Abstract
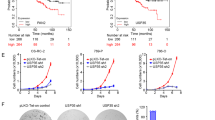
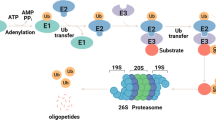
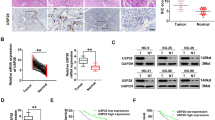
Deubiquitinating enzymes (DUBs) are promising targets for cancer therapy because of their pivotal roles in various physiological and pathological processes. Among these, ubiquitin-specific peptidase 26 (USP26) is a protease with crucial regulatory functions. Our study sheds light on the upregulation of USP26 in colorectal cancer (CRC), in which its increased expression correlates with an unfavorable prognosis. Herein, we evidenced the role of USP26 in promoting CRC tumorigenesis in a parkin RBR E3 ubiquitin-protein ligase (PRKN) protein-dependent manner. Our investigation revealed that USP26 directly interacted with PRKN protein, facilitating its deubiquitination, and subsequently reducing its activity. Additionally, we identified the K129 site on PRKN as a specific target for USP26-mediated deubiquitination. Our research highlights that a K-to-R mutation at the site on PRKN diminishes its potential for activation and ability to mediate mitophagy. In summary, our findings underscore the significance of USP26-mediated deubiquitination in restraining the activation of the PRKN-mediated mitophagy pathway, ultimately driving CRC tumorigenesis. This study not only elucidated the multifaceted role of USP26 in CRC but also introduced a promising avenue for therapeutic exploration through the development of small molecule inhibitors targeting USP26. This strategy holds promise as a novel therapeutic approach for CRC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All the data and materials used in this study are available from the corresponding author upon request.
References
Shaid S, Brandts CH, Serve H, Dikic I. Ubiquitination and selective autophagy. Cell Death Differ. 2013;20:21–30.
Ikeda F, Dikic I. Atypical ubiquitin chains: new molecular signals. ‘protein modifications: beyond the usual suspects’ review series. EMBO Rep. 2008;9:536–42.
Herrmann J, Lerman LO, Lerman A. Ubiquitin and ubiquitin-like proteins in protein regulation. Circ Res. 2007;100:1276–91.
Sun SC. Deubiquitylation and regulation of the immune response. Nat Rev Immunol. 2008;8:501–11.
Hu H, Sun SC. Ubiquitin signaling in immune responses. Cell Res. 2016;26:457–83.
Tang J, Luo Y, Xiao L. USP26 promotes anaplastic thyroid cancer progression by stabilizing TAZ. Cell Death Dis. 2022;13:326.
Li L, Zhou H, Zhu R, Liu Z. USP26 promotes esophageal squamous cell carcinoma metastasis through stabilizing Snail. Cancer Lett. 2019;448:52–60.
Kim I, Rodriguez-Enriquez S, Lemasters JJ. Selective degradation of mitochondria by mitophagy. Arch Biochem Biophys. 2007;462:245–53.
Pickles S, Vigié P, Youle RJ. Mitophagy and quality control mechanisms in mitochondrial maintenance. Curr Biol. 2018;28:R170–R185.
Porporato PE, Filigheddu N, Pedro JMB, Kroemer G, Galluzzi L. Mitochondrial metabolism and cancer. Cell Res. 2018;28:265–80.
Modica-Napolitano JS, Singh KK. Mitochondrial dysfunction in cancer. Mitochondrion. 2004;4:755–62.
Hsu CC, Tseng LM, Lee HC. Role of mitochondrial dysfunction in cancer progression. Exp Biol Med. 2016;241:1281–95.
Li Y, Liang R, Zhang X, Wang J, Shan C, Liu S, et al. Copper chaperone for superoxide dismutase promotes breast cancer cell proliferation and migration via ROS-mediated MAPK/ERK signaling. Front Pharmacol. 2019;10:356.
Youle RJ, Narendra DP. Mechanisms of mitophagy. Nat Rev Mol Cell Biol. 2011;12:9–14.
Poole LP, Macleod KF. Mitophagy in tumorigenesis and metastasis. Cell Mol Life Sci. 2021;78:3817–51.
Zhang T, Liu Q, Gao W, Sehgal SA, Wu H. The multifaceted regulation of mitophagy by endogenous metabolites. Autophagy. 2022;18:1216–39.
Saito S, Sirahama S, Matsushima M, Suzuki M, Sagae S, Kudo R, et al. Definition of a commonly deleted region in ovarian cancers to a 300-kb segment of chromosome 6q27. Cancer Res. 1996;56:5586–9.
Cesari R, Martin ES, Calin GA, Pentimalli F, Bichi R, McAdams H, et al. Parkin, a gene implicated in autosomal recessive juvenile parkinsonism, is a candidate tumor suppressor gene on chromosome 6q25-q27. Proc Natl Acad Sci USA. 2003;100:5956–61.
Denison SR, Wang F, Becker NA, Schüle B, Kock N, Phillips LA, et al. Alterations in the common fragile site gene Parkin in ovarian and other cancers. Oncogene. 2003;22:8370–8.
Picchio MC, Martin ES, Cesari R, Calin GA, Yendamuri S, Kuroki T, et al. Alterations of the tumor suppressor gene Parkin in non-small cell lung cancer. Clin Cancer Res. 2004;10:2720–4.
Liu B, Ruan J, Chen M, Li Z, Manjengwa G, Schlüter D, et al. Deubiquitinating enzymes (DUBs): decipher underlying basis of neurodegenerative diseases. Mol Psychiatry. 2022;27:259–68.
Yao RQ, Ren C, Xia ZF, Yao YM. Organelle-specific autophagy in inflammatory diseases: a potential therapeutic target underlying the quality control of multiple organelles. Autophagy. 2021;17:385–401.
Jeong YY, Jia N, Ganesan D, Cai Q. Broad activation of the PRKN pathway triggers synaptic failure by disrupting synaptic mitochondrial supply in early tauopathy. Autophagy. 2022;18:1472–4.
Li W, Li F, Zhang X, Lin HK, Xu C. Insights into the post-translational modification and its emerging role in shaping the tumor microenvironment. Signal Transduct Target Ther. 2021;6:422.
Wang H, Yang L, Liu M, Luo J. Protein post-translational modifications in the regulation of cancer hallmarks. Cancer Gene Ther. 2023;30:529–47.
Horn-Ghetko D, Krist DT, Prabu JR, Baek K, Mulder MPC, Klügel M, et al. Ubiquitin ligation to F-box protein targets by SCF-RBR E3-E3 super-assembly. Nature. 2021;590:671–6.
Wan Y, Yan C, Gao H, Liu T. Small-molecule PROTACs: novel agents for cancer therapy. Future Med Chem. 2020;12:915–38.
Gao H, Sun X, Rao Y. PROTAC technology: opportunities and challenges. ACS Med Chem Lett. 2020;11:237–40.
Harrigan JA, Jacq X, Martin NM, Jackson SP. Deubiquitylating enzymes and drug discovery: emerging opportunities. Nat Rev Drug Discov. 2018;17:57–78.
Lu Y, Li Z, Zhang S, Zhang T, Liu Y, Zhang L. Cellular mitophagy: mechanism, roles in diseases and small molecule pharmacological regulation. Theranostics. 2023;13:736–66.
Yamano K, Kikuchi R, Kojima W, Hayashida R, Koyano F, Kawawaki J. Critical role of mitochondrial ubiquitination and the OPTN-ATG9A axis in mitophagy. J Cell Biol. 2020;219:e201912144.
Jee SC, Cheong H. Autophagy/mitophagy regulated by ubiquitination: a promising pathway in cancer therapeutics. Cancers. 2023;15:1112.
Tsefou E, Ketteler R. Targeting deubiquitinating enzymes (DUBs) that regulate mitophagy via direct or indirect interaction with Parkin. Int J Mol Sci. 2022;23:12105.
Park GH, Park JH, Chung KC. Precise control of mitophagy through ubiquitin proteasome system and deubiquitin proteases and their dysfunction in Parkinson’s disease. BMB Rep. 2021;54:592–600.
Wang Y, Serricchio M, Jauregui M, Shanbhag R, Stoltz T, Di Paolo CT, et al. Deubiquitinating enzymes regulate PARK2-mediated mitophagy. Autophagy. 2015;11:595–606.
Sun Y, Lu F, Yu X, Wang B, Chen J, Lu F, et al. Exogenous H(2)S promoted USP8 sulfhydration to regulate mitophagy in the hearts of db/db mice. Aging Dis. 2020;11:269–85.
Bingol B, Tea JS, Phu L, Reichelt M, Bakalarski CE, Song Q, et al. The mitochondrial deubiquitinase USP30 opposes parkin-mediated mitophagy. Nature. 2014;510:370–5.
Niu K, Fang H, Chen Z, Zhu Y, Tan Q, Wei D, et al. USP33 deubiquitinates PRKN/parkin and antagonizes its role in mitophagy. Autophagy. 2020;16:724–34.
Lim KL, Chew KC, Tan JM, Wang C, Chung KK, Zhang Y, et al. Parkin mediates nonclassical, proteasomal-independent ubiquitination of synphilin-1: implications for Lewy body formation. J Neurosci. 2005;25:2002–9.
Matsuda N, Kitami T, Suzuki T, Mizuno Y, Hattori N, Tanaka K. Diverse effects of pathogenic mutations of Parkin that catalyze multiple monoubiquitylation in vitro. J Biol Chem. 2006;281:3204–9.
Zhong L, Tan Y, Zhou A, Yu Q, Zhou J. RING finger ubiquitin-protein isopeptide ligase Nrdp1/FLRF regulates parkin stability and activity. J Biol Chem. 2005;280:9425–30.
Chastagner P, Israël A, Brou C. Itch/AIP4 mediates Deltex degradation through the formation of K29-linked polyubiquitin chains. Embo Rep. 2006;7:1147–53.
Geisler S, Holmström KM, Skujat D, Fiesel FC, Rothfuss OC, Kahle PJ. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat Cell Biol. 2010;12:119–U70.
Ikeda H, Kerppola TK. Lysosomal localization of ubiquitinated Jun requires multiple determinants in a lysine-27-linked polyubiquitin conjugate. Mol Biol Cell. 2008;19:4588–601.
Tan JM, Wong ES, Kirkpatrick DS, Pletnikova O, Ko HS, Tay SP, et al. Lysine 63-linked ubiquitination promotes the formation and autophagic clearance of protein inclusions associated with neurodegenerative diseases. Hum Mol Genet. 2008;17:431–9.
Ben-Saadon R, Zaaroor D, Ziv T, Ciechanover A. The polycomb protein Ring1B generates self-atypical mixed ubiquitin chains required for its in vitro histone H2A ligase activity. Mol Cell. 2006;24:701–11.
Morris JR, Solomon E. BRCA1: BARD1 induces the formation of conjugated ubiquitin structures, dependent on K6 of ubiquitin, in cells during DNA replication and repair. Hum Mol Genet. 2004;13:807–17.
Acknowledgements
This work is supported by the National Key Research and Development Program of China (No. 2022YFA1105303 GW) and NSFC (GW, Nos. 81974432, 81922053, and 82330084; JH, No. 82273254).
Author information
Authors and Affiliations
Contributions
QW, GW, and JH conceived and designed the experiments; QW, ZW, and SC conducted biochemical and cellular experiments. XS, SZ, and PL performed the animal experiments. LL, KL, AL, and CH generated the gene knock-out cell lines. YC and FH provided help for bioinformatics analysis. JH and GW gave suggestions for many experiments. QW, GW, and JH organized and analyzed the data and wrote the manuscript, which was edited by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Tongji Hospital (TJ-IRB20220723). The clinical specimens randomly used in this study were obtained from the Department of Gastrointestinal Surgery, Tongji Hospital. Demographic information, including age and sex, is presented in Table S2. Animal experiments were performed strictly following the Animal Study Guideline of Huazhong University of Science and Technology (TJH-202210034).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Q., Wang, Z., Chen, S. et al. USP26 promotes colorectal cancer tumorigenesis by restraining PRKN-mediated mitophagy. Oncogene (2024). https://doi.org/10.1038/s41388-024-03009-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41388-024-03009-0