of nitric oxide synthesis as most distinctive markers. Int. J. Mol. Sci. 22, 3569 (2021).
Pretorius, C. J. et al. Metabolomics for biomarker discovery: key signatory metabolic profiles for the identification and discrimination of oat cultivars. Metabolites 11, 165 (2021).
Steuer, A. E. et al. Towards extending the detection window of gamma-hydroxybutyric acid-an untargeted metabolomics study in serum and urine following controlled administration in healthy men. Metabolites 11, 166 (2021).
Yamano, E., Watanabe, Y. & Kataoka, Y. Insights into metabolite diagnostic biomarkers for myalgic encephalomyelitis/chronic fatigue syndrome. Int. J. Mol. Sci. 22, 3423 (2021).
Yu, J. et al. Metabolic abnormalities in patients with chronic disorders of consciousness. Aging Dis. 12, 386–403 (2021).
Adav, S. S. & Wang, Y. Metabolomics signatures of aging: recent advances. Aging Dis. 12, 646–661 (2021).
He, Z., Liu, Z. & Gong, L. Biomarker identification and pathway analysis of rheumatoid arthritis based on metabolomics in combination with ingenuity pathway analysis. Proteomics 21, e2100037 (2021).
Baliga, M. M. et al. Metabolic profiling in children and young adults with autosomal dominant polycystic kidney disease. Sci. Rep. 11, 6629 (2021).
Aredo, J. V. et al. Metabolomic profiling for second primary lung cancer: A pilot case-control study. Lung cancer 155, 61–67 (2021).
Shao, F. et al. Plasma Metabolomics Reveals Systemic Metabolic Alterations of Subclinical and Clinical Hypothyroidism. J. Clin. Endocrinol. Metab. 108, 13–25 (2022).
Marino, C. et al. The metabolomic profile in amyotrophic lateral sclerosis changes according to the progression of the disease: An exploratory study. Metabolites 12, 837 (2022).
Amiri-Dashatan, N. et al. Metabolomic study of serum in patients with invasive ductal breast carcinoma with LC-MS/MS approach. Int J. Biol. Markers. https://doi.org/10.1177/03936155221123343 (2022).
Lu, C. et al. Comprehensive metabolomic characterization of atrial fibrillation. Front Cardiovasc Med. 9, 911845 (2022).
Wang, Z. et al. Serum untargeted metabolomics reveal potential biomarkers of progression of diabetic retinopathy in Asians. Front Mol. Biosci. 9, 871291 (2022).
Albóniga, O. E. et al. Metabolic snapshot of plasma samples reveals new pathways implicated in SARS-CoV-2 pathogenesis. J. Proteome Res. 21, 623–634 (2022).
Li, Z. et al. Analysis of the saliva metabolic signature in patients with primary Sjögren’s syndrome. PLoS One 17, e026927 (2022).
Zhu, C. et al. Distinct urinary metabolic biomarkers of human colorectal cancer. Dis. Markers 2022, 1758113 (2022).
Yue, L. et al. Nontargeted and targeted metabolomics approaches reveal the key amino acid alterations involved in multiple myeloma. PeerJ 10, e12918 (2022).
Feng, K. et al. Identification of biomarkers and the mechanisms of multiple trauma complicated with sepsis using metabolomics. Front Public Health 10, 923170 (2022).
Yu, F. et al. Phenylacetylglutamine, a novel biomarker in acute ischemic stroke. Front Cardiovasc Med. 8, 798765 (2021).
Sriwi, D. et al. Metabolomics profiling of cystic renal disease towards biomarker discovery. Biol. (Basel). 10, 770 (2021).
Gyawali, P. et al. A multi-platform metabolomics approach to identify possible biomarkers for human faecal contamination in Greenshell™ mussels (Perna canaliculus). Sci. total Environ. 771, 145363 (2021).
An, G. et al. Integrative analysis of vaginal microorganisms and serum metabolomics in rats with estrous cycle disorder induced by long-term heat exposure based on 16S rDNA gene sequencing and LC/MS-based metabolomics. Front. Cell. Infect. Microbiol. 11, 595716 (2021).
Jobard, E. et al. Investigation of circulating metabolites associated with breast cancer risk by untargeted metabolomics: a case-control study nested within the French E3N cohort. Br. J. Cancer 124, 1734–1743 (2021).
Abreu, A. C. et al. NMR-based metabolomics approach to explore brain metabolic changes induced by prenatal exposure to autism-inducing chemicals. ACS Chem. Biol. 16, 753–765 (2021).
Sampson, C. M. et al. Combined nicotinamide N-methyltransferase inhibition and reduced-calorie diet normalizes body composition and enhances metabolic benefits in obese mice. Sci. Rep. 11, 5637 (2021).
Irajizad, E. et al. Application of artificial intelligence to plasma metabolomics profiles to predict response to neoadjuvant chemotherapy in triple-negative breast cancer. Front Artif. Intell. 5, 876100 (2022).
Zhuang, J. et al. Metabolic profiling of bladder cancer patients' serum reveals their sensitivity to neoadjuvant chemotherapy. Metabolites 12, 558 (2022).
Yu, R. L. et al. Prediction of clinical efficacy of subcutaneous immunotherapy for Artemisia sieversiana pollen allergic rhinitis by serum metabolomics. J. Formos. Med. Assoc. S0929-6646, 00211 (2022).
Shen, Y. et al. Metabolomics study of treatment response to conbercept of patients with neovascular age-related macular degeneration and polypoidal choroidal vasculopathy. Front Pharmacol. 13, 991879 (2022).
Yan, J. et al. Decreased cerebrospinal fluid kynurenic acid in epileptic spasms: A biomarker of response to corticosteroids. EBioMedicine 84, 104280 (2022).
Guan, X. et al. Kynurenine pathway metabolites and therapeutic response to olanzapine in female patients with schizophrenia: A longitudinal study. CNS Neurosci. Ther. 28, 1539–1546 (2022).
Dai, D. et al. Toward personalized interventions for psoriasis vulgaris: molecular subtyping of patients by using a metabolomics approach. Front Mol. Biosci. 9, 945917 (2022).
Medcalf, M. R. et al. Plasma metabolomic profiling as a tool to identify predictive biomarkers of methotrexate efficacy in rheumatoid arthritis. Semin Arthritis Rheum. 56, 152056 (2022).
Zhong, Z. et al. Serum metabolic profiling analysis of gout patients based on UPLC-Q-TOF/MS. Clin. Chim. Acta; Int. J. Clin. Chem. 515, 52–60 (2022).
Baima, G. et al. Salivary metabolomics for the diagnosis of periodontal diseases: a systematic review with methodological quality assessment. Metabolomics 17, 1 (2022).
Gong, Y. et al. A serum metabolic profiling analysis during the formation of fatty liver in Landes Geese via GC-TOF/MS. Front. Physiol. 11, 581699 (2022).
Xi, M. et al. Discovery of urinary biomarkers of seaweed intake using untargeted LC-MS metabolomics in a three-way cross-over human study. Metabolites 11, 11 (2020).
Sun, X. et al. Sera and lungs metabonomics reveals key metabolites of resveratrol protecting against PAH in rats. Biomed. Pharmacother. 133, 110910 (2021).
Li, S. et al. Urinary metabolomic profiling reveals biological pathways and predictive signatures associated with childhood asthma. J. asthma allergy 13, 713–724 (2020).
Chatterjee, P. et al. Presymptomatic dutch-type hereditary cerebral amyloid angiopathy-related blood metabolite alterations. JAD 79, 895–903 (2021).
Yu, J. et al. Identification of potential serum biomarkers for simultaneously classifying lung adenocarcinoma, squamous cell carcinoma and small cell carcinoma. Cancer Biomark. 30, 331–342 (2021).
Liu, K. et al. Ultraperformance liquid chromatography coupled with quadrupole time-of-flight mass spectrometry-based metabolomics and lipidomics identify biomarkers for efficacy evaluation of mesalazine in a dextran sulfate sodium-induced ulcerative colitis mouse model. J. proteome Res. 20, 1371–1381 (2021).
Zarei, I. et al. Plasma and urine metabolite profiles impacted by increased dietary navy bean intake in colorectal cancer survivors: A randomized-controlled trial. Cancer Prev. Res. 14, 497–508 (2021).
Elpa, D. P. et al. Skin metabolomics. TEM 32, 66–75 (2021).
Wang, Q. et al. Metabolic profiling of angiopoietin-like protein 3 and 4 inhibition: a drug-target Mendelian randomization analysis. Eur. heart J. 42, 1160–1169 (2021).
Szczuko, M. et al. The role of arachidonic and linoleic acid derivatives in pathological pregnancies and the human reproduction process. Int. J. Mol. Sci. 21, 9628 (2020).
Nguyen, T. D. et al. Single-cell mass spectrometry enables insight into heterogeneity in infectious disease. Anal. Chem. 94, 10567–10572 (2022).
Suvannapruk, W. et al. Single-cell metabolic profiling of macrophages using 3D OrbiSIMS: Correlations with phenotype. Anal. Chem. 94, 9389–9398 (2022).
Izquierdo-Garcia, J. L. et al. Discovery and validation of an NMR-based metabolomic profile in urine as TB biomarker. Sci. Rep. 10, 22317 (2020).
Bouftas, M. A systematic review on the feasibility of salivary biomarkers for Alzheimer’s disease. J. Prev. Alzheimer’s Dis. 8, 84–91 (2021).
Guillamón, J. G. et al. Ascorbic acid and prunasin, two candidate biomarkers for endodormancy release in almond flower buds identified by a nontargeted metabolomic study. Horticulture Res. 7, 203 (2020).
Lee, H. S. et al. (2020). Identification of metabolic markers predictive of prediabetes in a Korean population. Sci. Rep. 10, 22009 (2020).
Hollowood-Jones, K. et al. Altered metabolism of mothers of young children with Autism Spectrum Disorder: a case control study. BMC Pediatr. 20, 557 (2020).
Khaliq, W. et al. Lipid metabolic signatures deviate in sepsis survivors compared to non-survivors. Comput. Struct. Biotechnol. J. 18, 3678–3691 (2020).
Kurbatova, N. et al. Urinary metabolic phenotyping for Alzheimer’s disease. Sci. Rep. 10, 21745 (2020).
Lin, G. et al. KCNK3 inhibits proliferation and glucose metabolism of lung adenocarcinoma via activation of AMPK-TXNIP pathway. Cell Death Discov. 8, 360 (2022).
Feng, Y. et al. Causal effects of genetically determined metabolites on cancers included lung, breast, ovarian cancer, and glioma: a Mendelian randomization study. Transl. Lung Cancer Res. 11, 1302–1314 (2022).
Zuo, L. et al. Integrative analysis of metabolomics and transcriptomics data identifies prognostic biomarkers associated with oral squamous cell carcinoma. Front Oncol. 11, 750794 (2021).
Peng, H. et al. Identification of metabolite markers associated with kidney function. J. Immunol. Res. 2022, 6190333 (2022).
Gu, M. et al. Sera metabolomics characterization of patients at different stages in wuhan identifies critical biomarkers of COVID-19. Front Cell Infect. Microbiol. 12, 882661 (2022).
Chen, X., Ye, J., Lei, H. & Wang, C. Novel potential diagnostic serum biomarkers of metabolomics in osteoarticular tuberculosis patients: A preliminary study. Front Cell Infect. Microbiol. 12, 827528 (2022).
Zong, Y. et al. Metabolomic alterations in the tear fluids of patients with superior limbic keratoconjunctivitis. Front Med (Lausanne). 8, 797630 (2022).
Okamoto, N. et al. A metabolomics study of serum in hospitalized patients with chronic schizophrenia. Front Psychiatry 12, 763547 (2021).
Zhu, Q. et al. Comprehensive metabolic profiling of inflammation indicated key roles of glycerophospholipid and arginine metabolism in coronary artery disease. Front Immunol. 13, 829425 (2022).
Tsoukalas, D. et al. Prediction of autoimmune diseases by targeted metabolomic assay of urinary organic acids. Metabolites 10, 502 (2020).
Alkhalil, A. et al. Cutaneous thermal injury modulates blood and skin metabolomes differently in a murine model. J. Burn Care Res. 42, 727–742 (2021).
Nishimura, M. et al. Upregulated kynurenine pathway enzymes in aortic atherosclerotic aneurysm: macrophage kynureninase downregulates inflammation. J. Atherosclerosis Thrombosis 28, 1214–1240 (2021).
Yang, F. et al. NMR-based plasma metabolomics of adult B-cell acute lymphoblastic leukemia. Mol. Omics. 17, 153–159 (2021).
Esperanza, M. G. et al. Liquid chromatography-mass spectrometry untargeted metabolomics reveals increased levels of tryptophan indole metabolites in urine of metabolic syndrome patients. Eur. J. Mass Spectrom. 26, 379–387 (2020).
Meng, F. et al. Serum biomarkers of the calcium-deficient rats identified by metabolomics based on UPLC/Q-TOF MS/MS. Nutr. Metab. 17, 99 (2020).
Shimizu, H. et al. Serum metabolomic profiling of patients with non-infectious uveitis. J. Clin. Med. 9, 3955 (2020).
Oz, O. et al. A pilot study for investigation of plasma amino acid profile in neurofibromatosis type 1 patients. Combinatorial Chem. high. throughput Screen. 25, 114–122 (2022).
Delarocque, J. et al. Metabolic changes induced by oral glucose tests in horses and their diagnostic use. J. Vet. Intern. Med. 35, 597–605 (2021).
Zaccherini, G. et al. Assessing the role of amino acids in systemic inflammation and organ failure in patients with ACLF. J. Hepatol. 74, 1117–1131 (2021).
Udo, R. et al. Urinary charged metabolite profiling of colorectal cancer using capillary electrophoresis-mass spectrometry. Sci. Rep. 10, 21057 (2020).
Liu, C. et al. A metabolomic study of cervical dystonia. Parkinsonism Relat. Disord. 82, 98–103 (2021).
Yang, J. et al. Non-targeted metabolomic analysis predicts the therapeutic effects of exenatide on endothelial injury in patients with type 2 diabetes. J. Diabetes Compl. 35, 107797 (2021).
Ishikawa, S. et al. Relationship between standard uptake values of positron emission tomography/computed tomography and salivary metabolites in oral cancer: A pilot study. J. Clin. Med. 9, 3958 (2020).
Li, X. K. et al. Dysregulation of glutamine/glutamate metabolism in COVID-19 patients: A metabolism study in African population and mini meta-analysis. J. Med Virol. https://doi.org/10.1002/jmv.28150 (2020).
Ozaki, T. et al. Metabolomic alterations in the blood plasma of older adults with mild cognitive impairment and Alzheimer’s disease (from the Nakayama Study). Sci. Rep. 12, 15205 (2022).
Li, Y. C. et al. Cerebrospinal fluid metabolic profiling reveals divergent modulation of pentose phosphate pathway by midazolam, propofol and dexmedetomidine in patients with subarachnoid hemorrhage: a cohort study. BMC Anesthesiol. 22, 34 (2022).
Thomas, I. et al. Serum metabolome associated with severity of acute traumatic brain injury. Nat. Commun. 13, 2545 (2022).
Zhu, Q. et al. Palmitic acid, a critical metabolite, aggravates cellular senescence through reactive oxygen species generation in kawasaki disease. Front Pharmacol. 13, 809157 (2022).
Scarale, M. G. et al. Circulating metabolites associate with and improve the prediction of all-cause mortality in type 2. Diabetes Diabetes 71, 1363–1370 (2022).
Yuan, Y. et al. Functional metabolome profiling may improve individual outcomes in colorectal cancer management implementing concepts of predictive, preventive, and personalized medical approach. EPMA J. 13, 39–55 (2022).
Shi, D. et al. USP14 promotes tryptophan metabolism and immune suppression by stabilizing IDO1 in colorectal cancer. Nat. Commun. 13, 5644 (2022).
Friedrich, M. et al. Tryptophan metabolism drives dynamic immunosuppressive myeloid states in IDH-mutant gliomas. Nat. Cancer 2, 723–740 (2021).
Platten, M. et al. Tryptophan metabolism as a common therapeutic target in cancer, neurodegeneration and beyond. Nat. Rev. Drug Discov. 18, 379–401 (2019).
Sulkowski, P. L. et al. Oncometabolites suppress DNA repair by disrupting local chromatin signalling. Nature 582, 586–591 (2020).
Shao, Y. et al. Alteration of metabolic profile and potential biomarkers in the plasma of Alzheimer’s Disease. Aging Dis. 11, 1459–1470 (2020).
Chen, D. et al. Effects of freeze-thaw cycles of blood samples on high-coverage quantitative metabolomics. Anal. Chem. 92, 9265–9272 (2020).
Foroutan, A. et al. Serum metabolite biomarkers for predicting residual feed intake (RFI) of young angus bulls. Metabolites 10, 491 (2020).
Li, H. et al. CMap analysis identifies Atractyloside as a potential drug candidate for type 2 diabetes based on integration of metabolomics and transcriptomics. J. Cell. Mol. Med. 24, 7417–7426 (2020).
Dutta, P. et al. Early detection of pancreatic intraepithelial neoplasias (PanINs) in transgenic mouse model by hyperpolarized 13C metabolic magnetic resonance spectroscopy. Int. J. Mol. Sci. 21, 3722 (2020).
Lin, Y. T. et al. Global plasma metabolomics to identify potential biomarkers of blood pressure progression. Arteriosclerosis Thrombosis Vasc. Biol. 40, e227–e237 (2020).
Castiglione Morelli, M. A. et al. Metabolic changes in follicular fluids of patients treated with recombinant versus urinary human chorionic gonadotropin for triggering ovulation in assisted reproductive technologies: a metabolomics pilot study. Arch. Gynecol. Obstet. 302, 741–751 (2020).
Wang, J. et al. Metabolomics window into the role of acute kidney injury after coronary artery bypass grafting in diabetic nephropathy progression. PeerJ 8, e9111 (2020).
Fernández-Ochoa, Á. et al. Metabolic disturbances in urinary and plasma samples from seven different systemic autoimmune diseases detected by HPLC-ESI-QTOF-MS. J. Proteome Res. 19, 3220–3229 (2020).
Benetti, E. et al. Sedentariness and urinary metabolite profile in type 2 diabetic patients, a cross-sectional study. Metabolites 10, 205 (2020).
Walker, M. E. et al. Proteomic and metabolomic correlates of healthy dietary patterns: The Framingham heart study. Nutrients 12, 1476 (2020).
Zhao, G. et al. A metabolomic study for chronic heart failure patients based on a dried blood spot mass spectrometry approach. RSC Adv. 10, 19621–19628 (2020).
Chen, Z. Z. & Gerszten, R. E. Metabolomics and proteomics in type 2 diabetes. Circulation Res. 126, 1613–1627 (2020).
Hopkins, A. L. Network pharmacology: the next paradigm in drug discovery. Nat. Chem. Biol. 4, 682–690 (2008).
Swaney, D. L. et al. A protein network map of head and neck cancer reveals PIK3CA mutant drug sensitivity. Science 374, 2911 (2021).
Theodoris, C. V. et al. Network-based screen in iPSC-derived cells reveals therapeutic candidate for heart valve disease. Science 371, 0724 (2021).
Emilsson, V. et al. Co-regulatory networks of human serum proteins link genetics to disease. Science 361, 769–773 (2018).
Li, Z. B. et al. Pyridoxal phosphate, pyridoxamine phosphate, and folic acid based on ceRNA regulatory network as potential biomarkers for the diagnosis of pulmonary tuberculosis. Infect. Genet Evol. 99, 105240 (2022).
Chang, R. et al. Predictive metabolic networks reveal sex- and APOE genotype-specific metabolic signatures and drivers for precision medicine in Alzheimer’s disease. Alzheimers Dement. https://doi.org/10.1002/alz.12675 (2022).
Guo, L. et al. Metabolic network-based identification of plasma markers for non-small cell lung cancer. Anal. Bioanal. Chem. 413, 7421–7430 (2021).
Ke, C. et al. Serum metabolic signatures of high myopia among older Chinese adults. Eye 35, 817–824 (2021).
Bennet, S. M. et al. Application of metabolomics to the study of irritable bowel syndrome. Neurogastroenterol. Motil. 32, e13884 (2020).
Ren, J. L. et al. Efficacy evaluation, active ingredients, and multitarget exploration of herbal medicine. Trends Endocrinol. Metab. 1043-2760, 00016–4 (2020).
Bereman, M. S. et al. Metabolite profiling reveals predictive biomarkers and the absence of β-methyl amino-l-alanine in plasma from individuals diagnosed with amyotrophic lateral sclerosis. J. Proteome Res. 19, 3276–3285 (2020).
Kamishikiryo, T. et al. Left DLPFC activity is associated with plasma kynurenine levels and can predict treatment response to escitalopram in major depressive disorder. Psychiatry Clin. Neurosci. 76, 367–376 (2022).
Suksawat, M. et al. Metabolic phenotyping predicts gemcitabine and cisplatin chemosensitivity in patients with cholangiocarcinoma. Front Public Health 10, 766023 (2022).
Medcalf, M. R. et al. Plasma metabolome normalization in rheumatoid arthritis following initiation of methotrexate and the identification of metabolic biomarkers of efficacy. Metabolites 11, 824 (2021).
Mao, C. et al. Circulating metabolites serve as diagnostic biomarkers for HER2-positive breast cancer and have predictive value for trastuzumab therapy outcomes. J. Clin. Lab Anal. 36, e24212 (2022).
Zhang, H. et al. Serum metabolomics reveals the intervention mechanism and compatible regularity of Chaihu Shu Gan San on chronic unpredictable mild stress-induced depression rat model. J. Pharm. Pharmacol. 72, 1133–1143 (2020).
Lu, Y. S. et al. A comprehensive analysis of metabolomics and transcriptomics reveals new biomarkers and mechanistic insights on DEHP exposures in MCF-7 cells. Chemosphere 255, 126865 (2020).
Liu, L. W. et al. Metabolomic insights into the synergistic effect of biapenem in combination with Xuebijing injection against sepsis. Front. Pharmacol. 11, 502 (2020).
Zhu, Y. L. et al. Metabolomics analysis of the antidepressant prescription Danzhi Xiaoyao Powder in a rat model of Chronic Unpredictable Mild Stress (CUMS). J. Ethnopharmacol. 260, 112832 (2020).
Saito, K. et al. Characterization of postprandial effects on CSF metabolomics: A pilot study with parallel comparison to plasma. Metabolites 10, 185 (2020).
Liu, Y. et al. Nuclear magnetic resonance-based plasma metabolomics revealed the protective effect of tea polyphenols on sulfur mustard-induced injury in rats. J. Pharm. Biomed. Anal. 186, 113278 (2020).
Chauhan, D. S. et al. Secondary metabolites in the treatment of diabetes mellitus: A paradigm Shift. Curr. drug Metab. 21, 493–511 (2020).
Yadav, N. et al. Novel archetype in cancer therapeutics: exploring prospective of phytonanocarriers. 3 Biotech 12, 324 (2022).
Zhang, A. et al. Recent developments and emerging trends of mass spectrometry for herbal ingredients analysis. TrAC 94, 70–76 (2017).
Guo, S. F. et al. Research advance in efficacy evaluation, active substances and action mechanism of traditional Chinese medicine based on metabonomics. Drug Evaluat. Res. 45, 2338–2445 (2022).
Kang, K. B. et al. Mass spectrometry data on specialized metabolome of medicinal plants used in East Asian traditional medicine. Sci. Data. 9, 528 (2022).
Yang, F. et al. Discovery of potential hypoglycemic metabolites in Cassiae Semen by coupling UHPLC-QTOF-MS/MS combined plant metabolomics and spectrum-effect relationship analyses. Food Funct. 13, 10291–10304 (2022).
Calabrese, V. et al. Molecular networking and collision cross section prediction for structural isomer and unknown compound identification in plant metabolomics: a case study applied to Zhanthoxylum heitzii extracts. Anal. Bioanal. Chem. 414, 4103–4118 (2022).
Zhang, K. et al. Integrated Strategy Drives Direct Infusion-Tandem Mass Spectrometry as an Eligible Tool for Shotgun Pseudo-Targeted Metabolomics of Medicinal Plants. Anal. Chem. 93, 2541–2550 (2021).
Crighton, E. et al. Exploring the application of the DSA-TOF, a direct, high-resolution time-of-flight mass spectrometry technique for the screening of potential adulterated and contaminated herbal medicines. J. Am. Soc. Mass Spectrom. 30, 1713–171 (2019).
Jang, A. K. et al. Metabolites identification for major active components of Agastache rugosa in rat by UPLC-Orbitap-MS: Comparison of the difference between metabolism as a single component and as a component in a multi-component extract. J. Pharm. Biomed. Anal. 220, 114976 (2022).
Wang, D. et al. Preliminary screening of the potential active ingredients in traditional Chinese medicines using the Ussing chamber model combined with HPLC-PDA-MS. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 1189, 123090 (2022).
Burico, M. et al. Metabolomic fingerprint of Hamamelis virginiana L. gallotannins by suspect screening analysis with UHPLC-qToF and their semiquantitative evaluation. J. Mass Spectrom. 57, e4878 (2022).
Vaou, N. et al. Interactions between medical plant-derived bioactive compounds: Focus on antimicrobial combination effects. Antibiotics (Basel). 11, 1014 (2022).
Lai J. et al. A deep clustering-based mass spectral data visualization strategy for anti-renal fibrotic lead compound identification from natural products. Analyst. (2022). https://doi.org/10.1039/d2an01185a.
Huang, W. et al. Rapid discovery of potentially vasodilative compounds from Uncaria by UHPLC/Q-Orbitrap-MS based metabolomics and correlation analysis. J. Pharm. Biomed. Anal. 206, 114384 (2021).
Xiong, H. et al. Discovery of quality-marker ingredients of Panax quinquefolius driven by high-throughput chinmedomics approach. Phytomedicine . https://doi.org/10.1016/j.phymed.2019.152928 (2020).
Han, L. et al. Candidate drug screen strategy: the discovery of oroxylin a in scutellariae radix against sepsis via the correlation analysis between plant metabolomics and pharmacodynamics. Front Pharmacol. 13, 861105 (2022).
Rai, A. et al. Mapping drug-target interactions and synergy in multi-molecular therapeutics for pressure-overload cardiac hypertrophy. NPJ Syst. Biol. Appl. 7, 11 (2021).
Lv, Y. et al. Screening and evaluation of anti-SARS-CoV-2 components from Ephedra sinica by ACE2/CMC-HPLC-IT-TOF-MS approach. Anal. Bioanal. Chem. 413, 2995–3004 (2021).
Abhyankar, M. M. et al. Optimizing a multi-component intranasal Entamoeba Histolytica vaccine formulation using a design of experiments strategy. Front Immunol. 12, 683157 (2021).
Zhou, K. et al. Targeted pharmacokinetics and bioinformatics screening strategy reveals JAK2 as the main target for Xin-Ji-Er-Kang in treatment of MIR injury. Biomed. Pharmacother. 155, 113792 (2022).
Hong, L. L. et al. Tentative exploration of pharmacodynamic substances: Pharmacological effects, chemical compositions, and multi-components pharmacokinetic characteristics of ESZWD in CHF-HKYd rats. Front Cardiovasc Med. 9, 913661 (2022).
Elbouzidi, A. et al. LC-MS/MS phytochemical profiling, antioxidant activity, and cytotoxicity of the ethanolic extract of Atriplex halimus L. against breast cancer cell lines: computational studies and experimental validation. Pharm. (Basel). 15, 1156 (2022).
Yang, Y. et al. Pharmacokinetic comparison of nine bioactive compounds of guanxinshutong capsule in normal and acute myocardial infarction rats. Eur. J. Drug Metab. Pharmacokinet. 47, 653–665 (2022).
Ji, L. et al. Characterization of the chemical constituents and metabolic profile of Polygonum cuspidatum Sieb. et Zucc. in rat plasma, urine, and feces by ultra-high performance liquid chromatography coupled with Quadrupole-Exactive Orbitrap mass spectrometry. J. Sep Sci. https://doi.org/10.1002/jssc.202200522 (2022).
He, Y. et al. Metabolic profiling and pharmacokinetic studies of Baihu-Guizhi decoction in rats by UFLC-Q-TOF-MS/MS and UHPLC-Q-TRAP-MS/MS. Chin. Med. 17, 117 (2022).
Du, Y. et al. LC-MS/MS combined with blood-brain dual channel microdialysis for simultaneous determination of active components of astragali radix-safflower combination and neurotransmitters in rats with cerebral ischemia reperfusion injury: Application in pharmacokinetic and pharmacodynamic study. Phytomedicine 106, 154432 (2022).
Hou, C. X. et al. Metabolomic analysis reveals that SPHK1 promotes oral squamous cell carcinoma progression through NF-κB activation. Ann. Surg. Oncol. 29, 7386–7399 (2022).
An, W. et al. Alpinia katsumadai Hayata induces growth inhibition and autophagy-related apoptosis by regulating the AMPK and Akt/mTOR/p70S6K signaling pathways in cancer cells. Oncol. Rep. 48, 142 (2022).
Bai, P. et al. Application of 2H stable isotope labelling methodology and ultra-high performance liquid chromatography/quadrupole time-of-flight mass spectrometry for the metabolite identification of dehydroandrographolide in rats. Anal. Sci. 38, 977–988 (2022).
Zhu, C. et al. Integrated approach toward absorption, distribution, metabolism, and excretion of Xiaoke pills in zebrafish based on UPLC-HRMS and DESI-MS techniques. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 1200, 123276 (2022).
Hu, J. et al. Cardioprotective effect of ginsenoside Rb1 via regulating metabolomics profiling and AMP-activated protein kinase-dependent mitophagy. J. Ginseng Res 46, 255–265 (2022).
Kiesel, B. F. et al. Dose-dependent bioavailability, absorption-rate limited elimination, and tissue distribution of the ATR inhibitor BAY-1895344 (elimusertib) in mice. Cancer Chemother. Pharmacol. 89, 795–807 (2022).
Feng, Z. et al. The inhibition of enterocyte proliferation by lithocholic acid exacerbates necrotizing enterocolitis through downregulating the Wnt/β-catenin signalling pathway. Cell Prolif. 55, e13228 (2022).
Bai, X. et al. Recent progress on mass spectrum based approaches for absorption, distribution, metabolism, and excretion characterization of traditional Chinese medicine. Curr. Drug Metab. 23, 99–112 (2022).
Liao, M. et al. Main active components of Si-Miao-Yong-An decoction (SMYAD) attenuate autophagy and apoptosis via the PDE5A-AKT and TLR4-NOX4 pathways in isoproterenol (ISO)-induced heart failure models. Pharm. Res. 176, 106077 (2022).
Jin, Z. et al. Protective effect of Qingre Huoxue decoction against myocardial infarction via PI3K/Akt autophagy pathway based on UPLC-MS, network pharmacology, and in vivo evidence. Pharm. Biol. 59, 1607–1618 (2022).
Yuan, R. et al. Hepatoprotective effect of Sophora moorcroftiana (Benth.) Benth.Ex baker seeds in vivo and in vitro. Drug Chem. Toxicol. 45, 2535–2544 (2022).
Uçkun, E. et al. BioID-screening identifies PEAK1 and SHP2 as components of the ALK proximitome in neuroblastoma cells. J. Mol. Biol. 433, 167158 (2021).
Tong, H. et al. Bioactive constituents and the molecular mechanism of Curcumae Rhizoma in the treatment of primary dysmenorrhea based on network pharmacology and molecular docking. Phytomedicine 86, 153558 (2021).
Kang, A. et al. Characterization of the chemical constituents and in vivo metabolic profile of Scutellaria barbata D. Don by ultra high performance liquid chromatography with high-resolution mass spectrometry. J. Sep Sci. 45, 1600–1609 (2022).
Murlanova, K. et al. Antidepressant-like effects of a chlorogenic acid- and cynarine-enriched fraction from Dittrichia viscosa root extract. Sci. Rep. 12, 3647 (2022).
Bian, Y. et al. Metabolites identification and species comparison of Oroxylin A, an anti-cancer flavonoid, in vitro and in vivo by HPLC-Q-TOF-MS/MS. Xenobiotica 52, 165–176 (2022).
Irfan, A. et al. Ultrasonic-assisted synthesis of benzofuran appended oxadiazole molecules as tyrosinase inhibitors: mechanistic approach through enzyme inhibition, molecular docking, chemoinformatics, ADMET and drug-likeness studies. Int J. Mol. Sci. 23, 10979 (2022).
Wang, Z. et al. Phytochemical drug discovery for COVID-19 using high-resolution computational docking and machine learning assisted binder prediction. J. Biomol. Struct. Dyn. 2022, 1–21 (2022).
Kumar, S. & Ayyannan, S. R. Identification of new small molecule monoamine oxidase-B inhibitors through pharmacophore-based virtual screening, molecular docking and molecular dynamics simulation studies. J. Biomol. Struct. Dyn. https://doi.org/10.1080/07391102.2022.2112082 (2022).
Adinortey, C. A. et al. Molecular structure-based screening of the constituents of Calotropis procera identifies potential inhibitors of diabetes mellitus target alpha glucosidase. Curr. Issues Mol. Biol. 44, 963–987 (2022).
Moshawih, S. et al. Synergy between machine learning and natural products cheminformatics: Application to the lead discovery of anthraquinone derivatives. Chem. Biol. Drug Des. 100, 185–217 (2022).
Bandyopadhyay, S. et al. Polypharmacology of some medicinal plant metabolites against SARS-CoV-2 and host targets: Molecular dynamics evaluation of NSP9 RNA binding protein. J. Biomol. Struct. Dyn. https://doi.org/10.1080/07391102.2021.1959401 (2021).
Jin, H. et al. A unique ligand-steered strategy for CC chemokine receptor 2 homology modeling to facilitate structure-based virtual screening. Chem. Biol. Drug Des. 97, 944–961 (2022).
Soares Rodrigues, G. C. et al. Computer-assisted discovery of compounds with insecticidal activity against Musca domestica and Mythimna separata. Food Chem. Toxicol. 147, 111899 (2021).
Grygorenko, O. O. et al. Generating multibillion chemical space of readily accessible screening compounds. iScience 23, 101681 (2020).
de Souza Neto, L. R. et al. In silico strategies to support fragment-to-lead optimization in drug discovery. Front Chem. 8, 93 (2020).
Zhang, X. et al. Identification and mechanism prediction of mulberroside A metabolites in vivo and in vitro of rats using an integrated strategy of UHPLC-Q-Exactive Plus Orbitrap MS and network pharmacology. Front Chem. 10, 981173 (2022).
Wang, C. et al. Discovery of metabolic markers for the discrimination of Helwingia species based on bioactivity evaluation, plant metabolomics and network pharmacology. Rapid Commun. Mass Spectrom. 2022, e9411 (2022).
Mahana, A. et al. Integrated serum pharmacochemistry and network pharmacology analyses reveal the bioactive metabolites and potential functional mechanism of ground cherry (Physalis pruinosa L.) in treatment of type 2 diabetes mellitus in rats. J. Ethnopharmacol. 300, 115750 (2023).
Jin, Y. et al. Ginseng total saponins and Fuzi total alkaloids exert antidepressant-like effects in ovariectomized mice through BDNF-mTORC1, autophagy and peripheral metabolic pathways. Phytomedicine 107, 154425 (2022).
Zhou, J. et al. Metabolomics and integrated network pharmacology analysis reveal that ginkgolides act as potential active anticancer components by regulating one-carbon metabolism. J. Ethnopharmacol. 298, 115609 (2022).
Le, H. H. et al. Characterization of interactions of dietary cholesterol with the murine and human gut microbiome. Nat. Microbiol. 7, 1390–1403 (2022).
Fawad, J. A. et al. Histone deacetylase inhibition by gut microbe-generated short-chain fatty acids entrains intestinal epithelial circadian rhythms. Gastroenterology 163, 1377–1390 (2022).
Taraskina, A. et al. Effects of traumatic brain injury on the gut microbiota composition and serum amino acid profile in rats. Cells 11, 1409 (2022).
Daniel, N. et al. Gut microbiota and fermentation-derived branched chain hydroxy acids mediate health benefits of yogurt consumption in obese mice. Nat. Commun. 13, 1343 (2022).
Cheney, A. M. et al. Gut microbiome dysbiosis drives metabolic dysfunction in Familial dysautonomia. Nat. Commun. 14, 218 (2023).
Si, Y. et al. Comprehensive 16S rDNA sequencing and LC-MS/MS-based metabolomics to investigate intestinal flora and metabolic profiles of the serum, hypothalamus and hippocampus in p-chlorophenylalanine-induced insomnia rats treated with lilium brownie. Neurochem. Res. 47, 574–589 (2022).
Shen, H. R. et al. Berberine improves the symptoms of DHEA-induced PCOS rats by regulating gut microbiotas and metabolites. Gynecologic Obstet. Investig. 86, 388–397 (2021).
Pattnaik, S. et al. Bioactive microbial metabolites in cancer therapeutics: mining, repurposing, and their molecular targets. Curr. Microbiol. 79, 300 (2022).
van der Lelie, D. et al. Rationally designed bacterial consortia to treat chronic immune-mediated colitis and restore intestinal homeostasis. Nat. Commun. 12, 3105 (2021).
Ziętek, M. et al. Short-chain fatty acids, maternal microbiota and metabolism in pregnancy. Nutrients 13, 1244 (2021).
Kindschuh, W. F. et al. Preterm birth is associated with xenobiotics and predicted by the vaginal metabolome. Nat. Microbiol. https://doi.org/10.1038/s41564-022-01293-8 (2023).
Lahiri, S. et al. The gut microbiota influences skeletal muscle mass and function in mice. Sci. Transl. Med. 11, 5662 (2019).
Wang, X. Q. et al. Gut microbiota as important modulator of metabolism in health and disease. RSC Adv. 8, 42380–42389 (2018).
Dong, F. et al. Intestinal microbiota-derived tryptophan metabolites are predictive of Ah receptor activity. Gut microbes 12, 1–24 (2020).
Apper, E. et al. Relationships between gut microbiota, metabolome, body weight, and glucose homeostasis of obese dogs fed with diets differing in prebiotic and protein content. Microorganisms 8, 513 (2020).
Vascellari, S. et al. Gut microbiota and metabolome alterations associated with Parkinson’s disease. mSystems 5, e00561 (2020).
Westfall, S. et al. Microbiota metabolites modulate the T helper 17 to regulatory T cell (Th17/Treg) imbalance promoting resilience to stress-induced anxiety- and depressive-like behaviors. Brain, Behav., Immun. 91, 350–368 (2021).
Malczewski, A. B. et al. Microbiome-derived metabolome as a potential predictor of response to cancer immunotherapy. J. Immunother. cancer 8, e001383 (2020).
Wang, X. et al. Altered gut bacterial and metabolic signatures and their interaction in gestational diabetes mellitus. Gut Microbes 12, 1–13 (2020).
Yoshimoto, S. et al. Enriched metabolites that potentially promote age-associated diseases in subjects with an elderly-type gut microbiota. Gut Microbes 13, 1–11 (2021).
Yang, W. & Cong, Y. Gut microbiota-derived metabolites in the regulation of host immune responses and immune-related inflammatory diseases. Cell. Mol. Immunol. 18, 866–877 (2021).
Hu, X. et al. Multi-omics study reveals that statin therapy is associated with restoration of gut microbiota homeostasis and improvement in outcomes in patients with acute coronary syndrome. Theranostics 11, 5778–5793 (2021).
Husted, A. S. et al. GPCR-mediated signaling of metabolites. Cell Metab. 25, 777–796 (2017).
Onyszkiewicz, M. et al. Short chain fatty acids and methylamines produced by gut microbiota as mediators and markers in the circulatory system. Exp. Biol. Med. 245, 166–175 (2020).
Frampton, J., Murphy, K. G., Frost, G. & Chambers, E. S. Short-chain fatty acids as potential regulators of skeletal muscle metabolism and function. Nat. Metab. 2, 840–848 (2020).
Martin, A. M. et al. Mechanisms controlling hormone secretion in human gut and its relevance to metabolism. J. Endocrinol. 244, R1–R15 (2019).
Sukkar, A. H. et al. Regulation of energy expenditure and substrate oxidation by short-chain fatty acids. J. Endocrinol. 242, R1–R8 (2019).
Noguchi, M. et al. Lactic acid bacteria-derived γ-linolenic acid metabolites are PPARδ ligands that reduce lipid accumulation in human intestinal organoids. J. Biol. Chem. 2022, 102534 (2022).
Fu, L. et al. Enhancement of liver mitochondrial complex I and energy metabolism induced by enteritis: The key role of gut microbiota derived endotoxins. Front Immunol. 13, 981917 (2022).
Canfora, E. E. & Blaak, E. E. Acetate: a diet-derived key metabolite in energy metabolism: good or bad in context of obesity and glucose homeostasis? Curr. Opin. Clin. Nutr. Metab. Care. 20, 477–483 (2017).
Blaut, M. Gut microbiota and energy balance: role in obesity. Proc. Nutr. Soc. 74, 227–234 (2015).
Madella, A. M. et al. Microbial-derived tryptophan catabolites, kidney disease and gut inflammation. Toxins 14, 645 (2022).
Cai, J., Sun, L. & Gonzalez, F. J. Gut microbiota-derived bile acids in intestinal immunity, inflammation, and tumorigenesis. Cell Host Microbe 30, 289–300 (2022).
Ikeda, T. et al. Short-chain fatty acid receptors and gut microbiota as therapeutic targets in metabolic, immune, and neurological diseases. Pharmacol. therapeutics. 239, 108273 (2022).
Mutalub, Y. B. et al. Gut microbiota modulation as a novel therapeutic strategy in cardiometabolic diseases. Foods 11, 2575 (2022).
Brial, F. et al. Human and preclinical studies of the host-gut microbiome co-metabolite hippurate as a marker and mediator of metabolic health. Gut 70, 2105–2114 (2021).
Wu, S. E. et al. Microbiota-derived metabolite promotes HDAC3 activity in the gut. Nature 586, 108–112 (2020).
Yuan, Y. et al. Airway microbiome and serum metabolomics analysis identify differential candidate biomarkers in allergic rhinitis. Front Immunol. 12, 771136 (2022).
Yu, D. et al. The gut microbiome and metabolites are altered and interrelated in patients with rheumatoid arthritis. Front Cell Infect. Microbiol. 11, 763507 (2022).
Kong, C. et al. Integrated metagenomic and metabolomic analysis reveals distinct gut-microbiome-derived phenotypes in early-onset colorectal cancer. Gut 2022, 327156 (2022).
Leyrolle, Q. et al. Microbiota and metabolite profiling as markers of mood disorders: a cross-sectional study in obese patients. Nutrients 14, 147 (2021).
Yu, J. S. et al. Lactobacillus lactis and Pediococcus pentosaceus-driven reprogramming of gut microbiome and metabolome ameliorates the progression of non-alcoholic fatty liver disease. Clin. Transl. Med. 11, e634 (2021).
Wang, Z. et al. The correlation between gut microbiota and serum metabolomic in elderly patients with chronic heart failure. Mediators Inflamm. 2021, 5587428 (2021).
Chen, L. et al. Influence of the microbiome, diet and genetics on inter-individual variation in the human plasma metabolome. Nat. Med. https://doi.org/10.1038/s41591-022-02014-8 (2022).
Dekkers, K. F. et al. An online atlas of human plasma metabolite signatures of gut microbiome composition. Nat. Commun. 13, 5370 (2022).
Ishizawa, S. et al. Integrated analysis of effect of daisaikoto, a traditional Japanese medicine, on the metabolome and gut microbiome in a mouse model of nonalcoholic fatty liver disease. Gene 846, 146856 (2022).
Liu, X. et al. Comprehensive 16S rRNA sequencing based microbiomes and 1H NMR based metabolomics reveal the relationships of aging and constipation. Exp. Gerontol. 166, 111882 (2022).
Ducarmon, Q. R. et al. Gut colonisation by extended-spectrum β-lactamase-producing Escherichia coli and its association with the gut microbiome and metabolome in Dutch adults: a matched case-control study. Lancet Microbe 3, e443–e451 (2022).
Andresen, C. et al. Comparison of extraction methods for intracellular metabolomics of human tissues. Front. Mol. Biosci. 9, 932261 (2022).
Hanafi, R. S. & Lämmerhofer, M. Quality-by-design approach for development of aqueous headspace microextraction GC-MS method for targeted metabolomics of small aldehydes in plasma of cardiovascular patients. Anal Chim. acta 1221, 340176 (2022).
Zhang, N. R. et al. Validation of a multiplexed and targeted lipidomics assay for accurate quantification of lipidomes. J. lipid Res. 63, 100218 (2022).
Ramos-Lopez, O. et al. Holistic integration of omics tools for precision nutrition in health and disease. Nutrients 14, 4074 (2022).
Cuypers, B. et al. Four layer multi-omics reveals molecular responses to aneuploidy in Leishmania. PLoS Pathog. 18, e1010848 (2022).
Louca, P. et al. Machine learning integration of multimodal data identifies key features of blood pressure regulation. EBioMedicine 84, 104243 (2022).
Chou, C. H. et al. Metabolomic and transcriptomic signatures of influenza vaccine response in healthy young and older adults. Aging cell. 21, e13682 (2022).
Avalon, N. E., Murray, A. E. & Baker, B. J. Integrated metabolomic-genomic workflows accelerate microbial natural product discovery. Anal. Chem. 94, 11959–11966 (2022).
Zhang, Q. et al. Integrative analysis of multi-omics data to detect the underlying molecular mechanisms for obesity in vivo in humans. Hum. genomics. 16, 15 (2022).
Lai, M. et al. Integrating serum proteomics and metabolomics to compare the common and distinct features between acute aggressive ischemic stroke (APIS) and acute non-aggressive ischemic stroke (ANPIS). J. Proteom. 261, 104581 (2022).
Yu, C. T. et al. An evaluation of the National Institutes of Health grants portfolio: identifying opportunities and challenges for multi-omics research that leverage metabolomics data. Metabolomics 18, 29 (2022).
Grant, C. W. et al. Multi-omics characterization of early- and adult-onset major depressive disorder. J. Personalized Med. 12, 412 (2022).
Di Filippo, M. et al. INTEGRATE: Model-based multi-omics data integration to characterize multi-level metabolic regulation. PLoS Comput. Biol. 18, e1009337 (2022).
Yazd, H. S. et al. LC-MS lipidomics of renal biopsies for the diagnosis of Fabry disease. J. Mass Spectrom. Adv. Clin. lab. 22, 71–78 (2021).
Yadav, C. B. et al. Metabolite diversity and metabolic genome-wide marker association studies (Mgwas) for health benefiting nutritional traits in pearl millet grains. Cells 10, 3076 (2021).
Zhang, J. et al. Diagnostic approach to thyroid cancer based on amino acid metabolomics in saliva by ultra-performance liquid chromatography with high resolution mass spectrometry. Talanta 235, 122729 (2021).
Zang, Q. et al. Spatially resolved metabolomics combined with multicellular tumor spheroids to discover cancer tissue relevant metabolic signatures. Anal Chim. Acta 1155, 338342 (2021).
Dekker, S. et al. Urinary metabolites associate with the rate of kidney function decline in patients with autosomal dominant polycystic kidney disease. PloS one 15, e0233213 (2020).
Wang, X. & Kadarmideen, H. N. Metabolite genome-wide association study (mGWAS) and gene-metabolite interaction network analysis reveal potential biomarkers for feed efficiency in pigs. Metabolites 10, 201 (2020).
Chen, L. et al. Plasma metabolomics study of Vogt-Koyanagi-Harada disease identifies potential diagnostic biomarkers. Exp. Eye Res. 196, 108070 (2020).
Hu, Y. et al. Disturbances in metabolic pathways and the identification of a potential biomarker panel for early cartilage degeneration in a rabbit anterior cruciate ligament transection model. Cartilage 13, 1376S–1387S (2021).
Wei, J. et al. A plasma metabolomics study suggests alteration of multiple metabolic pathways in patients with bipolar disorder. Psychiatry Res. 299, 113880 (2021).
BioRender.com. Available online: https://biorender.com/.
Metaboanalyst.ca. https://www.metaboanalyst.ca/.
Acknowledgements
We are grateful for the generous support from the Program of Natural Science Foundation of State (Grant No. 81973745, 82104733, 81302905), Talent Lift Engineering Project of China Association of TCM (QNRC2-B06), Natural Science Foundation of Heilongjiang Province (YQ2019H030). We also thank BioRender843 and MetaboAnalyst844 for the figure preparation.
Author information
Authors and Affiliations
Contributions
Conception and design: Y.X., S.T., A.Z.; Development of methodology: S.Q., Y.C., A.Z.; Analysis and interpretation of data: S.Q., Y.C., H.Y., C.L., A.Z.; Writing, review, and/or revision of the manuscript: S.Q., Y.C., H.Y., C.L., Y.X., S.T., A.Z.; Technical and reporting support: S.Q., Y.C., H.Y., C.L.; Study supervision: Y.X., A.Z. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qiu, S., Cai, Y., Yao, H. et al. Small molecule metabolites: discovery of biomarkers and therapeutic targets. Sig Transduct Target Ther 8, 132 (2023). https://doi.org/10.1038/s41392-023-01399-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-023-01399-3
This article is cited by
-
An integrated metabo-lipidomics profile of induced sputum for the identification of novel biomarkers in the differential diagnosis of asthma and COPD
Journal of Translational Medicine (2024)
-
Unveiling the metabolic landscape of pulmonary hypertension: insights from metabolomics
Respiratory Research (2024)
-
Plasma metabolites and risk of seven cancers: a two-sample Mendelian randomization study among European descendants
BMC Medicine (2024)
-
Unbiasedly decoding the tumor microenvironment with single-cell multiomics analysis in pancreatic cancer
Molecular Cancer (2024)
-
Transcending frontiers in prostate cancer: the role of oncometabolites on epigenetic regulation, CSCs, and tumor microenvironment to identify new therapeutic strategies
Cell Communication and Signaling (2024)