Abstract
Background
Poor placental function is a common cause of intrauterine growth restriction, which in turn is associated with increased risks of adverse health outcomes. Our prior work suggests that birthweight and childhood obesity-associated genetic variants functionally impact placental function and that placental microRNA are associated with birthweight. To address the influence of the placenta beyond birth, we assessed the relationship between placental microRNAs and early childhood growth.
Methods
Using the SITAR package, we generated two parameters that describe individual weight trajectories of children (0–5 years) in the New Hampshire Birth Cohort Study (NHBCS, n = 238). Using negative binomial generalized linear models, we identified placental microRNAs that relate to growth parameters (FDR < 0.1), while accounting for sex, gestational age at birth, and maternal parity.
Results
Genes targeted by the six growth trajectory-associated microRNAs are enriched (FDR < 0.05) in growth factor signaling (TGF/beta: miR-876; EGF/R: miR-155, Let-7c; FGF/R: miR-155; IGF/R: Let-7c, miR-155), calmodulin signaling (miR-216a), and NOTCH signaling (miR-629).
Conclusions
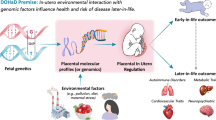
Growth-trajectory microRNAs target pathways affecting placental proliferation, differentiation and function. Our results suggest a role for microRNAs in regulating placental cellular dynamics and supports the Developmental Origins of Health and Disease hypothesis that fetal environment can have impacts beyond birth.
Impact
-
We found that growth trajectory associated placenta microRNAs target genes involved in signaling pathways central to the formation, maintenance and function of placenta; suggesting that placental cellular dynamics remain critical to infant growth to term and are under the control of microRNAs.
-
Our results contribute to the existing body of research suggesting that the placenta plays a key role in programming health in the offspring.
-
This is the first study to relate molecular patterns in placenta, specifically microRNAs, to early childhood growth trajectory.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RICHS total RNA-seq raw reads have been deposited at the NCBI database of Genotypes and Phenotypes (dbGaP) (phs001586). RICHS and NHBCS microRNA count matrices (https://doi.org/10.15139/S3/FUC5EW) and covariate data (https://doi.org/10.15139/S3/O9KYGB) are available at Dataverse.
References
Barker, D. J. P. & Thornburg, K. L. Placental programming of chronic diseases, cancer and lifespan: a review. Placenta 34, 841–845 (2013).
Red-Horse, K. et al. Trophoblast differentiation during embryo implantation and formation of the maternal-fetal interface. J. Clin. Invest. 114, 744–754 (2004).
Kaufmann, P., Mayhew, T. M. & Charnock-Jones, D. S. Aspects of human fetoplacental vasculogenesis and angiogenesis. II. Changes during normal pregnancy. Placenta 25, 114–126 (2004).
Bartel, D. P. Metazoan MicroRNAs. Cell 173, 20–51 (2018).
Quévillon Huberdeau, M. & Simard, M. J. A guide to microRNA-mediated gene silencing. FEBS J. 286, 642–652 (2019).
Pasquinelli, A. E. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat. Rev. Genet. 13, 271–282 (2012).
Hayder, H., O’Brien, J., Nadeem, U. & Peng, C. MicroRNAs: crucial regulators of placental development. Reproduction 155, R259–R271 (2018).
Kennedy, E. M. et al. Placental microRNA expression associates with birthweight through control of adipokines: results from two independent cohorts. Epigenetics 16, 770–782 (2021).
Peng, S. et al. Genetic regulation of the placental transcriptome underlies birth weight and risk of childhood obesity. PLoS Genet. 14, e1007799 (2018).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinforma. Oxf. Engl. 32, 3047–3048 (2016).
Friedländer, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 40, 37–52 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Griffiths-Jones, S., Saini, H. K., van Dongen, S. & Enright, A. J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–D158 (2008).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Cole, T. J., Donaldson, M. D. C. & Ben-Shlomo, Y. SITAR—a useful instrument for growth curve analysis. Int. J. Epidemiol. 39, 1558–1566 (2010).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinforma. Oxf. Engl. 28, 882–883 (2012).
Leek, J. T. svaseq: removing batch effects and other unwanted noise from sequencing data. Nucleic Acids Res. 42, e161 (2014).
Tokar, T. et al. mirDIP 4.1—integrative database of human microRNA target predictions. Nucleic Acids Res. 46, D360–D370 (2018).
Kamburov, A. et al. ConsensusPathDB: toward a more complete picture of cell biology. Nucleic Acids Res. 39, D712–D717 (2011).
Aplin, J. D. & Jones, C. J. P. Cell dynamics in human villous trophoblast. Hum. Reprod. Update 27, 904–922 (2021).
Gamage, T. K. et al. Side-population trophoblasts exhibit the differentiation potential of a trophoblast stem cell population, persist to term, and are reduced in fetal growth restriction. Stem Cell Rev. Rep. 16, 764–775 (2020).
Okae, H. et al. Derivation of human trophoblast stem cells. Cell Stem Cell 22, 50–63.e6 (2018).
Latos, P. A. & Hemberger, M. From the stem of the placental tree: trophoblast stem cells and their progeny. Development 143, 3650–3660 (2016).
Knöfler, M. et al. Human placenta and trophoblast development: key molecular mechanisms and model systems. Cell. Mol. Life Sci. 76, 3479–3496 (2019).
Gupta, S. K., Malhotra, S. S., Malik, A., Verma, S. & Chaudhary, P. Cell signaling pathways involved during invasion and syncytialization of trophoblast cells. Am. J. Reprod. Immunol. 75, 361–371 (2016).
Zhao, H.-J. et al. Bone morphogenetic protein 2 promotes human trophoblast cell invasion by upregulating N-cadherin via non-canonical SMAD2/3 signaling. Cell Death Dis. 9, 1–12 (2018).
Zhao, H.-J. et al. Bone morphogenetic protein 2 promotes human trophoblast cell invasion by inducing activin A production. Endocrinology 159, 2815–2825 (2018).
Haider, S. et al. Notch1 controls development of the extravillous trophoblast lineage in the human placenta. Proc. Natl Acad. Sci. U. S. A. 113, E7710–E7719 (2016).
Li, Y., Yan, J., Chang, H.-M., Chen, Z.-J. & Leung, P. C. K. Roles of TGF-β superfamily proteins in extravillous trophoblast invasion. Trends Endocrinol. Metab. 32, 170–189 (2021).
Nadeem, L. et al. Nodal signals through activin receptor-like kinase 7 to inhibit trophoblast migration and invasion: implication in the pathogenesis of preeclampsia. Am. J. Pathol. 178, 1177–1189 (2011).
Myatt, L. Placental adaptive responses and fetal programming. J. Physiol. 572, 25–30 (2006).
Soares, M. J., Chakraborty, D., Kubota, K., Renaud, S. J. & Rumi, M. A. K. Adaptive mechanisms controlling uterine spiral artery remodeling during the establishment of pregnancy. Int. J. Dev. Biol. 58, 247–259 (2014).
Karachaliou, M. et al. Association of trimester-specific gestational weight gain with fetal growth, offspring obesity, and cardiometabolic traits in early childhood. Am. J. Obstet. Gynecol. 212, 502.e1–502.e14 (2015).
Dai, Y. et al. MicroRNA-155 inhibits proliferation and migration of human extravillous trophoblast derived HTR-8/SVneo cells via down-regulating cyclin D1. Placenta 33, 824–829 (2012).
Ali, A., Bouma, G. J., Anthony, R. V. & Winger, Q. A. The role of LIN28-let-7-ARID3B pathway in placental development. Int. J. Mol. Sci. 21, 3637 (2020).
McWhorter, E. S. et al. LIN28B regulates androgen receptor in human trophoblast cells through Let-7c. Mol. Reprod. Dev. 86, 1086–1093 (2019).
Modi, B. P. et al. Expression patterns of the chromosome 21 MicroRNA cluster (miR-99a, miR-125b and let-7c) in chorioamniotic membranes. Placenta 49, 1–9 (2017).
van Rooij, J. et al. Evaluation of commonly used analysis strategies for epigenome- and transcriptome-wide association studies through replication of large-scale population studies. Genome Biol. 20, 235 (2019).
Funding
This work was supported by the National Institutes of Health (NIEHS R24ES028507, R01ES025145, P30ES019776, NIMHD R01MD011698 and NICHD K99HD104991).
Author information
Authors and Affiliations
Contributions
E.M.K., D.C.K., K.H., J.C., D.G.-D., M.R.K. and C.J.M. conceptualized and designed the study. K.H., A.B. and D.P. acquired data. E.M.K. analyzed and interpreted the data. E.M.K. Drafted the article. K.H., A.B., D.P., D.C.K., K.H., J.C., D.G.-D., U.R., M.R.K. and C.J.M. critically reviewed and carefully revised the article. All authors approved of the version to be published.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Ethics approval and consent to participate
All participants provided written, informed consent and all protocols were approved by the IRBs at the Women & Infants Hospital of Rhode Island, Dartmouth College and Emory University, respectively.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kennedy, E.M., Hermetz, K., Burt, A. et al. Placental microRNAs relate to early childhood growth trajectories. Pediatr Res 94, 341–348 (2023). https://doi.org/10.1038/s41390-022-02386-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-022-02386-0