Abstract
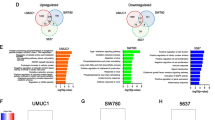
The urothelium is a stratified epithelium composed of basal cells, one or more layers of intermediate cells, and an upper layer of differentiated umbrella cells. Most bladder cancers (BLCA) are urothelial carcinomas. Loss of urothelial lineage fidelity results in altered differentiation, highlighted by the taxonomic classification into basal and luminal tumors. There is a need to better understand the urothelial transcriptional networks. To systematically identify transcription factors (TFs) relevant for urothelial identity, we defined highly expressed TFs in normal human bladder using RNA-Seq data and inferred their genomic binding using ATAC-Seq data. To focus on epithelial TFs, we analyzed RNA-Seq data from patient-derived organoids recapitulating features of basal/luminal tumors. We classified TFs as “luminal-enriched”, “basal-enriched” or “common” according to expression in organoids. We validated our classification by differential gene expression analysis in Luminal Papillary vs. Basal/Squamous tumors. Genomic analyses revealed well-known TFs associated with luminal (e.g., PPARG, GATA3, FOXA1) and basal (e.g., TP63, TFAP2) phenotypes and novel candidates to play a role in urothelial differentiation or BLCA (e.g., MECOM, TBX3). We also identified TF families (e.g., KLFs, AP1, circadian clock, sex hormone receptors) for which there is suggestive evidence of their involvement in urothelial differentiation and/or BLCA. Genomic alterations in these TFs are associated with BLCA. We uncover a TF network involved in urothelial cell identity and BLCA. We identify novel candidate TFs involved in differentiation and cancer that provide opportunities for a better understanding of the underlying biology and therapeutic intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hicks RM. The mammalian urinary bladder: an accommodating organ. Biol Rev Camb Philos Soc. 1975;50:215–46.
Truschel ST, Clayton DR, Beckel JM, Yabes JG, Yao Y, Wolf-Johnston A, et al. Age-related endolysosome dysfunction in the rat urothelium. PLoS One. 2018;13:e0198817.
Jost SP, Potten CS. Urothelial proliferation in growing mice. Cell Tissue Kinet. 1986;19:155–60.
Wang J, Batourina E, Schneider K, Souza S, Swayne T, Liu C, et al. Polyploid superficial cells that maintain the urothelial barrier are produced via incomplete cytokinesis and endoreplication. Cell Rep. 2019;25:464–.e4.
Hudoklin S, Jezernik K, Neumüller J, Pavelka M, Romih R. Electron tomography of fusiform vesicles and their organization in urothelial cells. PLoS One. 2012;7:e32935.
Varley CL, Garthwaite MAE, Cross W, Hinley J, Trejdosiewicz LK, Southgate J. PPARgamma-regulated tight junction development during human urothelial cytodifferentiation. J Cell Physiol. 2006;208:407–17.
Harnden P, Eardley I, Joyce AD, Southgate J. Cytokeratin 20 as an objective marker of urothelial dysplasia. Br J Urol. 1996;78:870–5.
Gandhi D, Molotkov A, Batourina E, Schneider K, Dan H, Reiley M, et al. Retinoid signaling in progenitors controls specification and regeneration of the urothelium. Dev Cell. 2013;26:469–82.
Papafotiou G, Paraskevopoulou V, Vasilaki E, Kanaki Z, Paschalidis N, Klinakis A. KRT14 marks a subpopulation of bladder basal cells with pivotal role in regeneration and tumorigenesis. Nat Commun. 2016;7:11914.
Colopy SA, Bjorling DE, Mulligan WA, Bushman W. A population of progenitor cells in the basal and intermediate layers of the murine bladder urothelium contributes to urothelial development and regeneration. Dev Dyn. 2014;243:988–98.
Jost SP. Cell cycle of normal bladder urothelium in developing and adult mice. Virchows Arch B Cell Pathol. 1989;57:27–36.
Shin K, Lee J, Guo N, Kim J, Lim A, Qu L, et al. Hedgehog/Wnt feedback supports regenerative proliferation of epithelial stem cells in bladder. Nature. 2011;472:110–4.
Hurst CD, Alder O, Platt FM, Droop A, Stead LF, Burns JE, et al. Genomic subtypes of non-invasive bladder cancer with distinct metabolic profile and female gender bias in KDM6A mutation frequency. Cancer Cell. 2017;32:701–.e7.
Robertson AG, Kim J, Al-Ahmadie H, Bellmunt J, Guo G, Cherniack AD, et al. Comprehensive molecular characterization of muscle-invasive bladder. Cancer Cell. 2017;171:540–.e25.
Wullweber A, Strick R, Lange F, Sikic D, Taubert H, Wach S, et al. Bladder tumor subtype commitment occurs in carcinoma in situ driven by key signaling pathways including ECM remodeling. Cancer Res. 2021;81:1552–66.
Bondaruk J, Jaksik R, Wang Z, Cogdell D, Lee S, Chen Y, et al. The origin of bladder cancer from mucosal field effects. iScience. 2022;25:104551.
Majewski T, Yao H, Bondaruk J, Chung W, Lee S, Lee JG, et al. Whole-organ genomic characterization of mucosal field effects initiating bladder carcinogenesis. Cell Rep. 2019;26:2241–.e4.
Lawson ARJ, Abascal F, Coorens THH, Hooks Y, O’Neill L, Latimer C, et al. Extensive heterogeneity in somatic mutation and selection in the human bladder. Science. 2020;370:75–82.
López-Knowles E, Hernández S, Malats N, Kogevinas M, Lloreta J, Carrato A, et al. PIK3CA mutations are an early genetic alteration associated with FGFR3 mutations in superficial papillary bladder tumors. Cancer Res. 2006;66:7401–4.
Taylor CF, Platt FM, Hurst CD, Thygesen HH, Knowles MA. Frequent inactivating mutations of STAG2 in bladder cancer are associated with low tumour grade and stage and inversely related to chromosomal copy number changes. Hum Mol Genet. 2014;23:1964–74.
Hedegaard J, Lamy P, Nordentoft I, Algaba F, Høyer S, Ulhøi BP, et al. Comprehensive transcriptional analysis of early-stage urothelial carcinoma. Cancer Cell. 2016;30:27–42.
Hartmann A, Schlake G, Zaak D, Hungerhuber E, Hofstetter A, Hofstaedter F, et al. Occurrence of chromosome 9 and p53 alterations in multifocal dysplasia and carcinoma in situ of human urinary bladder. Cancer Res. 2002;62:809–18.
Kamoun A, de Reyniès A, Allory Y, Sjödahl G, Robertson AG, Seiler R, et al. A consensus molecular classification of muscle-invasive bladder cancer. Eur Urol. 2020;77:420–33.
Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. The human transcription factors. Cell. 2018;172:650–65.
Lee SH, Hu W, Matulay JT, Silva MV, Owczarek TB, Kim K, et al. Tumor evolution and drug response in patient-derived organoid models of bladder. Cancer Cell. 2018;173:515–.e17.
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74.
Chen Z, Zhou L, Liu L, Hou Y, Xiong M, Yang Y, et al. Single-cell RNA sequencing highlights the role of inflammatory cancer-associated fibroblasts in bladder urothelial carcinoma. Nat Commun. 2020;11:5077.
Evans RM, Mangelsdorf DJ. Nuclear receptors, RXR, and the big bang. Cell. 2014;157:255–66.
Lehmann JM, Moore LB, Smith-Oliver TA, Wilkison WO, Willson TM, Kliewer SA. An antidiabetic thiazolidinedione is a high-affinity ligand for peroxisome proliferator-activated receptor gamma (PPAR gamma). J Biol Chem. 1995;270:12953–6.
Mamtani R, Haynes K, Bilker WB, Vaughn DJ, Strom BL, Glanz K, et al. Association between longer therapy with thiazolidinediones and risk of bladder cancer: a cohort study. J Natl Cancer Inst. 2012;104:1411–21.
Varley CL, Stahlschmidt J, Lee W-C, Holder J, Diggle C, Selby PJ, et al. Role of PPARgamma and EGFR signalling in the urothelial terminal differentiation programme. J Cell Sci. 2004;117:2029–36.
Santos CP, Lapi E, Martínez de Villarreal J, Álvaro-Espinosa L, Fernández-Barral A, Barbáchano A, et al. Urothelial organoids originating from Cd49fhigh mouse stem cells display Notch-dependent differentiation capacity. Nat Commun. 2019;10:4407.
Suzuki K, Koyanagi-Aoi M, Uehara K, Hinata N, Fujisawa M, Aoi T. Directed differentiation of human induced pluripotent stem cells into mature stratified bladder urothelium. Sci Rep. 2019;9:10506.
Weiss RM, Guo S, Shan A, Shi H, Romano R-A, Sinha S, et al. Brg1 determines urothelial cell fate during ureter development. J Am Soc Nephrol. 2013;24:618–26.
Liu C, Tate T, Batourina E, Truschel ST, Potter S, Adam M, et al. Pparg promotes differentiation and regulates mitochondrial gene expression in bladder epithelial cells. Nat Commun. 2019;10:4589.
Liang F-X, Bosland MC, Huang H, Romih R, Baptiste S, Deng F-M, et al. Cellular basis of urothelial squamous metaplasia: roles of lineage heterogeneity and cell replacement. J Cell Biol. 2005;171:835–44.
Pascual G, Fong AL, Ogawa S, Gamliel A, Li AC, Perissi V, et al. A SUMOylation-dependent pathway mediates transrepression of inflammatory response genes by PPAR-gamma. Nature. 2005;437:759–63.
Tate T, Xiang T, Wobker SE, Zhou M, Chen X, Kim H, et al. Pparg signaling controls bladder cancer subtype and immune exclusion. Nat Commun. 2021;12:6160.
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature. 2014;507:315–22.
Rochel N, Krucker C, Coutos-Thévenot L, Osz J, Zhang R, Guyon E, et al. Recurrent activating mutations of PPARγ associated with luminal bladder tumors. Nat Commun. 2019;10:253.
Halstead AM, Kapadia CD, Bolzenius J, Chu CE, Schriefer A, Wartman LD, et al. Bladder-cancer-associated mutations in RXRA activate peroxisome proliferator-activated receptors to drive urothelial proliferation. eLife. 2017;6:e30862.
Korpal M, Puyang X, Jeremy Wu Z, Seiler R, Furman C, Oo HZ, et al. Evasion of immunosurveillance by genomic alterations of PPARγ/RXRα in bladder cancer. Nat Commun. 2017;8:103.
Coutos-Thévenot L, Beji S, Neyret-Kahn H, Pippo Q, Fontugne J, Osz J, et al. PPARγ is a tumor suppressor in basal bladder tumors offering new potential therapeutic opportunities. BioRxiv. 2019; https://doi.org/10.1101/868190.
Warrick JI, Walter V, Yamashita H, Chung E, Shuman L, Amponsa VO, et al. FOXA1, GATA3 and ppar? cooperate to drive luminal subtype in bladder cancer: a molecular analysis of established human cell lines. Sci Rep. 2016;6:38531.
Tortora D, Roberts ME, Kumar G, Kotapalli SS, Ritch E, Scurll JM, et al. A genome-wide CRISPR screen maps endogenous regulators of PPARG gene expression in bladder cancer. iScience. 2023;26:106525.
Goldstein JT, Berger AC, Shih J, Duke FF, Furst L, Kwiatkowski DJ, et al. Genomic activation of PPARG reveals a candidate therapeutic axis in bladder cancer. Cancer Res. 2017;77:6987–98.
Sanchez DJ, Missiaen R, Skuli N, Steger DJ, Simon MC. Cell-intrinsic tumorigenic functions of PPARγ in bladder urothelial carcinoma. Mol Cancer Res. 2021;19:598–611.
Ko LJ, Engel JD. DNA-binding specificities of the GATA transcription factor family. Mol Cell Biol. 1993;13:4011–22.
van der Ven AT, Connaughton DM, Ityel H, Mann N, Nakayama M, Chen J, et al. Whole-exome sequencing identifies causative mutations in families with congenital anomalies of the kidney and urinary tract. J Am Soc Nephrol. 2018;29:2348–61.
Kouros-Mehr H, Slorach EM, Sternlicht MD, Werb Z. GATA-3 maintains the differentiation of the luminal cell fate in the mammary gland. Cell. 2006;127:1041–55.
Higgins JPT, Kaygusuz G, Wang L, Montgomery K, Mason V, Zhu SX, et al. Placental S100 (S100P) and GATA3: markers for transitional epithelium and urothelial carcinoma discovered by complementary DNA microarray. Am J Surg Pathol. 2007;31:673–80.
Ainoya K, Moriguchi T, Ohmori S, Souma T, Takai J, Morita M, et al. UG4 enhancer-driven GATA-2 and bone morphogenetic protein 4 complementation remedies the CAKUT phenotype in Gata2 hypomorphic mutant mice. Mol Cell Biol. 2012;32:2312–22.
Hoshino T, Shimizu R, Ohmori S, Nagano M, Pan X, Ohneda O, et al. Reduced BMP4 abundance in Gata2 hypomorphic mutant mice result in uropathies resembling human CAKUT. Genes Cells. 2008;13:159–70.
Fishwick C, Higgins J, Percival-Alwyn L, Hustler A, Pearson J, Bastkowski S, et al. Heterarchy of transcription factors driving basal and luminal cell phenotypes in human urothelium. Cell Death Differ. 2017;24:809–18.
Wang C, Yang S, Jin L, Dai G, Yao Q, Xiang H, et al. Biological and clinical significance of GATA3 detected from TCGA database and FFPE sample in bladder cancer patients. Onco Targets Ther. 2020;13:945–58.
Iyyanki T, Zhang B, Wang Q, Hou Y, Jin Q, Xu J, et al. Subtype-associated epigenomic landscape and 3D genome structure in bladder cancer. Genome Biol. 2021;22:105.
Lerner SP, McConkey DJ, Hoadley KA, Chan KS, Kim WY, Radvanyi F, et al. Bladder cancer molecular taxonomy: summary from a consensus meeting. Bladder Cancer. 2016;2:37–47.
Miyamoto H, Izumi K, Yao JL, Li Y, Yang Q, McMahon LA, et al. GATA binding protein 3 is down-regulated in bladder cancer yet strong expression is an independent predictor of poor prognosis in invasive tumor. Hum Pathol. 2012;43:2033–40.
Choi W, Porten S, Kim S, Willis D, Plimack ER, Hoffman-Censits J, et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell. 2014;25:152–65.
Eriksson P, Aine M, Veerla S, Liedberg F, Sjödahl G, Höglund M. Molecular subtypes of urothelial carcinoma are defined by specific gene regulatory systems. BMC Med Genom. 2015;8:25.
van Kessel KEM, van der Keur KA, Dyrskjøt L, Algaba F, Welvaart NYC, Beukers W, et al. Molecular markers increase precision of the European Association of Urology non-muscle-invasive bladder cancer progression risk groups. Clin Cancer Res. 2018;24:1586–93.
Clark KL, Halay ED, Lai E, Burley SK. Co-crystal structure of the HNF-3/fork head DNA-recognition motif resembles histone H5. Nature. 1993;364:412–20.
Cirillo LA, McPherson CE, Bossard P, Stevens K, Cherian S, Shim EY, et al. Binding of the winged-helix transcription factor HNF3 to a linker histone site on the nucleosome. EMBO J. 1998;17:244–54.
Cirillo LA, Lin FR, Cuesta I, Friedman D, Jarnik M, Zaret KS. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol Cell. 2002;9:279–89.
Li Z, Tuteja G, Schug J, Kaestner KH. Foxa1 and Foxa2 are essential for sexual dimorphism in liver cancer. Cell. 2012;148:72–83.
Tsuchiya H, da Costa K-A, Lee S, Renga B, Jaeschke H, Yang Z, et al. Interactions between nuclear receptor SHP and FOXA1 maintain oscillatory homocysteine homeostasis in mice. Gastroenterology. 2015;148:1012–.e14.
Belikov S, Astrand C, Wrange O. FoxA1 binding directs chromatin structure and the functional response of a glucocorticoid receptor-regulated promoter. Mol Cell Biol. 2009;29:5413–25.
Besnard V, Wert SE, Hull WM, Whitsett JA. Immunohistochemical localization of Foxa1 and Foxa2 in mouse embryos and adult tissues. Gene Expr Patterns. 2004;5:193–208.
Oottamasathien S, Wang Y, Williams K, Franco OE, Wills ML, Thomas JC, et al. Directed differentiation of embryonic stem cells into bladder tissue. Dev Biol. 2007;304:556–66.
Varley CL, Bacon EJ, Holder JC, Southgate J. FOXA1 and IRF-1 intermediary transcriptional regulators of PPARgamma-induced urothelial cytodifferentiation. Cell Death Differ. 2009;16:103–14.
DeGraff DJ, Clark PE, Cates JM, Yamashita H, Robinson VL, Yu X, et al. Loss of the urothelial differentiation marker FOXA1 is associated with high grade, late stage bladder cancer and increased tumor proliferation. PLoS One. 2012;7:e36669.
Guo Y, Yuan X, Li K, Dai M, Zhang L, Wu Y, et al. GABPA is a master regulator of luminal identity and restrains aggressive diseases in bladder cancer. Cell Death Differ. 2020;27:1862–77.
Reddy OL, Cates JM, Gellert LL, Crist HS, Yang Z, Yamashita H, et al. Loss of FOXA1 drives sexually dimorphic changes in urothelial differentiation and is an independent predictor of poor prognosis in bladder cancer. Am J Pathol. 2015;185:1385–95.
Osei-Amponsa V, Buckwalter JM, Shuman L, Zheng Z, Yamashita H, Walter V, et al. Hypermethylation of FOXA1 and allelic loss of PTEN drive squamous differentiation and promote heterogeneity in bladder cancer. Oncogene. 2020;39:1302–17.
Bernardo GM, Keri RA. FOXA1: a transcription factor with parallel functions in development and cancer. Biosci Rep. 2012;32:113–30.
Robinson D, Van Allen EM, Wu Y-M, Schultz N, Lonigro RJ, Mosquera J-M, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;161:1215–28.
Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell. 2015;163:506–19.
Sikic D, Eckstein M, Wirtz RM, Jarczyk J, Worst TS, Porubsky S, et al. FOXA1 gene expression for defining molecular subtypes of muscle-invasive bladder cancer after radical cystectomy. J Clin Med. 2020;9:994.
Oettgen P, Alani RM, Barcinski MA, Brown L, Akbarali Y, Boltax J, et al. Isolation and characterization of a novel epithelium-specific transcription factor, ESE-1, a member of the ets family. Mol Cell Biol. 1997;17:4419–33.
Ng AY-N, Waring P, Ristevski S, Wang C, Wilson T, Pritchard M, et al. Inactivation of the transcription factor Elf3 in mice results in dysmorphogenesis and altered differentiation of intestinal epithelium. Gastroenterology. 2002;122:1455–66.
Böck M, Hinley J, Schmitt C, Wahlicht T, Kramer S, Southgate J. Identification of ELF3 as an early transcriptional regulator of human urothelium. Dev Biol. 2014;386:321–30.
Na L, Wang Z, Bai Y, Sun Y, Dong D, Wang W, et al. WNT7B represses epithelial-mesenchymal transition and stem-like properties in bladder urothelial carcinoma. Biochim Biophys Acta Mol Basis Dis. 2022;1868:166271.
Gondkar K, Patel K, Krishnappa S, Patil A, Nair B, Sundaram GM, et al. E74 like ETS transcription factor 3 (ELF3) is a negative regulator of epithelial- mesenchymal transition in bladder carcinoma. Cancer Biomark. 2019;25:223–32.
Ting SB, Wilanowski T, Cerruti L, Zhao L-L, Cunningham JM, Jane SM. The identification and characterization of human Sister-of-Mammalian Grainyhead (SOM) expands the grainyhead-like family of developmental transcription factors. Biochem J. 2003;370:953–62.
Traylor-Knowles N, Hansen U, Dubuc TQ, Martindale MQ, Kaufman L, Finnerty JR. The evolutionary diversification of LSF and Grainyhead transcription factors preceded the radiation of basal animal lineages. BMC Evol Biol. 2010;10:101.
Yu Z, Mannik J, Soto A, Lin KK, Andersen B. The epidermal differentiation-associated Grainyhead gene Get1/Grhl3 also regulates urothelial differentiation. EMBO J. 2009;28:1890–903.
Osborn SL, Thangappan R, Luria A, Lee JH, Nolta J, Kurzrock EA. Induction of human embryonic and induced pluripotent stem cells into urothelium. Stem Cells Transl Med. 2014;3:610–9.
Wezel F, Lustig J, Azoitei A, Liu J, Meessen S, Najjar G, et al. Grainyhead-like 3 influences migration and invasion of urothelial carcinoma cells. Int J Mol Sci. 2021;22:2959.
Ghioni P, Bolognese F, Duijf PHG, Van Bokhoven H, Mantovani R, Guerrini L. Complex transcriptional effects of p63 isoforms: identification of novel activation and repression domains. Mol Cell Biol. 2002;22:8659–68.
Augustin M, Bamberger C, Paul D, Schmale H. Cloning and chromosomal mapping of the human p53-related KET gene to chromosome 3q27 and its murine homolog Ket to mouse chromosome 16. Mamm Genome. 1998;9:899–902.
Osada M, Ohba M, Kawahara C, Ishioka C, Kanamaru R, Katoh I, et al. Cloning and functional analysis of human p51, which structurally and functionally resembles p53. Nat Med. 1998;4:839–43.
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dötsch V, et al. p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell. 1998;2:305–16.
Dohn M, Zhang S, Chen X. p63alpha and DeltaNp63alpha can induce cell cycle arrest and apoptosis and differentially regulate p53 target genes. Oncogene. 2001;20:3193–205.
Lohrum MA, Vousden KH. Regulation and function of the p53-related proteins: same family, different rules. Trends Cell Biol. 2000;10:197–202.
Koster MI, Kim S, Mills AA, DeMayo FJ, Roop DR. p63 is the molecular switch for initiation of an epithelial stratification program. Genes Dev. 2004;18:126–31.
Guo X, Keyes WM, Papazoglu C, Zuber J, Li W, Lowe SW, et al. TAp63 induces senescence and suppresses tumorigenesis in vivo. Nat Cell Biol. 2009;11:1451–7.
Helton ES, Zhu J, Chen X. The unique NH2-terminally deleted (DeltaN) residues, the PXXP motif, and the PPXY motif are required for the transcriptional activity of the DeltaN variant of p63. J Biol Chem. 2006;281:2533–42.
Kouwenhoven EN, Oti M, Niehues H, van Heeringen SJ, Schalkwijk J, Stunnenberg HG, et al. Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation. EMBO Rep. 2015;16:863–78.
Somerville TDD, Xu Y, Miyabayashi K, Tiriac H, Cleary CR, Maia-Silva D, et al. TP63-mediated enhancer reprogramming drives the squamous subtype of pancreatic ductal adenocarcinoma. Cell Rep. 2018;25:1741–.e7.
Yang A, McKeon F. P63 and P73: P53 mimics, menaces and more. Nat Rev Mol Cell Biol. 2000;1:199–207.
Karni-Schmidt O, Castillo-Martin M, Shen TH, Gladoun N, Domingo-Domenech J, Sanchez-Carbayo M, et al. Distinct expression profiles of p63 variants during urothelial development and bladder cancer progression. Am J Pathol. 2011;178:1350–60.
Mills AA, Zheng B, Wang XJ, Vogel H, Roop DR, Bradley A. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature. 1999;398:708–13.
Yang A, Schweitzer R, Sun D, Kaghad M, Walker N, Bronson RT, et al. p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature. 1999;398:714–8.
Paris M, Rouleau M, Pucéat M, Aberdam D. Regulation of skin aging and heart development by TAp63. Cell Death Differ. 2012;19:186–93.
Urist MJ, Di Como CJ, Lu M-L, Charytonowicz E, Verbel D, Crum CP, et al. Loss of p63 expression is associated with tumor progression in bladder cancer. Am J Pathol. 2002;161:1199–206.
Signoretti S, Pires MM, Lindauer M, Horner JW, Grisanzio C, Dhar S, et al. p63 regulates commitment to the prostate cell lineage. Proc Natl Acad Sci USA. 2005;102:11355–60.
Cheng W, Jacobs WB, Zhang JJR, Moro A, Park J-H, Kushida M, et al. DeltaNp63 plays an anti-apoptotic role in ventral bladder development. Development. 2006;133:4783–92.
Celli J, Duijf P, Hamel BC, Bamshad M, Kramer B, Smits AP, et al. Heterozygous germline mutations in the p53 homolog p63 are the cause of EEC syndrome. Cell. 1999;99:143–53.
van Bokhoven H, Hamel BC, Bamshad M, Sangiorgi E, Gurrieri F, Duijf PH, et al. p63 Gene mutations in eec syndrome, limb-mammary syndrome, and isolated split hand-split foot malformation suggest a genotype-phenotype correlation. Am J Hum Genet. 2001;69:481–92.
Duijf PHG, Vanmolkot KRJ, Propping P, Friedl W, Krieger E, McKeon F, et al. Gain-of-function mutation in ADULT syndrome reveals the presence of a second transactivation domain in p63. Hum Mol Genet. 2002;11:799–804.
McGrath JA, Duijf PH, Doetsch V, Irvine AD, de Waal R, Vanmolkot KR, et al. Hay-Wells syndrome is caused by heterozygous missense mutations in the SAM domain of p63. Hum Mol Genet. 2001;10:221–9.
Kloesch B, Ionasz V, Paliwal S, Hruschka N, Martinez de Villarreal J, Öllinger R, et al. A GATA6-centred gene regulatory network involving HNFs and ΔNp63 controls plasticity and immune escape in pancreatic cancer. Gut. 2022;71:766–77.
Bankhead A, McMaster T, Wang Y, Boonstra PS, Palmbos PL. TP63 isoform expression is linked with distinct clinical outcomes in cancer. EBioMedicine. 2020;51:102561.
Papadimitriou M-A, Avgeris M, Levis PK, Tokas T, Stravodimos K, Scorilas A. ΔNp63 transcript loss in bladder cancer constitutes an independent molecular predictor of TaT1 patients post-treatment relapse and progression. J Cancer Res Clin Oncol. 2019;145:3075–87.
Guo CC, Majewski T, Zhang L, Yao H, Bondaruk J, Wang Y, et al. Dysregulation of EMT drives the progression to clinically aggressive sarcomatoid bladder cancer. Cell Rep. 2019;27:1781–.e4.
Eckert D, Buhl S, Weber S, Jäger R, Schorle H. The AP-2 family of transcription factors. Genome Biol. 2005;6:246.
Sinha S, Fuchs E. Identification and dissection of an enhancer controlling epithelial gene expression in skin. Proc Natl Acad Sci USA. 2001;98:2455–60.
Kaufman CK, Sinha S, Bolotin D, Fan J, Fuchs E. Dissection of a complex enhancer element: maintenance of keratinocyte specificity but loss of differentiation specificity. Mol Cell Biol. 2002;22:4293–308.
Leask A, Byrne C, Fuchs E. Transcription factor AP2 and its role in epidermal-specific gene expression. Proc Natl Acad Sci USA. 1991;88:7948–52.
McDade SS, Henry AE, Pivato GP, Kozarewa I, Mitsopoulos C, Fenwick K, et al. Genome-wide analysis of p63 binding sites identifies AP-2 factors as co-regulators of epidermal differentiation. Nucleic Acids Res. 2012;40:7190–206.
Yamashita H, Kawasawa YI, Shuman L, Zheng Z, Tran T, Walter V, et al. Repression of transcription factor AP-2 alpha by PPARγ reveals a novel transcriptional circuit in basal-squamous bladder cancer. Oncogenesis. 2019;8:69.
Davis AC, Wims M, Spotts GD, Hann SR, Bradley A. A null c-myc mutation causes lethality before 10.5 days of gestation in homozygotes and reduced fertility in heterozygous female mice. Genes Dev. 1993;7:671–82.
Watters AD, Latif Z, Forsyth A, Dunn I, Underwood MA, Grigor KM, et al. Genetic aberrations of c-myc and CCND1 in the development of invasive bladder cancer. Br J Cancer. 2002;87:654–8.
Li Y, Liu H, Lai C, Du X, Su Z, Gao S. The Lin28/let-7a/c-Myc pathway plays a role in non-muscle invasive bladder cancer. Cell Tissue Res. 2013;354:533–41.
Mahe M, Dufour F, Neyret-Kahn H, Moreno-Vega A, Beraud C, Shi M, et al. An FGFR3/MYC positive feedback loop provides new opportunities for targeted therapies in bladder cancers. EMBO Mol Med. 2018;10:e8163.
Zhuang C, Ma Q, Zhuang C, Ye J, Zhang F, Gui Y. LncRNA GClnc1 promotes proliferation and invasion of bladder cancer through activation of MYC. FASEB J. 2019;33:11045–59.
Jiang G, Huang C, Liao X, Li J, Wu X-R, Zeng F, et al. The RING domain in the anti-apoptotic protein XIAP stabilizes c-Myc protein and preserves anchorage-independent growth of bladder cancer cells. J Biol Chem. 2019;294:5935–44.
Robertson AG, Groeneveld CS, Jordan B, Lin X, McLaughlin KA, Das A, et al. Identification of differential tumor subtypes of T1 bladder cancer. Eur Urol. 2020;78:533–7.
Fontugne J, Wong J, Cabel L, Neyret-Kahn H, Karboul N, Maillé P, et al. Progression-associated molecular changes in basal/squamous and sarcomatoid bladder carcinogenesis. J Pathol. 2023;259:455–67.
Marquis L, Tran M, Choi W, Lee I-L, Huszar D, Siefker-Radtke A, et al. p63 expression correlates with sensitivity to the Eg5 inhibitor ZD4877 in bladder cancer cells. Cancer Biol Ther. 2012;13:477–86.
Greife A, Jankowiak S, Steinbring J, Nikpour P, Niegisch G, Hoffmann MJ, et al. Canonical Notch signalling is inactive in urothelial carcinoma. BMC Cancer. 2014;14:628.
Paraskevopoulou V, Bonis V, Dionellis VS, Paschalidis N, Melissa P, Chavdoula E, et al. Notch controls urothelial integrity in the mouse bladder. JCI Insight. 2020;5:e133232.
Rampias T, Vgenopoulou P, Avgeris M, Polyzos A, Stravodimos K, Valavanis C, et al. A new tumor suppressor role for the Notch pathway in bladder cancer. Nat Med. 2014;20:1199–205.
Maraver A, Fernandez-Marcos PJ, Cash TP, Mendez-Pertuz M, Dueñas M, Maietta P, et al. NOTCH pathway inactivation promotes bladder cancer progression. J Clin Investig. 2015;125:824–30.
Hayashi T, Gust KM, Wyatt AW, Goriki A, Jäger W, Awrey S, et al. Not all NOTCH is created equal: the oncogenic role of NOTCH2 in bladder cancer and its implications for targeted therapy. Clin Cancer Res. 2016;22:2981–92.
Zhang L, Sha J, Yang G, Huang X, Bo J, Huang Y. Activation of Notch pathway is linked with epithelial-mesenchymal transition in prostate cancer cells. Cell Cycle. 2017;16:999–1007.
Watabe T, Yoshida K, Shindoh M, Kaya M, Fujikawa K, Sato H, et al. The Ets-1 and Ets-2 transcription factors activate the promoters for invasion-associated urokinase and collagenase genes in response to epidermal growth factor. Int J Cancer. 1998;77:128–37.
Westermarck J, Seth A, Kähäri VM. Differential regulation of interstitial collagenase (MMP-1) gene expression by ETS transcription factors. Oncogene. 1997;14:2651–60.
Hashiya N, Jo N, Aoki M, Matsumoto K, Nakamura T, Sato Y, et al. In vivo evidence of angiogenesis induced by transcription factor Ets-1: Ets-1 is located upstream of angiogenesis cascade. Circulation. 2004;109:3035–41.
Sari A, Calli A, Gorgel SN, Altinboga AA, Kara C, Dincel C, et al. Immunohistochemical determination of ETS-1 oncoprotein expression in urothelial carcinomas of the urinary bladder. Appl Immunohistochem Mol Morphol. 2012;20:153–8.
Liu L, Liu Y, Zhang X, Chen M, Wu H, Lin M, et al. Inhibiting cell migration and cell invasion by silencing the transcription factor ETS-1 in human bladder cancer. Oncotarget. 2016;7:25125–34.
Lin S-R, Yeh H-C, Wang W-J, Ke H-L, Lin H-H, Hsu W-C, et al. MiR-193b mediates CEBPD-induced cisplatin sensitization through targeting ETS1 and cyclin D1 in human urothelial carcinoma cells. J Cell Biochem. 2017;118:1563–73.
Shin S-S, Park S-S, Hwang B, Kim WT, Choi YH, Kim W-J, et al. MicroRNA-106a suppresses proliferation, migration, and invasion of bladder cancer cells by modulating MAPK signaling, cell cycle regulators, and Ets-1-mediated MMP-2 expression. Oncol Rep. 2016;36:2421–9.
Bell SM, Zhang L, Mendell A, Xu Y, Haitchi HM, Lessard JL, et al. Kruppel-like factor 5 is required for formation and differentiation of the bladder urothelium. Dev Biol. 2011;358:79–90.
Zhang B, Li Y, Wu Q, Xie L, Barwick B, Fu C, et al. Acetylation of KLF5 maintains EMT and tumorigenicity to cause chemoresistant bone metastasis in prostate cancer. Nat Commun. 2021;12:1714.
Du C, Gao Y, Xu S, Jia J, Huang Z, Fan J, et al. KLF5 promotes cell migration by up-regulating FYN in bladder cancer cells. FEBS Lett. 2016;590:408–18.
Gao Y, Wu K, Chen Y, Zhou J, Du C, Shi Q, et al. Beyond proliferation: KLF5 promotes angiogenesis of bladder cancer through directly regulating VEGFA transcription. Oncotarget. 2015;6:43791–805.
Ohnishi S, Ohnami S, Laub F, Aoki K, Suzuki K, Kanai Y, et al. Downregulation and growth inhibitory effect of epithelial-type Krüppel-like transcription factor KLF4, but not KLF5, in bladder cancer. Biochem Biophys Res Commun. 2003;308:251–6.
Li H, Wang J, Xiao W, Xia D, Lang B, Wang T, et al. Epigenetic inactivation of KLF4 is associated with urothelial cancer progression and early recurrence. J Urol. 2014;191:493–501.
Ai X, Jia Z, Liu S, Wang J, Zhang X. Notch-1 regulates proliferation and differentiation of human bladder cancer cell lines by inhibiting expression of Krüppel-like factor 4. Oncol Rep. 2014;32:1459–64.
Xu X, Li J, Zhu Y, Xie B, Wang X, Wang S, et al. CRISPR-ON-Mediated KLF4 overexpression inhibits the proliferation, migration and invasion of urothelial bladder cancer in vitro and in vivo. Oncotarget. 2017;8:102078–87.
Suske G. The Sp-family of transcription factors. Gene. 1999;238:291–300.
Hagen G, Müller S, Beato M, Suske G. Sp1-mediated transcriptional activation is repressed by Sp3. EMBO J. 1994;13:3843–51.
Xu J, Hua X, Yang R, Jin H, Li J, Zhu J, et al. XIAP Interaction with E2F1 and Sp1 via its BIR2 and BIR3 domains specific activated MMP2 to promote bladder cancer invasion. Oncogenesis. 2019;8:71.
Huang H, Jin H, Zhao H, Wang J, Li X, Yan H, et al. RhoGDIβ promotes Sp1/MMP-2 expression and bladder cancer invasion through perturbing miR-200c-targeted JNK2 protein translation. Mol Oncol. 2017;11:1579–94.
Zhu J, Lu Z, Ke M, Cai X. Sp1 is overexpressed and associated with progression and poor prognosis in bladder urothelial carcinoma patients. Int Urol Nephrol. 2022;54:1505–12.
Malats N, Real FX. Epidemiology of bladder cancer. Hematol. Oncol. Clin. North Am. 2015;29:177–89.
Caliri AW, Tommasi S, Besaratinia A. Relationships among smoking, oxidative stress, inflammation, macromolecular damage, and cancer. Mutat Res Rev Mutat Res. 2021;787:108365.
Denison MS, Nagy SR. Activation of the aryl hydrocarbon receptor by structurally diverse exogenous and endogenous chemicals. Annu Rev Pharm Toxicol. 2003;43:309–34.
Pollenz RS, Sattler CA, Poland A. The aryl hydrocarbon receptor and aryl hydrocarbon receptor nuclear translocator protein show distinct subcellular localizations in Hepa 1c1c7 cells by immunofluorescence microscopy. Mol Pharm. 1994;45:428–38.
Hord NG, Perdew GH. Physicochemical and immunocytochemical analysis of the aryl hydrocarbon receptor nuclear translocator: characterization of two monoclonal antibodies to the aryl hydrocarbon receptor nuclear translocator. Mol Pharm. 1994;46:618–26.
Soshilov AA, Motta S, Bonati L, Denison MS. Transitional states in ligand-dependent transformation of the aryl hydrocarbon receptor into its DNA-binding form. Int J Mol Sci. 2020;21:2474.
Yu J, Lu Y, Muto S, Ide H, Horie S. The dual function of aryl hydrocarbon receptor in bladder carcinogenesis. Anticancer Res. 2020;40:1345–57.
Baker SC, Arlt VM, Indra R, Joel M, Stiborová M, Eardley I, et al. Differentiation-associated urothelial cytochrome P450 oxidoreductase predicates the xenobiotic-metabolizing activity of “luminal” muscle-invasive bladder cancers. Mol Carcinog. 2018;57:606–18.
Vlaar JM, Borgman A, Kalkhoven E, Westland D, Besselink N, Shale C, et al. Recurrent exon-deleting activating mutations in AHR act as drivers of urinary tract cancer. Sci Rep. 2022;12:10081.
Huang HC, Nguyen T, Pickett CB. Regulation of the antioxidant response element by protein kinase C-mediated phosphorylation of NF-E2-related factor 2. Proc Natl Acad Sci USA. 2000;97:12475–80.
Hayashi M, Guida E, Inokawa Y, Goldberg R, Reis LO, Ooki A, et al. GULP1 regulates the NRF2-KEAP1 signaling axis in urothelial carcinoma. Sci Signal. 2020;13:eaba0443.
Ihara T, Mitsui T, Nakamura Y, Kanda M, Tsuchiya S, Kira S, et al. The oscillation of intracellular Ca2+ influx associated with the circadian expression of Piezo1 and TRPV4 in the bladder urothelium. Sci Rep. 2018;8:5699.
Litlekalsoy J, Rostad K, Kalland K-H, Hostmark JG, Laerum OD. Expression of circadian clock genes and proteins in urothelial cancer is related to cancer-associated genes. BMC Cancer. 2016;16:549.
Honma S, Kawamoto T, Takagi Y, Fujimoto K, Sato F, Noshiro M, et al. Dec1 and Dec2 are regulators of the mammalian molecular clock. Nature. 2002;419:841–4.
Cobo I, Martinelli P, Flández M, Bakiri L, Zhang M, Carrillo-de-Santa-Pau E, et al. Transcriptional regulation by NR5A2 links differentiation and inflammation in the pancreas. Nature. 2018;554:533–7.
Bejjani F, Evanno E, Zibara K, Piechaczyk M, Jariel-Encontre I. The AP-1 transcriptional complex: Local switch or remote command? Biochim Biophys Acta Rev Cancer. 2019;1872:11–23.
Wang Y, Geng H, Zhao L, Zhang Z, Xie D, Zhang T, et al. Role of AP-1 in the tobacco smoke-induced urocystic abnormal cell differentiation and epithelial-mesenchymal transition in vivo. Int J Clin Exp Pathol. 2017;10:8243–52.
Barrows D, Feng L, Carroll TS, Allis CD. Loss of UTX/KDM6A and the activation of FGFR3 converge to regulate differentiation gene-expression programs in bladder cancer. Proc Natl Acad Sci USA. 2020;117:25732–41.
Qiu H, Makarov V, Bolzenius JK, Halstead A, Parker Y, Wang A, et al. KDM6A loss triggers an epigenetic switch that disrupts urothelial differentiation and drives cell proliferation in bladder cancer. Cancer Res. 2023;83:814–29.
Neyret-Kahn H, Fontugne J, Meng XY, Groeneveld CS, Cabel L, Ye T, et al. Epigenomic mapping identifies an enhancer repertoire that regulates cell identity in bladder cancer through distinct transcription factor networks. Oncogene. 2023;42:1524–42.
Cai Z, Chen H, Bai J, Zheng Y, Ma J, Cai X, et al. Copy number variations of CEP63, FOSL2 and PAQR6 serve as novel signatures for the prognosis of bladder cancer. Front Oncol. 2021;11:674933.
Iwata J, Suzuki A, Pelikan RC, Ho T-V, Sanchez-Lara PA, Urata M, et al. Smad4-Irf6 genetic interaction and TGFβ-mediated IRF6 signaling cascade are crucial for palatal fusion in mice. Development. 2013;140:1220–30.
Richardson RJ, Dixon J, Malhotra S, Hardman MJ, Knowles L, Boot-Handford RP, et al. Irf6 is a key determinant of the keratinocyte proliferation-differentiation switch. Nat Genet. 2006;38:1329–34.
Restivo G, Nguyen B-C, Dziunycz P, Ristorcelli E, Ryan RJH, Özuysal ÖY, et al. IRF6 is a mediator of Notch pro-differentiation and tumour suppressive function in keratinocytes. EMBO J. 2011;30:4571–85.
Weng H, Yuan S, Huang Q, Zeng X-T, Wang X-H. STAT1 is a key gene in a gene regulatory network related to immune phenotypes in bladder cancer: An integrative analysis of multi-omics data. J Cell Mol Med. 2021;25:3258–71.
Su Q, Sun Y, Zhang Z, Yang Z, Qiu Y, Li X, et al. Identification of prognostic immune genes in bladder urothelial carcinoma. Biomed Res Int. 2020;2020:7510120.
Kawahara T, Ishiguro Y, Ohtake S, Kato I, Ito Y, Ito H, et al. PD-1 and PD-L1 are more highly expressed in high-grade bladder cancer than in low-grade cases: PD-L1 might function as a mediator of stage progression in bladder cancer. BMC Urol. 2018;18:97.
Huang W-T, Yang S-F, Wu C-C, Chen W-T, Huang Y-C, Su Y-C, et al. Expression of signal transducer and activator of transcription 3 and suppressor of cytokine signaling 3 in urothelial carcinoma. Kaohsiung J Med Sci. 2009;25:640–6.
Hindupur SV, Schmid SC, Koch JA, Youssef A, Baur E-M, Wang D, et al. STAT3/5 inhibitors suppress proliferation in bladder cancer and enhance oncolytic adenovirus therapy. Int J Mol Sci. 2020;21:1106.
Gatta LB, Melocchi L, Bugatti M, Missale F, Lonardi S, Zanetti B, et al. Hyper-activation of STAT3 sustains progression of non-papillary basal-type bladder cancer via FOSL1 regulome. Cancers. 2019;11:1219.
Ching CB, Gupta S, Li B, Cortado H, Mayne N, Jackson AR, et al. Interleukin-6/Stat3 signaling has an essential role in the host antimicrobial response to urinary tract infection. Kidney Int. 2018;93:1320–9.
Schneidewind L, Neumann T, Plis A, Brückmann S, Keiser M, Krüger W, et al. Novel 3D organotypic urothelial cell culture model for identification of new therapeutic approaches in urological infections. J Clin Virol. 2020;124:104283.
Boorjian S, Ugras S, Mongan NP, Gudas LJ, You X, Tickoo SK, et al. Androgen receptor expression is inversely correlated with pathologic tumor stage in bladder cancer. Urology. 2004;64:383–8.
Tuygun C, Kankaya D, Imamoglu A, Sertcelik A, Zengin K, Oktay M, et al. Sex-specific hormone receptors in urothelial carcinomas of the human urinary bladder: a comparative analysis of clinicopathological features and survival outcomes according to receptor expression. Urol Oncol. 2011;29:43–51.
Miyamoto H, Yao JL, Chaux A, Zheng Y, Hsu I, Izumi K, et al. Expression of androgen and oestrogen receptors and its prognostic significance in urothelial neoplasm of the urinary bladder. BJU Int. 2012;109:1716–26.
Miyamoto H, Yang Z, Chen Y-T, Ishiguro H, Uemura H, Kubota Y, et al. Promotion of bladder cancer development and progression by androgen receptor signals. J Natl Cancer Inst. 2007;99:558–68.
Johnson AM, O’Connell MJ, Messing EM, Reeder JE. Decreased bladder cancer growth in parous mice. Urology. 2008;72:470–3.
Shen SS, Smith CL, Hsieh J-T, Yu J, Kim IY, Jian W, et al. Expression of estrogen receptors-alpha and -beta in bladder cancer cell lines and human bladder tumor tissue. Cancer. 2006;106:2610–6.
Hsu L-H, Liu K-J, Tsai M-F, Wu C-R, Feng A-C, Chu N-M, et al. Estrogen adversely affects the prognosis of patients with lung adenocarcinoma. Cancer Sci. 2015;106:51–9.
Ide H, Inoue S, Miyamoto H. Histopathological and prognostic significance of the expression of sex hormone receptors in bladder cancer: a meta-analysis of immunohistochemical studies. PLoS One. 2017;12:e0174746.
Warot X, Fromental-Ramain C, Fraulob V, Chambon P, Dollé P. Gene dosage-dependent effects of the Hoxa-13 and Hoxd-13 mutations on morphogenesis of the terminal parts of the digestive and urogenital tracts. Development. 1997;124:4781–91.
Mortlock DP, Innis JW. Mutation of HOXA13 in hand-foot-genital syndrome. Nat Genet. 1997;15:179–80.
Warrick JI, Knowles MA, Hurst CD, Shuman L, Raman JD, Walter V, et al. A transcriptional network of cell cycle dysregulation in noninvasive papillary urothelial carcinoma. Sci Rep. 2022;12:16538.
Marzouka N-A-D, Eriksson P, Bernardo C, Hurst CD, Knowles MA, Sjödahl G, et al. The lund molecular taxonomy applied to non-muscle-invasive urothelial carcinoma. J Mol Diagn. 2022;24:992–1008.
Lauss M, Aine M, Sjödahl G, Veerla S, Patschan O, Gudjonsson S, et al. DNA methylation analyses of urothelial carcinoma reveal distinct epigenetic subtypes and an association between gene copy number and methylation status. Epigenetics. 2012;7:858–67.
Aine M, Sjödahl G, Eriksson P, Veerla S, Lindgren D, Ringnér M, et al. Integrative epigenomic analysis of differential DNA methylation in urothelial carcinoma. Genome Med. 2015;7:23.
Bracken AP, Dietrich N, Pasini D, Hansen KH, Helin K. Genome-wide mapping of polycomb target genes unravels their roles in cell fate transitions. Genes Dev. 2006;20:1123–36.
Picchetti T, Chiquet J, Elati M, Neuvial P, Nicolle R, Birmelé E. A model for gene deregulation detection using expression data. BMC Syst Biol. 2015;9:S6.
Nicolle R, Radvanyi F, Elati M. CoRegNet: reconstruction and integrated analysis of co-regulatory networks. Bioinformatics. 2015;31:3066–8.
Champion M, Chiquet J, Neuvial P, Elati M, Radvanyi F, Birmelé E. Identification of deregulation mechanisms specific to cancer subtypes. J Bioinf Comput Biol. 2021;19:2140003.
Liang Y, Li L, Xin T, Li B, Zhang D. Superenhancer-transcription factor regulatory network in malignant tumors. Open Med (Wars). 2021;16:1564–82.
Sfakianos JP, Daza J, Hu Y, Anastos H, Bryant G, Bareja R, et al. Epithelial plasticity can generate multi-lineage phenotypes in human and murine bladder cancers. Nat Commun. 2020;11:2540.
Wang H, Mei Y, Luo C, Huang Q, Wang Z, Lu G-M, et al. Single-cell analyses reveal mechanisms of cancer stem cell maintenance and epithelial-mesenchymal transition in recurrent bladder cancer. Clin Cancer Res. 2021;27:6265–78.
Wang K-J, Wang C, Dai L-H, Yang J, Huang H, Ma X-J, et al. Targeting an autocrine regulatory loop in cancer stem-like cells impairs the progression and chemotherapy resistance of bladder cancer. Clin Cancer Res. 2019;25:1070–86.
Whitfield JR, Beaulieu M-E, Soucek L. Strategies to inhibit myc and their clinical applicability. Front Cell Dev Biol. 2017;5:10.
Beaulieu M-E, Jauset T, Massó-Vallés D, Martínez-Martín S, Rahl P, Maltais L, et al. Intrinsic cell-penetrating activity propels Omomyc from proof of concept to viable anti-MYC therapy. Sci Transl Med. 2019;11:eaar5012.
Lindskrog SV, Prip F, Lamy P, Taber A, Groeneveld CS, Birkenkamp-Demtröder K, et al. An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer. Nat Commun. 2021;12:2301.
Chamberlain PP, Hamann LG. Development of targeted protein degradation therapeutics. Nat Chem Biol. 2019;15:937–44.
Qi S-M, Dong J, Xu Z-Y, Cheng X-D, Zhang W-D, Qin J-J. PROTAC: an effective targeted protein degradation strategy for cancer therapy. Front Pharm. 2021;12:692574.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38:576–89.
Brooks DJ, Woodward S, Thompson FH, Dos Santos B, Russell M, Yang JM, et al. Expression of the zinc finger gene EVI-1 in ovarian and other cancers. Br J Cancer. 1996;74:1518–25.
Inoue Y, Kishida T, Kotani S-I, Akiyoshi M, Taga H, Seki M, et al. Direct conversion of fibroblasts into urothelial cells that may be recruited to regenerating mucosa of injured urinary bladder. Sci Rep. 2019;9:13850.
Primdahl H, von der Maase H, Christensen M, Wolf H, Orntoft TF. Allelic deletions of cell growth regulators during progression of bladder cancer. Cancer Res. 2000;60:6623–9.
Boorjian SA, Heemers HV, Frank I, Farmer SA, Schmidt LJ, Sebo TJ, et al. Expression and significance of androgen receptor coactivators in urothelial carcinoma of the bladder. Endocr Relat Cancer. 2009;16:123–37.
Dozmorov M, Stone R, Clifford JL, Sabichi AL, Engles CD, Hauser PJ, et al. System level changes in gene expression in maturing bladder mucosa. J Urol. 2011;185:1952–8.
Zhang Y, Dufau ML. Nuclear orphan receptors regulate transcription of the gene for the human luteinizing hormone receptor. J Biol Chem. 2000;275:2763–70.
Hermann-Kleiter N, Gruber T, Lutz-Nicoladoni C, Thuille N, Fresser F, Labi V, et al. The nuclear orphan receptor NR2F6 suppresses lymphocyte activation and T helper 17-dependent autoimmunity. Immunity. 2008;29:205–16.
Du X, Wang Q-R, Chan E, Merchant M, Liu J, French D, et al. FGFR3 stimulates stearoyl CoA desaturase 1 activity to promote bladder tumor growth. Cancer Res. 2012;72:5843–55.
Fu Y, Sun S, Bi J, Kong C, Yin L. Construction and analysis of a ceRNA network and patterns of immune infiltration in bladder cancer. Transl Androl Urol. 2021;10:1939–55.
Jiang A, Liu N, Bai S, Wang J, Gao H, Zheng X, et al. The construction and analysis of tumor-infiltrating immune cells and ceRNA networks in bladder cancer. Front Genet. 2020;11:605767.
Papaioannou VE. The T-box gene family: emerging roles in development, stem cells and cancer. Development. 2014;141:3819–33.
Lingbeek ME, Jacobs JJL, van Lohuizen M. The T-box repressors TBX2 and TBX3 specifically regulate the tumor suppressor gene p14ARF via a variant T-site in the initiator. J Biol Chem. 2002;277:26120–7.
Peres J, Davis E, Mowla S, Bennett DC, Li JA, Wansleben S, et al. The highly homologous T-box transcription Factors, TBX2 and TBX3, have distinct roles in the oncogenic process. Genes Cancer. 2010;1:272–82.
Ichijo R, Kobayashi H, Yoneda S, Iizuka Y, Kubo H, Matsumura S, et al. Tbx3-dependent amplifying stem cell progeny drives interfollicular epidermal expansion during pregnancy and regeneration. Nat Commun. 2017;8:508.
Ito A, Asamoto M, Hokaiwado N, Takahashi S, Shirai T. Tbx3 expression is related to apoptosis and cell proliferation in rat bladder both hyperplastic epithelial cells and carcinoma cells. Cancer Lett. 2005;219:105–12.
Aydogdu N, Rudat C, Trowe M-O, Kaiser M, Lüdtke TH, Taketo MM, et al. TBX2 and TBX3 act downstream of canonical WNT signaling in patterning and differentiation of the mouse ureteric mesenchyme. Development. 2018;145:dev171827.
Shi Z, Li X, Wu D, Tang R, Chen R, Xue S, et al. Silencing of HMGA2 suppresses cellular proliferation, migration, invasion, and epithelial-mesenchymal transition in bladder cancer. Tumour Biol. 2016;37:7515–23.
Chen Z, Li Q, Wang S, Zhang J. miR-485-5p inhibits bladder cancer metastasis by targeting HMGA2. Int J Mol Med. 2015;36:1136–42.
Ding X, Wang Y, Ma X, Guo H, Yan X, Chi Q, et al. Expression of HMGA2 in bladder cancer and its association with epithelial-to-mesenchymal transition. Cell Prolif. 2014;47:146–51.
Zhang Y, Luo G, You S, Zhang L, Liang C, Chen X. Exosomal LINC00355 derived from cancer-associated fibroblasts promotes bladder cancer cell proliferation and invasion by regulating miR-15a-5p/HMGA2 axis. Acta Biochim Biophys Sin. 2021;53:673–82.
Zhuang J, Shen L, Yang L, Huang X, Lu Q, Cui Y, et al. TGFβ1 promotes gemcitabine resistance through regulating the LncRNA-LET/NF90/miR-145 signaling axis in bladder cancer. Theranostics. 2017;7:3053–67.
Zheng Y, Izumi K, Yao JL, Miyamoto H. Dihydrotestosterone upregulates the expression of epidermal growth factor receptor and ERBB2 in androgen receptor-positive bladder cancer cells. Endocr Relat Cancer. 2011;18:451–64.
Xu C, Sun M, Zhang X, Xu Z, Miyamoto H, Zheng Y. Activation of glucocorticoid receptor inhibits the stem-like properties of bladder cancer via inactivating the β-catenin pathway. Front Oncol. 2020;10:1332.
Sun J, Hoshino H, Takaku K, Nakajima O, Muto A, Suzuki H, et al. Hemoprotein Bach1 regulates enhancer availability of heme oxygenase-1 gene. EMBO J. 2002;21:5216–24.
Kanno H, Ozawa H, Dohi Y, Sekiguchi A, Igarashi K, Itoi E. Genetic ablation of transcription repressor Bach1 reduces neural tissue damage and improves locomotor function after spinal cord injury in mice. J Neurotrauma. 2009;26:31–9.
Somerville TDD, Xu Y, Wu XS, Maia-Silva D, Hur SK, de Almeida LMN, et al. ZBED2 is an antagonist of interferon regulatory factor 1 and modifies cell identity in pancreatic cancer. Proc Natl Acad Sci USA. 2020;117:11471–82.
Finnegan A, Cho RJ, Luu A, Harirchian P, Lee J, Cheng JB, et al. Single-cell transcriptomics reveals spatial and temporal turnover of keratinocyte differentiation regulators. Front Genet. 2019;10:775.
Fossum SL, Mutolo MJ, Tugores A, Ghosh S, Randell SH, Jones LC, et al. Ets homologous factor (EHF) has critical roles in epithelial dysfunction in airway disease. J Biol Chem. 2017;292:10938–49.
Asai T, Morrison SL. The SRC family tyrosine kinase HCK and the ETS family transcription factors SPIB and EHF regulate transcytosis across a human follicle-associated epithelium model. J Biol Chem. 2013;288:10395–405.
Rubin AJ, Barajas BC, Furlan-Magaril M, Lopez-Pajares V, Mumbach MR, Howard I, et al. Lineage-specific dynamic and pre-established enhancer-promoter contacts cooperate in terminal differentiation. Nat Genet. 2017;49:1522–8.
Shi J, Qu Y, Li X, Sui F, Yao D, Yang Q, et al. Increased expression of EHF via gene amplification contributes to the activation of HER family signaling and associates with poor survival in gastric cancer. Cell Death Dis. 2016;7:e2442.
Taniue K, Oda T, Hayashi T, Okuno M, Akiyama T. A member of the ETS family, EHF, and the ATPase RUVBL1 inhibit p53-mediated apoptosis. EMBO Rep. 2011;12:682–9.
Lv Y, Sui F, Ma J, Ren X, Yang Q, Zhang Y, et al. Increased expression of EHF contributes to thyroid tumorigenesis through transcriptionally regulating HER2 and HER3. Oncotarget. 2016;7:57978–90.
Cheng Z, Guo J, Chen L, Luo N, Yang W, Qu X. Knockdown of EHF inhibited the proliferation, invasion and tumorigenesis of ovarian cancer cells. Mol Carcinog. 2016;55:1048–59.
Cangemi R, Mensah A, Albertini V, Jain A, Mello-Grand M, Chiorino G, et al. Reduced expression and tumor suppressor function of the ETS transcription factor ESE-3 in prostate cancer. Oncogene. 2008;27:2877–85.
Zhao T, Jiang W, Wang X, Wang H, Zheng C, Li Y, et al. ESE3 inhibits pancreatic cancer metastasis by upregulating E-cadherin. Cancer Res. 2017;77:874–85.
Wang L, Xing J, Cheng R, Shao Y, Li P, Zhu S, et al. Abnormal localization and tumor suppressor function of epithelial tissue-specific transcription factor ESE3 in esophageal squamous cell carcinoma. PLoS One. 2015;10:e0126319.
Fisher WG, Yang P-C, Medikonduri RK, Jafri MS. NFAT and NFkappaB activation in T lymphocytes: a model of differential activation of gene expression. Ann Biomed Eng. 2006;34:1712–28.
Lee JH, Kim M, Im YS, Choi W, Byeon SH, Lee HK. NFAT5 induction and its role in hyperosmolar stressed human limbal epithelial cells. Investig Ophthalmol Vis Sci. 2008;49:1827–35.
Johnsen O, Skammelsrud N, Luna L, Nishizawa M, Prydz H, Kolstø AB. Small Maf proteins interact with the human transcription factor TCF11/Nrf1/LCR-F1. Nucleic Acids Res. 1996;24:4289–97.
Johnsen O, Murphy P, Prydz H, Kolsto AB. Interaction of the CNC-bZIP factor TCF11/LCR-F1/Nrf1 with MafG: binding-site selection and regulation of transcription. Nucleic Acids Res. 1998;26:512–20.
Steffen J, Seeger M, Koch A, Krüger E. Proteasomal degradation is transcriptionally controlled by TCF11 via an ERAD-dependent feedback loop. Mol Cell. 2010;40:147–58.
Cui M, Atmanli A, Morales MG, Tan W, Chen K, Xiao X, et al. Nrf1 promotes heart regeneration and repair by regulating proteostasis and redox balance. Nat Commun. 2021;12:5270.
Xu Z, Chen L, Leung L, Yen TSB, Lee C, Chan JY. Liver-specific inactivation of the Nrf1 gene in adult mouse leads to nonalcoholic steatohepatitis and hepatic neoplasia. Proc Natl Acad Sci USA. 2005;102:4120–5.
Lu Q, Qiufang Y, Peng L, Xiaowen Z, Yonghui Y.Xiuman Z,et al. Function expansion of antitumor transcriptional activator NFE2L1 by the original discovery of its non-transcription factor activity. BioRxiv. 2020; https://doi.org/10.1101/2020.10.08.330597.
Xia W, Li Y, Wu Z, Wang Y, Xing N, Yang W, et al. Transcription factor YY1 mediates epithelial-mesenchymal transition through the TGFβ signaling pathway in bladder cancer. Med Oncol. 2020;37:93.
Chen W, Jiang T, Mao H, Gao R, Gao X, He Y, et al. Nodal promotes the migration and invasion of bladder cancer cells via regulation of snail. J Cancer. 2019;10:1511–9.
Li J, Xu X, Meng S, Liang Z, Wang X, Xu M, et al. MET/SMAD3/SNAIL circuit mediated by miR-323a-3p is involved in regulating epithelial-mesenchymal transition progression in bladder cancer. Cell Death Dis. 2017;8:e3010.
Acknowledgements
We thank David McConkey and Andrew Mason for critical review of a previous version of the manuscript, Jaime Martínez de Villarreal and other members of the Epithelial Carcinogenesis Group for valuable contributions.
Funding
This work was supported, in part, by a grant from Fundación Científica de la Asociación Española Contra el Cáncer to FXR and EL (PRYGN223005REAL). The project that gave rise to these results received the support of a fellowship from “la Caixa” Foundation (ID 100010434). The fellowship code is LCF/BQ/DR20/11790014. SC was supported by Fellowship PRE2018-085808 from Agencia Estatal de Investigación, co-financed by Fondo Social Europeo. CNIO is supported by Ministerio de Ciencia, Innovación y Universidades as a Centro de Excelencia Severo Ochoa SEV-2015-0510.
Author information
Authors and Affiliations
Contributions
All authors reviewed and summarized literature on the topic. MR also performed new bioinformatics analyses included in the manuscript. SC took the main responsibility for the illustrations.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ramal, M., Corral, S., Kalisz, M. et al. The urothelial gene regulatory network: understanding biology to improve bladder cancer management. Oncogene 43, 1–21 (2024). https://doi.org/10.1038/s41388-023-02876-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-023-02876-3