Abstract
Semaphorin 6A (SEMA6A), a membrane-bound protein, is downregulated in lung cancer tissue compared to its adjacent normal tissue. However, the functions of SEMA6A in lung cancer cells are still unclear. In the present study, full length SEMA6A and various truncations were transfected into lung cancer cells to investigate the role of the different domains of SEMA6A in cell proliferation and survival, apoptosis, and in vivo tumor growth. SEMA6A-induced cell signaling was explored using gene silencing, co-immunoprecipitation, and co-culture assays. Our results showed that overexpression of SEMA6A reduced the growth of lung cancer cells in vitro and in vivo, and silencing SEMA6A increased the proliferation of normal lung fibroblasts. Truncated SEMA6A lacking the SEMA domain or the extracellular region induced more apoptosis than full length SEMA6A, and reintroducing the SEMA domain attenuated the apoptosis. Fas-associated protein with death domain (FADD) bound to the cytosolic region of truncated SEMA6A and was involved in SEMA6A-associated cytosol-induced apoptosis. This study suggests a novel function of SEMA6A in inducing apoptosis via FADD binding in lung cancer cells.
Similar content being viewed by others
Introduction
Dysregulation of apoptotic pathways can lead to tumorigenesis through transformation of normal cells to malignant cells1. The apoptotic pathways are initiated by death receptors (DRs) on the cell membrane. Whereas, DRs are triggered by their extracellular ligands, molecules such as Fas-associated protein with death domain (FADD) or tumor necrosis factor receptor type 1-associated death domain (TRADD) are recruited to the cytosolic region of the DRs and subsequently activate the downstream death signaling2. The loss-of-function mutants of DRs are unable to recruit FADD and are thus inefficient in inducing apoptosis2. Furthermore, inactivation of DRs by mutations also shows a high association with some types of cancer, suggesting the importance of DRs in tumorigenesis3. However, DRs are not the only apoptosis-initiating proteins. Other membrane proteins like the semaphorin family may participate in regulation of apoptotic signaling4,5.
The semaphorin family includes seven subfamilies, all of which contain a characterized SEMA domain. The semaphorin family were initially identified as ligands that control the guidance of axons by directly binding to plexin, thus activating plexin-derived signaling6,7,8,9. Recently, several studies have demonstrated that the semaphorins have a role in tumor progression. For example, semaphorin 4D can act as a pro-tumorigenic factor that induces tumor angiogenesis in head and neck squamous cancer cells10. In contrast, several semaphorins, such as the semaphorin 3 subfamily, function as anti-tumorigenic factors by inducing apoptosis and inhibiting cell proliferation in lung cancer4,11,12, breast cancer5, and skin cancer cells13.
Previously, we observed that semaphorin 6A (SEMA6A), a single-pass transmembrane protein involved in the axonal guidance pathway14,15,16,17,18, was significantly downregulated in lung cancer tissues as compared to adjacent normal tissues19. Up until now, only a few studies have examined the role of SEMA6A in cancer biology19, and only one study reported that the extracellular region of SEMA6A could inhibit tumor formation via decreasing VEGF-induced xenograft vascularization20. Based on our prior results and another report mentioning that a somatic deletion in SEMA6A occurs at locus 5q23.1 in lung cancer cells21, in this study, it is hypothesized that SEMA6A might play a role in lung carcinogenesis.
Therefore, with the goal of better understanding the importance of SEMA6A in lung cancer cells, in vitro cell proliferation, clonogenic and apoptosis assays, and in vivo xenograft animal experiments were performed to examine the functions of SEMA6A. Moreover, different truncations of SEMA6A were overexpressed in lung cancer cells to study the functions of the parts of SEMA6A. Finally, SEMA6A-related signaling was delineated by co-immunoprecipitation, co-culture, and gene silencing assays. Through our study we aimed to identify the pleiotropic effects of SEMA6A in lung cancer cells, which could be regarded as a potential therapeutic target for lung cancer treatment.
Results
SEMA6A is downregulated in lung tumor tissues
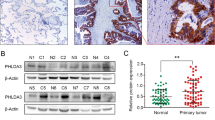
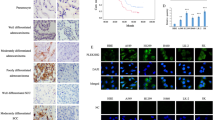
Our previous microarray results19 showed SEMA6A to be significantly downregulated in lung tumor tissue compared to adjacent normal tissue (Table S1). The results were consistently replicated by RT-qPCR (Fig. 1a). Among 172 lung cancer patients, lung cancer tissues from 73.26% of patients were failed to stain with SEMA6A antibody, whereas 26.74% patient samples were stained positively (Fig. 1b). Furthermore, three datasets from Gene Expression Omnibus (GEO)22,23,24 showed similar results (Fig. S1). In addition, endogenous SEMA6A was undetectable in several lung cancer cell lines, including A549, H1299, H1975, H441, and H520 (Fig. 1c).
a Validation of SEMA6A expression in lung adenocarcinoma samples (n = 60). b Quantitative analysis of lung cancer specimens (n = 172) for SEMA6A expression. c Endogenous expression levels of SEMA6A (6A) in different lung cell lines. 6A-FL was detected by custom manufactured anti-SEMA6A antibody from GenScript. Loading control: actin. Positive control: 293T and 6A-FL overexpressing 293T
SEMA6A decreases the growth of lung cancer cells
Given the low expression of SEMA6A in lung cancer cells, we studied the effects of SEMA6A overexpression on cell proliferation, colony formation, and apoptosis in A549 and H1299 cells. Cells with full length SEMA6A (6A-FL) overexpression had significantly lower cell proliferation (Fig. 2a), fewer colonies (Fig. 2b), and a higher rate of apoptosis (Fig. 2c) compared to cells transfected with empty vector. In addition, silencing of the SEMA6A gene in the normal lung fibroblast cell line, MRC5, by shSEMA6A (Fig. 2d) increased cell proliferation significantly (Fig. 2e).
a Proliferation of A549 and H1299 cells overexpressing 6A-FL or empty vector (Ctrl). Inset: western blot of His-tagged SEMA6A. Loading control: actin. **P < 0.01, n = 4. b Colony formation of A549 and H1299 cells overexpressing 6A-FL or Ctrl. Upper, quantitative analysis of colony numbers. **P < 0.01, n = 4. c Apoptosis analysis of A549 and H1299 cells overexpressing 6A-FL or Ctrl. **P < 0.01, n = 4. d Relative mRNA expression level of SEMA6A in MRC5 cells expressing SEMA6A shRNA and control shRNA lentiviral vectors. e The proliferation rate of MRC5 cells with or without SEMA6A knockdown. *P < 0.05, **P < 0.01, n = 3. f Tumor volume in mice transfected with 6A-FL- or Ctrl-overexpressing A549 (left) or H1299 (right) cells. *P < 0.05, **P < 0.01, n = 5. g Tumor image, left, and weight, right, from SEMA6A-overexpressing A549 and H1299 xenografts. **P < 0.01, n = 5
In nude mice, overexpression of SEMA6A decreased the growth of A549 and H1299 subcutaneous xenografts (Fig. 2f, g). These results indicate that SEMA6A induces apoptosis in lung cancer cells in vitro and reduces lung cancer cell growth in vivo.
SEMA6A induces apoptosis through its cytosolic domain
Structural and functional analyses were conducted to explore the effects of the extracellular and cytosolic domains and regions of SEMA6A in lung cancer cells. The constructs encoding the transmembrane domain with either the intracellular cytosolic domain (6Acyto, aa 575–1031) or the entire extracellular ectodomain (6Aect, aa 1–704) (Fig. 3a) were overexpressed at similar levels in H1299 and A549 cells (Fig. 3b). Compared to the empty vector cells, the 6Acyto-overexpressing A549 and H1299 cells significantly decreased cell proliferation (Fig. 3c, d), reduced colony formation (Fig. 3e), and induced more apoptosis (Fig. 3f). In contrast, overexpression of 6Aect did not significantly alter cell proliferation (Fig. 3c, d), colony numbers (Fig. 3e), and the percentage of apoptotic cells compared to the empty vector-transfected cells (Fig. 3f). H1299 cells with empty vector, 6Acyto, or 6Aect overexpression were injected subcutaneously into nude mice. The growth rates and sizes of tumors formed by 6Acyto-overexpressing H1299 cells were significantly decreased compared to the 6Aect- and empty vector-transfected H1299 cells in 30 days (Fig. 3g). The tumors with 6Acyto overexpression were much lighter than those formed from the empty vector- and 6Aect-overexpressing cells (Fig. 3g). These findings indicate that SEMA6A-induced in vivo apoptosis is carried out by its cytosolic domain.
a Schematic diagram of the SEMA6A constructs. 6A: full length. 6Aect: extracellular domain of SEMA6A. 6Acyto: cytosolic domain of SEMA6A. b Immunoblotting of lysates of A549 and H1299 cells overexpressing 6A-FL, 6Aect, 6Acyto, or empty vector (Ctrl). All constructs were histidine tagged. Loading control: actin. c and d Proliferation of A549 c and H1299 d cells overexpressing different truncations of SEMA6A. **P < 0.01, n = 4. e Colony formation of A549 and H1299 cells. **P < 0.01, n = 4. f Quantitative analysis of the apoptotic A549 and H1299. **P < 0.01, n = 4. g Tumor volume (left) and weight (right) in mice transfected with 6Aect-, 6Acyto-, or Ctrl-overexpressing H1299 cells. Inset: Tumor image from Ctrl-, 6Aect-, or 6Acyto-overexpressing H1299 xenografts. **P < 0.01, n = 5
Moreover, we found an unnamed EST clone in the NCBI database (http://www.ncbi.nlm.nih.gov/protein/bah13847.1), BAH13847.1, whose amino acid sequence is highly similar to the transmembrane and cytosolic regions of our 6Acyto construct (Fig. S2A). Our data demonstrated that this unnamed EST clone could also induce significant apoptosis in lung cancer cells (Fig. S2B). These results suggest that the cytosolic signaling of SEMA6A may exist in nature to induce apoptosis in cells.
The SEMA domain regulates the induction of apoptosis
The extracellular region of SEMA6A is dimerized via the SEMA domain to induce plexin-related signaling25; however, the other functions of the SEMA domain are still unclear. As 6Acyto triggered more effects on cell proliferation (Fig. 3c, d) and apoptosis (Fig. 3f) than 6A-FL, we tested whether the extracellular SEMA domain is required to modulate effects of SEMA6A cytosolic signaling in lung cancer cells. Therefore, a construct of SEMA6A lacking the SEMA domain (6AΔsema) (Fig. 4a) was overexpressed in both H1299 and A549 cells (Fig. 4b).
a Schematic diagram of the 6AΔsema construct in comparison to 6A-FL. b Immunoblotting analysis of A549 and H1299 cells overexpressing His-tagged 6AΔsema. Loading control: actin. c Percentage of apoptotic A549 and H1299 cells overexpressing 6AΔsema. **P < 0.01, n = 4. d Percentage of apoptotic A549 and H1299 cells overexpressing 6AΔsema after addition of purified SEMA domain. **P < 0.01, n = 4 per indicated concentration. e Percentage of apoptotic A549 and H1299 cells overexpressing 6Acyto after external addition of purified SEMA domain. n = 4 for each concentration. *P < 0.05, **P < 0.01
Overexpression of 6AΔsema-induced apoptosis in 25% of A549 and 29% of H1299 cells, which was comparable to the effect of 6Acyto overexpression (Fig. 4c), thus establishing that 6AΔsema and 6Acyto induce comparable levels of apoptosis. However, when purified SEMA domain (Fig. S3A and B) was added to the cells with 6AΔsema overexpression, apoptosis was significantly attenuated (Fig. 4d). In contrast, there was no difference in apoptosis observed in 6Acyto-overexpressing cells upon addition of purified SEMA domain (Fig. 4e). These results indicated that the degree of apoptosis induced by 6A-FL might be regulated by the SEMA domain.
The cytosolic domain of SEMA6A interacts with FADD
The results described so far are the first evidence that the cytosolic signaling mediated by the cytosolic region of SEMA6A can induce apoptosis and is regulated by the SEMA domain in lung cancer cells. We therefore investigated the mechanism by which apoptosis is activated during this signaling. Previous studies have indicated that the activation of DRs such as CD95 or TRAIL receptor (TRAILR)-1 leads to recruitment and activation of FADD, which in turn leads to cleavage of caspase-8, an apoptosis initiator26,27,28. We therefore proceeded to demonstrate that the overexpression of 6Acyto, but not 6Aect or 6A-FL, could activate the cleaved forms (p43/41 and p18) of caspase-8 in H1299 and A549 cells (Fig. 5a); p43/41 and p18 were detected in H1299 cells with 6AΔsema overexpression as well (Fig. 5b). These results suggest that the membrane-anchored cytosolic domain of SEMA6A might act as a DR by inducing cleavage of caspase-8.
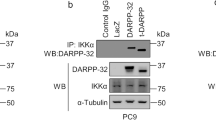
a Immunoblot of caspase-8 cleavage products (p43/41 and p18) in A549 and H1299 cells overexpressing constructs of SEMA6A. b Immunoblotting analysis of p43/41 and p18 in H1299 cells overexpressing 6AΔsema. Co-immunoprecipitation of FADD and 6Acyto c, 6AΔsema d, and 6Aect e in 293T cells. Input: 5% of non-immunoprecipitated cell lysate. IgG: control IP with isotype-matched IgG antibody from rabbit. The input and immunoprecipitates were subjected to western blot analysis using anti-FADD and anti-His antibodies
To investigate whether the apoptotic pathway induced by the cytosolic domain of SEMA6A was similar to these previously identified DRs, 6Acyto, 6AΔsema, and 6Aect were subjected to co-immunoprecipitation with FADD in 293T cells. 6Acyto was pulled down with FADD (Fig. 5c, left) and, conversely, FADD was also co-immunoprecipitated with 6Acyto (Fig. 5c, right). Similarly, 6AΔsema was co-immunoprecipitated with FADD (Fig. 5d, left) and vice versa (Fig. 5d, right). On the contrary, FADD failed to bind to 6Aect (Fig. 5e) and 6A-FL (Fig. S4). Similar results were also observed in H1299 cells (Fig. S5A–C).
The cytosolic domain of SEMA6A induces apoptosis via interaction with FADD
To demonstrate the requirement of FADD in 6Acyto-induced apoptosis, we co-transfected 6Acyto and shRNA targeting FADD in A549 and H1299 cells, which effectively knocksdown endogenous FADD levels (Fig. 6a). Under these conditions, a significant reduction of 6Acyto-induced apoptosis was observed (Fig. 6b). The same effect was observed when a dominant-negative mutant of FADD (DN-FADD, 80–205) that lacks the death domain29 was co-transfected with 6Acyto in A549 and H1299 cells (Fig. 6c, d). These data suggest that the cytosolic domain of SEMA6A attenuates apoptotic signaling through FADD in lung cancer cells.
a Immunoblotting analysis of silencing FADD in A549 and H1299 cells with two FADD shRNAs (shFADD #1 and #2) and empty lentiviral vector (Ctrl). b Percentage of apoptosis in A549 and H1299 cells overexpressing 6Acyto and transfected with shRNAs against FADD. **P < 0.01, n = 3. c Western blot analyses of A549 and H1299 cells overexpressing dominant-negative mutant of FADD (DN-FADD). The upper band is endogenous FADD and the lower band is DN-FADD. d Percentage of apoptosis in 6Acyto-overexpressing A549 and H1299 cells with DN-FADD overexpression. **P < 0.01, n = 3. Co-immunoprecipitation and quantification of FADD and 6AΔsema e and f or 6Acyto g and h in the presence of purified SEMA domain. **P < 0.01, n = 3
Finally, we examined whether the SEMA domain disrupts the interaction of 6Acyto with FADD. Co-immunoprecipitation results showed that the interaction between FADD and 6AΔsema (Fig. 6e, f), but not the interaction between FADD and 6Acyto (Fig. 6g, h), was diminished significantly by the addition of the SEMA domain. These results indicate that the SEMA domain attenuates apoptosis by disrupting the association between FADD and 6AΔsema.
Discussion
Our study has provided solid evidence supporting a novel role of SEMA6A in apoptosis, which can inhibit the growth of lung cancer cells in vitro and in vivo and is regulated by the integral SEMA domain. Previous studies showed that specific protease-induced shedding separates part of the extracellular domain from semaphorins on the plasma membrane, including semaphorin 4, and semaphorin 530,31,32. Therefore, the extracellular region of SEMA6A may act as a safety pin, which when removed through proteolytic shedding (“uncapped” SEMA6A), induces apoptosis. In addition, previous structure-based studies indicated that the SEMA6A ectodomain forms a homodimer arrangement to bind plexin A2, inducing the downstream signaling of plexins14. Based on such evidence, it can be asserted that the extracellular domain of SEMA6A not only modulates responses for plexin-related signaling, but also functions as a key regulator in SEMA6A cytosolic domain-induced apoptosis in lung cancer cells.
In the 6AΔsema and 6Acyto overexpression study, the FADD-associated pathway evidently contributes to SEMA6A-related apoptosis. FADD, a critical adaptor protein, has been indicated to activate caspase-8 through binding to the death domains (DDs) of DRs, including CD95/Fas, DR4/TRAIL-R1, and DR5/TRAIL-R228,33,34. Our data showed that 6Acyto was capable of interacting with FADD, thus leading to cleavage of caspase-8. Membrane proteins that induce apoptosis through FADD’s homotypic interaction with their DDs are generally considered to be DRs. As FADD binds to 6Acyto and not to 6A-FL, the 6A-FL could be regarded as a novel inactive type of DR. However, the potential mechanisms that can activate the DR-like SEMA6A need to be further investigated.
Although repression via shFADD and expression of DN-FADD both significantly reduced 6Acyto-induced apoptosis, the percentage of apoptotic cells remained at more than 15% in both A549 and H1299 cells. Residual endogenous FADD after gene silencing is one possible cause for sustained 6Acyto-induced apoptosis. In addition to residual FADD expression, the other DR-related apoptotic proteins, such as tumor necrosis factor receptor type 1-associated DEATH domain protein, might be involved in 6Acyto-induced apoptosis in lung cancer cells.
Overexpression of SEMA6A increased apoptosis in both lung cancer cells and normal lung fibroblasts. On the other hand, our data showed that silencing of the SEMA6A in MRC5 cells by shSEMA6A increased cell proliferation significantly. In the human body, the total number of cells remains a delicate balance between survival and cell death. A disruption of this balance can trigger the development of cancer35,36. Therefore, decreased expression of SEMA6A in normal lung fibroblasts might be a potential risk factor for tumorigenesis due to the insufficient activation of apoptosis. These findings suggest that the expression level of SEMA6A may be a prognostic biomarker for lung cancer patients.
In conclusion, we have provided the first evidence that the extracellular SEMA domain of 6A-FL appears to function as an attenuator of apoptosis via association of its cytosolic region with FADD in lung cancer cells. These data assert that wild-type SEMA6A is just like a “capped protein” whose ability to induce apoptosis is limited by the cap, i.e., the SEMA domain. Compared to the well-known DRs that can trigger autocrine death or paracrine suicide37, SEMA6A uses a novel mechanism to regulate ligand-independent apoptosis through its SEMA domain.
Materials and methods
Quantitative RT-PCR
After reverse transcription of total RNA from patients’ tissues, qPCR was processed using an ABI7300 system (Applied Biosystems, Foster, CA) with SYBR Green (Roche, Basel, Switzerland) according to the standard protocols. The forward and reverse primers of SEMA6A were 5′-GCTACACTTTGCTGGGGCTAA-3′ and 5′-CGGCTTTGGGTCTTTGGATTG-3′, respectively. The collection of patient samples was described in our previous study18. The institutional review boards of National Taiwan University Hospital and Taichung Veterans General Hospital both approved the sample acquisition and its subsequent use (IRB #200610015R).
Microarray data analysis
Microarray datasets from GSE19804, GSE18842, GSE4079, and GSE19188 were used to identify the expression level of SEMA6A in lung cancer patients. The intensity data of SEMA6A in these studies were analyzed by Partek (Partek, Chesterfield, MO) to obtain mRNA expression levels. Probe-level data of SEMA6A were preprocessed through background correction, quantile normalization, and summarization, using robust multi-array average analysis. The paired t-test (P-value<10−16) was used to identify differentially expressed SEMA6A.
Cell culture
Lung cancer cell lines (A549, H1299, H1975, H441, and H520) were purchased from Bioresource Collection and Research Center (BCRC, Hsinchu, Taiwan). Lung cancer cell lines H1299 and A549 with exogenous luciferase (Luc) activity were kindly provided by Dr. P.W. Hsiao. All cell lines were cultured following the protocols at BCRC and authenticated by STR-PCR profiling at BCRC.
Immunohistochemical staining
A total of 172 lung cancer samples collected from National Taiwan University Hospital (Taipei, Taiwan) and Taichung Veterans General Hospital (Taichung, Taiwan) were used for the examination of SEMA6A expression. The paraffin-embedded tissues were de-paraffinized and reactivated as described earlier38, after which the tissue sections were incubated with normal goat serum (dilution 1:500; Dako) at room temperature for 2 h followed by primary SEMA6A antibody (dilution 1:50; Cat. #AP2740b; Abgent, San Diego, CA) at 4 °C overnight. Streptavidin- and biotin-labeled secondary antibodies were used for staining.
SEMA6A, 6AΔsema, 6Acyto, and 6Aect constructs
The full length SEMA6A (6A-FL) cDNA was generated from normal foreskin fibroblasts, Hs68, using RT-PCR. The cDNAs of the extracellular ectodomain (6Aect, aa 1–704), the intracellular cytosolic domain (6Acyto, aa 575–1031), and the SEMA deletion mutant (6AΔsema, aa 513–1031) were produced from the 6A-FL sequence. The primers are listed in Table S2. The cDNA fragments were then sub-cloned into the lentiviral expression vector pCDH-CMV-MCS-EF1-puro (System Biosciences), and 6A-FL and all domain constructs were fused with a 6X His-tag at the C-terminus of the expressed protein.
Virus production and cell infection
HEK293T cells (4 × 106 cells) were co-transfected with transfer vector (pCDH-CMV-MCS-EF1-puro; 8 μg), packaging plasmid (psPAX2; 6 μg) and envelope plasmid (pMD2G; 2 μg) using TransIT®-2020 (Mirus Bio, Madison, WI). The supernatants containing infectious particles were collected, aliquoted, and stored at −80 °C at 48 h and 72 h post-transfection. The virus was added to 4 × 104 cells, and the cells were centrifuged at 2500 rpm for 60 min at room temperature.
Western blotting
Proteins were extracted using RIPA lysis buffer (Millipore, Billerica, MA) containing protease inhibitor cocktail, 10 mM β-GP, and 5 mM Na3VO4. Protein from lysate was separated by 10% SDS-PAGE and transferred onto a nitrocellulose membrane (Millipore). The blots were probed overnight at 4 °C with primary antibodies against SEMA6A (custom manufactured antibody from GenScript), His-tag (Cat. #05–949; Millipore), β-actin (Cat. # ABT264; Millipore), and cleaved caspase-8 (Cat. #9496S; Cell Signaling). After incubation with the appropriate HRP-conjugated secondary antibodies, HRP activity was visualized by an enhanced chemiluminescence system (UVP, Upland, CA).
MTT assay
Cells were seeded at 2 × 104 cells/well on a 24-well plates and then incubated at 37 °C for 3.5 h with 100 μL of 5 mg/mL MTT (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide, SIGMA, Saint Louis, MO) at specific time points. Subsequently, the cells were shaken with 400 μL MTT solvent (4 mM HCl mixed with isopropanol) at room temperature for 15 min. Absorbance was read at 590 nm with a reference filter of 620 nm by ELISA reader (PerkinElmer, Waltham, MA).
Annexin V analysis for apoptosis
Cells were stained with propidium iodide and annexin V following the manufacturer′s instructions (BD, San Jose, CA). A minimum of 10,000 cells in each sample were detected by a Cytomics FC500 cytometer (Beckman, Brea, CA). Cell apoptosis was analyzed by WinMDI software.
Clonogenic assay
Cells were seeded in a 6 cm dish at 200 cells/dish and incubated for 10 days. Colonies were fixed with 75% alcohol, stained with 20% crystal violet solution, and manually counted. Colony formation was described as follows: colony number/(number of cells seeded × plating efficiency), where plating efficiency was equivalent to the colony number divided by the number of cells with empty virus infection.
Xenograft tumor models
Male nude mice, 4–6 months old, were purchased from the National Laboratory Animal Center (Taipei, Taiwan). The lung cancer cells (3 × 106) were re-suspended in 100 μL medium and mixed with Matrigel at a 1:1 ratio by volume and then injected subcutaneously into the right flank of each mouse. Five mice were randomly divided into each group. After injection, a Vernier caliper was used to measure the tumor size weekly. Tumor volumes were calculated by the formula V = 1/2 (Length × Width2). At the sixth week, the mice were weighed and then killed. The number of mice was determined by pilot experiments between the mice injected with 6A-FL-overexpressing and control lung cancer cells. The experiments were performed in compliance with the protocol (IACUC-20130394) of the Research Ethics Office at National Taiwan University.
Purification of SEMA protein
HEK293T cells were transfected with the vector (pcDH-CMV-MCS-EF1-puro) for SEMA-His by the TransIT-2020 reagent (Mirus). After 3 days, the supernatants were applied to a 3 mL Ni-containing resin. Protein was eluted with 300 mM imidazole, 300 mM NaCl, and 20 mM sodium phosphate, pH 7.4. The eluted sample was passed through a 0.45 μm filter and applied to Sephacryl s-100 h gel filtration columns (GE Healthcare, Little Chalfont, UK) on an ÄKTA FPLC system (GE Healthcare) at a flow rate of 0.5 mL/min in PBS. Eluate was collected as 40 fractions and maintained at 4 °C. Each fraction was assessed for protein by immunoblotting analysis with anti-His mAb (1:5000, Millipore).
Co-immunoprecipitation assay
After 48 h overexpression of 6X His conjugated truncations of SEMA6A, cells were lysed with ice-cold Pierce® IP lysis buffer (Thermo Fisher, Waltham, MA). Cell lysates were incubated with rabbit anti-SEMA6A (custom manufactured antibody from GenScript) or rabbit anti-FADD antibodies at 4 °C for 16 h. Cell lysates were incubated with normal rabbit IgG (Millipore) as control. Then, magnetic beads (Thermo Fisher) were added to cell lysates and incubated overnight at 4 °C for immunoprecipitation. Beads were washed three times with wash buffer, boiled in sample buffer, and analyzed by western blotting.
Silencing of FADD
shFADD clones, TRCN0000332992, and TRCN0000332994, were obtained from the National RNAi Core Facility (Academia Sinica, Taiwan). The production of lentivirus for silencing of FADD was described above.
Statistical analysis
The investigators were randomly allocated and blinded to the different groups during collection of results of each experiment. A two-tailed Student’s t-test was applied to all the data in this study. The variance is similar between the groups in the same experiment. Differences were considered to be significant if the P-value was <0.05. All values in the text and figures are presented as mean ± standard deviation. The sample sizes are described in the figure legends.
Data availability
All data generated or analyzed during this study are included in this published article [and its supplementary information files].
References
Wong, R. S. Apoptosis in cancer: from pathogenesis to treatment. J. Exp. Clin. Cancer Res. 30, 87 (2011).
Lavrik, I., Golks, A. & Krammer, P. H. Death receptor signaling. J. Cell Sci. 118, 265–267 (2005).
OZOREN, N. & EL-DEIRY, W. S. Cell surface death receptor signaling in normal and cancer cells. Semin. Cancer Biol. 13, 135–147 (2003).
Tomizawa, Y. et al. Inhibition of lung cancer cell growth and induction of apoptosis after reexpression of 3p21.3 candidate tumor suppressor gene SEMA3B. Proc. Natl Acad. Sci. USA 98, 13954–13959 (2001).
Kigel, B., Varshavsky, A., Kessler, O. & Neufeld, G. Successful inhibition of tumor development by specific class-3 semaphorins is associated with expression of appropriate semaphorin receptors by tumor cells. PLoS ONE 3, e3287 (2008).
Tran, T. S., Kolodkin, A. L. & Bharadwaj, R. Semaphorin regulation of cellular morphology. Annu. Rev. Cell Dev. Biol. 23, 263–292 (2007).
Yaron, A., Huang, P. H., Cheng, H. J. & Tessier-Lavigne, M. Differential requirement for Plexin-A3 and -A4 in mediating responses of sensory and sympathetic neurons to distinct class 3 Semaphorins. Neuron 45, 513–523 (2005).
Winberg, M. L. et al. Plexin A is a neuronal semaphorin receptor that controls axon guidance. Cell 95, 903–916 (1998).
Li, X. & Lee, A. Y. Semaphorin 5A and plexin-B3 inhibit human glioma cell motility through RhoGDIalpha-mediated inactivation of Rac1 GTPase. J. Biol. Chem. 285, 32436–32445 (2010).
Basile, J. R., Castilho, R. M., Williams, V. P. & Gutkind, J. S. Semaphorin 4D provides a link between axon guidance processes and tumor-induced angiogenesis. Proc. Natl Acad. Sci. USA 103, 9017–9022 (2006).
Roche, J. et al. Distinct 3p21.3 deletions in lung cancer and identification of a new human semaphorin. Oncogene 12, 1289–1297 (1996).
Xiang, R. H. et al. Isolation of the human semaphorin III/F gene (SEMA3F) at chromosome 3p21, a region deleted in lung cancer. Genomics 32, 39–48 (1996).
Roodink, I. et al. Semaphorin 3E expression correlates inversely with Plexin D1 during tumor progression. Am. J. Pathol. 173, 1873–1881 (2008).
Xu, X. M. et al. The transmembrane protein semaphorin 6A repels embryonic sympathetic axons. J. Neurosci. 20, 2638–2648 (2000).
Runker, A. E. et al. Mutation of Semaphorin-6A disrupts limbic and cortical connectivity and models neurodevelopmental psychopathology. PLoS ONE 6, e26488 (2011).
Runker, A. E., Little, G. E., Suto, F., Fujisawa, H. & Mitchell, K. J. Semaphorin-6A controls guidance of corticospinal tract axons at multiple choice points. Neural Dev. 3, 34 (2008).
Suto, F. et al. Plexin-a4 mediates axon-repulsive activities of both secreted and transmembrane semaphorins and plays roles in nerve fiber guidance. J. Neurosci. 25, 3628–3637 (2005).
Suto, F. et al. Interactions between plexin-A2, plexin-A4, and semaphorin 6A control lamina-restricted projection of hippocampal mossy fibers. Neuron 53, 535–547 (2007).
Lu, T. P. et al. Identification of a novel biomarker, SEMA5A, for non-small cell lung carcinoma in nonsmoking women. Cancer Epidemiol. Biomark. Prev. 19, 2590–2597 (2010).
Dhanabal, M. et al. Recombinant semaphorin 6A-1 ectodomain inhibits in vivo growth factor and tumor cell line-induced angiogenesis. Cancer Biol. Ther. 4, 659–668 (2005).
Ueno, K., Kumagai, T., Kijima, T., Kishimoto, T. & Hosoe, S. Cloning and tissue expression of cDNAs from chromosome 5q21-22 which is frequently deleted in advanced lung cancer. Hum. Genet. 102, 63–68 (1998).
Sanchez-Palencia, A. et al. Gene expression profiling reveals novel biomarkers in nonsmall cell lung cancer. Int. J. Cancer 129, 355–364 (2011).
Zhang, Y. et al. USP44 regulates centrosome positioning to prevent aneuploidy and suppress tumorigenesis. J. Clin. Invest. 122, 4362–4374 (2012).
Hou, J. et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS ONE 5, e10312 (2010).
Nogi, T. et al. Structural basis for semaphorin signalling through the plexin receptor. Nature 467, 1123–1127 (2010).
McIlwain, D. R., Berger, T. & Mak, T. W. Caspase functions in cell death and disease. Cold Spring Harb. Perspect. Biol. 5, a008656 (2013).
Thorburn, A. Death receptor-induced cell killing. Cell Signal. 16, 139–144 (2004).
Chinnaiyan, A. M., O’Rourke, K., Tewari, M. & Dixit, V. M. FADD, a novel death domain-containing protein, interacts with the death domain of Fas and initiates apoptosis. Cell 81, 505–512 (1995).
Hsu, H., Shu, H. B., Pan, M. G. & Goeddel, D. V. TRADD-TRAF2 and TRADD-FADD interactions define two distinct TNF receptor 1 signal transduction pathways. Cell 84, 299–308 (1996).
Henningsen, J., Rigbolt, K. T. G., Blagoev, B., Pedersen, B. K. & Kratchmarova, I. Dynamics of the Skeletal Muscle Secretome during Myoblast Differentiation. Mol. Cell Proteom. 9, 2482–2496 (2010).
Gras, C. et al. Secreted semaphorin 5A activates immune effector cells and is a biomarker for rheumatoid arthritis. Arthritis Rheumatol. 66, 1461–1471 (2014).
Henningsen, J. et al. Quantitative proteomics for investigation of secreted factors: focus on muscle secretome. Clin. Proteom. 9, 2 (2012).
Lee, E. W., Seo, J., Jeong, M., Lee, S. & Song, J. The roles of FADD in extrinsic apoptosis and necroptosis. BMB Rep. 45, 496–508 (2012).
Kischkel, F. C. et al. Cytotoxicity-dependent APO-1 (Fas/CD95)-associated proteins form a death-inducing signaling complex (DISC) with the receptor. EMBO J. 14, 5579–5588 (1995).
Cotter, T. G. Apoptosis and cancer: the genesis of a research field. Nat. Rev. Cancer 9, 501–507 (2009).
Wu, F.-C. et al. Differential genomic alterations in normal and cancerous lung cells in response to chronic inflammation. Transl. Cancer Res. 2, 64–73 (2013).
Debatin, K. M. & Krammer, P. H. Death receptors in chemotherapy and cancer. Oncogene 23, 2950–2966 (2004).
Unoki, M. et al. UHRF1 is a novel diagnostic marker of lung cancer. Br. J. Cancer 103, 217–222 (2010).
Acknowledgements
We thank the Joint Centre for Instruments and Researches, National Taiwan University, for assistance with flow cytometry; and Melissa Stauffer, Ph.D., for editing the manuscript. This work was supported in part by Ministry of Science and Technology, Taiwan (103-2314-B-002-034-MY3); National Health Research Institutes (NHRI-EX105-10419BI); and Bioinformatics and Biostatistics Core, NTU Center of Genomic Medicine (Excellent Research Projects, 104R8400).
Author information
Authors and Affiliations
Contributions
C.-Y.S. designed and performed experiments, and wrote the paper. Y.-C.C., Y.-H.L., L.S.-T.Y. and H.-K.C. performed experiments. W.F. designed experiments. L.-H.C. and L.-C.L. wrote the paper. T.-P.L. performed bioinformatics analysis. C.-P.H. and J.-M.L. provided patient lung cancer samples. P.-W.H. provided cells and designed experiments. M.-H.T. and E.Y.C. designed experiments, supervised the project, and wrote the paper. All authors discussed the results and contributed to the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shen, CY., Chang, YC., Chen, LH. et al. The extracellular SEMA domain attenuates intracellular apoptotic signaling of semaphorin 6A in lung cancer cells. Oncogenesis 7, 95 (2018). https://doi.org/10.1038/s41389-018-0105-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-018-0105-z
This article is cited by
-
IER3IP1-mutations cause microcephaly by selective inhibition of ER-Golgi transport
Cellular and Molecular Life Sciences (2024)
-
A comprehensive review of tumor proliferative and suppressive role of semaphorins and therapeutic approaches
Biophysical Reviews (2020)
-
Semaphorin 6A Attenuates the Migration Capability of Lung Cancer Cells via the NRF2/HMOX1 Axis
Scientific Reports (2019)