Abstract
Background
It is well known that the gastrointestinal (GI) microbiota can influence the metabolism, pharmacokinetics, and toxicity of cancer therapies. Conversely, the effect of cancer treatments on the composition of the GI microbiota is poorly understood. We hypothesized that oral androgen receptor axis-targeted therapies (ATT), including bicalutamide, enzalutamide, and abiraterone acetate, may be associated with compositional differences in the GI microbiota.
Methods
We profiled the fecal microbiota in a cross-sectional study of 30 patients that included healthy male volunteers and men with different clinical states of prostate cancer (i.e., localized, biochemically recurrent, and metastatic disease) using 16S rDNA amplicon sequencing. Functional inference of identified taxa was performed using PICRUSt.
Results
We report a significant difference in alpha diversity in GI microbiota among men with versus without a prostate cancer diagnosis. Further analysis identified significant compositional differences in the GI microbiota of men taking ATT, including a greater abundance of species previously linked to response to anti-PD-1 immunotherapy such as Akkermansia muciniphila and Ruminococcaceae spp. In functional analyses, we found an enriched representation of bacterial gene pathways involved in steroid biosynthesis and steroid hormone biosynthesis in the fecal microbiota of men taking oral ATT.
Conclusions
There are measurable differences in the GI microbiota of men receiving oral ATT. We speculate that oral hormonal therapies for prostate cancer may alter the GI microbiota, influence clinical responses to ATT, and/or potentially modulate the antitumor effects of future therapies including immunotherapy. Given our findings, larger, longitudinal studies are warranted.
Similar content being viewed by others
Introduction
The gastrointestinal (GI) microbiota are known to influence the metabolism, pharmacokinetics, and toxicity of many drugs and xenobiotics [1], yet there are few mechanistic studies exploring this effect in relation to cancer therapies. Several compelling examples have emerged providing insight into the relationship between human-associated microbiota and cancer treatment. The bacterium Mycoplasma hyorhinis and species of Proteobacteria, when present within a tumor, may metabolize the chemotherapy drug, gemcitabine, into a deaminated inactive metabolite [2], resulting in drug resistance [3]. β-glucuronidases produced by bacterial species in the GI tract can reactivate the excreted, inactive metabolite of the topoisomerase I inhibitor, irinotecan, causing adverse drug toxicities, including severe diarrhea [4]. Likewise, although the mechanism is not fully understood, there is emerging evidence that the GI microbiota can influence the efficacy of immunotherapy [5,6,7,8,9,10,11].
Recent studies in animal models have demonstrated that intestinal microbiota are essential for therapeutic efficacy of agents such as cyclophosphamide [7], platinum chemotherapy [6], and both anti-CTLA-4 [5] and anti-PD-L1 [8] immunotherapies. Eradication of the commensal intestinal flora by antibiotic treatment or via use of germ-free mice eliminates therapeutic efficacy of these agents in different tumor models. A study in a melanoma model showed that the therapeutic benefit of anti-PD-L1 immunotherapy could be bolstered by feeding animals a strain of Bifidobacterium—a species commonly used in probiotic supplements—prior to initiating therapy [8]. Three recent human studies, two in melanoma patients [9, 11] and one in patients with epithelial tumors [10], reported that the presence of certain types of bacteria including Ruminococcaceae, Bifidobacteriaceae, and Akkermansia muciniphila are associated with response to anti-PD-1 immunotherapy. Fecal microbial transplant from human donors that were responders to anti-PD-1 immunotherapy into germ-free mouse allograft tumor models conferred antitumor efficacy of anti-PD-1 immunotherapy versus fecal samples transplanted from non-responders [9,10,11]. Collectively, these studies indicate that members of the intestinal microbiome may be essential for cancer drug efficacy and that modulating intestinal microbiome composition may enhance therapeutic response.
The relationship between the GI microbiota and cancer therapies in men with prostate cancer is underexplored. There is, however, compelling evidence that the GI microbiome is involved in multiple-related processes such as modulation of circulating hormone levels [12, 13], stimulation of antitumor immune responses [5, 6, 8], and induction of treatment-related toxicities (including immunotherapy-induced colitis [14] and radiation-induced bowel toxicity [15]), and/or morbidities including development of metabolic syndrome [16, 17]. Animal studies suggest that the GI microbiota may also be affected by circulating androgen levels [12, 13] and castration [16]. We hypothesize that hormonal therapy, particularly oral formulations of androgen axis-targeted drugs, used in the treatment of prostate cancer may promote changes in the GI microbiota. In this exploratory study, we determined compositional differences in GI microbiota in (1) men with and without prostate cancer and (2) men with localized prostate cancer, biochemical recurrence after primary treatment, and hormone-sensitive or castration-resistant metastatic disease. We also examined the relationship between GI microbiota composition and androgen deprivation therapies, with a focus on orally administered androgen receptor axis-targeted therapies (ATT).
Patients and methods
Study design and patient population
Specimens were obtained under a Johns Hopkins Medicine Institutional Review Board approved protocol with written informed consent. Rectal swabs from 30 patients were collected during routine Urology or Medical Oncology outpatient clinic visits at the Johns Hopkins Hospital and Sydney Kimmel Comprehensive Cancer Center. Patients who were currently taking an antibiotic were excluded. Patients designated as “controls” were being followed in the Urology clinic primarily for benign prostatic hyperplasia. Importantly, since the control patients did not undergo prostate biopsy, they cannot be definitively defined as cancer free. Men categorized as “benign” were being evaluated for suspicion of prostate cancer, but subsequently had a negative biopsy. For the benign group, the rectal swab was collected at the evaluation clinic visit, which was prior to the patient taking prophylactic antibiotics for the biopsy. For the seven men in the “cancer” group, three had swabs taken 1–2 months prior to their diagnostic biopsy, one had their swab taken 1 month after diagnostic biopsy, and three men had swabs taken >6 months after prior biopsy. For the purposes of our medication analyses, the designation “NoMeds” indicated men who were not undergoing androgen derivation therapy (ADT), “GNRH” were men only being treated with a gonadotropin-releasing hormone (GNRH) agonist/antagonist, and “oral ATT” were men being treated with oral androgen receptor axis-targeted therapies.
Sample collection and DNA isolation
The rectal swab procedure is detailed in the Supplemental Methods. Rectal swabs were immediately stored at −80 °C until DNA isolation. The investigators were blinded to group allocation until after all sequencing was completed. Swab contents were resuspended in 500 μl of 1× PBS and DNA was extracted with a phenol:chloroform method that incorporates multiple enzyme digest and bead beating as previously described [18]. A total of 16 “mock” (500 μl 1× PBS as starting material) DNA extractions were performed to control for contamination from DNA extraction through the full amplification and sequencing pipeline.
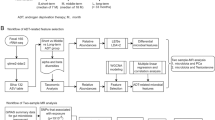
16S rDNA gene library generation, HiSeq sequencing, and analysis
Details of the sequencing and analysis can be found in the Supplemental Methods.
Statistical analysis
After contaminant removal, random subsampling to 80,000 sequences per sample was performed to provide even coverage prior to downstream statistical comparisons (rationale for subsampling described in ref. [19]). Differential abundance analysis was performed using the negative binomial test implemented in the DESeq R package. P values were adjusted for multiple hypothesis testing using the false discovery rate (FDR). Beta diversity analysis, including Bray-Curtis and UniFrac distance computation and principal coordinates analysis (PCoA), was performed in QIIME. Statistical comparisons of alpha diversity utilized generalized linear models (GLMs) and evaluated three different underlying response variable family distributions (Gaussian, Log-Normal, and Gamma).
Akkermansia muciniphila quantitative PCR
A. muciniphila genomic DNA was obtained from the American Type Culture Collection (BAA-835D-5) to develop a standard curve for quantitative PCR (qPCR). The A. muciniphila species-specific primer set was used as follows, Forward primer: 5′-CAGCACGTGAAGGTGGGGAC-3′, Reverse primer: 5′-CCTTGCGGTTGGCTTCAGAT-3′. The total estimated copies of A. muciniphila in each fecal sample were determined using this assay relative to the total number of estimated 16S rDNA copies assayed by qPCR using the universal 16S primer set that was also used for Illumina amplicon sequencing: Forward primer: 5′-CAACGCGWRGAACCTTACC-3′ and Reverse primer: 5′-CRRCACGAGCTGACGAC-3′.
Results
Differences in GI taxonomic profiles by disease status and medication
We characterized the bacterial composition of fecal samples from 30 men (n = 6 control, n = 3 benign (negative biopsy), n = 7 with localized prostate cancer, n = 7 with biochemically recurrent prostate cancer, and n = 7 with metastatic prostate cancer). Table 1 contains the clinical details of the men included in the study as well as the type of ATT administered at the time of sample collection. Most of the men included in this study were Caucasian (White) and, importantly, there was no statistical difference in patient age or body mass index (BMI) among the medication groups (p = 0.4 and p = 0.9, respectively, Kruskal–Wallis test). A complete list of all medications and supplements that the men were taking at the time of sample collection is provided in Supplementary Table S1.
The taxonomic profiles of each sample are depicted in Supplemental Figure S1. There was a significant difference in alpha diversity in samples from men with versus without prostate cancer, irrespective of medication status (p < 0.05 using multiple statistical models and measures of alpha diversity, Supplementary Table S2), meaning that the total number of species (or species “richness”) and the presence of rare individual species was greater in men without a prostate cancer diagnosis than in men with cancer. No significant differences in alpha diversity by medication status were observed (Supplementary Table S3).
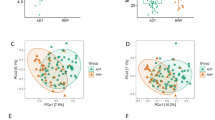
Comparison of taxonomic profiles indicated clustering that associated with medication status (ADT versus no ADT), but not with respect to cancer status (no cancer versus cancer, Fig. 1). Of interest, one patient in the recurrence group who had not yet started on ADT at the time of sample collection grouped with the “No ADT” samples (Fig. 1). In PCoA (a measure of relatedness between samples), we observed that samples from men undergoing ADT grouped separately from men who were not undergoing ADT (Fig. 2a). This was particularly pronounced for men taking oral ATT (Fig. 2b). Of interest, men with prostate cancer (localized, biochemically recurrent, or metastatic groups) separated distinctly from men without cancer (control and benign groups) in PCoA (Fig. 2c), a result that was also statistically significant in PERMANOVA (Adonis) model testing (p = 0.02). The between-sample distances (beta diversity) were smallest within the oral ATT group compared to the GNRH and NoMeds groups, meaning that the species profiles among the fecal samples within the oral ATT group were most similar to each other, and that a specific taxon or taxa distinguish them from the other groups (Fig. 2d). The greatest beta diversity was observed between the NoMeds group and the oral ATT group (Fig. 2d), meaning that the samples from these two groups were most dissimilar to each other.
Unsupervised clustering (log-transformed) of 16S rDNA Illumina-sequencing results from fecal samples by genus. The dendrogram was based on hierarchical clustering of the Euclidean distance between samples in the combined groups. L cancer localized prostate cancer, BCR biochemically recurrent prostate cancer, mHSPC metastatic hormone-sensitive prostate cancer, mCRPC metastatic castration-resistant prostate cancer. No cancer = no clinical and/or biopsy proven diagnosis of cancer (control and benign groups)
Principal coordinates analysis (PCoA) and beta diversity (unweighted UniFrac) of each fecal sample bacterial profile, analyzed by the indicated groups. a–c Principal coordinate axis 2 showed the most distinction between medication groups, so statistical comparisons were limited to this dimension. d Statistical comparison of beta diversity between the indicated groups (Mann–Whitney test). Shown is the mean unweighted UniFrac distance (+SEM)
Differentially abundant species in the GI microbiota of men taking oral ATT
Since we observed significant differences in beta diversity by ADT status and type in PCoA, we next determined if particular species of GI microbiota were differentially abundant between medication categories. As shown in Table 2, several species of bacteria were differentially abundant in terms of the proportion of sequencing reads that matched the species/OTU obtained from the samples across different treatment categories. Notably, species, such as Akkermansia muciniphila, Ruminococcaceae spp., and Lachnospiraceae spp., were significantly more abundant in the fecal samples of men taking oral ATT. When analyzed at the bacterial family level, we again observed a significant greater abundance of sequencing reads assigned to the bacterial families Verrucomicrobiaceae (of which Akkermansia muciniphila is one of the few members), Lachnospiraceae, and others in the oral ATT group (Table 2). There was also a significant decrease in the abundance of sequencing reads assigned to bacterial families such as Brevibacteriaceae, Erysipelotrichaceae, and Streptococcaceae in men receiving ADT versus no ADT and specifically in the oral ATT group versus men not undergoing ADT (Table 2).
As Akkermansia muciniphila is a species of particular recent interest with respect to treatment response to anti-PD-1 immunotherapy in patients with epithelial tumors, we verified the differential abundance of A. muciniphila in men taking oral ATT using an independent qPCR assay. These analyses confirmed that A. muciniphila was significantly more prevalent in the men who were taking oral ATT. Moreover, the results showed high correlation with our Illumina amplicon sequencing results (Fig. 3).
Quantitative PCR (qPCR) for Akkermansia muciniphila. a Confirmation of enriched abundance of this species in the GI microbiota of men taking oral ATT versus the other men included in the study. b The qPCR results were in strong correlation (R2 = 0.9938) with the results obtained by 16S rDNA Illumina amplicon sequencing
Enrichment of GI microbiota predicted to contain genes related to steroid/hormone biosynthesis in men taking oral ATT
We next performed functional inference analyses using PICRUSt [20]. Functional pathways involving steroid/hormone biosynthesis were significantly enriched within the oral ATT group compared to the no ADT (NoMeds) group (Table 3). Other pathways of interest that were more prevalent in the oral ATT group versus the no ADT group included caffeine metabolism and glycosaminoglycan degradation (Table 3).
Discussion
Our study aimed to assess the compositional profile of the GI microbiota in men with and without a diagnosis of prostate cancer and with and without treatment with ATT. We report initial evidence that the alpha diversity of the GI microbiota is greater in men without a prostate cancer diagnosis, and that there were measurable differences in the bacterial composition of the GI microbiota in men undergoing treatment with ATT.
Cancer therapies and GI microbiota
The ability of cancer therapies to affect and change the composition of the GI microbiota is not well studied. Interestingly, a screen of more than 1000 marketed non-antibiotic drugs against 40 representative GI bacterial strains found that nearly a quarter inhibited bacterial growth [21]. Chemotherapy and immunotherapy have been shown to induce dysbiosis (a pathogenic microbial imbalance) of the GI microbiota in rodent models [7, 22]. The most mature data involving longitudinal studies are in patients with non-Hodgkin’s lymphoma undergoing bone marrow transplant conditioning chemotherapy, which induced pathogenic shifts in the GI microbiota that were associated with treatment toxicities [23, 24]. A study in pediatric acute myeloid leukemia patients receiving chemotherapy demonstrated direct bacteriostatic effect of chemotherapeutics, as well as outgrowth of pathogenic enterococci that could not be fully explained by concurrent use of antibiotics [25]. Compositional changes to the GI microbiota induced by chemotherapy or immunotherapy could conceivably impact factors such as the local inflammatory environment in the intestinal tract, systemic inflammatory effects, and/or the efficacy of any subsequently administered cancer therapies.
The microbiome and systemic hormone levels
It has been reported that steroid biosynthesis occurs in prokaryotes [26, 27], and that certain species of bacteria are capable of metabolizing estrogen and androgen precursors and catabolizing estrogens and androgens thereby affecting systemic levels of these hormones [28,29,30]. Altering the gastrointestinal flora in a mouse model of type 1 diabetes impacted testosterone levels, as well as the development of type 1 diabetes [12]. In another study, mice consuming a diet rich in the commonly used probiotic strain Lactobacillus reuteri had a reduced systemic inflammatory state through reduction of IL-17, and an increase in serum testosterone levels [31]. On the converse, the microbiome can also be affected by hormone levels, as another mouse study showed that castrating mice induced alterations in GI microbiota composition, and subsequent development of abdominal obesity [16]. Intriguingly, this study by Harada et al. implies that the GI microbiota may mediate several of the side effects associated with ADT, including obesity and the metabolic syndrome. In our study, we found that men taking oral ATT had a different GI microbiota composition than men taking GNRH agonists/antagonists alone or men not undergoing ADT. Functional pathway inference of the species present in the fecal microbiota of men taking oral ATT indicated an intriguing possibility that the species capable of steroid/hormone biosynthesis are more abundant in the GI flora when men are taking these oral medications (Table 3). This finding, if confirmed, could have important implications and perhaps represent a mechanism for potential alternative pathways for production of steroid metabolites that could influence treatment response to oral ATT. Critical follow-up studies will correlate the presence of GI bacterial species capable of steroid/hormone biosynthesis to circulating hormone levels.
Oral ATT, GI microbiota, and immunotherapy
Much excitement has been generated after the publication of a series of human studies in melanoma patients [9, 11] and in patients with epithelial tumors [10], all indicating that the presence of certain types of bacteria including Ruminococcaceae, Bifidobacteriaceae, and Akkermansia muciniphila are associated with a positive response to anti-PD-1 immunotherapy. In our study, we observed overrepresentation of these same species (Ruminococcaceae and particularly Akkermansia muciniphila) in the fecal microbiota of men taking oral ATT (Table 2). Although preliminary, we speculate that our results might represent one potential explanation for the report of responses to anti-PD-1 immunotherapy observed in men with metastatic prostate cancer who have progressed on enzalutamide [32].
There are several limitations to our study including the relatively small sample size and the lack of longitudinal sampling. Important follow-up studies will include samples collected prior to start of therapy and then longitudinally after therapy initiation. Such studies will further strengthen our hypothesis that ATT is responsible for the compositional differences that we observed, as opposed to other factors that can influence the composition of the GI microbiome such as diet or stress levels. Furthermore, we observed a significant decrease in GI microbiota alpha diversity in patients with prostate cancer that was independent of medication status. Decreased diversity in GI microbiota has been reported as a risk factor for several other types of disease as well as “Western” lifestyle [33]. Our results should be taken with caution, however, as many of the men with prostate cancer in this study had undergone prior treatments that could have conceivably influenced the diversity of the GI microbiota. Our results prompt further examination of GI microbiota diversity as a risk factor for prostate cancer in larger patient cohorts.
In conclusion, our study provides preliminary evidence that the GI microbiota may be different in men undergoing treatment with androgen receptor axis-targeted therapies commonly used to treat prostate cancer. We hypothesize that these compositional differences may influence treatment response to oral ATT or to subsequent treatments such as immunotherapy. Future longitudinal studies pre-, during, and post-therapy are warranted to confirm the degree to which the GI microbiota are altered and to assess whether these alterations are correlated to prostate cancer treatment responses. Collectively, these studies could determine whether the GI microbiome is both essential for therapeutic efficacy and whether it could serve as a target that could be modulated to enhance treatment response.
References
Spanogiannopoulos P, Bess EN, Carmody RN, Turnbaugh PJ. The microbial pharmacists within us: a metagenomic view of xenobiotic metabolism. Nat Rev Microbiol. 2016;14:273–87.
Vande Voorde J, Sabuncuoğlu S, Noppen S, Hofer A, Ranjbarian F, Fieuws S, et al. Nucleoside-catabolizing enzymes in mycoplasma-infected tumor cell cultures compromise the cytostatic activity of the anticancer drug gemcitabine. J Biol Chem. 2014;289:13054–65.
Geller LT, Barzily-Rokni M, Danino T, Jonas OH, Shental N, Nejman D, et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357:1156–60.
Wallace BD, Wang H, Lane KT, Scott JE, Orans J, Koo JS, et al. Alleviating cancer drug toxicity by inhibiting a bacterial enzyme. Science. 2010;330:831–5.
Vétizou M, Pitt JM, Daillère R, Lepage P, Waldschmitt N, Flament C, et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science. 2015;350:1079–84.
Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA, et al. Commensal Bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342:967–70.
Viaud S, Saccheri F, Mignot G, Yamazaki T, Daillère R, Hannani D, et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science. 2013;342:971–6.
Sivan A, Corrales L, Hubert N, Williams JB, Aquino-Michaels K, Earley ZM, et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science. 2015;350:1084–9.
Matson V, Fessler J, Bao R, Chongsuwat T, Zha Y, Alegre M-L, et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science. 2018;359:104–8.
Routy B, Le Chatelier E, Derosa L, Duong CPM, Alou MT, Daillere R, et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359:91–97.
Gopalakrishnan V, Spencer CN, Nezi L, Reuben A, Andrews MC, Karpinets TV, et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359:97–103.
Markle JGM, Frank DN, Mortin-Toth S, Robertson CE, Feazel LM, Rolle-Kampczyk U, et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science. 2013;339:1084–8.
Neuman H, Debelius JW, Knight R, Koren O. Microbial endocrinology: the interplay between the microbiota and the endocrine system. FEMS Microbiol Rev. 2015;39:509–21.
Dubin K, Callahan MK, Ren B, Khanin R, Viale A, Ling L, et al. Intestinal microbiome analyses identify melanoma patients at risk for checkpoint-blockade-induced colitis. Nat Commun. 2016;7.
Ferreira MR, Muls A, Dearnaley DP, Andreyev HJN. Microbiota and radiation-induced bowel toxicity: lessons from inflammatory bowel disease for the radiation oncologist. Lancet Oncol. 2014;15:e139–e147.
Harada N, Hanaoka R, Horiuchi H, Kitakaze T, Mitani T, Inui H, et al. Castration influences intestinal microflora and induces abdominal obesity in high-fat diet-fed mice. Sci Rep. 2016;6:23001.
Festi D, Schiumerini R, Eusebi LH, Marasco G, Taddia M, Colecchia A. Gut microbiota and metabolic syndrome. World J Gastroenterol: Wjg. 2014;20:16079–94.
Shrestha E, White JR, Yu S-H, Kulac I, Ertunc O, De Marzo AM, et al. Profiling the urinary microbiome in men with positive versus negative biopsies for prostate cancer. J Urol. 2018;199:161–71.
Weiss S, Xu ZZ, Peddada S, Amir A, Bittinger K, Gonzalez A, et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome. 2017;5:27.
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol. 2013;31:814.
Maier L, Pruteanu M, Kuhn M, Zeller G, Telzerow A, Anderson EE, et al. Extensive impact of non-antibiotic drugs on human gut bacteria. Nature. 2018;555:623.
Fijlstra M, Ferdous M, Koning AM, Rings EH, Harmsen HJ, Tissing WJ. Substantial decreases in the number and diversity of microbiota during chemotherapy-induced gastrointestinal mucositis in a rat model. Support Care Cancer. 2015;23:1513–22.
Montassier E, Batard E, Massart S, Gastinne T, Carton T, Caillon J, et al. 16S rRNA gene pyrosequencing reveals shift in patient faecal microbiota during high-dose chemotherapy as conditioning regimen for bone marrow transplantation. Microb Ecol. 2014;67:690–9.
Montassier E, Gastinne T,Vangay P, Al-Ghalith GA, Bruley des Varannes S, Massart S, et al. Chemotherapy-driven dysbiosis in the intestinal microbiome. Aliment Pharmacol Ther. 2015;42:515–28.
van Vliet MJ,Tissing WJE,Dun CAJ,Meessen NEL,Kamps WA, de Bont ESJM, et al. Chemotherapy treatment in pediatric patients with acute myeloid leukemia receiving antimicrobial prophylaxis leads to a relative increase of colonization with potentially pathogenic bacteria in the gut. Clin Infect Dis. 2009;49:262–70.
Bode HB, Zeggel B, Silakowski B, Wenzel SC, Reichenbach H, Müller R. Steroid biosynthesis in prokaryotes: identification of myxobacterial steroids and cloning of the first bacterial 2,3(S)-oxidosqualene cyclase from the myxobacterium Stigmatella aurantiaca. Mol Microbiol. 2003;47:471–81.
Wei JH, Yin X, Welander PV. Sterol synthesis in diverse bacteria. Front Microbiol. 2016;7:990.
Kwa M, Plottel CS, Blaser MJ, Adams S. The intestinal microbiome and estrogen receptor-positive female breast cancer. J Natl Cancer Inst. 2016;108:djw029.
Ridlon JM, Ikegawa S, Alves JMP, Zhou B, Kobayashi A, Iida T, et al. Clostridium scindens: a human gut microbe with a high potential to convert glucocorticoids into androgens. J Lipid Res. 2013;54:2437–49.
Gorbach SL. Estrogens, breast cancer, and intestinal flora. Rev Infect Dis. 1984;6(Suppl 1):S85–S90.
Poutahidis T, Springer A, Levkovich T, Qi P, Varian BJ, Lakritz JR, et al. Probiotic microbes sustain youthful serum testosterone levels and testicular size in aging mice. PLoS ONE. 2014;9:e84877.
Graff JN, Alumkal JJ, Drake CG, Thomas GV, Redmond WL, Farhad M, et al. Early evidence of anti-PD-1 activity in enzalutamide-resistant prostate cancer. Oncotarget. 2016;7:52810–17.
Mosca A, Leclerc M, Hugot JP. Gut microbiota diversity and human diseases: should we reintroduce key predators in our ecosystem? Front Microbiol. 2016;7:455.
Acknowledgements
We thank the members of the SKCCC Next Generation Sequencing Core, supported by NCI grant P30CA006973, for assistance with sequencing. This work was supported by Prostate Cancer Foundation Challenge Award 16CHAL13 to KSS, KJP, ESA, and AER. ESA is partially funded by Department of Defense grant W81XWH-13-PCRP-CCA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
JRW has financial and/or other relationship with Resphera Biosciences. The remaining authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sfanos, K.S., Markowski, M.C., Peiffer, L.B. et al. Compositional differences in gastrointestinal microbiota in prostate cancer patients treated with androgen axis-targeted therapies. Prostate Cancer Prostatic Dis 21, 539–548 (2018). https://doi.org/10.1038/s41391-018-0061-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-018-0061-x
This article is cited by
-
Gut microbiota in patients with prostate cancer: a systematic review and meta-analysis
BMC Cancer (2024)
-
Association between concomitant proton pump inhibitor use and survival of patients with metastatic prostate cancer receiving abiraterone acetate: a post-hoc analysis of pooled data from three randomized controlled trials
Prostate Cancer and Prostatic Diseases (2024)
-
Associations between gut microbiota and three prostate diseases: a bidirectional two-sample Mendelian randomization study
Scientific Reports (2024)
-
Unresolved Pyuria
Current Bladder Dysfunction Reports (2024)
-
Current Trends and Challenges of Microbiome Research in Prostate Cancer
Current Oncology Reports (2024)