Abstract
Background
Chronic lung disease (CLD) is the most common pulmonary morbidity in extremely preterm infants. It is unclear to what extent prenatal exposures influence the risk of CLD. Epigenetic variation in placenta DNA methylation may be associated with differential risk of CLD, and these associations may be dependent upon sex.
Methods
Data were obtained from a multi-center cohort of infants born extremely preterm (<28 weeks’ gestation) and an epigenome-wide approach was used to identify associations between placental DNA methylation and CLD (n = 423). Associations were evaluated using robust linear regression adjusting for covariates, with a false discovery rate of 0.05. Analyses stratified by sex were used to assess differences in methylation-CLD associations.
Results
CLD was associated with differential methylation at 49 CpG sites representing 46 genes in the placenta. CLD was associated with differential methylation of probes within genes related to pathways involved in fetal lung development, such as p53 signaling and myo-inositol biosynthesis. Associations between CpG methylation and CLD differed by sex.
Conclusions
Differential placental methylation within genes with key roles in fetal lung development may reflect complex cell signaling between the placenta and fetus which mediate CLD risk. These pathways appear to be distinct based on fetal sex.
Impact
-
In extremely preterm infants, differential methylation of CpG sites within placental genes involved in pathways related to cell signaling, oxidative stress, and trophoblast invasion is associated with chronic lung disease of prematurity.
-
DNA methylation patterns associated with chronic lung disease were distinctly based on fetal sex, suggesting a potential mechanism underlying dimorphic phenotypes.
-
Mechanisms related to fetal hypoxia and placental myo-inositol signaling may play a role in fetal lung programming and the developmental origins of chronic lung disease.
-
Continued research of the relationship between the placental epigenome and chronic lung disease could inform efforts to ameliorate or prevent this condition.
Similar content being viewed by others
Introduction
Chronic lung disease of prematurity (CLD; also called bronchopulmonary dysplasia) is the most common pulmonary morbidity in extremely preterm infants. Generally, clinical factors have been found to be more predictive of CLD than genetic markers.1 For example, fetal growth restriction (FGR) is highly predictive of CLD.2 It is unclear to what extent prenatal exposures, including chronic hypoxia, maternal socioeconomic status, and placental inflammation, influence the risk of postnatal outcomes, such as CLD. Placental epigenetic programming, as measured by placental DNA methylation, has been proposed as an intermediate linking prenatal exposures to later health outcomes in premature infants.3
Consistent with the placenta’s critical role in fetal development through both the exchange of nutrients and mediation of environmental influences, the fetal origins hypothesis of CLD posits that an adverse intrauterine milieu alters lung development and increases the risk of CLD.4 Several epidemiological studies demonstrating an association between placental disorders and CLD support this hypothesis.2,5 Still, a knowledge gap remains concerning whether the placental epigenome plays a role in fetal origins of CLD in extremely preterm infants.
Preclinical studies suggest that epigenetic mechanisms underlying fetal lung development may occur in a sex-specific manner. For example, sex-specific impairments in alveolarization were found in rats in response to FGR via alterations in the transcription of peroxisome proliferator-activated receptor-gamma (PPARγ).6 Likewise, in murine models, mechanical ventilation and hyperoxia lead to epigenetic changes in the lung related to histone deacetylase activity, which inhibits the alveolar formation and promotes lung remodeling.7 While sexual dimorphism exists for CLD with a higher incidence in males compared to females,8 whether placental epigenetic variation contributes to the pathogenesis of CLD in a sex-specific manner in humans remains unclear.
The objective of this study was to assess the relationship between placental CpG methylation and CLD in extremely preterm infants. We hypothesized that genes involved in pathways related to placental hypoxia and oxidative stress, which may adversely affect fetal lung and pulmonary vascular development,9 would be differentially methylated in placentas collected from infants who later went on to develop CLD versus those who did not. Furthermore, based on significant sex-based differences in placental methylation in response to prenatal exposures as documented by Martin et al.10 and sex-based differences in the incidence of CLD,8,11 we hypothesized that patterns of placental DNA methylation would differ by sex.
Methods
Study sample
Participants in this study represent a subset of individuals enrolled in the Extremely Low Gestational Age Newborn (ELGAN) Study. The ELGAN cohort of newborns delivered prior to 28 weeks of gestation, enrolled at 14 United States hospitals located in five states (Connecticut, Massachusetts, North Carolina, Michigan, and Illinois).12 Of the 1506 infants enrolled, 1251 infants survived to 36 weeks postmenstrual age (PMA) and 1241 (99.2%) were evaluated at 36 weeks PMA for CLD. Epigenetic data were available from the placentas of 423 (34%) of the infants evaluated for CLD. The institutional review boards of all participating institutions approved enrollment and consent procedures for the ELGAN Study.
Classification of chronic lung disease
We classified infants as having CLD if they were receiving either supplemental oxygen or mechanical ventilation, or both, at 36 weeks PMA. In addition, we recorded the type of respiratory support infants were receiving at 36 weeks PMA, classified as follows: none, increased ambient oxygen, nasal cannula, nasal continuous positive airway pressure, conventional mechanical ventilation, or high-frequency ventilation.
Placenta tissue collection
As previously described, placental biopsies were taken shortly after delivery.13 Briefly, a tissue sample from the fetal side of the placenta was collected by applying traction to the chorion and the underlying trophoblast tissue and cutting a sample out at the base of this tissue structure. The tissue sample was immediately frozen in liquid nitrogen and stored at −80 °C until further processing.
DNA extraction and assessment of epigenome-wide DNA methylation
DNA was extracted and epigenome-wide DNA methylation assessed using the EZ DNA Methylation kit (Zymo Research, Irvine, CA) and Infinium MethylationEPIC BeadChip (Illumina, San Diego, CA), as previously described.13 The Infinium MethylationEPIC array measures more than 850,000 CpG loci at single-nucleotide resolution. Samples were randomly allocated to different plates and chips to minimize confounding variables. Quality control was performed at the sample level using the minfi package.14 Functional normalization was performed with a preliminary step of normal-exponential out-of-band (noob) correction method15 for background subtraction and dye normalization, followed by functional normalization to the top two principal components of the control matrix.16 Quality control was performed on individual probes by computing a detection p-value and excluded 806 (0.09%) probes with non-significant detection (p > 0.01) for 5% or more of the samples. The ComBat function was used from the sva package to adjust for batch effects from the sample plate.17 A total of 856,832 CpG probes were retained for analyses. Average methylation levels at each CpG probe were measured and expressed as β-values (β = intensity of the methylated allele (M))/(intensity of the unmethylated allele (U) + intensity of the methylated allele (M) + 100). β-values were logit transformed to M-values for statistical analyses.18
Statistical analysis
The demographic and clinical characteristics of the participants are reported using means, with standard deviations, or proportions. A directed acyclic graph was created using the web-based software DAGitty (www.dagitty.net) to delineate relationships between a priori selected covariates with placental CpG methylation as the exposure and CLD as the outcome (Supplemental Figure).13,19 Based on this diagram, the minimally sufficient set of covariates for adjustment included: gestational age, birth weight Z-score, maternal age, race, infant sex, inflammation level in the placenta, cell-type heterogeneity, and maternal socioeconomic status, based on marital status, eligibility for public insurance (Medicaid), and the highest level of education completed. Maternal age was treated as a continuous variable. The race was dichotomized as white and non-white. Linear mixed models were used to assess the impact of family structure (e.g., twins) and results were found to be consistent with the standard, linear models. To account for the possibility that methylation levels were affected by the infiltration of inflammatory cells within the placenta, acute inflammation status of the chorion/decidua was used as an adjustment factor in all models. Inflammation of the placenta was defined as greater than 10,000 neutrophils per cubic millimeter in the chorion or decidua.20 To control for cell-type heterogeneity, the top 10 component variables were selected based on a surrogate variable analysis (SVA), a reference-free method that efficiently estimates cell-type mixture in heterogeneous tissues using iteratively re-weighted least squares.21 Missing covariates, both continuous and categorical, were simultaneously imputed using a random forest trained on observed values of the data matrix, assessing for complex interactions and non-linear relations (missForest package).22 Imputation was assessed with an out-of-bag imputation error estimate.
An epigenome-wide association study (EWAS) was performed by fitting a robust linear regression model for each CpG probe adjusted for a priori selected covariates with DNA methylation as the response variable on the M-value scale and CLD as the main predictor. Robust linear regression was used to protect against potential heteroscedasticity.23 In these models, test statistics were modified using the Phipson’s robust empirical Bayes procedure, shrinking probe-wise sample variances towards a common value and controlling for test-statistic inflation.24 Statistically significant associations were identified based on Benjamini–Hochberg false discovery rate (FDR) q-values using a significance level of <0.05.25 As a secondary analysis, sex-specific associations focusing on DNA methylation in autosomes were assessed via stratification of adjusted robust linear regression models, excluding sex of the infant as covariates.
Manhattan plots and QQ-plots were used to report results. QQ-plots show potential test-statistic inflation at tails of the inferential distribution by plotting theoretical quantiles of the t-distribution on the x axis against the empirical modified t-statistics estimated using the empirical Bayes procedure. The interpretation of these QQ-plots is equivalent to those that plot theoretical and sample p-values. Traditional inflation factors are not interpretable in this empirical Bayes setting, often underrepresenting inflation in test statistics.
Pathway analysis
Ingenuity Pathway Analysis (IPA), a web-based software application (available at http://www.ingenuity.com), was used to determine whether the genes that contained the differentially methylated probes are enriched for specific biological functions. IPA relies on a repository of biological pathways based on curated findings from the research literature. For this analysis, we focused on genes associated with CLD identified in both the overall and sex-stratified analyses at FDR values < 0.05. IPA provided associated pathways for each focus gene, as well as p-values calculated using the right-tailed Fisher exact test.
Results
Study cohort
Maternal and newborn characteristics are presented for the ELGAN subjects used in this analysis (n = 217 subjects with CLD and n = 206 subjects without CLD) and the overall ELGAN cohort evaluated for CLD at 36 weeks PMA (n = 1241) in Table 1. Infants with CLD had lower mean gestational age, lower mean birth weight, and were more likely to be white, male, and growth-restricted (i.e., birth weight Z-score < −1) compared to infants without CLD. Mothers of ELGANs with placental tissue available (n = 423) were slightly older and more likely to be non-Hispanic white, have higher educational attainment, and have multiple gestation pregnancy, and less likely to have public insurance and smoke during pregnancy compared to mothers without placentas available. CLD frequency was similar between infants with and without placental samples available, and similar to CLD frequency in the overall sample (Supplemental Table).
Differences in placental DNA methylation based on CLD diagnoses
EWAS analyses for CLD, after adjusting for confounders, identified 49 differentially methylated CpG probes, corresponding to 46 unique genes, at FDR adjusted q-values < 0.05 (Table 2). CLD was associated with increased methylation levels in 3 probes (6%) and decreased methylation levels in 46 probes (94%). The Manhattan plot displays the genomic location of CpG methylation probes based on their association with CLD using threshold FDR values of 0.05 and 0.01 (Fig. 1). A comprehensive list of all significant CpG probes is included as a Supplemental File. Analysis was performed on the genes associated with CLD to explore their involvement/enrichment in canonical pathways. Among the top significant CpG probes associated with top canonical pathways, we identified cg24455359 located on chromosome 17 within the 1st exon of the Acetyl-CoA Carboxylase Alpha (ACACA) gene, cg09529537 located on chromosome 5 within the TSS200 region of the Junction Mediating and Regulatory Protein, P53 Cofactor (JMY) gene, cg214664091 located on chromosome 20 within the TSS220 region of the Proliferating Cell Nuclear Antigen (PCNA) gene, cg09089417 located on chromosome 12 within the body of the Phosphoglycerate Mutase Family Member 5 (PGAM5) gene, and cg13564825 located on chromosome 19 within the TSS200 region of the Protein Phosphatase 1 Regulatory Subunit 14A (PPP1R14A) gene (Table 3).
The chromosomal location of CpG probes are presented on the x axis and the −log10(p-value) associated with CLD is shown on the y axis. The models are adjusted for gestational age, birth weight Z-score, maternal age, race, infant sex, inflammation level in the placenta, cell-type heterogeneity, and maternal socioeconomic status. We added lines to indicate FDR thresholds at 0.05 (bottom) and 0.01 (top). On the right side of the figure, we include QQ-plots to indicate theoretical quantiles of the t-distribution on the x axis against the empirical modified t-statistics estimated using the empirical Bayes procedure.
Table 4 shows the top canonical pathways associated with differentially methylated genes in infants with CLD. The top canonical pathways enriched within the CLD-associated CpGs include biotin-carboxyl carrier protein assembly (p = 3.94 × 10−3), p53 signaling (p = 7.34 × 10−3), mismatch repair in eukaryotes (2.08 × 10−2), and d-myo-inositol (1,4,5,6)- and (3,4,5,6)-tetrakisphosphate biosynthesis (p = 2.23 × 10−2).
Sex-stratified analyses of differentially methylated genes
EWAS analyses stratified by the sex of the infant revealed distinct CpG methylation patterns, with male-derived placentas having more differentially methylated CpG probes than female-derived placentas in relation to CLD. In the male-derived placentas (n = 222), there were 518 differentially methylated CpG probes corresponding to 414 unique genes that were associated with CLD. Decreased methylation levels were observed in 440 (85%) CpG sites that corresponded to 358 genes. Increased methylation levels were observed in 78 (15%) CpG sites that corresponded to 56 genes (Table 2). Among the female-derived placentas (n = 201), placental CpG methylation of 12 sites was associated with CLD. Decreased methylation was observed in 7 (58%) CpG probes, corresponding to five genes, and increased methylation was observed in 5 (42%) CpG sites, corresponding to three genes (Table 2). Manhattan and QQ-plots for the sex-stratified EWAS are shown in Fig. 2. There were no overlapping CpG methylation probes between the sexes that displayed significance at q < 0.05. In male placentas, the top canonical pathways included AMP-activated protein kinase signaling pathway (p = 9.25 × 10−4), spliceosomal cycle (p = 1.59 × 10−3), apelin cardiac fibroblast signaling pathway (p = 1.62 × 10−3), Huntington’s disease signaling (p = 2.58 × 10−3), and pyridoxal 5′-phosphate salvage pathway (p = 4.73 × 10−3). In female placentas, the top canonical pathways identified were the role of BRCA1 in DNA damage response (p = 6.77 × 10−3) and hereditary breast cancer signaling (p = 1.20 × 10−2) (Table 4).
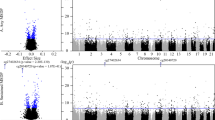
Manhattan plots of CpG methylation probes associated between placental DNA methylation and CLD in males (a) and females (b). The chromosomal location of CpG probes are presented on the x axis and the −log10(p-value) associated with CLD is shown on the y axis. The models are adjusted for gestational age, birth weight Z-score, maternal age, race, infant sex, inflammation level in the placenta, cell-type heterogeneity, and maternal socioeconomic status. We added lines to indicate FDR thresholds at 0.05 (bottom) and 0.01 (top). On the right side of the figure, we include QQ-plots to indicate theoretical quantiles of the t-distribution on the x axis against the empirical modified t-statistics estimated using the empirical Bayes procedure.
Discussion
This study is among the first to examine CpG methylation levels in the placenta at more than 850,000 sites in relation to CLD in extremely preterm infants. The placenta is the key organ for this study due to its critical role in the development of the fetus.26 In addition, DNA methylation in the placenta has been associated with childhood cognitive impairment and body mass index, suggesting a potential role as a mediator in the developmental origins of health and disease.27,28 Based on evidence that oxidative stress may adversely affect fetal lung development, we hypothesized that genes involved in pathways related to placental hypoxia would be differentially methylated in infants based on CLD status, and that the patterns of methylation would differ based on fetal sex.
A total of 49 differentially methylated CpG probes corresponding to 46 unique genes were associated with CLD at FDR adjusted q-value < 0.05. CLD was associated with decreased methylation at a CpG site within the PCNA gene and increased methylation at a CpG site within the JMY gene. The PCNA gene encodes a cofactor for DNA polymerase and the JMY gene encodes an important nuclear cofactor for p53 signaling, which regulates cell growth.29 Both of these genes are involved in p53 signaling, regulating syncytiotrophoblast apoptosis and cell-cycle arrest in the placenta in response to hypoxia. Placental p53 expression is upregulated by increased concentrations of hypoxia-inducible factors in response chronic hypoxia.30,31 Chronic fetal hypoxia leads to impaired fetal lung and vascular development through altered expression of fibroblast growth factor and vascular endothelial growth factors.32 Impaired fetal lung development leads to alveolar simplification, impaired pulmonary angiogenesis, and pulmonary vascular remodeling, which can increase the need for invasive respiratory support following birth.33 Prolonged exposure to mechanical ventilation and supplemental oxygen causes additional lung injury, further increasing the risk of CLD.
CLD was associated with differential methylation levels of CpG sites within genes that are part of the canonical pathway myo-inositol (1,4,5,6)-tetrakisphosphate biosynthesis based on decreased methylation at CpG sites within the PGAM5 and PPP1R14A genes. Myo-inositol is an important cell-signaling molecule that in the placenta acts via abundant inositol transporters to regulate fetal growth.34 In the fetus, myo-inositol plays a major role in lung development by promoting the maturation of phospholipids involved in surfactant production.35 In premature infants, low serum inositol levels are associated with more severe respiratory symptoms.36 This observation has led to multiple randomized control trials of supplemental inositol in infants with respiratory distress syndrome. However, a Cochrane meta-analysis found that inositol supplementation did not reduce death, CLD, or other clinically important outcomes.37
We found unique CpG methylation patterns associated with CLD based on infant sex. CLD in male placentas was associated with differential methylation levels at CpG sites within 9 genes in the AMP-activated protein kinase (AMPK) signaling pathway: decreased methylation at CpG sites within the AK2, ARID2, CHRNA10, GNG7, PFKL, PRKAA1, RAB39, and TSC2 genes, and increased methylation at CpG sites within the PIK3CD gene. Interestingly, single nucleotide polymorphisms leading to increased maternal expression of PRKAA1 in response to altitude-related hypoxia, resulting in increased uterine artery blood flow via AMPK signaling activation, are thought to be an adaptive response to protect against FGR in populations living in the Andean and Tibetan mountains.38 In our study, decreased methylation at CpG sites within 8 genes in the AMPK signaling pathway, a pattern typically indicative of increased gene expression,39 may reflect simultaneous compensatory mechanisms occurring in the maternal circulation to promote uterine blood flow in response to hypoxia in pregnancy.40 Activation of the AMPK signaling pathway has also been associated with impaired trophoblast invasion in placentas from pregnancies complicated by pre-eclampsia.41 Our observation of distinct genes and pathways associated with CLD based on fetal sex is consistent with evidence that sex influences gene expression in human placental villi, which may contribute to worse outcomes in male fetuses.42
While this study is among the first to highlight the relationship between placental CpG methylation and CLD, a few limitations should be noted when interpreting the results. First, the study may not be sufficiently powered to draw definite conclusions using an EWAS approach. While the placenta plays a critical role in fetal development, we are mindful that tissue-specific patterns of CpG methylation exist, and that changes that are observed within the placenta may not reflect patterns in fetal lung tissue.43 Despite this limitation, studies examining fetal and postnatal samples from multiple tissues over time demonstrate that limited patterns of differentially methylated regulatory regions of select genes may be highly conserved across placental, perinatal (i.e., umbilical cord blood), and postnatal human tissues.44,45 An additional limitation of the study is that CpG methylation does not directly indicate the direction or extent of gene expression change.46 Further studies incorporating messenger RNA levels could enhance understanding of the regulatory mechanisms involved. We used the SVA approach to address cell-type heterogeneity. However, this approach may not have adequately controlled for the mix of villous and vascular structures present in fetal-side placenta samples and alternative methods could be used in future studies to address cell-type heterogeneity. Our analyses would also be more robust with the inclusion of ancestry data. In our EWAS analyses, we considered FDR adjusted values < 0.05 as significant. However, we acknowledge that no CpG sites in the overall sample were significant at the more stringent level of <0.01. Our definition of CLD was operationally defined by the ELGAN Study as supplemental oxygen therapy with or without mechanical ventilation at 36 weeks PMA, which is not consistent with the most recent NIH consensus definition of CLD.47
It is important to note that several mechanisms may explain the association of placental methylation and CLD. CpG methylation is correlated with gene expression,48 and altered gene expression may be an indicator of placental health. Therefore, changes in gene expression involved in pathways related to trophoblast migration and implantation may reflect reduced nutrient and oxygen delivery to the developing fetus, resulting in altered fetal lung programming affecting alveolar and pulmonary vascular development. Altered placental methylation may also be an indicator of disrupted inflammation and signaling pathways that lead to cytokine-mediated deleterious effects on the fetal lung, or an indicator of FGR. We included placental inflammation and FGR (as defined by birth weight Z-score) as covariates in our model due to their independent associations with altered placental methylation and CLD.49,50,51 Finally, our results may reflect residual confounding of covariates related to CLD that were not included in the model.
Our findings suggest that prenatal factors may play a role in the development of CLD, and, if replicated in future studies, might explain why CLD incidence has not decreased despite multiple intervention trials to reduce its frequency.52 Further analyses measuring gene expression in the placenta and correlation with blood sampling from infants may provide further insights into the complex interaction between the placenta and fetal lung development.
References
Laughon, M. M. et al. Prediction of bronchopulmonary dysplasia by postnatal age in extremely premature infants. Am. J. Respir. Crit. Care Med. 183, 1715–1722 (2011).
Bose, C. et al. Fetal growth restriction and chronic lung disease among infants born before the 28th week of gestation. Pediatrics 124, e450–e458 (2009).
Bangma, J. T., Hartwell, H., Santos, H. P., O’Shea, T. M. & Fry, R. C. Placental programming, perinatal inflammation, and neurodevelopment impairment among those born extremely preterm. Pediatr. Res. 89, 326–335 (2021).
Mestan, K. K. & Steinhorn, R. H. Fetal origins of neonatal lung disease: understanding the pathogenesis of bronchopulmonary dysplasia. Am. J. Physiol. Lung Cell Mol. Physiol. 301, L858–L859 (2011).
Lal, M. K., Manktelow, B. N., Draper, E. S. & Field, D. J. Chronic lung disease of prematurity and intrauterine growth retardation: a population-based study. Pediatrics 111, 483–487 (2003).
Joss-Moore, L. A. et al. Iugr differentially alters Mecp2 expression and H3k9me3 of the Ppargamma gene in male and female rat lungs during alveolarization. Birth Defects Res. A Clin. Mol. Teratol. 91, 672–681 (2011).
Londhe, V. A. et al. Hyperoxia impairs alveolar formation and induces senescence through decreased histone deacetylase activity and up-regulation of P21 in neonatal mouse lung. Pediatr. Res. 69, 371–377 (2011).
Binet, M. E., Bujold, E., Lefebvre, F., Tremblay, Y. & Piedboeuf, B. Role of gender in morbidity and mortality of extremely premature neonates. Am. J. Perinatol. 29, 159–166 (2012).
Rozance, P. J. et al. Intrauterine growth restriction decreases pulmonary alveolar and vessel growth and causes pulmonary artery endothelial cell dysfunction in vitro in fetal sheep. Am. J. Physiol. Lung Cell Mol. Physiol. 301, L860–L871 (2011).
Martin, E. et al. Sexual epigenetic dimorphism in the human placenta: implications for susceptibility during the prenatal period. Epigenomics 9, 267–278 (2017).
Stevenson, D. K. et al. Sex differences in outcomes of very low birthweight infants: the newborn male disadvantage. Arch. Dis. Child Fetal Neonatal Ed. 83, F182–F185 (2000).
O’Shea, T. M. et al. The Elgan study of the brain and related disorders in extremely low gestational age newborns. Early Hum. Dev. 85, 719–725 (2009).
Santos, H. P., Jr. et al. Epigenome-wide DNA methylation in placentas from preterm infants: association with maternal socioeconomic status. Epigenetics 14, 751–765 (2019).
Aryee, M. J. et al. Minfi: a flexible and comprehensive bioconductor package for the analysis of infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Triche, T. J. Jr., Weisenberger, D. J., Van Den Berg, D., Laird, P. W. & Siegmund, K. D. Low-level processing of illumina infinium DNA methylation beadarrays. Nucleic Acids Res. 41, e90 (2013).
Fortin, J. P., Triche, T. J. & Hansen, K. D. Preprocessing, normalization and integration of the illumina humanmethylationepic array with minfi. Bioinformatics 33, 558–560 (2017).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Du, P. et al. Comparison of beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinforma. 11, 587 (2010).
Jackson, W. M. et al. Risk factors for chronic lung disease and asthma differ among children born extremely preterm. Pediatr. Pulmonol. 53, 1533–1540 (2018).
Hecht, J. L. et al. Characterization of chorioamnionitis in 2nd-trimester c-section placentas and correlation with microorganism recovery from subamniotic tissues. Pediatr. Dev. Pathol. 11, 15–22 (2008).
Chen, J. et al. Fast and robust adjustment of cell mixtures in epigenome-wide association studies with Smartsva. BMC Genomics 18, 413 (2017).
Stekhoven, D. J. & Bühlmann, P. Missforest-non-parametric missing value imputation for mixed-type data. Bioinformatics 28, 112–118 (2012).
Teschendorff, A. E. & Zheng, S. C. Cell-type deconvolution in epigenome-wide association studies: a review and recommendations. Epigenomics 9, 757–768 (2017).
Phipson, B., Lee, S., Majewski, I. J., Alexander, W. S. & Smyth, G. K. Robust hyperparameter estimation protects against hypervariable genes and improves power to detect differential expression. Ann. Appl. Stat. 10, 946–963 (2016).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Manokhina, I., Del Gobbo, G. F., Konwar, C., Wilson, S. L. & Robinson, W. P. Review: placental biomarkers for assessing fetal health. Hum. Mol. Genet. 26, R237–r245 (2017).
Meakin, C. J. et al. Placental Cpg methylation of Hpa-axis genes is associated with cognitive impairment at age 10 among children born extremely preterm. Horm. Behav. 101, 29–35 (2018).
Clark, J. et al. Associations between placental Cpg methylation of metastable epialleles and childhood body mass index across ages one, two and ten in the extremely low gestational age newborns (Elgan) cohort. Epigenetics 14, 1102–1111 (2019).
Adighibe, O. & Pezzella, F. The role of Jmy in P53 regulation. Cancers 10, 173 (2018).
Levy, R. et al. Trophoblast apoptosis from pregnancies complicated by fetal growth restriction is associated with enhanced P53 expression. Am. J. Obstet. Gynecol. 186, 1056–1061 (2002).
Scifres, C. M. & Nelson, D. M. Intrauterine growth restriction, human placental development and trophoblast cell death. J. Physiol. 587, 3453–3458 (2009).
Alvira, C. M. Aberrant pulmonary vascular growth and remodeling in bronchopulmonary dysplasia. Front. Med. 3, 21 (2016).
Baker, C. D. & Abman, S. H. Impaired pulmonary vascular development in bronchopulmonary dysplasia. Neonatology 107, 344–351 (2015).
Gallo, L. A., Barrett, H. L. & Dekker Nitert, M. Review: placental transport and metabolism of energy substrates in maternal obesity and diabetes. Placenta 54, 59–67 (2017).
Bizzarri, M., Laganà, A. S., Aragona, D. & Unfer, V. Inositol and pulmonary function. could myo-inositol treatment downregulate inflammation and cytokine release syndrome in Sars-Cov-2? Eur. Rev. Med. Pharm. Sci. 24, 3426–3432 (2020).
Hallman, M., Saugstad, O. D., Porreco, R. P., Epstein, B. L. & Gluck, L. Role of myoinositol in regulation of surfactant phospholipids in the newborn. Early Hum. Dev. 10, 245–254 (1985).
Howlett, A., Ohlsson, A. & Plakkal, N. Inositol in preterm infants at risk for or having respiratory distress syndrome. Cochrane Database Syst. Rev. 7, Cd000366 (2019).
Bigham, A. et al. Identifying signatures of natural selection in tibetan and andean populations using dense genome scan data. PLoS Genet. 6, e1001116 (2010).
Luo, R. et al. DNA methylation subpatterns at distinct regulatory regions in human early embryos. Open Biol. 8 (2018).
Skeffington, K. L. et al. Hypoxia, Ampk activation and uterine artery vasoreactivity. J. Physiol. 594, 1357–1369 (2016).
Yang, X. et al. Ampk hyper-activation alters fatty acids metabolism and impairs invasiveness of trophoblasts in preeclampsia. Cell Physiol. Biochem. 49, 578–594 (2018).
Cvitic, S. et al. The human placental sexome differs between trophoblast epithelium and villous vessel endothelium. PLoS ONE 8, e79233–e79233 (2013).
Maccani, J. Z. & Maccani, M. A. Altered placental DNA methylation patterns associated with maternal smoking: current perspectives. Adv. Genomics Genet. 2015, 205–214 (2015).
Murphy, S. K., Huang, Z. & Hoyo, C. Differentially methylated regions of imprinted genes in prenatal, perinatal and postnatal human tissues. PLoS ONE 7, e40924 (2012).
Slieker, R. C. et al. DNA methylation landscapes of human fetal development. PLoS Genet. 11, e1005583 (2015).
van Eijk, K. R. et al. Genetic analysis of DNA methylation and gene expression levels in whole blood of healthy human subjects. BMC Genomics 13, 636 (2012).
Higgins, R. D. et al. Bronchopulmonary dysplasia: executive summary of a workshop. J. Pediatr. 197, 300–308 (2018).
Cardenas, A. et al. Placental DNA methylation adaptation to maternal glycemic response in pregnancy. Diabetes 67, 1673–1683 (2018).
Bourque, D. K., Avila, L., Penaherrera, M., von Dadelszen, P. & Robinson, W. P. Decreased placental methylation at the H19/Igf2 imprinting control region is associated with normotensive intrauterine growth restriction but not preeclampsia. Placenta 31, 197–202 (2010).
Xiao, X. et al. Fetal growth restriction and methylation of growth-related genes in the placenta. Epigenomics 8, 33–42 (2016).
Czamara, D. et al. Betamethasone administration during pregnancy is associated with placental epigenetic changes with implications for inflammation. Clin. Epigenet. 13, 165 (2021).
Stoll, B. J. et al. Trends in care practices, morbidity, and mortality of extremely preterm neonates, 1993-2012. JAMA 314, 1039–1051 (2015).
Acknowledgements
We would like to acknowledge the ELGAN study participants for their time, effort, and generous provision of biosamples. As well, we acknowledge the clinical staff for the collection of placentas. This research was made possible through funding from the NIH including R01HD092374, UH3OD023348, and UG3OD023348.
Author information
Authors and Affiliations
Contributions
Each author made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data. All authors have approved the final version of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent statement
Mothers provided informed consent for the ELGAN Study following admission to the hospital, before birth, or immediately following birth. Enrollment and consent procedures were approved by the institutional review boards at all participating sites.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Jackson, W.M., Santos, H.P., Hartwell, H.J. et al. Differential placental CpG methylation is associated with chronic lung disease of prematurity. Pediatr Res 91, 1428–1435 (2022). https://doi.org/10.1038/s41390-021-01868-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-021-01868-x
This article is cited by
-
Epigenetic associations in HPA axis genes related to bronchopulmonary dysplasia and antenatal steroids
Pediatric Research (2024)
-
Advocacy for research starting early in the life course
Pediatric Research (2022)