Abstract
Background
Lower respiratory tract infection (LRTI) including pneumonia, bronchitis, and bronchiolitis is the sixth leading cause of mortality around the world and leading cause of death in children under 5 years. Systemic immune response to viral infection is well characterized. However, there is little data regarding the immune response at the upper respiratory tract mucosa. The upper respiratory mucosa is the site of viral entry, initial replication and the first barrier against respiratory infections. Lower respiratory tract samples can be challenging to obtain and require more invasive procedures. However, nasal wash (NW) samples from the upper respiratory tract can be obtained with minimal discomfort to the patient.
Method
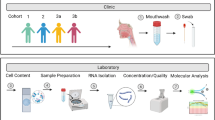
In a pilot study, we developed a protocol using NW samples obtained from hospitalized children with LRTI that enables single cell RNA sequencing (scRNA-seq) after the NW sample is methanol-fixed.
Results
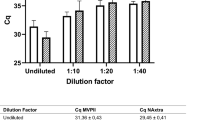
We found no significant changes in scRNA-seq qualitative and quantitative parameters between methanol-fixed and fresh NW samples.
Conclusions
We present a novel protocol to enable scRNA-seq in NW samples from children admitted with LRTI. With the inherent challenges associated with clinical samples, the protocol described allows for processing flexibility as well as multicenter collaboration.
Impact
-
There are no significant differences in scRNA-seq qualitative and quantitative parameters between methanol fixed and fresh Pediatric Nasal wash samples.
-
The study demonstrates the effectiveness of methanol fixation process on preserving respiratory samples for single cell sequencing.
-
This enables Pediatric Nasal wash specimen for single cell RNA sequencing in pediatric patients with respiratory tract infection and allows processing flexibility and multicenter collaboration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Collaborators, G. L. Estimates of the global, regional, and national morbidity, mortality, and aetiologies of lower respiratory tract infections in 195 countries: a systematic analysis for the global burden of disease study 2015. Lancet Infect. Dis. 17, 1133–1161 (2017).
Welliver, R. C. Sr. et al. Fatality rates in published reports of Rsv hospitalizations among high-risk and otherwise healthy children. Curr. Med. Res. Opin. 26, 2175–2181 (2010).
Mauskopf, J., Margulis, A. V., Samuel, M. & Lohr, K. N. Respiratory syncytial virus hospitalizations in healthy preterm infants: systematic review. Pediatr. Infect. Dis. J. 35, e229–e238 (2016).
Melero, J. A. & Moore, M. L. Influence of respiratory syncytial virus strain differences on pathogenesis and immunity. Curr. Top. Microbiol. Immunol. 372, 59–82 (2013).
Mazur, N. I. et al. The respiratory syncytial virus vaccine landscape: lessons from the graveyard and promising candidates. Lancet Infect. Dis. 18, e295–e311 (2018).
Heinonen, S. et al. Infant immune response to respiratory viral infections. Immunol. Allergy Clin. North Am. 39, 361–376 (2019).
Newton, A. H., Cardani, A. & Braciale, T. J. The host immune response in respiratory virus infection: balancing virus clearance and immunopathology. Semin. Immunopathol. 38, 471–482 (2016).
Spyridaki, I. S. et al. Comparison of four nasal sampling methods for the detection of viral pathogens by Rt-Pcr-a Ga(2)Len project. J. Virol. Methods 156, 102–106 (2009).
Principi, N. & Esposito, S. Nasal irrigation: an imprecisely defined medical procedure. Int J. Environ. Res Public Health 14, 516 (2017).
Kurono, Y. The mucosal immune system of the upper respiratory tract and recent progress in mucosal vaccines. Auris Nasus Larynx 49, 1–10 (2022).
Hewitt, R. J. & Lloyd, C. M. Regulation of immune responses by the airway epithelial cell landscape. Nat. Rev. Immunol. 21, 347–362 (2021).
Tang, X., Huang, Y., Lei, J., Luo, H. & Zhu, X. The single-cell sequencing: new developments and medical applications. Cell Biosci. 9, 53 (2019).
Grun, D. et al. Single-cell messenger Rna sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015).
genomics, X. Guidelines for Accurate Target Cell Counts Using 10x Genomics® Single Cell Solutions, https://www.10xgenomics.com/support/single-cell-gene-expression/documentation/steps/sample-prep/guidelines-for-accurate-target-cell-counts-using-10-x-genomics-r-single-cell-solutions (2022).
Haque, A., Engel, J., Teichmann, S. A. & Lonnberg, T. A practical guide to single-cell Rna-Sequencing for Biomedical Research and Clinical Applications. Genome Med 9, 75 (2017).
Chen, J. et al. Pbmc Fixation and Processing for Chromium Single-Cell Rna Sequencing. J. Transl. Med 16, 198 (2018).
Genomics, X. Chromium Next Gem Single Cell 3ʹ Lt Reagent Kits (V3.1 Dual Index) with Feature Barcode Technology for Cell Surface Protein https://www.10xgenomics.com/support/single-cell-gene-expression/documentation/steps/library-prep/chromium-next-gem-single-cell-3-lt-reagent-kits-v-3-1-dual-index-with-feature-barcode-technology-for-cell-surface-protein (2021).
Hwang, B., Lee, J. H. & Bang, D. Author correction: single-cell RNA sequencing technologies and bioinformatics pipelines. Exp. Mol. Med. 53, 1005 (2021).
Sage, S. E. et al. Single-cell gene expression analysis of cryopreserved equine bronchoalveolar cells. Front. Immunol. 13, 929922 (2022).
Andrews S. FastQC: a quality control tool for high throughput sequence data, http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Ilicic, T. et al. Classification of low quality cells from single-cell Rna-seq data. Genome Biol. 17, 29 (2016).
Pollen, A. A. et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat. Biotechnol. 32, 1053–1058 (2014).
Diaz-Papkovich, A., Anderson-Trocme, L. & Gravel, S. A review of Umap in population genetics. J. Hum. Genet 66, 85–91 (2021).
Dal Molin, A., Baruzzo, G. & Di Camillo, B. Single-cell Rna-sequencing: assessment of differential expression analysis methods. Front. Genet 8, 62 (2017).
Genomics, X. Chromium Next Gem Single Cell 3ʹ Lt Reagent Kits (V3.1 Dual Index) with Feature Barcode Technology for Cell Surface Protein, https://cdn.10xgenomics.com/image/upload/v1668017706/supportdocuments/CG000315_ChromiumNextGEMSingleCell3-_GeneExpression_v3.1_DualIndex__RevE.pdf (2022).
Pliner, H. A., Shendure, J. & Trapnell, C. Supervised classification enables rapid annotation of cell atlases. Nat. Methods 16, 983–986 (2019).
Bancroft, J. D. & A. S. Theory and Practice of Histological Techniques, Vol. 3rd ed (Churchill Livingstone, Edinburgh, 1990).
Srinivasan, M., Sedmak, D. & Jewell, S. Effect of fixatives and tissue processing on the content and integrity of nucleic acids. Am. J. Pathol. 161, 1961–1971 (2002).
Alles, J. et al. Cell fixation and preservation for droplet-based single-cell transcriptomics. BMC Biol. 15, 44 (2017).
Lee, J. S., Yi, K., Ju, Y. S. & Shin, E. C. Effects of cryopreservation and thawing on single-cell transcriptomes of human T cells. Immune Netw. 20, e34 (2020).
Guillaumet-Adkins, A. et al. Single-cell transcriptome conservation in cryopreserved cells and tissues. Genome Biol. 18, 45 (2017).
Verheijen, M. et al. Dmso induces drastic changes in human cellular processes and epigenetic landscape in vitro. Sci. Rep. 9, 4641 (2019).
Wu, S. Z. et al. Cryopreservation of human cancers conserves tumour heterogeneity for single-cell multi-omics analysis. Genome Med. 13, 81 (2021).
Wagh, V. et al. Effects of cryopreservation on the transcriptome of human embryonic stem cells after thawing and culturing. Stem Cell Rev. Rep. 7, 506–517 (2011).
Acknowledgements
This project is supported by an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health under grant number 5P20GM12134.
Author information
Authors and Affiliations
Contributions
Each author has met the Pediatric Research authorship requirements. S.M.C.L. and K.A.I. completed the conception and design. K.A.I. and M.M. carried out the experiment. M.S.K., S.M.C.L. and K.A.I. completed acquisition of data, analysis and interpretation of data. S.M.C.L., K.A.I., M.M. and M.S.K. completed drafting the article and revising it critically for important intellectual content. S.M.C.L. completed the final approval of the version to be published.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent to participate
Patient’s parental consent was required for the paper.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ismail, K.A., Mukherjee, M., Kareta, M.S. et al. Enabling methanol fixation of pediatric nasal wash during respiratory illness for single cell sequencing in comparison with fresh samples. Pediatr Res 95, 835–842 (2024). https://doi.org/10.1038/s41390-023-02780-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-023-02780-2