Abstract
Background
Tumors with mutations associated with homologous recombination deficiency (HRD) are uncommon in prostate cancer (PCa) and variably responsive to PARP inhibition. To better identify tumors with HRD, we developed a transcriptomic signature for HRD in PCa (HRD-P).
Methods
By using an established mutational signature, we created and validated HRD-P in six independent PCa cohorts (primary PCa, n = 8224; metastatic castration-resistant PCa [mCRPC], n = 328). Molecular and clinical features were compared between HRD-P+ tumors and those with single HR-gene mutations.
Results
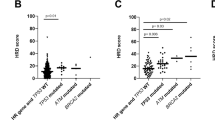
HRD-P+ tumors were more common than tumors with single HR-gene mutations in primary (201/491, 41% vs 32/491 6.5%) and mCRPC (126/328, 38% vs 82/328, 25%) cases, and HRD-P+ was more predictive of genomic instability suggestive of HRD. HRD-P+ was associated with a shorter time to recurrence following surgery and shorter overall survival in men with mCRPC. In a prospective trial of mCRPC treated with olaparib (n = 10), all three men with HRD-P+ experienced prolonged (>330 days) PSA progression-free survival.
Conclusion
These results suggest transcriptomics can identify more patients that harbor phenotypic HRD than single HR-gene mutations and support further exploration of transcriptionally defined HRD tumors perhaps in conjunction with genomic markers for therapeutic application.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 4 print issues and online access
$259.00 per year
only $64.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Castro E, Goh C, Leongamornlert D, Saunders E, Tymrakiewicz M, Dadaev T, et al. Effect of BRCA mutations on metastatic relapse and cause-specific survival after radical treatment for localised prostate cancer. Eur Urol. 2015;68:186–93.
Castro E, Romero-Laorden N, Del Pozo A, Lozano R, Medina A, Puente J, et al. PROREPAIR-B: a prospective cohort study of the impact of germline DNA repair mutations on the outcomes of patients with metastatic castration-resistant prostate cancer. J Clin Oncol. 2019;37:490–503.
Mateo J, Porta N, Bianchini D, McGovern U, Elliott T, Jones R, et al. Olaparib in patients with metastatic castration-resistant prostate cancer with DNA repair gene aberrations (TOPARP-B): a multicentre, open-label, randomised, phase 2 trial. Lancet Oncol. 2020;21:162–74.
Abeshouse A, Ahn J, Akbani R, Ally A, Amin S, Andry CD, et al. The molecular taxonomy of primary prostate cancer. Cell. 2015;163:1011–25.
de Bono J, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, et al. Olaparib for metastatic castration-resistant prostate cancer. N. Engl J Med. 2020;382:2091–102.
Lord CJ, Ashworth A. BRCAness revisited. Nat Rev Cancer. 2016;16:110–20.
Knijnenburg TA, Wang L, Zimmermann MT, Chambwe N, Gao GF, Cherniack AD, et al. Genomic and molecular landscape of dna damage repair deficiency across the cancer genome atlas. Cell Rep. 2018;23:239–254. e6
Spratt DE, Alshalalfa M, Fishbane N, Weiner AB, Mehra R, Mahal BA et al. Transcriptomic heterogeneity of androgen receptor (AR) activity defines a de novo low AR-active subclass in treatment naïve primary prostate cancer. Clin Cancer Res. 2019. https://doi.org/10.1158/1078-0432.CCR-19-1587.
Sztupinszki Z, Diossy M, Krzystanek M, Borcsok J, Pomerantz MM, Tisza V, et al. Detection of molecular signatures of homologous recombination deficiency in prostate cancer with or without BRCA1/2 mutations. Clin Cancer Res. 2020;26:2673–80.
Roychowdhury S, Iyer MK, Robinson DR, Lonigro RJ, Wu Y-M, Cao X, et al. Personalized oncology through integrative high-throughput sequencing: a pilot study. Sci Transl Med. 2011;3:111ra121–111ra121.
Davies H, Glodzik D, Morganella S, Yates LR, Staaf J, Zou X, et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat Med. 2017;23:517–25.
Peng G, Chun-Jen Lin C, Mo W, Dai H, Park Y-Y, Kim SM, et al. Genome-wide transcriptome profiling of homologous recombination DNA repair. Nat Commun. 2014;5:3361.
Hastie T, Tibshirani R, Friedman J. The Elements of Statistical Learning. 1st ed. New York: Springer; 2001.
Huang DW, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37:1–13.
Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The molecular signatures database hallmark gene set collection. Cell Syst. 2015;1:417–25.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Disco. 2012;2:401–4.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1–pl1.
Telli ML, Timms KM, Reid J, Hennessy B, Mills GB, Jensen KC, et al. Homologous recombination deficiency (HRD) score predicts response to platinum-containing neoadjuvant chemotherapy in patients with triple-negative breast cancer. Clin Cancer Res. 2016;22:3764–73.
Karnes RJ, Bergstralh EJ, Davicioni E, Ghadessi M, Buerki C, Mitra AP, et al. Validation of a genomic classifier that predicts metastasis following radical prostatectomy in an at risk patient population. J Urol. 2013;190:2047–53.
Abida W, Cyrta J, Heller G, Prandi D, Armenia J, Coleman I, et al. Genomic correlates of clinical outcome in advanced prostate cancer. Proc Natl Acad Sci USA. 2019;116:11428–36.
Konstantinopoulos PA, Spentzos D, Karlan BY, Taniguchi T, Fountzilas E, Francoeur N, et al. Gene expression profile of BRCAness that correlates with responsiveness to chemotherapy and with outcome in patients with epithelial ovarian cancer. JCO. 2010;28:3555–61.
Karnes RJ, Sharma V, Choeurng V, Ashab HA-D, Erho N, Alshalalfa M, et al. Development and validation of a prostate cancer genomic signature that predicts early ADT treatment response following radical prostatectomy. Clin Cancer Res. 2018;24:3908–16.
Larsen MJ, Kruse TA, Tan Q, Lænkholm A-V, Bak M, Lykkesfeldt AE et al. Classifications within Molecular Subtypes Enables Identification of BRCA1/BRCA2 Mutation Carriers by RNA Tumor Profiling. PLoS ONE 2013;8. https://doi.org/10.1371/journal.pone.0064268.
Yuan J, Hu Z, Mahal BA, Zhao SD, Kensler KH, Pi J, et al. Integrated analysis of genetic ancestry and genomic alterations across cancers. Cancer Cell. 2018;34:549–560.e9.
Ren B, Cam H, Takahashi Y, Volkert T, Terragni J, Young RA, et al. E2F integrates cell cycle progression with DNA repair, replication, and G2/M checkpoints. Genes Dev. 2002;16:245–56.
Abida W, Patnaik A, Campbell D, Shapiro J, Bryce AH, McDermott R et al. Rucaparib in men with metastatic castration-resistant prostate cancer harboring a BRCA1 or BRCA2 gene alteration. JCO 2020; JCO.20.01035.
Taylor RA, Fraser M, Livingstone J, Espiritu SMG, Thorne H, Huang V, et al. Germline BRCA2 mutations drive prostate cancers with distinct evolutionary trajectories. Nat Commun. 2017;8:13671.
Xu X, Qiao W, Linke SP, Cao L, Li W-M, Furth PA, et al. Genetic interactions between tumor suppressors Brca1 and p53 in apoptosis, cell cycle and tumorigenesis. Nat Genet. 2001;28:266–71.
Na R, Zheng SL, Han M, Yu H, Jiang D, Shah S, et al. Germline mutations in ATM and BRCA1/2 distinguish risk for lethal and indolent prostate cancer and are associated with early age at death. Eur Urol. 2017;71:740–7.
Lalonde E, Ishkanian AS, Sykes J, Fraser M, Ross-Adams H, Erho N, et al. Tumour genomic and microenvironmental heterogeneity for integrated prediction of 5-year biochemical recurrence of prostate cancer: a retrospective cohort study. Lancet Oncol. 2014;15:1521–32.
Funding
This work was supported in part by the National Institutes of Health grant 5U01CA196390 (EMS), the Prostate Cancer Foundation (EMS), Department of Defense grant W81XWH-15-1-0661 (EMS and TLL), the 2019 Urology Care Foundation Residency Research Award Program and the Russell Scott, Jr., MD Urology Research Fund (ABW), as well as the University of Michigan Prostate Specialized Program of Research Excellence (SPORE) P50 CA186786-05 and the Early Detection Research Network grant UO1 CA111275 (AMC). The sponsors had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
Conception and design: ABW, ED, AMC, TLL, DES. Acquisition of data: ABW, YL, MMcF, PSB, XZ, ZL, TH, MH, RJK, ED, AMC, TLL. Analysis and interpretation of data: ABW, YL, MMcF, XZ, ZL. Drafting of manuscript: ABW, YL, MMcF, PSB, EVL, XZ, ZL, TH, MH, DES, EMS. Critical revision of the manuscript for important intellectual content: ABW, R. JK, ED, ZRR, AMC, TLL, DES, EMS. Statistical analysis: ABW, YL. Obtaining funding: ABW, AMC, TLL, EMS. Administrative, technical, or material support: TLL, AMC, TLL, DES, EMS. Supervision: TLL, AMC, TLL, DES, EMS. Other: none.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The study was performed in accordance with the Declaration of Helsinki.
Competing interests
YL, XZ, ZL, ED are employees of Decipher Biosciences. AMC currently serves on the scientific advisory board of Tempus. The remaining authors declare no potential conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Weiner, A.B., Liu, Y., McFarlane, M. et al. A transcriptomic model for homologous recombination deficiency in prostate cancer. Prostate Cancer Prostatic Dis 25, 659–665 (2022). https://doi.org/10.1038/s41391-021-00416-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-021-00416-2
This article is cited by
-
High intratumoral plasma cells content in primary prostate cancer defines a subset of tumors with potential susceptibility to immune-based treatments
Prostate Cancer and Prostatic Diseases (2023)
-
Validation of genomic and transcriptomic models of homologous recombination deficiency in a real-world pan-cancer cohort
BMC Cancer (2022)