Abstract
Background
Biallelic deleterious variants in RTTN, which encodes rotatin, are associated with primary microcephaly, polymicrogyria, seizures, intellectual disability, and primordial dwarfism in human infants.
Methods and results
We performed exome sequencing of an infant with primary microcephaly, pontocerebellar hypoplasia, and intractable seizures and his healthy, unrelated parents. We cultured the infant’s fibroblasts to determine primary ciliary phenotype.
Results
We identified biallelic variants in RTTN in the affected infant: a novel missense variant and a rare, intronic variant that results in aberrant transcript splicing. Cultured fibroblasts from the infant demonstrated reduced length and number of primary cilia.
Conclusion
Biallelic variants in RTTN cause primary microcephaly in infants. Functional characterization of primary cilia length and number can be used to determine pathogenicity of RTTN variants.
Similar content being viewed by others
Introduction
Autosomal recessive primary microcephaly (MCPH) is a rare, heterogeneous neurodevelopmental disorder characterized by developmental disruption of brain growth including reduced cerebral cortex, simplified gyri, reduced white matter volume, abnormalities of the corpus callosum, and intellectual disability.1,2 At least 18 genes have been linked to MCPH,3 with variants in ASPM4 and WDR625 identified most frequently.3,6 Most of these genes encode components of basal bodies and centrosomes,7 illustrating the significant role of primary cilia in normal brain development.
RTTN encodes rotatin, a centrosome-associated protein that co-localizes to the basal bodies of primary cilia8 and is required for appropriate expression of nodal, lefty2, and pitx2 in the left lateral plate mesoderm of the developing mouse embryo.9 Mouse embryos lacking rotatin demonstrate abnormal heart looping, delayed neural tube closure, and alterations of left–right-sidedness.9,10 Biallelic RTTN variants have been identified among infants and children with primary microcephaly, polymicrogyria, seizures, intellectual disability, and somatic growth impairment (Table 1),7,8,11 and fibroblasts from these individuals demonstrate shortened cilia.8 We present a male infant with primary microcephaly, simplified gyri, pontocerebellar hypoplasia, contractures, and intractable epilepsy with a novel missense variant and a rare, intronic variant in RTTN that results in an aberrantly spliced transcript, and reduced length and number of cilia in fibroblasts. Our data expand the genotypic and phenotypic spectrum for MCPH that results from genetic disruption of RTTN and demonstrate the usefulness of ciliary length and number for functional characterization of RTTN missense variants.
Methods
Clinical report
A male infant of European-descent was born at 33 weeks gestation to a 38-year-old primigravid mother whose pregnancy was complicated by the antenatal detection of calcifications of the fetal liver and cardiac intraventricular septum with non-diagnostic maternal serum Toxoplasma gondii and cytomegalovirus studies, shortening of the fetal long bones, polyhydramnios, pregnancy-induced hypertension, pre-pregnancy maternal hyperthyroidism treated with thyroidectomy and thyroid hormone replacement, and preterm, premature rupture of membranes for which mother received antenatal corticosteroids and magnesium. The infant was delivered via cesarean section due to non-reassuring fetal surveillance. The infant required resuscitation at birth including intubation and mechanical ventilation. Family history was significant for a paternal grandfather with childhood seizures and a maternal grandfather with young-onset Parkinson’s disease.
birth measurements were: weight 1710 g (−0.8 standard deviation (SD) below mean), length 38 cm (−2.1 SD), and occipitofrontal circumference (OFC) 28 cm (−1.5 SD). His measurements at 6 weeks of age were: weight 1950 g (−3.4 SD), length 42.5 cm (−3.4 SD), and OFC 27.5 cm (−4.9 SD). Of note, OFC measurement at 6 weeks (27.5 cm) was decreased from birth (28 cm), possibly related to neonatal caput succedaneum or inter-individual differences in measurement. OFC measurement at 6 weeks was confirmed by a clinical geneticist and a pediatric neurologist. Physical findings included relative microcephaly with metopic ridging, occipital prominence, bilateral microophthalmia, reactive pupils, microstomia, microretrognathia, smooth philtrum, relatively large, cupped, low-set ears, bilateral contractures of knees and ankles, mild camptodactyly with contractures of interphalangeal joints, bilateral syndactyly of fourth and fifth fingers and second to fifth toes, microphallus, cryptorchidism, and appendicular hypertonia with normal deep tendon reflexes.
Shortly after birth, the infant developed clinical seizures with head turning and extension of all extremities. His electroencephalogram demonstrated suppressed background and intermittent burst suppression that arose independently from both hemispheres. Despite aggressive anti-epileptic treatment with phenobarbital, fosphenytoin, levetiracetam, lorazepam, midazolam, pyridoxine, leucovorin, vigabatrin, and topiramate, his seizures persisted with multiple events per day. He died at 4 months of age after developing acute, progressive respiratory failure.
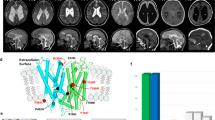
His diagnostic evaluation included magnetic resonance imaging of his brain that was notable for cerebral hypoplasia with simplified gyral pattern, pontocerebellar hypoplasia, bilateral frontal cortical dysplasia, agenesis of the corpus callosum, thinning of the periventricular white matter with ex vacuo dilatation of the occipital and temporal horns, misshapen orbital globes, and optic nerve hypoplasia (Fig. 1). Ophthalmologic evaluation demonstrated rudimentary retinal vasculature, hypoplastic optic nerves, and pale optic disks. Skeletal radiographs demonstrated gracile appearing bones with thin ribs, hypoplastic mandible, increased density of the temporal bones, and soft tissue syndactyly. Renal ultrasound showed bilateral pyelocaliectasis. Cytomegalovirus and Toxoplasma gondii studies, serum amino acids, urine organic acids, lactate, pyruvate, thyroid studies, 7-dehydrocholesterol (7-DHC) reductase (to exclude Smith-Lemli Opitz syndrome), routine newborn screening, and chromosomal microarray analysis were non-diagnostic. Autopsy was not performed.
MRI findings at term equivalent age. T1-weighted sagittal view (a) demonstrates cerebellopontine hypoplasia and micrognathia. T2-weighted transverse view (b) demonstrates enlarged occipital horns and agenesis of the corpus callosum with large third ventricle. T2-weighted coronal view (c) demonstrates typical ventricular configuration for agenesis of the corpus callosum with superior extension of an enlarged third ventricle. All three views demonstrate microcephaly and markedly delayed folding pattern for age
Exome sequencing
This study was approved by the Human Research Protection Office at Washington University. After parental informed consent was obtained, genomic DNA was isolated from the proband’s skin fibroblasts and from parental saliva. Exome capture was performed using the Nimblegen VCRome v2.1 Exome kit (Roche, Madison, WI) with paired-end sequencing (2 × 125 bp) on an Illumina HiSeq 2500 instrument (Illumina, San Diego, CA). Sequence reads were aligned to the human reference genome sequence (GRCh37/hg19) with 90% of the exome having at least 20× coverage. Variants were annotated with Annovar (http://annovar.openbioinformatics.org/en/latest/).12 Variants in coding regions and near exon-intron junctions that were novel or rare (minor allele frequency less than 0.01 in the Exome Aggregation Consortium (ExAC) database (exac.broadinstitute.org))13 were assessed for predicted pathogenicity using Combined Annotation Dependent Depletion (CADD, cadd.gs.washington.edu),14 SIFT (sift.jcvi.org),15 Polyphen2 (genetics.bwh.harvard.edu/pph2/),16 LRT (genetics.wustl.edu/jflab/lrt_query.html),17 MutationTaster (www.mutationtaster.org),18 GERP++ (mendel.stanford.edu/SidowLab/downloads/gerp/),19 and PhyloP (http://compgen.cshl.edu/phast/help-pages/phyloP.txt).20 Exonic variants were classified as deleterious if predicted to be pathogenic by the majority of these programs. We used the dbscSNV database21 within Annovar to assess variants in splicing consensus regions. We evaluated for de novo, autosomal recessive, and X-linked recessive transmission, and candidate genes were reviewed for possible associations with the clinical phenotype.
Exon level oligo array comparative genomic hybridization (CGH) (ExonArrayDx) for the coding exons of the candidate gene was performed in a clinical laboratory (GeneDx, Gaithersburg, MD) on DNA obtained from the proband. Probe sequences and location were based on human genome build 19. Array CGH alterations were reported according to the International System for Human Cytogenetic Nomenclature (ISCN) guidelines.
Transcript characterization
We extracted RNA from the proband’s skin fibroblasts (RNeasy RNA extraction kit (Qiagen, Germantown, MD)) and from parental peripheral blood (PaxGene RNA tubes and blood RNA kit (Qiagen, Germantown, MD)) and synthesized cDNA using SuperScript III (Invitrogen, Carlsbad, CA). To assess RNA splicing, we designed PCR primers that would specifically amplify cDNA that includes exons 1 through 5, spanning several splice junctions. To test for possible aberrant splicing from a cryptic splice site in intron 1, we designed a primer pair in which the forward primer was located in intron 1, and the reverse primer in exon 3. To characterize RNA splicing further, we ligated Illumina adaptors to our PCR products, and performed deep resequencing on an Illumina Miseq instrument (Illumina, Carlsbad, CA).
To evaluate for aberrant splicing in a larger cohort, we queried the GTex database (gtexportal.org) which contains 8812 RNA-seq BAM files from 551 individuals for 55 tissue types to tabulate raw read counts for splice junctions. Briefly, data from each RNA-seq experiment were filtered to obtain spliced reads (containing the ‘N’ CIGAR operation) with mapping quality >20 that overlap the genomic interval of interest. Splice junctions were calculated from read mapping positions and the gapped alignment information contained with the CIGAR string. The number of reads supporting each unique junction was counted for each sample. Splice junctions were compared to Refseq gene annotations to define canonical versus non-canonical splicing events (e.g., exon skipping, alternate donor/acceptor sites, or alternate transcription start sites).
Ciliary staining and characterization
We cultured fibroblasts from the proband and a healthy control on coverslips maintained with 10% fetal bovine serum (FBS)/Dulbecco’s minimal essential medium (DMEM). After fibroblasts reached 90% confluence, we arrested cell growth and facilitated ciliogenesis by reducing FBS/DMEM concentration (0.5%) for 48 h. To assess cilia length and number, we performed immunofluorescent staining with anti-γ tubulin (to mark centrosomes) and anti-acetylated tubulin (to mark cilia). Briefly, after cell fixation with pre-chilled methanol (−20 °C for 10 min) and washing with PBS and 0.1% Triton X-100 in PBS (PBS-T), cells were stained with monoclonal IgG1 anti-γ tubulin (1:1000, Sigma, St. Louis, MO) and monoclonal IgG2b anti-acetylated tubulin (1:10,000, Sigma, St. Louis, MO) for 1 h. After washing cells with PBS-T, cells were incubated with goat anti-mouse IgG1 Alexa Fluor 488 and goat anti-mouse IgG2b Alexa Fluor 594 (Invitrogen, Carlsbad, CA) for 1 h. Images were captured with a Nikon Eclipse Ti-E inverted microscope and a wide-field immunofluorescence microscope. At least 100 cells were scored for the presence or absence of cilia. We used Chi square testing to compare the percentage of ciliated cells. We used Image J (https://imagej.net) to measure cilia length and Student’s t-test to compare cilia length in the proband and control fibroblasts.
Magnetic resonance imaging scanning
Images were collected on a Siemens Magnetom Trio 3T scanner. Structural images were collected with an axial magnetization-prepared rapid gradient-echo (MP-RAGE) T1-weighted sequence (time of repetition/echo time [TR/TE] 1550/3.05 ms and voxel size 1 mm3) and a turbo spin-echo T2-weighted sequence (TR/TE 8950/161 ms, voxel size 1 mm3, and echo train length 15).
Results
Exome sequencing
We identified two rare variants in RTTN in the proband inherited in trans: a novel, predicted deleterious (CADD score = 33)14 missense variant, c.190G>T; p.Val64Phe, inherited from mother that is not present in the ExAC database (accessed May, 2018) and a rare, intronic variant (c.32-3C>T) inherited from father (frequency of 0.0073 in ExAC).(Supplemental Figure S1) Clinical exon level oligo array CGH did not detect any deletions or duplications that include the RTTN locus.
Transcript characterization
We speculate that the c.32-3C>T variant results in leaky splicing22 that concurrently produces canonical and aberrantly spliced (exon 2 skipping and activation of a cryptic splice site with partial intron 1 retention) cDNA transcripts. To assess exon 2 skipping, we designed primers to amplify the cDNA region from exon 1 to exon 5 of RTTN (primers listed in Supplement). Agarose gel electrophoresis (Supplemental Figure S2a) revealed an expected band size from the canonical transcript of 601 base pairs (bp) (Supplemental Figure S3a), but also a much lighter intensity , smaller band at 407 bp in all samples. Gel extraction and Sanger sequencing of individual bands confirmed that the 601 bp band was the expected canonical transcript, and the 407 bp lower band lacked exon 2 (Supplemental Figure S3b). The identification of the exon 2 missense variant from the proband’s 601 bp band indicates that some canonical splicing is occurring from both alleles. Since the sample from the mother, who is not a carrier of the c.32-3C>T splice variant, also showed a much lighter intensity 407 bp band consistent with exon 2 skipping, we performed deep next-generation sequencing on the PCR products to assess the relative amounts of canonical and aberrantly spliced alleles in each member of the family and in four unaffected control samples. We detected much more aberrantly spliced cDNA in the proband and father than in the mother and controls (Supplemental Table).
To assess the possibility of leaky activation of a cryptic splice site upstream in intron 1, we designed a primer pair in which the 5′ (forward) primer was located in intron 1, and the 3′ (reverse) primer was located in exon 3. PCR amplification of cDNA produced a band only in the samples from the subjects who carried the c.32-3C>T variant (proband and father) and not from the mother’s sample (Supplemental Fig. 2b). Gel extraction and Sanger sequencing of this amplicon revealed intron 1 sequence, but canonical exon2–exon3 splicing, suggesting this variant also results in retention of intron 1 (Supplemental Figure S3c). This finding indicates that the c.32-3>T variant also results in an aberrantly spliced transcript from an upstream alternate splice site. However, the very small size of the exon 1–intron 1 region (~60 bp) precluded sufficient amplification and genomic resolution to confirm this additional aberrant transcript sequence.
As we detected some aberrant transcript in the samples from the mother and three controls and an alternate splice site in the samples from the proband and the father, we queried the GTex database to tabulate raw read counts for splice junctions involving exons 1–3 of RTTN. We found 146 (0.8%) reads in which exon 2 was skipped compared to 18,330 reads which demonstrated canonical exon 1–exon 2 splicing, demonstrating a low level of aberrant splicing in unrelated individuals. Of note, the canonical junction is supported by a mean of only 2 reads per sample in the GTex database, in contrast to our NextGen data with greater than 300× coverage per sample. In addition, the aberrantly spliced transcript is smaller than the canonical transcript (407 bp vs. 601 bp) and is likely to be preferentially amplified.
Immunofluorescent staining
Biallelic variants in RTTN have been shown to cause defects in ciliary assembly.8 We identified fewer ciliated cells (28% ciliated vs. 89% ciliated, p < 0.0001) and shorter cilia (2.4 ± 0.8 µm vs. 3.7 ± 1.5 µm, p < 0.0001) from the proband fibroblasts as compared to the control fibroblasts (Fig. 2) in three independent experiments.
Comparison of confocal immunofluorescence microscopy images of control (a) and proband fibroblasts (b) demonstrate fewer ciliated cells and shortened cilia in the proband fibroblasts. Green fluorescent-labeled anti-gamma tubulin was used to stain centrosomes, red fluorescent-labeled anti-acetylated tubulin was used to stain primary cilia, and DAPI was used to stain nuclei
Discussion
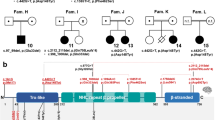
Rotatin, a 2226 amino acid protein, contains two highly conserved, armadillo-type fold domains that mediate protein–protein interaction.8,9,10 Both the c.190G>T (p.Val64Phe) and the c.32-3C>T variants are located in the genomic region that encodes the first armadillo-type fold domain. Other variants (p.Cys27Tyr, p.Ala578Pro, p.His865Arg) within this region have been previously identified among individuals with microcephaly, abnormal gyri, and seizures6,7,8 and suggest the importance of this domain for normal function and formation of primary cilia and for subsequent neurodevelopment. The RTTN variants identified among infants and children with primary microcephaly have been missense or splicing variants, suggesting that biallelic null variants may be embryonic lethal as described for another microcephalic primordial dwarfism gene DONSON.23
In silico splicing software predicts that the native 3′ splice site near exon 2 of the RTTN gene is relatively weak, and that the c.32-3C>T variant further reduces the strength of this site (Alamut®). Our experimental results suggest that a very low rate of exon 2 aberrant splicing occurs in unrelated, healthy individuals, but the c.32-3C>T variant substantially increases the rate of aberrant splicing. Unlike the novel p.Val64Phe variant, the c.32-3C>T variant is present in 882 heterozygous individuals in ExAC including four homozygous individuals without available clinical phenotype information. We demonstrate that the c.32-3C>T variant reduces the strength of the native splice site and results in leaky splicing22 in which some canonically spliced product is produced. We speculate that the individuals homozygous for this variant produce sufficient canonical transcript for neurodevelopmental viability. However, when in trans with a deleterious variant, a single c.32-3C>T variant is insufficient to achieve a threshold of expression to permit normal neurodevelopment. Shortened ciliary length of our patient’s fibroblasts has been previously identified among other patients with biallelic RTTN variants8 and supports the pathogenicity of these RTTN variants. The decreased number of ciliated fibroblasts from our patient has not been previously observed among fibroblasts from patients with biallelic RTTN variants.
Murine embryos with genetically abrogated rotatin expression demonstrate randomized heart looping, delayed neural tube closure, and abnormalities of left–right-sidedness.9,10 The lack of congenital heart disease or situs abnormalities in our patient and in previously reported MCPH patients with RTTN variants suggests species-specific differences in the role of RTTN, fetal lethality associated with RTTN-mediated cardiac abnormalities, genetic redundancy for RTTN encoded functions during human cardiac development, or differences in phenotypic consequences between complete null alleles (murine model) and biallelic missense/splice variants observed among the infants and children. Genetic disruption of RTTN and its Drosophila homolog Ana3 expression in murine and fly models, respectively, demonstrates the importance of rotatin in brain development.9,24 Ana3 also localizes to centrioles and basal bodies, and Ana3-deficient flies are severely uncoordinated and die soon after larval emergence, a phenotype similar to mutants of other centriolar and basal body structural proteins which result in defective cilia of type I neurons.24
The neurologic (microcephaly, simplified gyral pattern, agenesis of the corpus callosum, cortical dysplasia, reduced white matter, and contractures) and extra-central nervous system (bilateral cryptorchidism, microphallus, and renal pyelocaliectasis) findings in our patient are similar to that of the previously reported patients with biallelic RTTN variants.7,8,11 However, his ophthalmologic findings (misshapen orbital globes, optic nerve hypoplasia, rudimentary retinal vasculature, and pale optic disks) and skeletal findings (gracile appearing bones, thin ribs, hypoplastic mandible, increased density of the temporal bones, and syndactyly) have not been previously reported.
In conclusion, our results support and extend the association of biallelic RTTN variants with severe primary microcephaly and emphasize the usefulness of functional characterization of primary cilia length and number for assessment of pathogenicity of RTTN variants. While some individuals with RTTN-related microcephaly have survived into adolescence or even early adulthood,7,8,11 our patient had intractable epilepsy and died at 4 months of age. While earlier identification of a candidate gene to account for his phenotype would likely not have changed his anti-epileptic management or his disease course, genomic diagnosis permitted prediction of recurrence risk and prenatal or pre-implantation genetic diagnosis.
References
Mochida, G. H. & Walsh, C. A. Molecular genetics of human microcephaly. Curr. Opin. Neurol. 14, 151–156 (2001).
Adachi, Y. et al. Congenital microcephaly with a simplified gyral pattern: associated findings and their significance. AJNR Am. J. Neuroradiol. 32, 1123–1129 (2011).
Zaqout, S., Morris-Rosendahl, D. & Kaindl, A. M. Autosomal recessive primary microcephaly (MCPH): an update. Neuropediatrics 48, 135–142 (2017).
Desir, J., Cassart, M., David, P., Van Bogaert, P. & Abramowicz, M. Primary microcephaly with ASPM mutation shows simplified cortical gyration with antero-posterior gradient pre- and post-natally. Am. J. Med. Genet. A 146A, 1439–1443 (2008).
Bhat, V. et al. Mutations in WDR62, encoding a centrosomal and nuclear protein, in Indian primary microcephaly families with cortical malformations. Clin. Genet. 80, 532–540 (2011).
Rump, P. et al. Whole-exome sequencing is a powerful approach for establishing the etiological diagnosis in patients with intellectual disability and microcephaly. BMC Med. Genomics 9, 7 (2016).
Shamseldin, H. et al. RTTN mutations cause primary microcephaly and primordial dwarfism in humans. Am. J. Hum. Genet. 97, 862–868 (2015).
Kheradmand Kia, S. et al. RTTN mutations link primary cilia function to organization of the human cerebral cortex. Am. J. Hum. Genet. 91, 533–540 (2012).
Faisst, A. M., Alvarez-Bolado, G., Treichel, D. & Gruss, P. Rotatin is a novel gene required for axial rotation and left-right specification in mouse embryos. Mech. Dev. 113, 15–28 (2002).
Chatterjee, B., Richards, K., Bucan, M. & Lo, C. Nt mutation causing laterality defects associated with deletion of rotatin. Mamm. Genome 18, 310–315 (2007).
Grandone, A. et al. Expanding the phenotype of RTTN variations: a new family with primary microcephaly, severe growth failure, brain malformations and dermatitis. Clin. Genet. 90, 445–450 (2016).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Ng, P. C. & Henikoff, S. Predicting deleterious amino acid substitutions. Genome Res. 11, 863–874 (2001).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Chun, S. & Fay, J. C. Identification of deleterious mutations within three human genomes. Genome Res. 19, 1553–1561 (2009).
Schwarz, J. M., Cooper, D. N., Schuelke, M. & Seelow, D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat. Methods 11, 361–362 (2014).
Davydov, E. V. et al. Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLoS Comput. Biol. 6, e1001025 (2010).
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
Liu, X., Wu, C., Li, C. & Boerwinkle, E. dbNSFPv3.0: a one-stop database of functional predictions and annotations for human nonsynonymous and splice-site SNVs. Hum. Mutat. 37, 235–241 (2016).
Rogan, P. K., Faux, B. M. & Schneider, T. D. Information analysis of human splice site mutations. Hum. Mutat. 12, 153–171 (1998).
Reynolds, J. J. et al. Mutations in DONSON disrupt replication fork stability and cause microcephalic dwarfism. Nat. Genet. 49, 537–549 (2017).
Stevens, N. R., Dobbelaere, J., Wainman, A., Gergely, F. & Raff, J. W. Ana3 is a conserved protein required for the structural integrity of centrioles and basal bodies. J. Cell Biol. 187, 355–363 (2009).
Acknowledgements
The authors thank the Exome Aggregation Consortium Database; a full list of contributing groups can be found at http://exac.broadinstitute.org/about. The authors thank the Genotype-Tissue Expression (GTEx) Project, which was supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS. The data used for the analyses described in this manuscript were obtained from the GTEx Portal on 1 May 2018. This work was supported by grants from the National Institutes of Health (K08 HL105891 (J.A.W.), K12 HL120002 (F.S.C.), R21/33 HL120760 (F.S.C.)), R01 HL128370 (M.R.M.), the Eunice Kennedy Shriver National Institute of Child Health & Human Development (U54 HD087011 (J.S.S.), the Children’s Discovery Institute (F.S.C., J.A.W., M.R.M.), and the Saigh Foundation (F.S.C.).
Author information
Authors and Affiliations
Contributions
Conception and design: J.A.W., D.J.W., M.S., D.B., and F.S.C. Acquisition, analysis, or interpretation of data: J.A.W., D.J.W., P.Y., M.S., D.B., M.V.A., E.B., J.S.S., D.S., M.R.M., and F.S.C. Drafting and revising the manuscript for important intellectual content: J.A.W., D.J.W., P.Y., M.S., D.B., E.B., J.S.S., D.S., B.P.H., M.V.A., T.F., S.K.D., M.R.M., and F.S.C. All authors have approved the final version and agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All authors have seen and approved the submission and take full responsibility for the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Wambach, J.A., Wegner, D.J., Yang, P. et al. Functional characterization of biallelic RTTN variants identified in an infant with microcephaly, simplified gyral pattern, pontocerebellar hypoplasia, and seizures. Pediatr Res 84, 435–441 (2018). https://doi.org/10.1038/s41390-018-0083-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-018-0083-z
This article is cited by
-
Pontocerebellar Hypoplasia: a Pattern Recognition Approach
The Cerebellum (2020)