Abstract
Background
Biliary atresia (BA) is a severe immune-related disease that is characterized by biliary obstruction and cholestasis. The etiology of BA is unclear, our aim was to explore the relationship between biliary tract inflammation and immune-related genes.
Methods
We selected 14 SNPs in 13 immune-related genes and investigated their associations with BA by using a large case‒control cohort with a total of 503 cases and 1473 controls from southern China.
Results
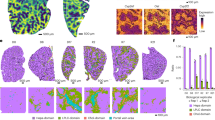
SNP rs1518111 in interleukin10 (IL10) was identified as associated with BA (P = 5.79E-03; OR: 0.80; 95% CI: 0.68–0.94). The epistatic effects of the following pairwise interactions among these SNPs were associated with BA: signal transducer and activator of transcription 4 (STAT4) and chemokine (C-X-C motif) ligand 3 (CXCL3); STAT4 and damage-regulated autophagy modulator1 (DRAM1); CXCL3 and RAD51 paralog B (RAD51B); and interferon gamma (IFNG) and interleukin26 (IL26). Furthermore, we explored the potential role of IL-10 in the pathogenesis of the neonatal mouse model of BA. IL-10 effectively prevented biliary epithelial cell injury and biliary obstruction in murine BA as well as inhibit the activation of BA-related immune cells.
Conclusions
In conclusion, this study provided strong evidence implicating IL10 as a susceptibility gene for BA in the southern Chinese population.
Impact
-
This study provided strong evidence implicating IL10 as a susceptibility gene for BA in the southern Chinese population.
-
This study could infer that IL-10 may play a protective role in BA mouse model.
-
We found that four SNPs (rs7574865, rs352038, rs4622329, and rs4902562) have genetic interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data included in this manuscript are available upon request by contacting with the corresponding author and will be freely available to any researcher wishing to use them for non-commercial purposes, without breaching participant confidentiality.
References
Lendahl, U., Lui, V., Chung, P. & Tam, P. Biliary atresia - emerging diagnostic and therapy opportunities. EBioMedicine 74, 103689 (2021).
Antala, S. & Taylor, S. A. Biliary atresia in children: update on disease mechanism, therapies, and patient outcomes. Clin. Liver Dis. 26, 341–354 (2022).
Petersen, C. et al. European biliary atresia registries: summary of a symposium. Eur. J. Pediatr. Surg. 18, 111–116 (2008).
Nio, M. et al. Five- and 10-year survival rates after surgery for biliary atresia: a report from the Japanese Biliary Atresia Registry. J. Pediatr. Surg. 38, 997–1000 (2003).
Gallo, A. & Esquivel, C. O. Current options for management of biliary atresia. Pediatr. Transpl. 17, 95–98 (2013).
Asai, A., Miethke, A. & Bezerra, J. A. Pathogenesis of biliary atresia: defining biology to understand clinical phenotypes. Nat. Rev. Gastroenterol. Hepatol. 12, 342–352 (2015).
Wang, J. et al. Liver immune profiling reveals pathogenesis and therapeutics for biliary atresia. Cell 183, 1867.e26–1883.e26 (2020).
Sundaram, S. S., Mack, C. L., Feldman, A. G. & Sokol, R. J. Biliary atresia: indications and timing of liver transplantation and optimization of pretransplant care. Liver Transpl. 23, 96–109 (2017).
Hartley, J. L., Davenport, M. & Kelly, D. A. Biliary atresia. Lancet 374, 1704–1713 (2009).
Cheng, G. et al. Common genetic variants regulating ADD3 gene expression alter biliary atresia risk. J. Hepatol. 59, 1285–1291 (2013).
Silva, C. E. et al. [Search for antibodies against human T-cell lymphotropic virus type I (HTLV-I) in blood donors and risk groups]. Rev. Cubana Med. Trop. 49, 24–27 (1997).
Ningappa, M. et al. The role of ARF6 in biliary atresia. PLoS ONE 10, e0138381 (2015).
Ningappa, M. et al. Genome-wide association studies in biliary atresia. Wiley Interdiscip. Rev. Syst. Biol. Med. 7, 267–273 (2015).
Chen, Y. et al. A genome-wide association study identifies a susceptibility locus for biliary atresia on 2p16.1 within the gene EFEMP1. PLoS Genet. 14, e1007532 (2018).
Zhao, R. et al. Polymorphism of ITGB2 gene 3’-UTR+145C/A is associated with biliary atresia. Digestion 88, 65–71 (2013).
Yang, Y. et al. MicroRNA-29b/142-5p contribute to the pathogenesis of biliary atresia by regulating the IFN-gamma gene. Cell Death Dis. 9, 545 (2018).
Liang, J. et al. Association of IL18 genetic polymorphisms with increased risk of Biliary atresia susceptibility in Southern Chinese children. Gene 677, 228–231 (2018).
Verma, A. et al. Human-disease phenotype map derived from PheWAS across 38,682 individuals. Am. J. Hum. Genet. 104, 55–64 (2019).
Sivakumaran, S. et al. Abundant pleiotropy in human complex diseases and traits. Am. J. Hum. Genet. 89, 607–618 (2011).
Cotsapas, C. et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet. 7, e1002254 (2011).
Dong, R. et al. Development and validation of novel diagnostic models for biliary atresia in a large cohort of Chinese patients. EBioMedicine 34, 223–230 (2018).
Zhang, R. et al. The role of neonatal Gr-1(+) myeloid cells in a murine model of rhesus-rotavirus-induced biliary atresia. Am. J. Pathol. 188, 2617–2628 (2018).
Li, P. et al. BATF-JUN is critical for IRF4-mediated transcription in T cells. Nature 490, 543–546 (2012).
Ramirez, K. et al. Gene deregulation and chronic activation in natural killer cells deficient in the transcription factor ETS1. Immunity 36, 921–932 (2012).
Kwon, S. J. et al. KLF13 cooperates with c-Maf to regulate IL-4 expression in CD4+ T cells. J. Immunol. 192, 5703–5709 (2014).
Halstrom, S. et al. Susceptibility to non-tuberculous mycobacterial disease is influenced by rs1518111 in IL10. Hum. Immunol. 78, 391–393 (2017).
El, H. R. et al. Genetic association between interleukin-10 gene rs1518111 and rs3021094 polymorphisms and risk of type 1 diabetes and diabetic nephropathy in Egyptian children and adolescents. Pediatr. Diabetes 22, 567–576 (2021).
Remmers, E. F. et al. Genome-wide association study identifies variants in the MHC class I, IL10, and IL23R-IL12RB2 regions associated with Behcet’s disease. Nat. Genet. 42, 698–702 (2010).
Wang, Z. et al. The intragenic epistatic association of ADD3 with biliary atresia in Southern Han Chinese population. Biosci. Rep. 38, BSR20171688 (2018).
Saraiva, M., Vieira, P. & O’Garra, A. Biology and therapeutic potential of interleukin-10. J. Exp. Med. 217, e20190418 (2020).
Couper, K. N., Blount, D. G. & Riley, E. M. IL-10: the master regulator of immunity to infection. J. Immunol. 180, 5771–5777 (2008).
de Waal, M. R. et al. Interleukin 10 (IL-10) and viral IL-10 strongly reduce antigen-specific human T cell proliferation by diminishing the antigen-presenting capacity of monocytes via downregulation of class II major histocompatibility complex expression. J. Exp. Med. 174, 915–924 (1991).
An, Q., Yan, W., Zhao, Y. & Yu, K. Enhanced neutrophil autophagy and increased concentrations of IL-6, IL-8, IL-10 and MCP-1 in rheumatoid arthritis. Int. Immunopharmacol. 65, 119–128 (2018).
Yang, Y., Dong, R., Zheng, C., Zheng, S. & Chen, G. Infiltration of polarized macrophages associated with liver fibrosis in infants with biliary atresia. J. Pediatr. Surg. 52, 1984–1988 (2017).
Acknowledgements
The authors would like to thank the Clinical Biological Resource Bank of Guangzhou Women and Children’s Medical Center for providing the clinical samples.
Funding
This study was supported by grant from the Medical Scientific Research Foundation of Guangdong Province grant A2017406 (L.L.), the Science and Technology Project of Guangzhou grant 202102020196 (Z.L.), 202201020616 (Z.L.), and the National Natural Science Foundation of China grant 82101808 (Z.L.).
Author information
Authors and Affiliations
Contributions
Z.L., Y.T.(Yan Tian), and C.C. participated in analyzing data and wrote the manuscript. L.T., L.L., X.G., Z.W., and H.W., collected clinical samples and information. M.F., J.Z., and Q.W., revised the manuscript for important intellectual content. R.Z., and Y.Z., coordinated the study over the entire time. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This study was approved by the Medical Ethics Committee of Guangzhou Women and Children’s Medical Center (NO. 2018030327). And all animal experimental protocols were approved by the Institutional Animal Care and Use Committee of Guangzhou Forevergen Biosciences Medical Laboratory Animal Center (SYXK (yue) 2018-0186). Consent was provided by parents or legal guardians (carers) of all patients via written, informed consent ahead of study initiation. Within this informed consent, they agreed to allow analysis of the research data and publication of the paper.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, Z., Tian, Y., Chai, C. et al. The association of immune-related genes and the potential role of IL10 with biliary atresia. Pediatr Res 94, 1659–1666 (2023). https://doi.org/10.1038/s41390-023-02626-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-023-02626-x