Abstract
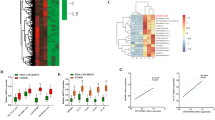
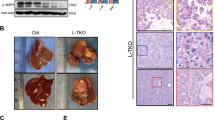
Non-alcoholic steatohepatitis (NASH) is rapidly surpassing viral hepatitis as the primary cause of hepatocellular carcinoma (HCC). However, understanding of NASH-progressed HCC remains poor, which might impede HCC diagnosis and therapy. In this study, we aim to identify shared transcriptional changes between NASH and HCC, of which we focused on E3 ligase TRIM45. We found TRIM45 exacerbates HCC cells proliferation and metastasis in vitro and in vivo. Further transcriptome analysis revealed TRIM45 predominantly affects fatty acid metabolism and oleic acid restored impaired proliferation and metastasis of TRIM45-deficient HCC cells. IP-tandem mass spectrum and FABP5 depriving experiment indicated that TRIM45 enhance fatty acid synthesis depending on FABP5 presence. Interestingly, we found TRIM45 directly added K33-type and K63-type poly-ubiquitin chains to FABP5 NLS domain, which ultimately promoted FABP5 nuclear translocation. Nuclear FABP5 interacted with PPARγ to facilitate downstream lipid synthesis gene expression. We observed TRIM45 accelerated NASH-to-HCC transition and exacerbated both NASH and NASH-HCC with the enhanced fatty acid production in vivo. Moreover, high concentration of fatty acid increased TRIM45 expression. The established mechanism was substantiated by gene expression correlation in TCGA-LIHC. Collectively, our research revealed a common lipid reprograming process in NASH and HCC and identified the cyclical amplification of the TRIM45-FABP5-PPARγ-fatty acid axis. This signaling pathway offers potential therapeutic targets for therapeutic intervention in NASH and NASH-progressed HCC.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Full data will be available from the corresponding author upon reasonable request. The RNA-sequencing data for this study are deposited in GEO database (GSE261075).
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209–49.
Huang DQ, El-Serag HB, Loomba R. Global epidemiology of NAFLD-related HCC: trends, predictions, risk factors and prevention. Nat Rev Gastroenterol Hepatol. 2021;18:223–38.
Rumgay H, Arnold M, Ferlay J, Lesi O, Cabasag CJ, Vignat J, et al. Global burden of primary liver cancer in 2020 and predictions to 2040. J Hepatol. 2022;77:1598–606.
Asrani SK, Devarbhavi H, Eaton J, Kamath PS. Burden of liver diseases in the world. J HEPATOL. 2019;70:151–71.
Yang JD, Hainaut P, Gores GJ, Amadou A, Plymoth A, Roberts LR. A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat Rev Gastroenterol Hepatol. 2019;16:589–604.
Younossi ZM. Non-alcoholic fatty liver disease–a global public health perspective. J HEPATOL. 2019;70:531–44.
Organization, GWH. Global health sector strategies on HIV, viral hepatitis and sexually transmitted infections for the period 2022–2030. 2022.
Hindson J. Molecular landscape of NASH-HCC. Nature Reviews Gastroenterology & Hepatology. 2021;18:456–456.
Nakagawa H, Hayata Y, Kawamura S, Yamada T, Fujiwara N, Koike K. Lipid metabolic reprogramming in hepatocellular carcinoma. Cancers. 2018;10:447.
Xu K, Xia P, Chen X, Ma W, Yuan Y. ncRNA-mediated fatty acid metabolism reprogramming in HCC. Trends Endocrinol Metab. 2023;34:278–91.
Xiong L, Wu F, Wu Q, Xu L, Cheung OK, Kang W, et al. Aberrant enhancer hypomethylation contributes to hepatic carcinogenesis through global transcriptional reprogramming. Nat Commun. 2019;10:335.
Horisawa K, Udono M, Ueno K, Ohkawa Y, Nagasaki M, Sekiya S, et al. The dynamics of transcriptional activation by hepatic reprogramming factors. MOL CELL. 2020;79:660–676.e668.
Jühling F, Hamdane N, Crouchet E, Li S, El Saghire H, Mukherji A, et al. Targeting clinical epigenetic reprogramming for chemoprevention of metabolic and viral hepatocellular carcinoma. Gut. 2021;70:157–69.
Schuster S, Cabrera D, Arrese M, Feldstein AE. Triggering and resolution of inflammation in NASH. Nat Rev Gastroenterol Hepatol. 2018;15:349–64.
Hall Z, Chiarugi D, Charidemou E, Leslie J, Scott E, Pellegrinet L, et al. Lipid remodeling in hepatocyte proliferation and hepatocellular carcinoma. Hepatol (Baltim, Md). 2021;73:1028–44.
Chen Z, Tian R, She Z, Cai J, Li H. Role of oxidative stress in the pathogenesis of nonalcoholic fatty liver disease. Free Radic Biol Med. 2020;152:116–41.
Arroyave-Ospina JC, Wu Z, Geng Y, Moshage H. Role of oxidative stress in the pathogenesis of non-alcoholic fatty liver disease: Implications for prevention and therapy. Antioxidants. 2021;10:174.
Ma Y, Lee G, Heo S-Y, Roh Y-S. Oxidative stress is a key modulator in the development of nonalcoholic fatty liver disease. Antioxidants. 2021;11:91.
Geng Y, Faber KN, de Meijer VE, Blokzijl H, Moshage H. How does hepatic lipid accumulation lead to lipotoxicity in non-alcoholic fatty liver disease? Hepatol Int. 2021;15:21–35.
Rada P, González-Rodríguez Á, García-Monzón C, Valverde ÁM. Understanding lipotoxicity in NAFLD pathogenesis: is CD36 a key driver? Cell death Dis. 2020;11:802.
Sangineto M, Villani R, Cavallone F, Romano A, Loizzi D, Serviddio G. Lipid metabolism in development and progression of hepatocellular carcinoma. Cancers. 2020;12:1419.
Kim H, Rodriguez-Navas C, Kollipara RK, Kapur P, Pedrosa I, Brugarolas J, et al. Unsaturated fatty acids stimulate tumor growth through stabilization of β-catenin. Cell Rep. 2015;13:495–503.
Ishida T, Ciulli A. E3 ligase ligands for PROTACs: how they were found and how to discover new ones. SLAS Discov: Advancing Sci Drug Discov. 2021;26:484–502.
Anstee QM, Reeves HL, Kotsiliti E, Govaere O, Heikenwalder M. From NASH to HCC: current concepts and future challenges. Nat Rev Gastroenterol Hepatol. 2019;16:411–28.
Pickart CM. Mechanisms underlying ubiquitination. Annu Rev Biochem. 2001;70:503–33.
Ge M-K, Zhang N, Xia L, Zhang C, Dong S-S, Li Z-M, et al. FBXO22 degrades nuclear PTEN to promote tumorigenesis. Nat Commun. 2020;11:1720.
Lu F, Ye M, Hu C, Chen J, Yan L, Gu D, et al. FABP5 regulates lipid metabolism to facilitate pancreatic neuroendocrine neoplasms progression via FASN mediated Wnt/β‐catenin pathway. Cancer Sci. 2023;114:3553–67.
Matsumoto M, Han S, Kitamura T, Accili D. Dual role of transcription factor FoxO1 in controlling hepatic insulin sensitivity and lipid metabolism. J Clin Investig. 2006;116:2464–72.
Li Y, Ma Z, Jiang S, Hu W, Li T, Di S, et al. A global perspective on FOXO1 in lipid metabolism and lipid-related diseases. Prog lipid Res. 2017;66:42–49.
Younossi Z, Anstee QM, Marietti M, Hardy T, Henry L, Eslam M, et al. Global burden of NAFLD and NASH: trends, predictions, risk factors and prevention. Nat Rev Gastroenterol Hepatol. 2018;15:11–20.
Shah PA, Patil R, Harrison SA. NAFLD‐related hepatocellular carcinoma: The growing challenge. Hepatol (Baltim, Md). 2023;77:323–38.
Xia SW, Wang ZM, Sun SM, Su Y, Li ZH, Shao JJ, et al. Endoplasmic reticulum stress and protein degradation in chronic liver disease. Pharmacol Res. 2020;161:105218.
Park JS, Ma H, Roh YS. Ubiquitin pathways regulate the pathogenesis of chronic liver disease. Biochemical Pharmacol. 2021;193:114764.
Herhaus L, Dikic I. Expanding the ubiquitin code through post-translational modification. EMBO Rep 2015;16:1071–83.
Wang Y, Wang F. Post-translational modifications of deubiquitinating enzymes: expanding the ubiquitin code. Front Pharm. 2021;12:685011.
Hunter T. The age of crosstalk: phosphorylation, ubiquitination, and beyond. Mol Cell. 2007;28:730–8.
Deng L, Meng T, Chen L, Wei W, Wang P. The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct Target Ther. 2020;5:11.
Sun T, Liu Z, Yang Q. The role of ubiquitination and deubiquitination in cancer metabolism. MOL Cancer. 2020;19:146.
Yao F, Zhou Z, Kim J, Hang Q, Xiao Z, Ton BN, et al. SKP2- and OTUD1-regulated non-proteolytic ubiquitination of YAP promotes YAP nuclear localization and activity. NAT COMMUN. 2018;9:2269.
Trotman LC, Wang X, Alimonti A, Chen Z, Teruya-Feldstein J, Yang H, et al. Ubiquitination regulates PTEN nuclear import and tumor suppression. Cell. 2007;128:141–56.
Seo J, Jeong DW, Park JW, Lee KW, Fukuda J, Chun YS. Fatty-acid-induced FABP5/HIF-1 reprograms lipid metabolism and enhances the proliferation of liver cancer cells. Commun Biol. 2020;3:638.
Pan L, Xiao H, Liao R, Chen Q, Peng C, Zhang Y, et al. Fatty acid binding protein 5 promotes tumor angiogenesis and activates the IL6/STAT3/VEGFA pathway in hepatocellular carcinoma. Biomedicine Pharmacother. 2018;106:68–76.
Armstrong EH, Goswami D, Griffin PR, Noy N, Ortlund EA. Structural basis for ligand regulation of the fatty acid-binding protein 5, peroxisome proliferator-activated receptor β/δ (FABP5-PPARβ/δ) signaling pathway. J Biol Chem. 2014;289:14941–54.
Suppresses H. Genetic ablation of the fatty acid. Cancer Res. 2013;73:4770.
Chen, H, Tan, H, Wan, J, Zeng, Y, Wang, J, Wang, H et al. PPAR-γ signaling in nonalcoholic fatty liver disease: Pathogenesis and therapeutic targets. Pharmacology & therapeutics, 108391 (2023).
Xiong Z, Chan SL, Zhou J, Vong JS, Kwong TT, Zeng X, et al. Targeting PPAR-gamma counteracts tumour adaptation to immune-checkpoint blockade in hepatocellular carcinoma. Gut. 2023;72:1758–73.
Peiseler M, Schwabe R, Hampe J, Kubes P, Heikenwaelder M, Tacke F. Immune mechanisms linking metabolic injury to inflammation and fibrosis in fatty liver disease–novel insights into cellular communication circuits. J HEPATOL. 2022;77:1136–60.
Pfister D, Núñez NG, Pinyol R, Govaere O, Pinter M, Szydlowska M, et al. NASH limits anti-tumour surveillance in immunotherapy-treated HCC. Nature. 2021;592:450–6.
Hirsova P, Ibrabim SH, Gores GJ, Malhi H. Lipotoxic lethal and sublethal stress signaling in hepatocytes: relevance to NASH pathogenesis [S]. J lipid Res. 2016;57:1758–70.
Lebeaupin C, Vallée D, Hazari Y, Hetz C, Chevet E, Bailly-Maitre B. Endoplasmic reticulum stress signalling and the pathogenesis of non-alcoholic fatty liver disease. J HEPATOL. 2018;69:927–47.
Flessa C-M, Kyrou I, Nasiri-Ansari N, Kaltsas G, Papavassiliou AG, Kassi E, et al. Endoplasmic reticulum stress and autophagy in the pathogenesis of non-alcoholic fatty liver disease (NAFLD): current evidence and perspectives. Curr Obes Rep. 2021;10:134–61.
Vincenzi M, Mercurio FA, Leone M. Protein Interaction domains: structural features and drug discovery applications (Part 2). Curr Med Chem. 2021;28:854–92.
Goggins S, Marsh BJ, Lubben AT, Frost CG. Signal transduction and amplification through enzyme-triggered ligand release and accelerated catalysis. Chem Sci. 2015;6:4978–85.
Scott DE, Bayly AR, Abell C, Skidmore J. Small molecules, big targets: drug discovery faces the protein–protein interaction challenge. Nat Rev Drug Discov. 2016;15:533–50.
Tward AD, Jones KD, Yant S, Cheung ST, Fan ST, Chen X, et al. Distinct pathways of genomic progression to benign and malignant tumors of the liver. Proc Natl Acad Sci USA. 2007;104:14771–6.
Stewart SA, Dykxhoorn DM, Palliser D, Mizuno H, Yu EY, An DS, et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA (N. Y). 2003;9:493–501.
Chow S-C, Shao J, Wang H & Lokhnygina Y. Sample size calculations in clinical research. (chapman and hall/CRC, 2017).
Acknowledgements
We would like to acknowledge and thank the teachers from the Scientific Research Center of Zhongnan Hospital of Wuhan University for their help in this study.
Funding
This work was supported by Key Research and Development Program of Hubei Province (Grant No. 2021BCA114) and National Natural Science Foundation of China (Grant No. 82300675).
Author information
Authors and Affiliations
Contributions
XL, WH, XC, TW and YY devised and coordinated the project. WH, XC, TW and YY supervised the project. XL, HW and FL collected clinical HCC samples, XL, FL, JZ, DZ and YZ performed most of the experiments, XL, WH analyzed data. XL, PX, and WM provided significant intellectual input. XL, WH and WM wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
The study was conducted in accordance with the principles of the Declaration of Helsinki principles. All animal experiments were approved by the Animal Use and Care Committees at Zhongnan Hospital, Wuhan University (License No. WP20220494). The human samples and data usage was reviewed and approved by the Scientific Research Ethics Committee of Zhongnan Hospital, Wuhan University (License No. 2022076 K).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., He, W., Chen, X. et al. TRIM45 facilitates NASH-progressed HCC by promoting fatty acid synthesis via catalyzing FABP5 ubiquitylation. Oncogene (2024). https://doi.org/10.1038/s41388-024-03056-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41388-024-03056-7