Abstract
Background
The androgen receptor (AR) is a key prostate cancer drug target. Suppression of AR signaling mediated by the full-length AR (AR-FL) is the therapeutic goal of all existing AR-directed therapies. AR-targeting agents impart therapeutic benefit, but lead to AR aberrations that underlie disease progression and therapeutic resistance. Among the AR aberrations specific to castration-resistant prostate cancer (CRPC), AR variants (AR-Vs) have emerged as important indicators of disease progression and therapeutic resistance.
Methods
We conducted a systemic review of the literature focusing on recent laboratory studies on AR-Vs following our last review article published in 2016. Topics ranged from measurement and detection, molecular origin, regulation, genomic function, and preclinical therapeutic targeting of AR-Vs. We provide expert opinions and perspectives on these topics.
Results
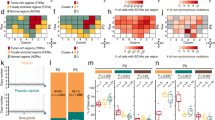
Transcript sequences for 22 AR-Vs have been reported in the literature. Different AR-Vs may arise through different mechanisms, and can be regulated by splicing factors and dictated by genomic rearrangements, but a low-androgen environment is a prerequisite for generation of AR-Vs. The unique transcript structures allowed development of in situ and in-solution measurement and detection methods, including mRNA and protein detection, in both tissue and blood specimens. AR-V7 remains the main measurement target and the most extensively characterized AR-V. Although AR-V7 coexists with AR-FL, genomic functions mediated by AR-V7 do not require the presence of AR-FL. The distinct cistromes and transcriptional programs directed by AR-V7 and their coregulators are consistent with genomic features of progressive disease in a low-androgen environment. Preclinical development of AR-V-directed agents currently focuses on suppression of mRNA expression and protein degradation as well as targeting of the amino-terminal domain.
Conclusions
Current literature continues to support AR-Vs as biomarkers and therapeutic targets in prostate cancer. Laboratory investigations reveal both challenges and opportunities in targeting AR-Vs to overcome resistance to current AR-directed therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 4 print issues and online access
$259.00 per year
only $64.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Huggins C, Hodges CV. Studies on prostatic cancer: I. The effect of castration, of estrogen and of androgen injection on serum phosphatases in metastatic carcinoma of the prostate. J Urol. 2002;168:9–12.
Chen Y, Sawyers CL, Scher HI. Targeting the androgen receptor pathway in prostate cancer. Curr Opin Pharm. 2008;8:440–8.
Attard G, Cooper CS, de Bono JS. Steroid hormone receptors in prostate cancer: a hard habit to break? Cancer Cell. 2009;16:458–62.
Tran C, Ouk S, Clegg NJ, Chen Y, Watson PA, Arora V, et al. Development of a second-generation antiandrogen for treatment of advanced prostate cancer. Science. 2009;324:787–90.
de Bono JS, Logothetis CJ, Molina A, Fizazi K, North S, Chu L, et al. Abiraterone and increased survival in metastatic prostate cancer. N Engl J Med. 2011;364:1995–2005.
Scher HI, Fizazi K, Saad F, Taplin ME, Sternberg CN, Miller K, et al. Increased survival with enzalutamide in prostate cancer after chemotherapy. N Engl J Med. 2012;367:1187–97.
Ryan CJ, Smith MR, de Bono JS, Molina A, Logothetis CJ, de Souza P, et al. Abiraterone in metastatic prostate cancer without previous chemotherapy. N Engl J Med. 2013;368:138–48.
Beer TM, Armstrong AJ, Rathkopf DE, Loriot Y, Sternberg CN, Higano CS, et al. Enzalutamide in metastatic prostate cancer before chemotherapy. N Engl J Med. 2014;371:424–33.
Fizazi K, Tran N, Fein L, Matsubara N, Rodriguez-Antolin A, Alekseev BY, et al. Abiraterone plus prednisone in metastatic, castration-sensitive prostate cancer. N Engl J Med. 2017;377:352–60.
Hussain M, Fizazi K, Saad F, Rathenborg P, Shore N, Ferreira U, et al. Enzalutamide in men with nonmetastatic, castration-resistant prostate cancer. N Engl J Med. 2018;378:2465–74.
Fizazi K, Shore N, Tammela TL, Ulys A, Vjaters E, Polyakov S, et al. Darolutamide in nonmetastatic, castration-resistant prostate cancer. N Engl J Med. 2019;380:1235–46.
Smith MR, Saad F, Chowdhury S, Oudard S, Hadaschik BA, Graff JN, et al. Apalutamide treatment and metastasis-free survival in prostate cancer. N Engl J Med. 2018;378:1408–18.
Armstrong AJ, Halabi S, Luo J, Nanus DM, Giannakakou P, Szmulewitz RZ, et al. Prospective multicenter validation of androgen receptor splice variant 7 and hormone therapy resistance in high-risk castration-resistant prostate cancer: the PROPHECY Study. J Clin Oncol. 2019;37:1120–9.
Khalaf DJ, Annala M, Taavitsainen S, Finch DL, Oja C, Vergidis J, et al. Optimal sequencing of enzalutamide and abiraterone acetate plus prednisone in metastatic castration-resistant prostate cancer: a multicentre, randomised, open-label, phase 2, crossover trial. Lancet Oncol. 2019;20:1730–9.
Penson DF, Armstrong AJ, Concepcion R, Agarwal N, Olsson C, Karsh L, et al. Enzalutamide versus bicalutamide in castration-resistant prostate cancer: the STRIVE Trial. J Clin Oncol. 2016;34:2098–106.
Smith MR, Antonarakis ES, Ryan CJ, Berry WR, Shore ND, Liu G, et al. Phase 2 study of the safety and antitumor activity of apalutamide (ARN-509), a potent androgen receptor antagonist, in the High-risk Nonmetastatic Castration-resistant Prostate Cancer Cohort. Eur Urol. 2016;70:963–70.
Fizazi K, Massard C, Bono P, Kataja V, James N, Tammela TL, et al. Safety and antitumour activity of ODM-201 (BAY-1841788) in castration-resistant, CYP17 Inhibitor-naive prostate cancer: results from extended follow-up of the ARADES Trial. Eur Urol focus. 2017;3:606–14.
James ND, de Bono JS, Spears MR, Clarke NW, Mason MD, Dearnaley DP, et al. Abiraterone for prostate cancer not previously treated with hormone therapy. N Engl J Med. 2017;377:338–51.
Fizazi K, Tran N, Fein L, Matsubara N, Rodriguez-Antolin A, Alekseev BY, et al. Abiraterone acetate plus prednisone in patients with newly diagnosed high-risk metastatic castration-sensitive prostate cancer (LATITUDE): final overall survival analysis of a randomised, double-blind, phase 3 trial. Lancet Oncol. 2019;20:686–700.
Ryan CJ, Crawford ED, Shore ND, Underwood W 3rd, Taplin ME, Londhe A, et al. The IMAAGEN Study: effect of abiraterone acetate and prednisone on prostate specific antigen and radiographic disease progression in patients with nonmetastatic castration resistant prostate cancer. J Urol. 2018;200:344–52.
Small EJ, Saad F, Chowdhury S, Oudard S, Hadaschik BA, Graff JN, et al. Apalutamide and overall survival in non-metastatic castration-resistant prostate cancer. Ann Oncol. 2019;30:1813–20.
Tombal B, Saad F, Penson D, Hussain M, Sternberg CN, Morlock R, et al. Patient-reported outcomes following enzalutamide or placebo in men with non-metastatic, castration-resistant prostate cancer (PROSPER): a multicentre, randomised, double-blind, phase 3 trial. Lancet Oncol. 2019;20:556–69.
Lu C, Luo J. Decoding the androgen receptor splice variants. Transl Androl Urol. 2013;2:178–86.
Antonarakis ES, Armstrong AJ, Dehm SM, Luo J. Androgen receptor variant-driven prostate cancer: clinical implications and therapeutic targeting. Prostate Cancer Prostatic Dis. 2016;19:231–41.
Luo J, Attard G, Balk SP, Bevan C, Burnstein K, Cato L, et al. Role of androgen receptor variants in prostate cancer: report from the 2017 Mission Androgen Receptor Variants Meeting. Eur Urol. 2018;73:715–23.
Ware KE, Garcia-Blanco MA, Armstrong AJ, Dehm SM. Biologic and clinical significance of androgen receptor variants in castration resistant prostate cancer. Endocr Relat Cancer. 2014;21:T87–T103.
Antonarakis ES, Lu C, Luber B, Wang H, Chen Y, Nakazawa M, et al. Androgen receptor splice variant 7 and efficacy of taxane chemotherapy in patients with metastatic castration-resistant prostate cancer. JAMA Oncol. 2015;1:582–91.
Lokhandwala PM, Riel SL, Haley L, Lu C, Chen Y, Silberstein J, et al. Analytical validation of androgen receptor splice variant 7 detection in a Clinical Laboratory Improvement Amendments (CLIA) Laboratory Setting. J Mol Diagn. 2017;19:115–25.
Markowski MC, Silberstein JL, Eshleman JR, Eisenberger MA, Luo J, Antonarakis ES. Clinical utility of CLIA-Grade AR-V7 testing in patients with metastatic castration-resistant prostate cancer. JCO Precis Oncol. 2017;1:1–9.
Taplin ME, Antonarakis ES, Ferrante KJ, Horgan K, Blumenstein B, Saad F, et al. Androgen receptor modulation optimized for response-splice variant: a phase 3, randomized trial of galeterone versus enzalutamide in androgen receptor splice variant-7-expressing metastatic castration-resistant prostate cancer. Eur Urol. 2019;76:843–51.
Miyamoto DT, Lee RJ, Kalinich M, LiCausi JA, Zheng Y, Chen T, et al. An RNA-based digital circulating tumor cell signature is predictive of drug response and early dissemination in prostate cancer. Cancer Discov. 2018;8:288–303.
Ma Y, Luk A, Young FP, Lynch D, Chua W, Balakrishnar B, et al. Droplet digital PCR based androgen receptor variant 7 (AR-V7) detection from prostate cancer patient blood biopsies. Int J Mol Sci. 2016;17:E1264.
Markou A, Lazaridou M, Paraskevopoulos P, Chen S, Swierczewska M, Budna J, et al. Multiplex gene expression profiling of in vivo isolated circulating tumor cells in high-risk prostate cancer patients. Clin Chem. 2018;64:297–306.
El-Heliebi A, Hille C, Laxman N, Svedlund J, Haudum C, Ercan E, et al. In situ detection and quantification of AR-V7, AR-FL, PSA, and KRAS point mutations in circulating tumor cells. Clin Chem. 2018;64:536–46.
Theil G, Fischer K, Weber E, Medek R, Hoda R, Lucke K, et al. The use of a new CellCollector to isolate circulating tumor cells from the blood of patients with different stages of prostate cancer and clinical outcomes—a proof-of-concept study. PLoS ONE. 2016;11:e0158354.
Vermesh O, Aalipour A, Ge TJ, Saenz Y, Guo Y, Alam IS, et al. An intravascular magnetic wire for the high-throughput retrieval of circulating tumour cells in vivo. Nat Biomed Eng. 2018;2:696–705.
Skerenova M, Mikulova V, Capoun O, Svec D, Kolostova K, Soukup V, et al. Gene expression analysis of immunomagnetically enriched circulating tumor cell fraction in castration-resistant prostate cancer. Mol Diagn Ther. 2018;22:381–90.
Seitz AK, Thoene S, Bietenbeck A, Nawroth R, Tauber R, Thalgott M, et al. AR-V7 in peripheral whole blood of patients with castration-resistant prostate cancer: association with treatment-specific outcome under abiraterone and enzalutamide. Eur Urol. 2017;72:828–34.
Todenhofer T, Azad A, Stewart C, Gao J, Eigl BJ, Gleave ME, et al. AR-V7 transcripts in whole blood RNA of patients with metastatic castration resistant prostate cancer correlate with response to abiraterone acetate. J Urol. 2017;197:135–42.
Liu X, Ledet E, Li D, Dotiwala A, Steinberger A, Feibus A, et al. A whole blood assay for AR-V7 and AR(v567es) in patients with prostate cancer. J Urol. 2016;196:1758–63.
Takeuchi T, Okuno Y, Hattori-Kato M, Zaitsu M, Mikami K. Detection of AR-V7 mRNA in whole blood may not predict the effectiveness of novel endocrine drugs for castration-resistant prostate cancer. Res Rep Urol. 2016;8:21–25.
Walecki M, Eisel F, Klug J, Baal N, Paradowska-Dogan A, Wahle E, et al. Androgen receptor modulates Foxp3 expression in CD4+CD25+Foxp3+ regulatory T-cells. Mol Biol Cell. 2015;26:2845–57.
Ashcroft GS, Mills SJ. Androgen receptor-mediated inhibition of cutaneous wound healing. f. 2002;110:615–24.
Del ReM, Biasco E, Crucitta S, Derosa L, Rofi E, Orlandini C, et al. The Detection of Androgen Receptor Splice Variant 7 in Plasma-derived Exosomal RNA Strongly Predicts Resistance to Hormonal Therapy in Metastatic Prostate Cancer Patients. Eur Urol. 2017;71:680–7.
Woo HK, Park J, Ku JY, Lee CH, Sunkara V, Ha HK, et al. Urine-based liquid biopsy: non-invasive and sensitive AR-V7 detection in urinary EVs from patients with prostate cancer. Lab Chip. 2018;19:87–97.
Guedes LB, Morais CL, Almutairi F, Haffner MC, Zheng Q, Isaacs JT, et al. Analytic Validation of RNA In Situ Hybridization (RISH) for AR and AR-V7 Expression in Human Prostate Cancer. Clin Cancer Res. 2016;22:4651–63.
Saylor PJ, Lee RJ, Arora KS, Deshpande V, Hu R, Olivier K, et al. Branched Chain RNA In Situ Hybridization for Androgen Receptor Splice Variant AR-V7 as a Prognostic Biomarker for Metastatic Castration-Sensitive Prostate Cancer. Clin Cancer Res. 2017;23:363–9.
Zhu Y, Sharp A, Anderson CM, Silberstein JL, Taylor M, Lu C, et al. Novel Junction-specific and Quantifiable In Situ Detection of AR-V7 and its Clinical Correlates in Metastatic Castration-resistant Prostate Cancer. Eur Urol. 2018;73:727–35.
Scher HI, Graf RP, Schreiber NA, McLaughlin B, Lu D, Louw J, et al. Nuclear-specific AR-V7 Protein Localization is Necessary to Guide Treatment Selection in Metastatic Castration-resistant Prostate Cancer. Eur Urol. 2017;71:874–82.
Sharp A, Coleman I, Yuan W, Sprenger C, Dolling D, Rodrigues DN, et al. Androgen receptor splice variant-7 expression emerges with castration resistance in prostate cancer. J Clin Investig. 2019;129:192–208.
Welti J, Rodrigues DN, Sharp A, Sun S, Lorente D, Riisnaes R, et al. Analytical validation and clinical qualification of a new immunohistochemical assay for androgen receptor splice variant-7 protein expression in metastatic castration-resistant prostate cancer. Eur Urol. 2016;70:599–608.
Zhao P, Zhu Y, Cheng L, Luo J. Detection of androgen receptor (AR) and AR-V7 in small cell prostate carcinoma: diagnostic and therapeutic implications. Asian J Urol. 2019;6:109–13.
Li H, Wang Z, Xiao W, Yan L, Guan W, Hu Z, et al. Androgen-receptor splice variant-7-positive prostate cancer: a novel molecular subtype with markedly worse androgen-deprivation therapy outcomes in newly diagnosed patients. Mod Pathol. 2018;31:198–208.
Scher HI, Graf RP, Schreiber NA, Jayaram A, Winquist E, McLaughlin B, et al. Assessment of the validity of nuclear-localized androgen receptor splice variant 7 in circulating tumor cells as a predictive biomarker for castration-resistant prostate cancer. JAMA Oncol. 2018;4:1179–86.
Sharp A, Welti JC, Lambros MBK, Dolling D, Rodrigues DN, Pope L, et al. Clinical utility of circulating tumour cell androgen receptor splice variant-7 status in metastatic castration-resistant prostate cancer. Eur Urol. 2019;76:676–85.
Yang RK, Zhao P, Lu C, Luo J, Hu R. Expression pattern of androgen receptor and AR-V7 in androgen-deprivation therapy-naive salivary duct carcinomas. Hum Pathol. 2019;84:173–82.
Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;162:454.
Hu R, Dunn TA, Wei S, Isharwal S, Veltri RW, Humphreys E, et al. Ligand-independent androgen receptor variants derived from splicing of cryptic exons signify hormone-refractory prostate cancer. Cancer Res. 2009;69:16–22.
Cancer Genome Atlas Research N. The molecular taxonomy of primary prostate cancer. Cell. 2015;163:1011–25.
Henzler C, Li Y, Yang R, McBride T, Ho Y, Sprenger C, et al. Truncation and constitutive activation of the androgen receptor by diverse genomic rearrangements in prostate cancer. Nat Commun. 2016;7:13668.
De Laere B, van Dam PJ, Whitington T, Mayrhofer M, Diaz EH, Van den Eynden G, et al. Comprehensive profiling of the androgen receptor in liquid biopsies from castration-resistant prostate cancer reveals novel intra-AR structural variation and splice variant expression patterns. Eur Urol. 2017;72:192–200.
Kallio HML, Hieta R, Latonen L, Brofeldt A, Annala M, Kivinummi K, et al. Constitutively active androgen receptor splice variants AR-V3, AR-V7 and AR-V9 are co-expressed in castration-resistant prostate cancer metastases. Br J Cancer. 2018;119:347–56.
Kohli M, Ho Y, Hillman DW, Van Etten JL, Henzler C, Yang R, et al. Androgen receptor variant AR-V9 is coexpressed with AR-V7 in prostate cancer metastases and predicts abiraterone resistance. Clin Cancer Res. 2017;23:4704–15.
Hu R, Lu C, Mostaghel EA, Yegnasubramanian S, Gurel M, Tannahill C, et al. Distinct transcriptional programs mediated by the ligand-dependent full-length androgen receptor and its splice variants in castration-resistant prostate cancer. Cancer Res. 2012;72:3457–62.
Liu LL, Xie N, Sun S, Plymate S, Mostaghel E, Dong X. Mechanisms of the androgen receptor splicing in prostate cancer cells. Oncogene. 2014;33:3140–50.
Nakata D, Nakayama K, Masaki T, Tanaka A, Kusaka M, Watanabe T. Growth inhibition by testosterone in an androgen receptor splice variant-driven prostate cancer model. Prostate. 2016;76:1536–45.
Teply BA, Wang H, Luber B, Sullivan R, Rifkind I, Bruns A, et al. Bipolar androgen therapy in men with metastatic castration-resistant prostate cancer after progression on enzalutamide: an open-label, phase 2, multicohort study. Lancet Oncol. 2018;19:76–86.
Watson PA, Chen YF, Balbas MD, Wongvipat J, Socci ND, Viale A, et al. Constitutively active androgen receptor splice variants expressed in castration-resistant prostate cancer require full-length androgen receptor. Proc Natl Acad Sci USA. 2010;107:16759–65.
Yu Z, Chen S, Sowalsky AG, Voznesensky OS, Mostaghel EA, Nelson PS, et al. Rapid induction of androgen receptor splice variants by androgen deprivation in prostate cancer. Clin Cancer Res. 2014;20:1590–1600.
Zhu Y, Luo J. Regulation of androgen receptor variants in prostate cancer. Asian J Urol. 2020. https://doi.org/10.1016/j.ajur.2020.01.001.
Nadiminty N, Tummala R, Liu C, Lou W, Evans CP, Gao AC. NF-kappaB2/p52:c-Myc:hnRNPA1 pathway regulates expression of androgen receptor splice variants and enzalutamide sensitivity in prostate cancer. Mol Cancer Ther. 2015;14:1884–95.
Zhang Z, Zhou N, Huang J, Ho TT, Zhu Z, Qiu Z, et al. Regulation of androgen receptor splice variant AR3 by PCGEM1. Oncotarget. 2016;7:15481–91.
Ho TT, Huang J, Zhou N, Zhang Z, Koirala P, Zhou X, et al. Regulation of PCGEM1 by p54/nrb in prostate cancer. Sci Rep. 2016;6:34529.
Wang B, Lo UG, Wu K, Kapur P, Liu X, Huang J, et al. Developing new targeting strategy for androgen receptor variants in castration resistant prostate cancer. Int J Cancer. 2017;141:2121–30.
Fan L, Zhang F, Xu S, Cui X, Hussain A, Fazli L, et al. Histone demethylase JMJD1A promotes alternative splicing of AR variant 7 (AR-V7) in prostate cancer cells. Proc Natl Acad Sci USA. 2018;115:E4584–93.
Yamane K, Toumazou C, Tsukada Y, Erdjument-Bromage H, Tempst P, Wong J, et al. JHDM2A, a JmjC-containing H3K9 demethylase, facilitates transcription activation by androgen receptor. Cell. 2006;125:483–95.
Fan L, Peng G, Sahgal N, Fazli L, Gleave M, Zhang Y, et al. Regulation of c-Myc expression by the histone demethylase JMJD1A is essential for prostate cancer cell growth and survival. Oncogene. 2016;35:2441–52.
Wilson S, Fan L, Sahgal N, Qi J, Filipp FV. The histone demethylase KDM3A regulates the transcriptional program of the androgen receptor in prostate cancer cells. Oncotarget. 2017;8:30328–43.
Nakata D, Nakao S, Nakayama K, Araki S, Nakayama Y, Aparicio S, et al. The RNA helicase DDX39B and its paralog DDX39A regulate androgen receptor splice variant AR-V7 generation. Biochem Biophys Res Commun. 2017;483:271–6.
Tummala R, Nadiminty N, Lou W, Evans CP, Gao AC. Lin28 induces resistance to anti-androgens via promotion of AR splice variant generation. Prostate. 2016;76:445–55.
Wang R, Sun Y, Li L, Niu Y, Lin W, Lin C, et al. Preclinical study using Malat1 small interfering RNA or androgen receptor splicing variant 7 degradation enhancer ASC-J9((R)) to suppress enzalutamide-resistant prostate cancer progression. Eur Urol. 2017;72:835–44.
Szafran AT, Stephan C, Bolt M, Mancini MG, Marcelli M, Mancini MA. High-content screening identifies Src family kinases as potential regulators of AR-V7 expression and androgen-independent cell growth. Prostate. 2017;77:82–93.
Mahajan K, Malla P, Lawrence HR, Chen Z, Kumar-Sinha C, Malik R, et al. ACK1/TNK2 regulates histone H4 Tyr88-phosphorylation and AR gene expression in castration-resistant prostate cancer. Cancer Cell. 2017;31:790–803.e798.
Shiota M, Fujimoto N, Imada K, Yokomizo A, Itsumi M, Takeuchi A, et al. Potential role for YB-1 in castration-resistant prostate cancer and resistance to enzalutamide through the androgen receptor V7. J Natl Cancer Inst. 2016;108:1–10.
Udayakumar TS, Stoyanova R, Shareef MM, Mu Z, Philip S, Burnstein KL, et al. Edelfosine promotes apoptosis in androgen-deprived prostate tumors by increasing ATF3 and inhibiting androgen receptor activity. Mol Cancer Ther. 2016;15:1353–63.
Wang J, Zou JX, Xue X, Cai D, Zhang Y, Duan Z, et al. ROR-gamma drives androgen receptor expression and represents a therapeutic target in castration-resistant prostate cancer. Nat Med. 2016;22:488–96.
Wu G, Sun Y, Xiang Z, Wang K, Liu B, Xiao G, et al. Preclinical study using circular RNA 17 and micro RNA 181c-5p to suppress the enzalutamide-resistant prostate cancer progression. Cell Death Dis. 2019;10:37.
Cai L, Tsai YH, Wang P, Wang J, Li D, Fan H, et al. ZFX mediates non-canonical oncogenic functions of the androgen receptor splice variant 7 in castrate-resistant prostate cancer. Mol Cell. 2018;72:341–54.e346.
Chen Z, Wu D, Thomas-Ahner JM, Lu C, Zhao P, Zhang Q, et al. Diverse AR-V7 cistromes in castration-resistant prostate cancer are governed by HoxB13. Proc Natl Acad Sci USA. 2018;115:6810–5.
Cato L, de Tribolet-Hardy J, Lee I, Rottenberg JT, Coleman I, Melchers D, et al. ARv7 represses tumor-suppressor genes in castration-resistant prostate cancer. Cancer Cell. 2019;35:401–13.e6.
He Y, Lu J, Ye Z, Hao S, Wang L, Kohli M, et al. Androgen receptor splice variants bind to constitutively open chromatin and promote abiraterone-resistant growth of prostate cancer. Nucleic Acids Res. 2018;46:1895–911.
Cai C, He HH, Chen S, Coleman I, Wang H, Fang Z, et al. Androgen receptor gene expression in prostate cancer is directly suppressed by the androgen receptor through recruitment of lysine-specific demethylase 1. Cancer Cell. 2011;20:457–71.
Cao B, Qi Y, Zhang G, Xu D, Zhan Y, Alvarez X, et al. Androgen receptor splice variants activating the full-length receptor in mediating resistance to androgen-directed therapy. Oncotarget. 2014;5:1646–56.
Xu D, Zhan Y, Qi Y, Cao B, Bai S, Xu W, et al. Androgen receptor splice variants dimerize to transactivate target genes. Cancer Res. 2015;75:3663–71.
Purushottamachar P, Kwegyir-Afful AK, Martin MS, Ramamurthy VP, Ramalingam S, Njar VC. Identification of novel steroidal androgen receptor degrading agents inspired by galeterone 3beta-imidazole carbamate. ACS Med Chem Lett. 2016;7:708–13.
Yang YC, Banuelos CA, Mawji NR, Wang J, Kato M, Haile S, et al. Targeting androgen receptor activation function-1 with EPI to overcome resistance mechanisms in castration-resistant prostate cancer. Clin Cancer Res. 2016;22:4466–77.
Brand LJ, Olson ME, Ravindranathan P, Guo H, Kempema AM, Andrews TE, et al. EPI-001 is a selective peroxisome proliferator-activated receptor-gamma modulator with inhibitory effects on androgen receptor expression and activity in prostate cancer. Oncotarget. 2015;6:3811–24.
Shimizu Y, Tamada S, Kato M, Hirayama Y, Takeyama Y, Iguchi T, et al. Androgen receptor splice variant 7 drives the growth of castration resistant prostate cancer without being involved in the efficacy of taxane chemotherapy. J Clin Med. 2018;7:E444.
Moigne RL, Banuelos CA, Mawji NR, Tam T, Wang J, Jian K, et al. EPI-7386 is a novel N-terminal domain androgen receptor inhibitor for the treatment of prostate cancer. Ann Oncol. 2019;30(Suppl 5):v159–93.
Goicochea NL, Garnovskaya M, Blanton MG, Chan G, Weisbart R, Lilly MB. Development of cell-penetrating bispecific antibodies targeting the N-terminal domain of androgen receptor for prostate cancer therapy. Protein Eng Des Sel. 2017;30:785–93.
Lai KP, Huang CK, Chang YJ, Chung CY, Yamashita S, Li L, et al. New therapeutic approach to suppress castration-resistant prostate cancer using ASC-J9 via targeting androgen receptor in selective prostate cells. Am J Pathol. 2013;182:460–73.
Cheng MA, Chou FJ, Wang K, Yang R, Ding J, Zhang Q, et al. Androgen receptor (AR) degradation enhancer ASC-J9((R)) in an FDA-approved formulated solution suppresses castration resistant prostate cancer cell growth. Cancer Lett. 2018;417:182–91.
Li H, Hassona MD, Lack NA, Axerio-Cilies P, Leblanc E, Tavassoli P, et al. Characterization of a new class of androgen receptor antagonists with potential therapeutic application in advanced prostate cancer. Mol Cancer Ther. 2013;12:2425–35.
Dalal K, Morin H, Ban F, Shepherd A, Fernandez M, Tam KJ, et al. Small molecule-induced degradation of the full length and V7 truncated variant forms of human androgen receptor. Eur J Med Chem. 2018;157:1164–73.
Ponnusamy S, Coss CC, Thiyagarajan T, Watts K, Hwang DJ, He Y, et al. Novel selective agents for the degradation of androgen receptor variants to treat castration-resistant prostate cancer. Cancer Res. 2017;77:6282–98.
Hwang DJ, He Y, Ponnusamy S, Mohler ML, Thiyagarajan T, McEwan IJ, et al. New generation of selective androgen receptor degraders: our initial design, synthesis, and biological evaluation of new compounds with enzalutamide-resistant prostate cancer activity. J Med Chem. 2019;62:491–511.
Neklesa TK, Winkler JD, Crews CM. Targeted protein degradation by PROTACs. Pharm Ther. 2017;174:138–44.
Rodriguez-Gonzalez A, Cyrus K, Salcius M, Kim K, Crews CM, Deshaies RJ, et al. Targeting steroid hormone receptors for ubiquitination and degradation in breast and prostate cancer. Oncogene. 2008;27:7201–11.
Han X, Wang C, Qin C, Xiang W, Fernandez-Salas E, Yang CY, et al. Discovery of ARD-69 as a highly potent proteolysis targeting chimera (PROTAC) degrader of androgen receptor (AR) for the treatment of prostate cancer. J Med Chem. 2019;62:941–64.
Salami J, Alabi S, Willard RR, Vitale NJ, Wang J, Dong H, et al. Androgen receptor degradation by the proteolysis-targeting chimera ARCC-4 outperforms enzalutamide in cellular models of prostate cancer drug resistance. Commun Biol. 2018;1:100.
Neklesa T, Snyder LB, Willard RR, Vitale N, Pizzano J, Gordon DA, et al. ARV-110: an oral androgen receptor PROTAC degrader for prostate cancer. J Clin Oncol. 2019;37(Suppl 7):259–259.
Raina K, Lu J, Qian Y, Altieri M, Gordon D, Rossi AM, et al. PROTAC-induced BET protein degradation as a therapy for castration-resistant prostate cancer. Proc Natl Acad Sci USA. 2016;113:7124–9.
Zhang X, Castanotto D, Nam S, Horne D, Stein C. 6BIO enhances oligonucleotide activity in cells: a potential combinatorial anti-androgen receptor therapy in prostate cancer cells. Mol Ther. 2017;25:79–91.
Wadosky KM, Shourideh M, Goodrich DW, Koochekpour S. Riluzole induces AR degradation via endoplasmic reticulum stress pathway in androgen-dependent and castration-resistant prostate cancer cells. Prostate. 2019;79:140–50.
Singh KB, Ji X, Singh SV. Therapeutic potential of leelamine, a novel inhibitor of androgen receptor and castration-resistant prostate cancer. Mol Cancer Ther. 2018;17:2079–90.
Moses MA, Kim YS, Rivera-Marquez GM, Oshima N, Watson MJ, Beebe KE, et al. Targeting the Hsp40/Hsp70 chaperone axis as a novel strategy to treat castration-resistant prostate cancer. Cancer Res. 2018;78:4022–35.
Kita K, Shiota M, Tanaka M, Otsuka A, Matsumoto M, Kato M, et al. Heat shock protein 70 inhibitors suppress androgen receptor expression in LNCaP95 prostate cancer cells. Cancer Sci. 2017;108:1820–7.
Ferraldeschi R, Welti J, Powers MV, Yuan W, Smyth T, Seed G, et al. Second-generation HSP90 inhibitor onalespib blocks mRNA splicing of androgen receptor variant 7 in prostate cancer cells. Cancer Res. 2016;76:2731–42.
Welti J, Sharp A, Yuan W, Dolling D, Nava Rodrigues D, Figueiredo I, et al. Targeting bromodomain and extra-terminal (BET) family proteins in castration-resistant prostate cancer (CRPC). Clin Cancer Res. 2018;24:3149–62.
Asangani IA, Dommeti VL, Wang X, Malik R, Cieslik M, Yang R, et al. Therapeutic targeting of BET bromodomain proteins in castration-resistant prostate cancer. Nature. 2014;510:278–82.
Yang L, Zhang Y, Shan W, Hu Z, Yuan J, Pi J, et al. Repression of BET activity sensitizes homologous recombination-proficient cancers to PARP inhibition. Sci Transl Med. 2017;9:eaal1645.
Zhang P, Wang D, Zhao Y, Ren S, Gao K, Ye Z, et al. Intrinsic BET inhibitor resistance in SPOP-mutated prostate cancer is mediated by BET protein stabilization and AKT-mTORC1 activation. Nat Med. 2017;23:1055–62.
Dai X, Gan W, Li X, Wang S, Zhang W, Huang L, et al. Prostate cancer-associated SPOP mutations confer resistance to BET inhibitors through stabilization of BRD4. Nat Med. 2017;23:1063–71.
Malvaez M, McQuown SC, Rogge GA, Astarabadi M, Jacques V, Carreiro S, et al. HDAC3-selective inhibitor enhances extinction of cocaine-seeking behavior in a persistent manner. Proc Natl Acad Sci USA. 2013;110:2647–52.
McLeod AB, Stice JP, Wardell SE, Alley HM, Chang CY, McDonnell DP. Validation of histone deacetylase 3 as a therapeutic target in castration-resistant prostate cancer. Prostate. 2018;78:266–77.
Varambally S, Dhanasekaran SM, Zhou M, Barrette TR, Kumar-Sinha C, Sanda MG, et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature. 2002;419:624–9.
Xu K, Wu ZJ, Groner AC, He HH, Cai C, Lis RT, et al. EZH2 oncogenic activity in castration-resistant prostate cancer cells is Polycomb-independent. Science. 2012;338:1465–9.
Kim J, Lee Y, Lu X, Song B, Fong KW, Cao Q, et al. Polycomb- and methylation-independent roles of EZH2 as a transcription activator. Cell Rep. 2018;25:2808–20.e2804.
Wu C, Jin X, Yang J, Yang Y, He Y, Ding L, et al. Inhibition of EZH2 by chemo- and radiotherapy agents and small molecule inhibitors induces cell death in castration-resistant prostate cancer. Oncotarget. 2016;7:3440–52.
Murashima A, Shinjo K, Katsushima K, Onuki T, Kondoh Y, Osada H, et al. Identification of a chemical modulator of EZH2-mediated silencing by cell-based high-throughput screening assay. J Biochem. 2019;166:41–50.
Ku SY, Rosario S, Wang Y, Mu P, Seshadri M, Goodrich ZW, et al. Rb1 and Trp53 cooperate to suppress prostate cancer lineage plasticity, metastasis, and antiandrogen resistance. Science. 2017;355:78–83.
Ma L, Yan Y, Bai Y, Yang Y, Pan Y, Gang X, et al. Overcoming EZH2 inhibitor resistance by taxane in PTEN-Mutated cancer. Theranostics. 2019;9:5020–34.
Liu Q, Wang G, Li Q, Jiang W, Kim JS, Wang R, et al. Polycomb group proteins EZH2 and EED directly regulate androgen receptor in advanced prostate cancer. Int J Cancer. 2019;145:415–26.
Lai CJ, Bao R, Tao X, Wang J, Atoyan R, Qu H, et al. CUDC-101, a multitargeted inhibitor of histone deacetylase, epidermal growth factor receptor, and human epidermal growth factor receptor 2, exerts potent anticancer activity. Cancer Res. 2010;70:3647–56.
Sun H, Mediwala SN, Szafran AT, Mancini MA, Marcelli M. CUDC-101, a novel inhibitor of full-length androgen receptor (flAR) and androgen receptor variant 7 (AR-V7) activity: mechanism of action and in vivo efficacy. Horm Cancer. 2016;7:196–210.
Ianculescu I, Wu DY, Siegmund KD, Stallcup MR. Selective roles for cAMP response element-binding protein binding protein and p300 protein as coregulators for androgen-regulated gene expression in advanced prostate cancer cells. J Biol Chem. 2012;287:4000–13.
Heemers HV, Sebo TJ, Debes JD, Regan KM, Raclaw KA, Murphy LM, et al. Androgen deprivation increases p300 expression in prostate cancer cells. Cancer Res. 2007;67:3422–30.
Jin L, Garcia J, Chan E, de la Cruz C, Segal E, Merchant M, et al. Therapeutic targeting of the CBP/p300 bromodomain blocks the growth of castration-resistant prostate cancer. Cancer Res. 2017;77:5564–75.
Pegg N, Brooks N, Worthington J, Young B, Prosser A, Lane J, et al. Characterisation of CCS1477: a novel small molecule inhibitor of p300/CBP for the treatment of castration resistant prostate cancer. J Clin Oncol. 2017;35(Suppl 15):11590–11590.
Yan Y, Ma J, Wang D, Lin D, Pang X, Wang S, et al. The novel BET-CBP/p300 dual inhibitor NEO2734 is active in SPOP mutant and wild-type prostate cancer. EMBO Mol Med. 2019;11:e10659.
Butler L, Irani S, Centenera M, Ryan N, Pegg N, Brooks AN. Preclinical investigation of a small molecule inhibitor of p300/CBP reveals efficacy in patient-derived prostate tumor explants. J Clin Oncol. 2019;37 (Suppl 15):e16534–e16534.
Stice JP, Wardell SE, Norris JD, Yllanes AP, Alley HM, Haney VO, et al. CDK4/6 therapeutic intervention and viable alternative to taxanes in CRPC. Mol Cancer Res. 2017;15:660–9.
Kivinummi K, Urbanucci A, Leinonen K, Tammela TLJ, Annala M, Isaacs WB, et al. The expression of AURKA is androgen regulated in castration-resistant prostate cancer. Sci Rep. 2017;7:17978.
Jones D, Noble M, Wedge SR, Robson CN, Gaughan L. Aurora A regulates expression of AR-V7 in models of castrate resistant prostate cancer. Sci Rep. 2017;7:40957.
Sarkar S, Brautigan DL, Larner JM. Aurora kinase A promotes AR degradation via the E3 ligase CHIP. Mol Cancer Res. 2017;15:1063–72.
Beltran H, Oromendia C, Danila DC, Montgomery B, Hoimes C, Szmulewitz RZ, et al. A phase II trial of the aurora kinase A inhibitor alisertib for patients with castration-resistant and neuroendocrine prostate cancer: efficacy and biomarkers. Clin Cancer Res. 2019;25:43–51.
Titov DV, Gilman B, He QL, Bhat S, Low WK, Dang Y, et al. XPB, a subunit of TFIIH, is a target of the natural product triptolide. Nat Chem Biol. 2011;7:182–8.
Chugh R, Sangwan V, Patil SP, Dudeja V, Dawra RK, Banerjee S, et al. A preclinical evaluation of minnelide as a therapeutic agent against pancreatic cancer. Sci Transl Med. 2012;4:156ra139.
Isharwal S, Modi S, Arora N, Uhlrich C 3rd, Giri B, Barlass U, et al. Minnelide inhibits androgen dependent, castration resistant prostate cancer growth by decreasing expression of androgen receptor full length and splice variants. Prostate. 2017;77:584–96.
Rosati R, Patki M, Chari V, Dakshnamurthy S, McFall T, Saxton J, et al. The amino-terminal domain of the androgen receptor co-opts extracellular signal-regulated kinase (ERK) docking sites in ELK1 protein to induce sustained gene activation that supports prostate cancer cell growth. J Biol Chem. 2016;291:25983–98.
Rosati R, Polin L, Ducker C, Li J, Bao X, Selvakumar D, et al. Strategy for tumor-selective disruption of androgen receptor function in the spectrum of prostate cancer. Clin Cancer Res. 2018;24:6509–22.
Gaur S, Gross ME, Liao CP, Qian B, Shih JC. Effect of monoamine oxidase A (MAOA) inhibitors on androgen-sensitive and castration-resistant prostate cancer cells. Prostate. 2019;79:667–77.
Zadra G, Ribeiro CF, Chetta P, Ho Y, Cacciatore S, Gao X, et al. Inhibition of de novo lipogenesis targets androgen receptor signaling in castration-resistant prostate cancer. Proc Natl Acad Sci USA. 2019;116:631–40.
Nakata D, Koyama R, Nakayama K, Kitazawa S, Watanabe T, Hara T. Glycogen synthase kinase-3 inhibitors suppress the AR-V7-mediated transcription and selectively inhibit cell growth in AR-V7-positive prostate cancer cells. Prostate. 2017;77:955–61.
Magani F, Bray ER, Martinez MJ, Zhao N, Copello VA, Heidman L, et al. Identification of an oncogenic network with prognostic and therapeutic value in prostate cancer. Mol Syst Biol. 2018;14:e8202.
Recouvreux MV, Wu JB, Gao AC, Zonis S, Chesnokova V, Bhowmick N, et al. Androgen receptor regulation of local growth hormone in prostate cancer cells. Endocrinology. 2017;158:2255–68.
Ramamurthy VP, Ramalingam S, Gediya LK, Njar VCO. The retinamide VNLG-152 inhibits f-AR/AR-V7 and MNK-eIF4E signaling pathways to suppress EMT and castration-resistant prostate cancer xenograft growth. FEBS J. 2018;285:1051–63.
Ramamurthy VP, Ramalingam S, Gediya L, Kwegyir-Afful AK, Njar VC. Simultaneous targeting of androgen receptor (AR) and MAPK-interacting kinases (MNKs) by novel retinamides inhibits growth of human prostate cancer cell lines. Oncotarget. 2015;6:3195–210.
Li D, Tian G, Wang J, Zhao LY, Co O, Underill ZC, et al. Inhibition of androgen receptor transactivation function by adenovirus type 12 E1A undermines prostate cancer cell survival. Prostate. 2018;78:1140–56.
Nunes JJ, Pandey SK, Yadav A, Goel S, Ateeq B. Targeting NF-kappa B signaling by artesunate restores sensitivity of castrate-resistant prostate cancer cells to antiandrogens. Neoplasia. 2017;19:333–45.
Dyshlovoy SA, Menchinskaya ES, Venz S, Rast S, Amann K, Hauschild J, et al. The marine triterpene glycoside frondoside A exhibits activity in vitro and in vivo in prostate cancer. Int J Cancer. 2016;138:2450–65.
Acknowledgements
ESA has received funding from the Prostate Cancer Foundation, the Patrick C. Walsh Fund, and NIH grants R01 CA185297 and P30 CA006973. AJA has received funding from a Prostate Cancer Foundation and Movember Global Treatment Sciences Challenge Award and the NIH under a P30 CA014236 and 1R01CA233585-01 grant. JL is currently funded by a Prostate Cancer Foundation grant, NIH grant R01 CA185297, and US Department of Defense Prostate Cancer Research Program grant W81XWH-19-1-0686.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
ESA has served as a paid consultant/advisor for Janssen, Pfizer, Sanofi, Dendreon, Essa, Merck, Bristol-Myers Squibb, AstraZeneca, Clovis, Eli Lilly, and Amgen; has received research funding to his institution from Janssen, Johnson & Johnson, Sanofi, Dendreon, Genentech, Novartis, Tokai, Merck, Bristol-Myers Squibb, AstraZeneca, and Constellation; and is a co-inventor of an AR-V7 biomarker technology that has been licensed to Qiagen. AJA has served as a paid consultant for AstraZeneca, Merck, Dendreon, Janssen, Clovis, Bayer, and Medivation/Astellas; is on the speaker’s bureau for Bayer and Dendreon; and receives research funding to his institution from Janssen, Medivation/Astellas, Sanofi-Aventis, Active Biotech, Bayer, Dendreon, Merck, AstraZeneca, Genentech/Roche, BMS, Constellation, Novartis, and Pfizer. JL has served as a paid consultant/advisor for Sun Pharma, Janssen, Tolero, and Sanofi; has received research funding to his institution from Orion, Mirati, Astellas, Sanofi, Constellation, Calibr, Pandomedx, and Gilead; and is a co-inventor of a technology that has been licensed to Tokai, Qiagen, and A&G. CL is a co-inventor of a technology that has been licensed to Tokai and Qiagen.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lu, C., Brown, L.C., Antonarakis, E.S. et al. Androgen receptor variant-driven prostate cancer II: advances in laboratory investigations. Prostate Cancer Prostatic Dis 23, 381–397 (2020). https://doi.org/10.1038/s41391-020-0217-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-020-0217-3
This article is cited by
-
The CDK7 inhibitor CT7001 (Samuraciclib) targets proliferation pathways to inhibit advanced prostate cancer
British Journal of Cancer (2023)
-
Targeting the radiation-induced ARv7-mediated circNHS/miR-512-5p/XRCC5 signaling with Quercetin increases prostate cancer radiosensitivity
Journal of Experimental & Clinical Cancer Research (2022)
-
Resistance to the Androgen Receptor Centred Therapies: Biology and Management
SN Comprehensive Clinical Medicine (2021)
-
Isolation and characterization of castration-resistant prostate cancer LNCaP95 clones
Human Cell (2021)
-
Role of androgen receptor splice variant-7 (AR-V7) in prostate cancer resistance to 2nd-generation androgen receptor signaling inhibitors
Oncogene (2020)